

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133780.8 - phase: 0 /pseudo
(287 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
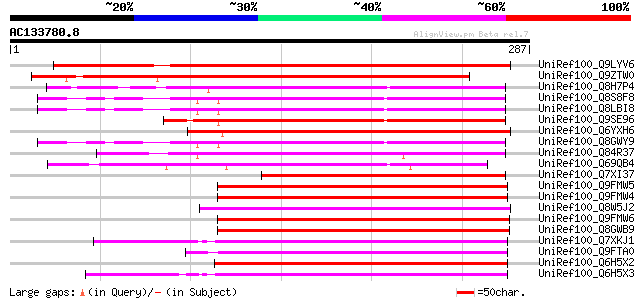
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LYV6 ABA-responsive protein-like [Arabidopsis thaliana] 313 4e-84
UniRef100_Q9ZTW0 ABA-responsive protein [Hordeum vulgare] 241 2e-62
UniRef100_Q8H7P4 Putative FH protein interacting protein FIP1 [O... 189 5e-47
UniRef100_Q8S8F8 Expressed protein [Arabidopsis thaliana] 187 2e-46
UniRef100_Q8LBI8 Hypothetical protein [Arabidopsis thaliana] 187 3e-46
UniRef100_Q9SE96 FH protein interacting protein FIP1 [Arabidopsi... 187 4e-46
UniRef100_Q6YXH6 FH protein interacting protein FIP1-like [Oryza... 185 1e-45
UniRef100_Q8GWY9 Hypothetical protein At2g22475/F14M13.2 [Arabid... 184 3e-45
UniRef100_Q84R37 Putative ABA-responsive protein [Oryza sativa] 176 5e-43
UniRef100_Q69QB4 Putative ABA-responsive protein [Oryza sativa] 162 1e-38
UniRef100_Q7XI37 Putative FH protein interacting protein FIP1 [O... 151 2e-35
UniRef100_Q9FMW5 Similarity to ABA-responsive protein [Arabidops... 141 2e-32
UniRef100_Q9FMW4 Similarity to ABA-responsive protein [Arabidops... 140 3e-32
UniRef100_Q8W5J2 Putative ABA-responsive protein [Oryza sativa] 140 5e-32
UniRef100_Q9FMW6 Similarity to ABA-repsonsive protein [Arabidops... 138 1e-31
UniRef100_Q8GWB9 Hypothetical protein At5g23350/MKD15_21 [Arabid... 138 1e-31
UniRef100_Q7XKJ1 OSJNBa0038O10.17 protein [Oryza sativa] 138 2e-31
UniRef100_Q9FTA0 Hypothetical protein F8L15_80 [Arabidopsis thal... 137 3e-31
UniRef100_Q6H5X2 ABA-responsive protein-like [Oryza sativa] 134 2e-30
UniRef100_Q6H5X3 ABA-responsive protein-like [Oryza sativa] 128 2e-28
>UniRef100_Q9LYV6 ABA-responsive protein-like [Arabidopsis thaliana]
Length = 272
Score = 313 bits (801), Expect = 4e-84
Identities = 148/254 (58%), Positives = 189/254 (74%), Gaps = 9/254 (3%)
Query: 25 PPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYY 84
P + + P P + + WGTH+MGAPA P +HPDN++AA A + Q Q Q
Sbjct: 18 PKTLETEHQPEPSSSSPDQKKWGTHVMGAPAAPVAHPDNQQAAAWVAGDNQQTQYQ---- 73
Query: 85 QQDHPYVQHSPVDKPSSS-PMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKM 143
PYV +SPV+ P+++ P+E ++ MF +WS+KAE A N+WHNLKTGPS+S A GK+
Sbjct: 74 ----PYVIYSPVEHPTTNNPLEPVIGMFHTWSRKAETVARNLWHNLKTGPSMSETAWGKV 129
Query: 144 NLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDR 203
NLT KAI++GGFESL++QIF T PNE LKKTFACYLSTTTGPVAGT+YLS+ +AFCSDR
Sbjct: 130 NLTAKAITKGGFESLFRQIFGTEPNETLKKTFACYLSTTTGPVAGTVYLSNARVAFCSDR 189
Query: 204 PLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNY 263
PL FTAPSGQ +WSYY+V+VPL + TVNPV+++E E+YIQ+ TVDGHDFWFMGFVNY
Sbjct: 190 PLYFTAPSGQESWSYYRVVVPLANVATVNPVVVKETPPEKYIQLTTVDGHDFWFMGFVNY 249
Query: 264 DKAVKNLSEGISHF 277
+KA +L +S F
Sbjct: 250 EKATHHLLTSVSDF 263
>UniRef100_Q9ZTW0 ABA-responsive protein [Hordeum vulgare]
Length = 326
