

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134049.2 - phase: 0
(170 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
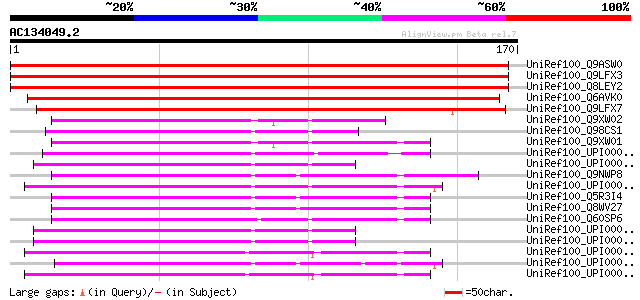
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ASW0 At1g27150/T7N9_21 [Arabidopsis thaliana] 225 4e-58
UniRef100_Q9LFX3 T7N9.21 [Arabidopsis thaliana] 225 4e-58
UniRef100_Q8LEY2 Hypothetical protein [Arabidopsis thaliana] 219 2e-56
UniRef100_Q6AVK0 Expressed protein, having alternative splicing ... 205 3e-52
UniRef100_Q9LFX7 T7N9.17 [Arabidopsis thaliana] 192 2e-48
UniRef100_Q9XW02 Hypothetical protein Y54G11A.4 [Caenorhabditis ... 78 8e-14
UniRef100_Q98CS1 Mlr5032 protein [Rhizobium loti] 78 1e-13
UniRef100_Q9XW01 Hypothetical protein Y54G11A.7 [Caenorhabditis ... 77 2e-13
UniRef100_UPI00002A8B75 UPI00002A8B75 UniRef100 entry 75 7e-13
UniRef100_UPI00003AF447 UPI00003AF447 UniRef100 entry 72 4e-12
UniRef100_Q9NWP8 Hypothetical protein FLJ20699 [Homo sapiens] 72 6e-12
UniRef100_UPI00003615B8 UPI00003615B8 UniRef100 entry 72 7e-12
UniRef100_Q5R3I4 OTTHUMP00000028646 [Homo sapiens] 71 1e-11
UniRef100_Q8WV27 FLJ20699 protein [Homo sapiens] 71 1e-11
UniRef100_Q60SP6 Hypothetical protein CBG20808 [Caenorhabditis b... 71 1e-11
UniRef100_UPI00003AFC7F UPI00003AFC7F UniRef100 entry 70 2e-11
UniRef100_UPI00003AF61D UPI00003AF61D UniRef100 entry 70 2e-11
UniRef100_UPI000024C387 UPI000024C387 UniRef100 entry 70 2e-11
UniRef100_UPI0000318DFB UPI0000318DFB UniRef100 entry 69 4e-11
UniRef100_UPI000024C3C1 UPI000024C3C1 UniRef100 entry 69 6e-11
>UniRef100_Q9ASW0 At1g27150/T7N9_21 [Arabidopsis thaliana]
Length = 468
Score = 225 bits (573), Expect = 4e-58
Identities = 102/167 (61%), Positives = 133/167 (79%)
Query: 1 MGINHHVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQ 60
+G+ VLP N+ E++I+G+LAFPLLELG+M+EA A+++G+EIN +D W+ H CHVLQ
Sbjct: 144 LGLVQQVLPANQEESYIHGLLAFPLLELGRMEEAAAASRKGYEINKEDAWAHHCLCHVLQ 203
Query: 61 YECRFREAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKEL 120
+ECRF+EAVEFME + +W S SFM THNWWHVALCYLEG +PM +V E+YD++IWKEL
Sbjct: 204 HECRFKEAVEFMEALAGTWPSCSSFMYTHNWWHVALCYLEGGSPMSKVEEIYDHHIWKEL 263
Query: 121 DKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQVSY 167
+K DA PEVYLNA+ LL+RL VRD L+ F DRLK LA RL +Q ++
Sbjct: 264 EKDDAVPPEVYLNALGLLIRLDVRDALDGFEDRLKNLAVRLTNQANW 310
>UniRef100_Q9LFX3 T7N9.21 [Arabidopsis thaliana]
Length = 468
Score = 225 bits (573), Expect = 4e-58
Identities = 102/167 (61%), Positives = 133/167 (79%)
Query: 1 MGINHHVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQ 60
+G+ VLP N+ E++I+G+LAFPLLELG+M+EA A+++G+EIN +D W+ H CHVLQ
Sbjct: 144 LGLVQQVLPANQEESYIHGLLAFPLLELGRMEEAAAASRKGYEINKEDAWAHHCLCHVLQ 203
Query: 61 YECRFREAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKEL 120
+ECRF+EAVEFME + +W S SFM THNWWHVALCYLEG +PM +V E+YD++IWKEL
Sbjct: 204 HECRFKEAVEFMEALAGTWPSCSSFMYTHNWWHVALCYLEGGSPMSKVEEIYDHHIWKEL 263
Query: 121 DKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQVSY 167
+K DA PEVYLNA+ LL+RL VRD L+ F DRLK LA RL +Q ++
