

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133341.5 - phase: 1 /pseudo
(808 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
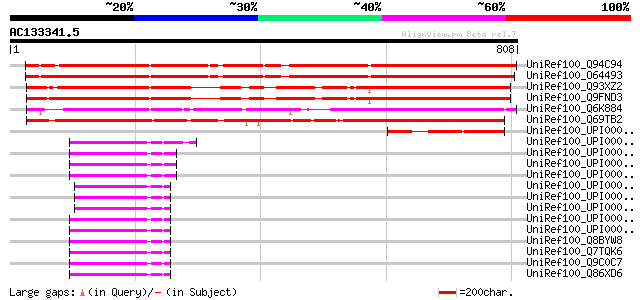
Sequences producing significant alignments: (bits) Value
UniRef100_Q94C94 Hypothetical protein At1g04140 [Arabidopsis tha... 823 0.0
UniRef100_O64493 F20D22.9 protein [Arabidopsis thaliana] 822 0.0
UniRef100_Q93XZ2 Hypothetical protein At5g43930; MRH10.2 [Arabid... 742 0.0
UniRef100_Q9FND3 Similarity to unknown protein [Arabidopsis thal... 740 0.0
UniRef100_Q6K884 Transducin-like [Oryza sativa] 716 0.0
UniRef100_Q69TB2 Transducin family protein-like [Oryza sativa] 665 0.0
UniRef100_UPI000034CC12 UPI000034CC12 UniRef100 entry 189 2e-46
UniRef100_UPI00003ADD70 UPI00003ADD70 UniRef100 entry 123 2e-26
UniRef100_UPI00003ADD73 UPI00003ADD73 UniRef100 entry 121 9e-26
UniRef100_UPI00003ADD72 UPI00003ADD72 UniRef100 entry 121 9e-26
UniRef100_UPI00003ADD71 UPI00003ADD71 UniRef100 entry 121 9e-26
UniRef100_UPI000024A8F2 UPI000024A8F2 UniRef100 entry 120 2e-25
UniRef100_UPI00002490D0 UPI00002490D0 UniRef100 entry 120 2e-25
UniRef100_UPI00002478E6 UPI00002478E6 UniRef100 entry 120 2e-25
UniRef100_UPI000021ECEF UPI000021ECEF UniRef100 entry 119 5e-25
UniRef100_UPI000021D4D3 UPI000021D4D3 UniRef100 entry 119 5e-25
UniRef100_Q8BYW8 Mus musculus 16 days neonate thymus cDNA, RIKEN... 119 5e-25
UniRef100_Q7TQK6 Hypothetical protein D030051N19Rik [Mus musculus] 119 5e-25
UniRef100_Q9C0C7 KIAA1736 protein [Homo sapiens] 119 5e-25
UniRef100_Q86XD6 Hypothetical protein FLJ20294 [Homo sapiens] 119 5e-25
>UniRef100_Q94C94 Hypothetical protein At1g04140 [Arabidopsis thaliana]
Length = 790
Score = 823 bits (2127), Expect = 0.0
Identities = 445/785 (56%), Positives = 535/785 (67%), Gaps = 35/785 (4%)
Query: 26 CSNVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILRV 85
C +VF LL +REISP TK V R KR S S T S + L +I V
Sbjct: 31 CRSVFKLLVQREISPNTKFVPR--KRWGESRCDADSSCGTTSEPVREQ----GLNLISWV 84
Query: 86 EADSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGH 145
EA+SL+HLSAKYCPL+P PRSTIAAAFS DG+ LASTHGDHTVKIIDCETG+CLK+L GH
Sbjct: 85 EAESLQHLSAKYCPLVPPPRSTIAAAFSSDGRTLASTHGDHTVKIIDCETGKCLKILTGH 144
Query: 146 MRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAV 205
RTPWVVRFHP H +I+ASGSLD EVRLW+A T ECI +H FYRPIASIAFHA GE++AV
Sbjct: 145 RRTPWVVRFHPRHSEIVASGSLDHEVRLWNAKTGECIRTHDFYRPIASIAFHAGGELLAV 204
Query: 206 ASGHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMT 265
ASGHKL+IWHY+K G+ S +P VLKTRRSLRAVHFHPH P LLTAEV D+DSSDS+MT
Sbjct: 205 ASGHKLHIWHYNKGGDDS-APAIVLKTRRSLRAVHFHPHGVPLLLTAEVTDIDSSDSAMT 263
Query: 266 EATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHD 325
+TS GYL+YPPPA+F TN +L++E V LP+ L+P Y+ D+ R + ++S
Sbjct: 264 RSTSPGYLRYPPPAIFFTNTQSGSRTSLAAELPLVPLPYLLLPSYSADDPR--ILYSSGT 321
Query: 326 AGSGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTA---FSN 382
G Q +FQ++ +S E T+SP + P ++A F+
Sbjct: 322 TGPRNAQ------TRFQSNQSSVEHGSRTISPSPLPMATSADLSGSYHVPDNSASNTFAT 375
Query: 383 GMGIGIGNLTMDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGHPF 442
G +D M+ DE +P ++R P+ SS +L Q H