Score = 241 bits (615), Expect = 2e-62
Identities = 125/249 (50%), Positives = 163/249 (65%), Gaps = 10/249 (4%)
Query: 13 SSPDENPFASPTPPSSSS----SSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAAL 68
++P A+P+P + + + P PP A + WGT MG PA P +HP+N++AA
Sbjct: 81 AAPQPGTGATPSPLTQQAVDGKDAAPTPP---AESARWGTRQMGPPAAPGAHPENQQAAQ 137
Query: 69 QTASADGQGQPQ--VHYYQQDHPYVQHSPVDKPS-SSPMESILHMFDSWSKKAEATANNI 125
TA+ Q P + + P DK + SPME IL F++WS+KAE ++NI
Sbjct: 138 WTATRGDQELPPYVIMGAPEPAPAASARRTDKEARDSPMEHILDFFNTWSRKAEELSSNI 197
Query: 126 WHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGP 185
W NLKT PS+S AAMGK++L KAI+ GGFE LYKQ F + P+E +KKTFACYLST TGP
Sbjct: 198 WLNLKTAPSMSDAAMGKLSLGAKAITGGGFEKLYKQTFGSGPDEHVKKTFACYLSTATGP 257
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
VAGTLYL++ ++AFCSDRPLSF APSGQ WSYYKVM+PL K+ V PV +E+ ERYI
Sbjct: 258 VAGTLYLTNTNVAFCSDRPLSFAAPSGQTAWSYYKVMIPLAKLAAVEPVTAKESPPERYI 317
Query: 246 QIVTVDGHD 254
IV ++
Sbjct: 318 HIVAAPAYE 326
>UniRef100_Q8H7P4 Putative FH protein interacting protein FIP1 [Oryza sativa]
Length = 264
Score = 189 bits (481), Expect = 5e-47
Identities = 102/258 (39%), Positives = 149/258 (57%), Gaps = 19/258 (7%)
Query: 21 ASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQ 80
A+P P + ++++ ++A + P PS H A +++ G P
Sbjct: 14 AAPAPATGAAAATA---SDYAHYPRLSPEDVAPPPPPSYH------AAASSAPPYSGNPY 64
Query: 81 VHYYQQDHPYVQHSPVDKPSSSPMESIL----HMFDSWSKKAEATANNIWHNLKTGPSVS 136
V P +P K + ++ +L F ++K E N W +LKTGPS++
Sbjct: 65 V-----SSPAGGVAPASKNTMDTVKDVLGKMGKRFGEAARKTETLTGNFWQHLKTGPSIT 119
Query: 137 SAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIH 196
AAMG+++ K I+EGG++ ++ Q F P+EKLKK +ACYLST+ GPV G LYLS+
Sbjct: 120 DAAMGRVSQITKVIAEGGYDKIFHQTFDVLPDEKLKKPYACYLSTSAGPVMGVLYLSNKK 179
Query: 197 LAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFW 256
LAFCSD PL++ + WSYYKV++P ++ +VNP R N SE+YIQ+V+VD H+FW
Sbjct: 180 LAFCSDNPLAYKV-GDKDEWSYYKVVIPHTQLRSVNPSTSRTNASEKYIQVVSVDNHEFW 238
Query: 257 FMGFVNYDKAVKNLSEGI 274
FMGFV YD AVKNL E +
Sbjct: 239 FMGFVYYDSAVKNLQEAL 256
>UniRef100_Q8S8F8 Expressed protein [Arabidopsis thaliana]
Length = 299
Score = 187 bits (476), Expect = 2e-46
Identities = 100/269 (37%), Positives = 149/269 (55%), Gaps = 39/269 (14%)
Query: 16 DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADG 75
D P +P+ SS S + +W ++ S PD K A+ +A ++
Sbjct: 52 DSTPVKAPSRTSSGSKK----------SVHWSPELVSG----SQEPDQKAASSSSAGSN- 96
Query: 76 QGQPQVHYYQQDHPYVQHSPVDKPSSS---PMESILHMFDSW-------SKKAEATANNI 125
PY+ SP + +S ME++ + W +KK E+ A N
Sbjct: 97 -------------PYIARSPAETSDASLKDTMETVKGVLGRWGKRVAEAAKKTESLAGNT 143
Query: 126 WHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGP 185
W +L+T PS + AAMG++ + K +EGG+E +++Q F T P E+L +FACYLST+ GP
Sbjct: 144 WQHLRTAPSFADAAMGRIAQSTKVFAEGGYEKIFRQTFETDPEEQLLNSFACYLSTSAGP 203
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
V G LY+S LA+CSD PLS+ Q WSYYKV++PL ++ VNP N +E+YI
