

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133571.7 + phase: 0
(79 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
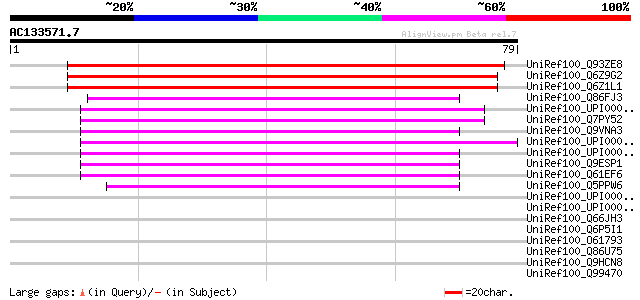
Sequences producing significant alignments: (bits) Value
UniRef100_Q93ZE8 Stromal cell-derived factor 2-like protein prec... 129 2e-29
UniRef100_Q6Z9G2 Putative stromal cell-derived factor 2 [Oryza s... 116 2e-25
UniRef100_Q6Z1L1 Stromal cell-derived factor 2-like protein [Ory... 115 3e-25
UniRef100_Q86FJ3 Clone ZZD1313 mRNA sequence [Schistosoma japoni... 55 6e-07
UniRef100_UPI00002441DE UPI00002441DE UniRef100 entry 50 1e-05
UniRef100_Q7PY52 ENSANGP00000013320 [Anopheles gambiae str. PEST] 50 1e-05
UniRef100_Q9VNA3 CG11999-PA [Drosophila melanogaster] 47 2e-04
UniRef100_UPI00003AB68F UPI00003AB68F UniRef100 entry 46 2e-04
UniRef100_UPI000017F603 UPI000017F603 UniRef100 entry 46 3e-04
UniRef100_Q9ESP1 Stromal cell-derived factor 2-like protein 1 pr... 46 3e-04
UniRef100_Q61EF6 Hypothetical protein CBG12091 [Caenorhabditis b... 45 3e-04
UniRef100_Q5PPW6 Hypothetical protein [Xenopus laevis] 44 8e-04
UniRef100_UPI00001CB008 UPI00001CB008 UniRef100 entry 44 0.001
UniRef100_UPI000036C832 UPI000036C832 UniRef100 entry 44 0.001
UniRef100_Q66JH3 MGC79547 protein [Xenopus tropicalis] 44 0.001
UniRef100_Q6P5I1 Stromal cell-derived factor 2 precursor [Mus mu... 44 0.001
UniRef100_O61793 Hypothetical protein R12E2.13 [Caenorhabditis e... 44 0.001
UniRef100_Q86U75 Dihydropyrimidinase-like 2 [Homo sapiens] 44 0.001
UniRef100_Q9HCN8 Stromal cell-derived factor 2-like protein 1 pr... 44 0.001
UniRef100_Q99470 Stromal cell-derived factor 2 precursor [Homo s... 44 0.001
>UniRef100_Q93ZE8 Stromal cell-derived factor 2-like protein precursor [Arabidopsis
thaliana]
Length = 218
Score = 129 bits (323), Expect = 2e-29
Identities = 57/68 (83%), Positives = 63/68 (91%)
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQR RLQHIDTSGY +SHDKKY RIAGGQQEVCG+REK+ADN+WLAAEGVY
Sbjct: 151 EGSGKTWKQDQRVRLQHIDTSGYLHSHDKKYQRIAGGQQEVCGIREKKADNIWLAAEGVY 210
Query: 70 LPVTESKQ 77
LP+ ES +
Sbjct: 211 LPLNESSK 218
>UniRef100_Q6Z9G2 Putative stromal cell-derived factor 2 [Oryza sativa]
Length = 217
Score = 116 bits (290), Expect = 2e-25
Identities = 49/67 (73%), Positives = 59/67 (87%)
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EG GK WKQDQ+ RL+H+DT GY +SH+KKY+R+ GGQQEVCGVREKRA+N+WLA EGVY
Sbjct: 151 EGGGKLWKQDQKVRLRHVDTGGYLHSHNKKYNRLGGGQQEVCGVREKRAENIWLATEGVY 210
Query: 70 LPVTESK 76
LPV +SK
Sbjct: 211 LPVNKSK 217
>UniRef100_Q6Z1L1 Stromal cell-derived factor 2-like protein [Oryza sativa]
Length = 217
Score = 115 bits (288), Expect = 3e-25
Identities = 49/67 (73%), Positives = 60/67 (89%)
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGK+W+Q+Q+ RL+H+DT GY +SHD+KY+RIAGGQQEVCGV +KR DNVWLAAEGVY
Sbjct: 151 EGSGKSWRQNQKIRLRHVDTGGYLHSHDRKYTRIAGGQQEVCGVGDKRPDNVWLAAEGVY 210
Query: 70 LPVTESK 76
LPV + K
Sbjct: 211 LPVNQQK 217
>UniRef100_Q86FJ3 Clone ZZD1313 mRNA sequence [Schistosoma japonicum]
Length = 216
Score = 54.7 bits (130), Expect = 6e-07
Identities = 26/58 (44%), Positives = 31/58 (52%)