Sbjct: 264 EKDDAVPPEVYLNALGLLIRLDVRDALDGFEDRLKNLAVRLTNQANW 310
>UniRef100_Q8LEY2 Hypothetical protein [Arabidopsis thaliana]
Length = 468
Score = 219 bits (559), Expect = 2e-56
Identities = 101/167 (60%), Positives = 132/167 (78%)
Query: 1 MGINHHVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQ 60
+G+ VLP N+ E++I+G+LAFPLLELG+M+EA A+++G+EIN +D W+ H CHVLQ
Sbjct: 144 LGLVQQVLPANQEESYIHGLLAFPLLELGRMEEAAAASRKGYEINKEDAWAHHCLCHVLQ 203
Query: 61 YECRFREAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKEL 120
+ECRF+EAVEFME + +W S SFM THNW HVALCYLEG +PM +V E+YD++IWKEL
Sbjct: 204 HECRFKEAVEFMEALAGTWPSCSSFMYTHNWRHVALCYLEGGSPMSKVEEIYDHHIWKEL 263
Query: 121 DKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQVSY 167
+K DA PEVYLNA+ LL+RL VRD L+ F DRLK LA RL +Q ++
Sbjct: 264 EKDDAVPPEVYLNALGLLIRLDVRDALDGFEDRLKNLAVRLTNQANW 310
>UniRef100_Q6AVK0 Expressed protein, having alternative splicing products [Oryza
sativa]
Length = 424
Score = 205 bits (522), Expect = 3e-52
Identities = 93/158 (58%), Positives = 118/158 (73%)
Query: 7 VLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFR 66
VLP+N+ +N+IYGMLAFPLLELG+M +AE+AA++G IN D WSQH CHV Q EC F+
Sbjct: 105 VLPENQDQNYIYGMLAFPLLELGRMDDAEKAARKGLAINKNDCWSQHNLCHVFQQECHFK 164
Query: 67 EAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDAT 126
EA EFM+ CSPSW + SFMLTHNWWHVA+CYLEG P +VLE+YD+ EL+K+D
Sbjct: 165 EATEFMKSCSPSWAACSSFMLTHNWWHVAVCYLEGEFPTSKVLEIYDHNFMTELEKSDCE 224
Query: 127 VPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQ 164
EVYLNA+ LLLRL +R +++ DRL L D L ++
Sbjct: 225 AAEVYLNALGLLLRLHIRGQVDLAKDRLAALLDALTNE 262
>UniRef100_Q9LFX7 T7N9.17 [Arabidopsis thaliana]
Length = 519
Score = 192 bits (489), Expect = 2e-48
Identities = 92/158 (58%), Positives = 120/158 (75%), Gaps = 1/158 (0%)
Query: 10 QNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAV 69
+NEG+ ++ GMLAF L+ELG ++EAEEAA++G EIN D W+ HA CHVLQ ECRF+EAV
Sbjct: 132 KNEGQVYVNGMLAFCLIELGHLREAEEAARKGCEINENDSWAHHALCHVLQTECRFKEAV 191
Query: 70 EFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPE 129
+FMEE S SW+S S +HNWWHVA+CYLEG + + +V EVYD+ +WKEL+K DA +
Sbjct: 192 KFMEEHSDSWDSCSSLRFSHNWWHVAVCYLEGGSHISKVEEVYDHQMWKELEKDDAVARD 251
Query: 130 VYLNAVALLLRLCVRDEL-EFFGDRLKMLADRLADQVS 166
VY +A+ LLLRL R +L + F DRL+ LAD L D+VS
Sbjct: 252 VYTDALGLLLRLDTRGKLDDGFQDRLEKLADSLTDKVS 289
>UniRef100_Q9XW02 Hypothetical protein Y54G11A.4 [Caenorhabditis elegans]
Length = 497
Score = 78.2 bits (191), Expect = 8e-14
Identities = 42/113 (37%), Positives = 63/113 (55%), Gaps = 4/113 (3%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
++++GM AF L E G +AE+ A R ++N D W+ HA HVL+ R +E EFM +
Sbjct: 180 SYLHGMYAFGLEECGIYGDAEKQADRALQLNRFDCWASHAKAHVLEMNGRHKEGKEFMYK 239
Query: 75 CSPSWNSFLSFML-THNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDAT 126
W +ML HN+WH AL ++E +A + LE++D I K K D +
Sbjct: 240 TEDDWRQ--GWMLAAHNYWHTALFHIE-SAEYEPALEIFDREIVKRSSKVDGS 289
>UniRef100_Q98CS1 Mlr5032 protein [Rhizobium loti]
Length = 440
Score = 77.8 bits (190), Expect = 1e-13