Sbjct: 204 VMGVLYISSAKLAYCSDNPLSY-KNGDQTEWSYYKVVIPLHQLKAVNPSASIVNPAEKYI 262
Query: 246 QIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
Q+++VD H+FWFMGF+NYD AV +L + +
Sbjct: 263 QVISVDNHEFWFMGFLNYDGAVTSLQDSL 291
>UniRef100_Q8LBI8 Hypothetical protein [Arabidopsis thaliana]
Length = 299
Score = 187 bits (475), Expect = 3e-46
Identities = 100/269 (37%), Positives = 148/269 (54%), Gaps = 39/269 (14%)
Query: 16 DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADG 75
D P +P+ SS S + +W ++ S PD K A+ +A ++
Sbjct: 52 DSTPVKAPSRTSSGSKK----------SVHWSPELVSG----SQEPDQKAASSSSAGSN- 96
Query: 76 QGQPQVHYYQQDHPYVQHSPVDKPSSS---PMESILHMFDSW-------SKKAEATANNI 125
PY+ SP + +S ME++ + W +KK E+ A N
Sbjct: 97 -------------PYIARSPAETSDASLKDTMETVKGVLGRWGKRVAEAAKKTESLAGNT 143
Query: 126 WHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGP 185
W L+T PS + AAMG++ + K +EGG+E +++Q F T P E+L +FACYLST+ GP
Sbjct: 144 WQQLRTAPSFADAAMGRIAQSTKVFAEGGYEKIFRQTFETDPEEQLLNSFACYLSTSAGP 203
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
V G LY+S LA+CSD PLS+ Q WSYYKV++PL ++ VNP N +E+YI
Sbjct: 204 VMGVLYISSAKLAYCSDNPLSY-KNGDQTEWSYYKVVIPLHQLKAVNPSASIVNPAEKYI 262
Query: 246 QIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
Q+++VD H+FWFMGF+NYD AV +L + +
Sbjct: 263 QVISVDNHEFWFMGFLNYDGAVTSLQDSL 291
>UniRef100_Q9SE96 FH protein interacting protein FIP1 [Arabidopsis thaliana]
Length = 259
Score = 187 bits (474), Expect = 4e-46
Identities = 91/196 (46%), Positives = 127/196 (64%), Gaps = 11/196 (5%)
Query: 86 QDHPYVQHSPVDKPSSSPMESILHMFDSW-------SKKAEATANNIWHNLKTGPSVSSA 138
+ +PYV SP + + M+S+ W +KKAE A N W +LKTGPSV+ A
Sbjct: 63 ESNPYVSPSPAPR---NTMDSVKDTLGKWGKMAADATKKAEDLAGNFWQHLKTGPSVADA 119
Query: 139 AMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLA 198
A+ ++ K ++EGG+E ++KQ F P+EKL KT+ACYLST+ GPV G +YLS LA
Sbjct: 120 AVSRIAQGTKILAEGGYEKVFKQTFDCLPDEKLLKTYACYLSTSAGPVLGVMYLSTHKLA 179
Query: 199 FCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFM 258
F SD PLS+ Q WSYYKV++P ++ VNP R N S++YIQ++++D H+FWFM
Sbjct: 180 FSSDNPLSY-KEGEQTLWSYYKVVLPANQLKAVNPSTSRVNTSDKYIQVISIDNHEFWFM 238
Query: 259 GFVNYDKAVKNLSEGI 274
GFV Y+ AVK+L E +
Sbjct: 239 GFVTYESAVKSLQEAV 254
>UniRef100_Q6YXH6 FH protein interacting protein FIP1-like [Oryza sativa]
Length = 192
Score = 185 bits (469), Expect = 1e-45
Identities = 84/186 (45%), Positives = 126/186 (67%), Gaps = 7/186 (3%)
Query: 99 PSSSPMESILHMFDSWSK-------KAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAIS 151
PSSS E++ + W++ KAE + N W +L+T PS+ AA+G++ K ++
Sbjct: 2 PSSSRTETVKNALSRWARRVGETTRKAEDLSRNTWQHLRTAPSIGEAAVGRIAQGTKVLA 61
Query: 152 EGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPS 211
EGG + +++Q F+ P+E+L+K++ACYLST+ GPV G LYLS +AFCSD PLS+ A
Sbjct: 62 EGGHDRIFRQAFSAPPDEQLRKSYACYLSTSAGPVMGILYLSTARVAFCSDSPLSYEAGG 121
Query: 212 GQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLS 271