Query: 13 GKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
G WKQ RL+HI T GY + K+YSR GQ EV + W AAEGVY+
Sbjct: 140 GAYWKQSSNIRLKHISTEGYLHLSGKRYSRPISGQYEVSSTPKLTNAITWTAAEGVYI 197
>UniRef100_UPI00002441DE UPI00002441DE UniRef100 entry
Length = 196
Score = 50.1 bits (118), Expect = 1e-05
Identities = 24/63 (38%), Positives = 34/63 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
SG +W + RLQHIDT Y + + R GQ EV G+ + W+AAEG+++
Sbjct: 123 SGDSWVRTNPVRLQHIDTDMYLGVSGRTFGRPINGQMEVIGLPNPYSGTEWIAAEGLFIH 182
Query: 72 VTE 74
TE
Sbjct: 183 PTE 185
>UniRef100_Q7PY52 ENSANGP00000013320 [Anopheles gambiae str. PEST]
Length = 231
Score = 50.1 bits (118), Expect = 1e-05
Identities = 24/63 (38%), Positives = 34/63 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
SG +W + RLQHIDT Y + + R GQ EV G+ + W+AAEG+++
Sbjct: 158 SGDSWVRTNPVRLQHIDTDMYLGVSGRTFGRPINGQMEVIGLPNPYSGTEWIAAEGLFIH 217
Query: 72 VTE 74
TE
Sbjct: 218 PTE 220
>UniRef100_Q9VNA3 CG11999-PA [Drosophila melanogaster]
Length = 216
Score = 46.6 bits (109), Expect = 2e-04
Identities = 20/59 (33%), Positives = 30/59 (49%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
S + W + RL+HIDT Y + Y R GQ E+ GV + + W AEG+++
Sbjct: 141 SNENWMRSAHVRLRHIDTGMYLGMSGRSYGRPISGQMEIVGVHKPQHGTRWTTAEGLFI 199
>UniRef100_UPI00003AB68F UPI00003AB68F UniRef100 entry
Length = 158
Score = 46.2 bits (108), Expect = 2e-04
Identities = 22/68 (32%), Positives = 33/68 (48%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
SG W +D R QH T + ++Y R GQ+EV G+ +N W EG+++
Sbjct: 85 SGTYWARDSEVRFQHASTDVFLSVTGEQYGRPINGQREVHGMATSSQNNYWKVMEGIFMQ 144
Query: 72 VTESKQAE 79
E +AE
Sbjct: 145 PGEMFKAE 152
>UniRef100_UPI000017F603 UPI000017F603 UniRef100 entry
Length = 223
Score = 45.8 bits (107), Expect = 3e-04
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ A N W A EG+++
Sbjct: 152 SGQHWEREASVRFQHVGTSVFLSVTGEQYGNPIRGQHEVHGMPSANAHNTWKAMEGIFI 210
>UniRef100_Q9ESP1 Stromal cell-derived factor 2-like protein 1 precursor [Mus
musculus]
Length = 221
Score = 45.8 bits (107), Expect = 3e-04
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ A N W A EG+++
Sbjct: 150 SGQHWEREASVRFQHVGTSVFLSVTGEQYGNPIRGQHEVHGMPSANAHNTWKAMEGIFI 208
>UniRef100_Q61EF6 Hypothetical protein CBG12091 [Caenorhabditis briggsae]
Length = 206
Score = 45.4 bits (106), Expect = 3e-04
Identities = 18/59 (30%), Positives = 32/59 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
+G W + ++F+L+H+ T Y +++ R GQ+EV G + W AEG+Y+
Sbjct: 141 NGDEWLESEQFKLRHVVTGSYLSLSGQQFGRPIHGQREVVGTDSITGGSAWKVAEGIYI 199
>UniRef100_Q5PPW6 Hypothetical protein [Xenopus laevis]
Length = 218
Score = 44.3 bits (103), Expect = 8e-04
Identities = 20/55 (36%), Positives = 30/55 (54%)
Query: 16 WKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
W+++ R +HI T+ Y ++Y GQ+EV G+ A N W A EGV+L
Sbjct: 150 WEREDTVRFKHIGTNVYLTITGEQYGHPIRGQREVHGITNPNAHNYWKAMEGVFL 204
>UniRef100_UPI00001CB008 UPI00001CB008 UniRef100 entry
Length = 219
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/68 (30%), Positives = 35/68 (50%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
+G W +D R +H T ++Y R GQ+EV G+ + +N W A EG+++