Identities = 39/105 (37%), Positives = 60/105 (57%), Gaps = 2/105 (1%)
Query: 13 GENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFM 72
G + I GM AF L E+G AE+ + EI +DGW+QHA HV++ + R R+ + +M
Sbjct: 158 GYHAILGMQAFGLEEMGDYTRAEKLGRTAVEIEPRDGWAQHAVAHVMEMQSRQRDGIVWM 217
Query: 73 EECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIW 117
+W SF+ HNWWH+AL + + + +VL +YD I+
Sbjct: 218 RANPEAWTK-ESFLQVHNWWHLALFHYD-LGEIDQVLALYDGPIY 260
>UniRef100_Q9XW01 Hypothetical protein Y54G11A.7 [Caenorhabditis elegans]
Length = 467
Score = 76.6 bits (187), Expect = 2e-13
Identities = 46/128 (35%), Positives = 71/128 (54%), Gaps = 6/128 (4%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
++++GM AF L E G +AE A R ++N D W+ HA HVL+ R +E EFM +
Sbjct: 180 SYLHGMYAFGLEECGIYDDAETQADRALQLNRFDCWASHAKAHVLEMNGRHKEGKEFMYK 239
Query: 75 CSPSWNSFLSFML-THNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLN 133
W +ML +HN+WH AL ++E A + L ++D I +KT++ + V +
Sbjct: 240 TEDDWRQ--GWMLASHNYWHTALFHIE-YAEYESALGIFDREIANRFNKTNSLLDMV--D 294
Query: 134 AVALLLRL 141
A +LL RL
Sbjct: 295 ASSLLWRL 302
>UniRef100_UPI00002A8B75 UPI00002A8B75 UniRef100 entry
Length = 464
Score = 75.1 bits (183), Expect = 7e-13
Identities = 42/130 (32%), Positives = 66/130 (50%), Gaps = 6/130 (4%)
Query: 12 EGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEF 71
+G F+ G F L E G +EAE ++ +IN D W+ HA HVL+ + R E +++
Sbjct: 159 DGYGFVLGCRCFALEEAGHYEEAEPIGRKAVDINENDIWAGHAVAHVLEMQGRRVEGIDW 218
Query: 72 MEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVY 131
+ E +W H WWH AL YLE N ++ V +DN W E + + +
Sbjct: 219 ITEHEEAWRK-RGLFSKHLWWHRALHYLELN-DIEAVKASFDNEFWAEPSEDNIDI---- 272
Query: 132 LNAVALLLRL 141
N+ ++L+RL
Sbjct: 273 CNSSSMLMRL 282
>UniRef100_UPI00003AF447 UPI00003AF447 UniRef100 entry
Length = 188
Score = 72.4 bits (176), Expect = 4e-12
Identities = 34/108 (31%), Positives = 56/108 (51%), Gaps = 2/108 (1%)
Query: 9 PQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREA 68
P+ +++ G AF L+E AEE A+ +IN D WS H H+ + + +
Sbjct: 56 PEVPLSSYVKGYYAFGLMESNFFDRAEELAREALDINRTDAWSVHTIAHINEMKAEVEKG 115
Query: 69 VEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYI 116
+ FM+E +W ++THN+WH AL ++E + L +YDN+I
Sbjct: 116 LAFMKETEDNWKD-SDMLITHNYWHWALGFIE-KGEYEAALTIYDNHI 161
>UniRef100_Q9NWP8 Hypothetical protein FLJ20699 [Homo sapiens]
Length = 336
Score = 72.0 bits (175), Expect = 6e-12
Identities = 41/143 (28%), Positives = 68/143 (46%), Gaps = 4/143 (2%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
+++ G+ +F L+E +AE+ AK IN D WS H H+ + + ++ +EFM+
Sbjct: 180 SYVKGIYSFGLMETNFYDQAEKLAKEALSINPTDAWSVHTVAHIHEMKAEIKDGLEFMQH 239
Query: 75 CSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNA 134
W + HN+WH AL YL + L +YD +I L DA + V ++
Sbjct: 240 SETLWKD-SDMLACHNYWHWAL-YLIEKGEYEAALTIYDTHILPSLQANDAMLDVV--DS 295
Query: 135 VALLLRLCVRDELEFFGDRLKML 157
++L RL + G R+ L
Sbjct: 296 CSMLYRLQMEGVSVGHGGRMSCL 318
>UniRef100_UPI00003615B8 UPI00003615B8 UniRef100 entry
Length = 456
Score = 71.6 bits (174), Expect = 7e-12