G WSYYKV +PL ++ + +P ++ +E++IQ+V+VD H+FW MGFVNYD AVK+L
Sbjct: 122 GSKEWSYYKVAIPLHRLRSASPSASKQRPAEKFIQLVSVDRHEFWLMGFVNYDSAVKHLQ 181
Query: 272 EGISHF 277
E +S F
Sbjct: 182 EALSGF 187
>UniRef100_Q8GWY9 Hypothetical protein At2g22475/F14M13.2 [Arabidopsis thaliana]
Length = 299
Score = 184 bits (466), Expect = 3e-45
Identities = 99/269 (36%), Positives = 148/269 (54%), Gaps = 39/269 (14%)
Query: 16 DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADG 75
D P +P+ SS S + +W ++ S PD K A+ +A ++
Sbjct: 52 DSTPVKAPSRTSSGSKK----------SVHWSPELVSG----SQEPDQKAASSSSAGSN- 96
Query: 76 QGQPQVHYYQQDHPYVQHSPVDKPSSS---PMESILHMFDSW-------SKKAEATANNI 125
PY+ SP + +S ME++ + W +KK E+ A N
Sbjct: 97 -------------PYIARSPAETSDASLKDTMETVKGVLGRWGKRVAEAAKKTESLAGNT 143
Query: 126 WHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGP 185
W +L+T PS + AAMG++ + K +EGG+E +++Q F T P E+L +FACYLST+ G
Sbjct: 144 WQHLRTAPSFADAAMGRIAQSTKVFAEGGYEKIFRQTFETDPEEQLLNSFACYLSTSAGL 203
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
V G LY+S LA+CSD PLS+ Q WSYYKV++PL ++ VNP N +E+YI
Sbjct: 204 VMGVLYISSAKLAYCSDNPLSY-KNGDQTEWSYYKVVIPLHQLKAVNPSASIVNPAEKYI 262
Query: 246 QIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
Q+++VD H+FWFMGF+NYD AV +L + +
Sbjct: 263 QVISVDNHEFWFMGFLNYDGAVTSLQDSL 291
>UniRef100_Q84R37 Putative ABA-responsive protein [Oryza sativa]
Length = 247
Score = 176 bits (447), Expect = 5e-43
Identities = 93/240 (38%), Positives = 135/240 (55%), Gaps = 24/240 (10%)
Query: 49 HMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSS------ 102
+ G HP A T ++G G +PYV +P S+
Sbjct: 14 YAQGQGGAQPPHPPQSTAVAVTPVSNGVG----------NPYVIVTPASASPSTCQSLRK 63
Query: 103 PMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQI 162
+E + ++KA T NIWH+L+T P+++ AA+ ++ K +EGG + ++ Q
Sbjct: 64 ALERYGRKLEDGTRKAADTTGNIWHHLRTAPNMADAAVARLAQGTKVYAEGGHDRVFTQA 123
Query: 163 FTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTW------ 216
F P E+L+K +ACYLST++GPV GTLY+S LAFCSD P+S+ AP+ V
Sbjct: 124 FGVVPGEQLRKAYACYLSTSSGPVIGTLYISTARLAFCSDSPISYHAPAVAVAGAAPAHP 183
Query: 217 --SYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
+ YKV++PL ++ +VNP N ERYIQI+T D H+FWFMGFV+YDKA+KNL E +
Sbjct: 184 PEAIYKVVLPLNQVKSVNPSASMTNRGERYIQIMTTDNHEFWFMGFVSYDKALKNLYEAL 243
>UniRef100_Q69QB4 Putative ABA-responsive protein [Oryza sativa]
Length = 409
Score = 162 bits (409), Expect = 1e-38
Identities = 88/261 (33%), Positives = 132/261 (49%), Gaps = 24/261 (9%)
Query: 22 SPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQV 81
S PP++S S P PP A P + P +K + + V
Sbjct: 23 SDDPPAASDPSSPPPPPPPVAVA-----AATADPPPPAQPQGQKTVTWSEKLTSESPTYV 77
Query: 82 HYYQ----QDHPYVQHSPVDKPSSSPMESILHMFDSWSKKA-------EATANNIWHNLK 130
+ YV P S +E++ W K E+ + + W + K
Sbjct: 78 AAATAEAAESSQYVSRGPASSSSKGAVEAMKETLSRWGKSVGETTKMVESLSRDTWQHFK 137
Query: 131 TGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTL 190