Sbjct: 146 NGPYWVRDGEVRFKHSSTDVLLSVTGEQYGRPISGQKEVHGMAQPSQNNYWKAMEGIFMK 205
Query: 72 VTESKQAE 79
+E +AE
Sbjct: 206 PSELLRAE 213
>UniRef100_UPI000036C832 UPI000036C832 UniRef100 entry
Length = 221
Score = 43.9 bits (102), Expect = 0.001
Identities = 19/59 (32%), Positives = 31/59 (52%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ N W A EG+++
Sbjct: 150 SGQHWEREAAVRFQHVGTSVFLSVTGEQYGSPIRGQHEVHGMPSANTHNTWKAMEGIFI 208
>UniRef100_Q66JH3 MGC79547 protein [Xenopus tropicalis]
Length = 218
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/59 (33%), Positives = 31/59 (51%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
S W++++ R +HI T+ Y ++Y GQ+EV G+ A N W EGV+L
Sbjct: 146 SDTLWEREESVRFKHIGTNVYLTITGEQYGHPIRGQREVHGITNPNAHNYWKVMEGVFL 204
>UniRef100_Q6P5I1 Stromal cell-derived factor 2 precursor [Mus musculus]
Length = 217
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/68 (30%), Positives = 35/68 (50%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
+G W +D R +H T ++Y R GQ+EV G+ + +N W A EG+++
Sbjct: 144 NGPYWVRDGEVRFKHSSTDVLLSVTGEQYGRPISGQKEVHGMAQPSQNNYWKAMEGIFMK 203
Query: 72 VTESKQAE 79
+E +AE
Sbjct: 204 PSELLRAE 211
>UniRef100_O61793 Hypothetical protein R12E2.13 [Caenorhabditis elegans]
Length = 206
Score = 43.9 bits (102), Expect = 0.001
Identities = 18/59 (30%), Positives = 31/59 (52%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
+G W + ++F+L+H T Y +++ R GQ+EV G + W AEG+Y+
Sbjct: 141 NGDEWLESEQFKLRHAVTGSYLSLSGQQFGRPIHGQREVVGTDSITGGSAWKVAEGIYI 199
>UniRef100_Q86U75 Dihydropyrimidinase-like 2 [Homo sapiens]
Length = 619
Score = 43.9 bits (102), Expect = 0.001
Identities = 19/59 (32%), Positives = 31/59 (52%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ N W A EG+++
Sbjct: 548 SGQHWEREAAVRFQHVGTSVFLSVTGEQYGSPIRGQHEVHGMPSANTHNTWKAMEGIFI 606
>UniRef100_Q9HCN8 Stromal cell-derived factor 2-like protein 1 precursor [Homo
sapiens]
Length = 221
Score = 43.9 bits (102), Expect = 0.001
Identities = 19/59 (32%), Positives = 31/59 (52%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ N W A EG+++
Sbjct: 150 SGQHWEREAAVRFQHVGTSVFLSVTGEQYGSPIRGQHEVHGMPSANTHNTWKAMEGIFI 208
>UniRef100_Q99470 Stromal cell-derived factor 2 precursor [Homo sapiens]
Length = 211
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/68 (30%), Positives = 35/68 (50%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
+G W +D R +H T ++Y R GQ+EV G+ + +N W A EG+++
Sbjct: 138 NGPYWVRDGEVRFKHSSTEVLLSVTGEQYGRPISGQKEVHGMAQPSQNNYWKAMEGIFMK 197
Query: 72 VTESKQAE 79
+E +AE
Sbjct: 198 PSELLKAE 205
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.131 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 151,415,682
Number of Sequences: 2790947
Number of extensions: 5235351
Number of successful extensions: 7069
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 7027
Number of HSP's gapped (non-prelim): 46
length of query: 79
length of database: 848,049,833
effective HSP length: 55
effective length of query: 24
effective length of database: 694,547,748
effective search space: 16669145952
effective search space used: 16669145952
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC133571.7