Identities = 44/145 (30%), Positives = 71/145 (48%), Gaps = 9/145 (6%)
Query: 6 HVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRF 65
H P N ++ GML+F LLE +AE+ A G + D W+ H+ HV +
Sbjct: 161 HWKPHNPLYRYLNGMLSFGLLETRLYDQAEKVAMEGLALTPDDAWAVHSLAHVYEMRAEV 220
Query: 66 REAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDA 125
+++ F E W S + +HN WH AL ++E + LE++D+ I++ + A
Sbjct: 221 DKSLSFFERTEKDWQS-ADMLASHNNWHWALNFIE-KGQYEAALEIFDSKIFRFCKSSGA 278
Query: 126 TVPEVYLNAVALLLRL-----CVRD 145
+ V +A +LL RL CV+D
Sbjct: 279 MLDIV--DACSLLYRLEMEGVCVKD 301
>UniRef100_Q5R3I4 OTTHUMP00000028646 [Homo sapiens]
Length = 469
Score = 71.2 bits (173), Expect = 1e-11
Identities = 38/127 (29%), Positives = 63/127 (48%), Gaps = 4/127 (3%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
+++ G+ +F L+E +AE+ AK IN D WS H H+ + + ++ +EFM+
Sbjct: 180 SYVKGIYSFGLMETNFYDQAEKLAKEALSINPTDAWSVHTVAHIHEMKAEIKDGLEFMQH 239
Query: 75 CSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNA 134
W + HN+WH AL YL + L +YD +I L DA + V ++
Sbjct: 240 SETFWKD-SDMLACHNYWHWAL-YLIEKGEYEAALTIYDTHILPSLQANDAMLDVV--DS 295
Query: 135 VALLLRL 141
++L RL
Sbjct: 296 CSMLYRL 302
>UniRef100_Q8WV27 FLJ20699 protein [Homo sapiens]
Length = 469
Score = 71.2 bits (173), Expect = 1e-11
Identities = 38/127 (29%), Positives = 63/127 (48%), Gaps = 4/127 (3%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
+++ G+ +F L+E +AE+ AK IN D WS H H+ + + ++ +EFM+
Sbjct: 180 SYVKGIYSFGLMETNFYDQAEKLAKEALSINPTDAWSVHTVAHIHEMKAEIKDGLEFMQH 239
Query: 75 CSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNA 134
W + HN+WH AL YL + L +YD +I L DA + V ++
Sbjct: 240 SETLWKD-SDMLACHNYWHWAL-YLIEKGEYEAALTIYDTHILPSLQANDAMLDVV--DS 295
Query: 135 VALLLRL 141
++L RL
Sbjct: 296 CSMLYRL 302
>UniRef100_Q60SP6 Hypothetical protein CBG20808 [Caenorhabditis briggsae]
Length = 467
Score = 70.9 bits (172), Expect = 1e-11
Identities = 41/127 (32%), Positives = 67/127 (52%), Gaps = 4/127 (3%)
Query: 15 NFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEE 74
++++GM AF L E G +AE A++ +N D W+ HA HVL+ R +E EFM
Sbjct: 180 SYLHGMYAFGLEECGLYGDAETEAEKALSLNRFDCWASHAKAHVLEMNGRHKEGKEFMYR 239
Query: 75 CSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNA 134
W + THN+WH AL ++E + L ++D I K ++++ + V +A
Sbjct: 240 TEDDWRKGW-MIATHNYWHTALFHIE-FGEYEDALSIFDREISKRFLRSNSLLDMV--DA 295
Query: 135 VALLLRL 141
++L RL
Sbjct: 296 SSILWRL 302
>UniRef100_UPI00003AFC7F UPI00003AFC7F UniRef100 entry
Length = 404
Score = 70.5 bits (171), Expect = 2e-11
Identities = 34/108 (31%), Positives = 54/108 (49%), Gaps = 2/108 (1%)
Query: 9 PQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREA 68
P+ +++ G +F L+E AEE A+ +IN D WS H HV + + +
Sbjct: 112 PEVPLSSYVKGFYSFGLMETNFFDRAEELAREALDINRTDAWSVHTIAHVNEMKAEVEKG 171
Query: 69 VEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYI 116
+ FM+E +W + HN+WH AL Y+E + L +YDN+I
Sbjct: 172 LAFMKETEDNWKD-SDMLACHNYWHWALYYVE-KGEYEAALTIYDNHI 217
>UniRef100_UPI00003AF61D UPI00003AF61D UniRef100 entry
Length = 244