TGPS + AAMG++ K ++EGG+E +++Q F P E+LK ++ACYLST+ GPV G +
Sbjct: 138 TGPSFTEAAMGRLAQGTKVLAEGGYEKIFRQTFEVLPEEQLKISYACYLSTSAGPVMGVM 197
Query: 191 YLSDIHLAFCSDRPLSFTAPSGQVTWSYYK-------VMVPLGKIGTVNPVIMRENHSER 243
Y+S +AFCSD PLS+ A + WSYYK V++PL ++ NP + + N +E+
Sbjct: 198 YISTAKIAFCSDNPLSYKA-GNKTEWSYYKARIVHFLVVIPLHQLRAANPSVSKVNPAEK 256
Query: 244 YIQIVTVDGHDFWFMGFVNYD 264
YIQ+V+V+GH+FWFM V D
Sbjct: 257 YIQVVSVEGHEFWFMAKVIRD 277
>UniRef100_Q7XI37 Putative FH protein interacting protein FIP1 [Oryza sativa]
Length = 149
Score = 151 bits (382), Expect = 2e-35
Identities = 67/135 (49%), Positives = 96/135 (70%)
Query: 140 MGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAF 199
MG++ K I+EGG++ ++ Q F P+EKLKK +ACYLST+ GP+ G LY+S +AF
Sbjct: 1 MGRIAQISKVIAEGGYDKVFHQTFECLPDEKLKKAYACYLSTSHGPIMGVLYISTAKIAF 60
Query: 200 CSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMG 259
CSD P+++ + S YKV+VP+ ++ +V P ++N +ERYIQ+V+VD HDFWFMG
Sbjct: 61 CSDSPVAYVTEDNKNQSSIYKVVVPVAQLRSVTPTASQQNPAERYIQVVSVDNHDFWFMG 120
Query: 260 FVNYDKAVKNLSEGI 274
FVNYD AVK+L E +
Sbjct: 121 FVNYDGAVKSLQEAV 135
>UniRef100_Q9FMW5 Similarity to ABA-responsive protein [Arabidopsis thaliana]
Length = 210
Score = 141 bits (355), Expect = 2e-32
Identities = 67/160 (41%), Positives = 98/160 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N K GP ++ K++L K + GG E +YK++F EKL K +
Sbjct: 47 KKTDSFTNGARDQDKLGPKLTETVKRKLSLGAKILQMGGLEKIYKRLFKVCDKEKLFKAY 106
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT G +AG L++S +AFCS+R + T+P G +T +YKV +PL KI VN
Sbjct: 107 QCYLSTTEGSIAGLLFISSKKIAFCSERSIKVTSPQGDLTRVHYKVSIPLCKINGVNQSQ 166
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+ S+RY+++VTVD +DFWFMGFV+Y KA L + ++
Sbjct: 167 NTKKPSQRYLEVVTVDNYDFWFMGFVSYQKAFNCLEKALN 206
>UniRef100_Q9FMW4 Similarity to ABA-responsive protein [Arabidopsis thaliana]
Length = 219
Score = 140 bits (354), Expect = 3e-32
Identities = 65/160 (40%), Positives = 98/160 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N + K GP ++ K++L + + GG E +YK++F EKL K +
Sbjct: 55 KKNDSFTNGVRDQDKLGPKLTETVKRKLSLGARILQMGGLEKIYKRLFKVSDEEKLFKAY 114
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT GP+AG L++S +AFCS+R + +P G++ +YKV +PL KI VN
Sbjct: 115 QCYLSTTAGPIAGLLFISSKKIAFCSERSIKVASPQGELNRVHYKVSIPLCKINGVNQSQ 174
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
S++Y+++VTVDG DFWFMGF++Y KA L + +S
Sbjct: 175 NTTKPSQKYLEVVTVDGFDFWFMGFLSYQKAFNCLEQALS 214
>UniRef100_Q8W5J2 Putative ABA-responsive protein [Oryza sativa]
Length = 214
Score = 140 bits (352), Expect = 5e-32
Identities = 68/172 (39%), Positives = 101/172 (58%)
Query: 106 SILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTT 165
SI++ S+K ++ ++ GP +S GK++L K + G + +++Q F
Sbjct: 42 SIIYRMSKLSQKTDSYVQGFKEHITLGPKISDTLKGKLSLGAKVLQAGSIDKVFRQYFQV 101
Query: 166 YPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPL 225
+EKL K F CYLSTT GP+AG L++S +AF SDRPL T+P G +T YKV++P
Sbjct: 102 DKDEKLLKAFQCYLSTTAGPIAGMLFISTEKIAFHSDRPLDLTSPKGGITRVPYKVLIPA 161
Query: 226 GKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISHF 277
+I + N E+YI +VTVDG DFWFMGF+++ K+ + L IS F
Sbjct: 162 KRIKSAAVRENLYNPDEKYIDVVTVDGFDFWFMGFISHTKSFEYLQRVISEF 213
>UniRef100_Q9FMW6 Similarity to ABA-repsonsive protein [Arabidopsis thaliana]
Length = 280
Score = 138 bits (348), Expect = 1e-31
Identities = 65/161 (40%), Positives = 98/161 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N K GP ++ K++L K + GG E +YK++F EKL K +
Sbjct: 117 KKTDSFTNGARDQDKLGPKLTETVKRKLSLGAKILQMGGLEKIYKRLFKVCDQEKLFKAY 176
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT GP+AG L++S +AFCS+R + +P G ++ +YKV +PL KI VN
Sbjct: 177 QCYLSTTAGPIAGLLFISSKKIAFCSERSIKVASPQGVLSRVHYKVSIPLCKINGVNQSQ 236
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISH 276
+ S++Y++IVT+D DFWFMGFV+Y KA L + +++
Sbjct: 237 NTKKPSQKYLEIVTIDNFDFWFMGFVSYQKAFNCLEKALNN 277
>UniRef100_Q8GWB9 Hypothetical protein At5g23350/MKD15_21 [Arabidopsis thaliana]
Length = 218
Score = 138 bits (348), Expect = 1e-31
Identities = 65/161 (40%), Positives = 98/161 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N K GP ++ K++L K + GG E +YK++F EKL K +
Sbjct: 55 KKTDSFTNGARDQDKLGPKLTETVKRKLSLGAKILQMGGLEKIYKRLFKVCDQEKLFKAY 114
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT GP+AG L++S +AFCS+R + +P G ++ +YKV +PL KI VN
Sbjct: 115 QCYLSTTAGPIAGLLFISSKKIAFCSERSIKVASPQGVLSRVHYKVSIPLCKINGVNQSQ 174
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISH 276
+ S++Y++IVT+D DFWFMGFV+Y KA L + +++
Sbjct: 175 NTKKPSQKYLEIVTIDNFDFWFMGFVSYQKAFNCLEKALNN 215
>UniRef100_Q7XKJ1 OSJNBa0038O10.17 protein [Oryza sativa]
Length = 233
Score = 138 bits (347), Expect = 2e-31
Identities = 77/229 (33%), Positives = 117/229 (50%), Gaps = 6/229 (2%)
Query: 47 GTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
G H++G P + ++ +L + + HP S K SS
Sbjct: 8 GVHVIGVPVTAKAFGIEEEVSLARGQSFRKADGDHLAVSLSHPSPYTSFGYKHSSKGQ-- 65
Query: 107 ILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTY 166
++H W K A ++ GP +S GK++L K + GG E ++++ F+
Sbjct: 66 VIH----WVSKLSRRAQGFREHVTLGPKLSETVKGKLSLGAKILQAGGIERVFRKAFSAE 121
Query: 167 PNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLG 226
E+L K CYL TT GP+AG L++S +AF SDRP++ T+ G V YKV+VPL
Sbjct: 122 KGERLVKALQCYLYTTGGPIAGMLFVSTKKVAFRSDRPVTVTSAKGDVARVPYKVVVPLR 181
Query: 227 KIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+I V P + E+YI +VTVDG +FWFMGFV+Y ++ K + + IS
Sbjct: 182 RIAQVRPSENADKPEEKYIHVVTVDGFEFWFMGFVSYQRSCKYMQQAIS 230
>UniRef100_Q9FTA0 Hypothetical protein F8L15_80 [Arabidopsis thaliana]
Length = 222
Score = 137 bits (345), Expect = 3e-31
Identities = 68/178 (38%), Positives = 103/178 (57%), Gaps = 5/178 (2%)
Query: 98 KPSSSPMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFES 157
K S ++SIL KK + N + K P ++ K++L + + GG E
Sbjct: 41 KSEQSNVKSILKR-----KKTDGFTNGVRDQSKIRPKLTETVKRKLSLGARILQVGGLEK 95