Score = 70.5 bits (171), Expect = 2e-11
Identities = 34/108 (31%), Positives = 54/108 (49%), Gaps = 2/108 (1%)
Query: 9 PQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREA 68
P+ +++ G +F L+E AEE A+ +IN D WS H HV + + +
Sbjct: 112 PEVPLSSYVKGFYSFGLMETNFFDRAEELAREALDINRTDAWSVHTIAHVNEMKAEVEKG 171
Query: 69 VEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYI 116
+ FM+E +W + HN+WH AL Y+E + L +YDN+I
Sbjct: 172 LAFMKETEDNWKD-SDMLACHNYWHWALYYVE-KGEYEAALTIYDNHI 217
>UniRef100_UPI000024C387 UPI000024C387 UniRef100 entry
Length = 458
Score = 70.1 bits (170), Expect = 2e-11
Identities = 43/137 (31%), Positives = 70/137 (50%), Gaps = 6/137 (4%)
Query: 6 HVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRF 65
H P ++I GM +F LLE EAE+ AK + +DGWS HA HV + +
Sbjct: 163 HWKPHMPLGSYIKGMYSFGLLETRLYDEAEKMAKEALSLTPEDGWSVHAVAHVHEMKAEV 222
Query: 66 REAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLE-GNAPMQRVLEVYDNYIWKELDKTD 124
+ + FM +W + + HN+WH AL ++E GN + L+++D + + K+
Sbjct: 223 EKGLNFMASTEKNW-TVCDMLACHNYWHWALYHIEKGN--YEAALKIFDEQVSQRCVKSG 279
Query: 125 ATVPEVYLNAVALLLRL 141
A + V ++ +LL RL
Sbjct: 280 AMLDIV--DSCSLLYRL 294
>UniRef100_UPI0000318DFB UPI0000318DFB UniRef100 entry
Length = 466
Score = 69.3 bits (168), Expect = 4e-11
Identities = 40/135 (29%), Positives = 72/135 (52%), Gaps = 9/135 (6%)
Query: 16 FIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEEC 75
++ GML+F LLE +AE+AA G + D W+ HA HV + + + F +
Sbjct: 180 YLNGMLSFGLLETRLYDQAEKAAMAGLALTPDDAWAVHALAHVYEMRAEVDKGLSFFQRT 239
Query: 76 SPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNAV 135
W S + +HN+WH AL YL + LE++D+ +++ L ++ ++ ++ +++
Sbjct: 240 EKDWQS-ADILASHNYWHWAL-YLVEKGQYEEALEIFDSQVFR-LCRSSGSMLDM-VDSC 295
Query: 136 ALLLRL-----CVRD 145
+LL RL CV+D
Sbjct: 296 SLLYRLEMEGVCVKD 310
>UniRef100_UPI000024C3C1 UPI000024C3C1 UniRef100 entry
Length = 456
Score = 68.6 bits (166), Expect = 6e-11
Identities = 43/137 (31%), Positives = 69/137 (49%), Gaps = 6/137 (4%)
Query: 6 HVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRF 65
H P ++I GM +F LLE EAE+ AK + +DGWS HA HV + +
Sbjct: 161 HWKPHMPLGSYIKGMYSFGLLETRLYDEAEKMAKEALSLTPEDGWSVHAVAHVHEMKAEV 220
Query: 66 REAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLE-GNAPMQRVLEVYDNYIWKELDKTD 124
+ + FM +W + HN+WH AL ++E GN + L+++D + + K+
Sbjct: 221 DKGLNFMASTEKNW-MVCDMLACHNYWHWALYHIEKGN--YEAALKIFDEQVSQRCVKSG 277
Query: 125 ATVPEVYLNAVALLLRL 141
A + V ++ +LL RL
Sbjct: 278 AMLDIV--DSCSLLYRL 292
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 293,282,302
Number of Sequences: 2790947
Number of extensions: 11654560
Number of successful extensions: 24840
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 24747
Number of HSP's gapped (non-prelim): 76
length of query: 170
length of database: 848,049,833
effective HSP length: 118
effective length of query: 52
effective length of database: 518,718,087
effective search space: 26973340524
effective search space used: 26973340524
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC134049.2