Query: 158 LYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWS 217
++K++F EKL K + CYLSTT GP+AG L++S +AFCS+R + +P G +
Sbjct: 96 IFKRLFRVSEGEKLFKMYQCYLSTTAGPIAGLLFISSKKMAFCSERSIKVDSPQGDIIRV 155
Query: 218 YYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+YKV +PL KI VN + S++Y+++VTVDG DFWFMGF++Y KA L + +S
Sbjct: 156 HYKVSIPLCKIDRVNQSQNTKKPSQKYLEVVTVDGFDFWFMGFLSYQKAFNCLEKALS 213
>UniRef100_Q6H5X2 ABA-responsive protein-like [Oryza sativa]
Length = 232
Score = 134 bits (338), Expect = 2e-30
Identities = 62/162 (38%), Positives = 98/162 (60%)
Query: 114 WSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKK 173
+ +K + A I ++ GP +S GK+ L + + GG E +++Q F+ NEKL +
Sbjct: 67 YGRKGDKIAQGIKEHVTLGPKLSETVKGKLTLGARILQAGGVEKVFRQWFSVDKNEKLLR 126
Query: 174 TFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNP 233
CYLSTT GP+AG L++S +AF SDRPL+ +AP G YKV +PL K+ P
Sbjct: 127 ASQCYLSTTAGPIAGMLFVSTERVAFRSDRPLAVSAPGGDKVRVPYKVTIPLRKVKAAKP 186
Query: 234 VIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+ ++YI++VT DG +FWFMGFV+Y +++ +L + ++
Sbjct: 187 SENKHKPEQKYIEVVTNDGFEFWFMGFVSYHRSLHHLEQAVA 228
>UniRef100_Q6H5X3 ABA-responsive protein-like [Oryza sativa]
Length = 229
Score = 128 bits (322), Expect = 2e-28
Identities = 74/234 (31%), Positives = 117/234 (49%), Gaps = 9/234 (3%)
Query: 43 TENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSS 102
T++ H++G P ++ + + G PY SS
Sbjct: 2 TKSSCAHVVGVPVTSKAYAIEEATTARDGGKKVDGDRLAVSLTHPSPYTSFG---YKHSS 58
Query: 103 PMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQI 162
++ ++H W K A ++ GP +S GK++L + + GG E +++Q
Sbjct: 59 KLQ-VIH----WVNKLGRRAQGFRDHVTLGPKLSETVRGKLSLGARILQAGGVERVFRQA 113
Query: 163 FTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSY-YKV 221
F+ E+L K CYL TT GP+AG L++S+ +AF SDR L+ T+P+G V YKV
Sbjct: 114 FSAEKGERLVKALQCYLYTTGGPIAGMLFVSNRKIAFRSDRSLAVTSPAGDVVARVPYKV 173
Query: 222 MVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+VPL +I V P + ++YI + TVDG +FWFMGFV+Y + K + + IS
Sbjct: 174 VVPLRRIKRVRPSENADKPEQKYIHVATVDGFEFWFMGFVSYQRCCKYMQQVIS 227
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.127 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 539,253,087
Number of Sequences: 2790947
Number of extensions: 23890867
Number of successful extensions: 226118
Number of sequences better than 10.0: 883
Number of HSP's better than 10.0 without gapping: 360
Number of HSP's successfully gapped in prelim test: 552
Number of HSP's that attempted gapping in prelim test: 199878
Number of HSP's gapped (non-prelim): 12064
length of query: 287
length of database: 848,049,833
effective HSP length: 126
effective length of query: 161
effective length of database: 496,390,511
effective search space: 79918872271
effective search space used: 79918872271
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 74 (33.1 bits)
Medicago: description of AC133780.8