Sbjct: 376 QAGARNSTTAVDAMDVDEAQPV--GRNRVPSQVSSQPDLLEFGQLQQLFH---------- 423
Query: 443 VSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADV 502
RD WELPFLQGW+M QSQAG S+ TG S + S A+ T++ +
Sbjct: 424 --FRDRGSWELPFLQGWLMAQSQAGANSVALPTGSSGHVNSTPYMGSSSASHSSTASLEA 481
Query: 503 AMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVSESGNLAASINTPHDGSDIQTIMSRIQ 562
+ S + G +N+ G S R R SR S +S NT H+G+D Q +++RI
Sbjct: 482 GVASLEIPGGVNLYGVSARGDSRDRILQSRFAGSGLAEGRSSRNTQHEGADAQPVVNRIP 541
Query: 563 SELATSVAAAAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEMGAHFSPC 622
SELA+S+AAA ELPCTVKLRVWSHDIK+PCS L +D+CRL I HAVLCSEMGAHFSPC
Sbjct: 542 SELASSIAAA---ELPCTVKLRVWSHDIKDPCSILKSDKCRLTIHHAVLCSEMGAHFSPC 598
Query: 623 GRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYSLEEATFG 682
GR+LAACVAC++PH E DP LQT V Q+SG+ATSPTRHP++AHQVMYELR+YSLE+ +FG
Sbjct: 599 GRYLAACVACVIPHAETDPSLQTLVQQDSGLATSPTRHPVTAHQVMYELRVYSLEKESFG 658
Query: 683 LVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRV 742
VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIV DGETT +TVLE+YRV
Sbjct: 659 SVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVSDGETTSHFFTVLEIYRV 718
Query: 743 SDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEE 802
SDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N T P+ P+E
Sbjct: 719 SDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFRYNTAAASNLTAPNSSPDE 778
Query: 803 TIVGV 807
+ V
Sbjct: 779 NLAEV 783
>UniRef100_O64493 F20D22.9 protein [Arabidopsis thaliana]
Length = 838
Score = 822 bits (2123), Expect = 0.0
Identities = 443/782 (56%), Positives = 534/782 (67%), Gaps = 32/782 (4%)
Query: 26 CSNVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILRV 85
C +VF LL +REISP TK V R KR S S T S + + + L V
Sbjct: 31 CRSVFKLLVQREISPNTKFVPR--KRWGESRCDADSSCGTTSEPVREQGLNL-ISWYLSV 87
Query: 86 EADSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGH 145
EA+SL+HLSAKYCPL+P PRSTIAAAFS DG+ LASTHGDHTVKIIDCETG+CLK+L GH
Sbjct: 88 EAESLQHLSAKYCPLVPPPRSTIAAAFSSDGRTLASTHGDHTVKIIDCETGKCLKILTGH 147
Query: 146 MRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAV 205
RTPWVVRFHP H +I+ASGSLD EVRLW+A T ECI +H FYRPIASIAFHA GE++AV
Sbjct: 148 RRTPWVVRFHPRHSEIVASGSLDHEVRLWNAKTGECIRTHDFYRPIASIAFHAGGELLAV 207
Query: 206 ASGHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMT 265
ASGHKL+IWHY+K G+ S +P VLKTRRSLRAVHFHPH P LLTAEV D+DSSDS+MT
Sbjct: 208 ASGHKLHIWHYNKGGDDS-APAIVLKTRRSLRAVHFHPHGVPLLLTAEVTDIDSSDSAMT 266
Query: 266 EATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHD 325
+TS GYL+YPPPA+F TN +L++E V LP+ L+P Y+ D+ R + ++S
Sbjct: 267 RSTSPGYLRYPPPAIFFTNTQSGSRTSLAAELPLVPLPYLLLPSYSADDPR--ILYSSGT 324
Query: 326 AGSGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTA---FSN 382
G Q +FQ++ +S E T+SP + P ++A F+
Sbjct: 325 TGPRNAQ------TRFQSNQSSVEHGSRTISPSPLPMATSADLSGSYHVPDNSASNTFAT 378
Query: 383 GMGIGIGNLTMDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGHPF 442
G +D M+ DE +P ++R P+ SS +L Q H
Sbjct: 379 QAGARNSTTAVDAMDVDEAQPV--GRNRVPSQVSSQPDLLEFGQLQQLFH---------- 426
Query: 443 VSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADV 502
RD WELPFLQGW+M QSQAG S+ TG S + S A+ T++ +
Sbjct: 427 --FRDRGSWELPFLQGWLMAQSQAGANSVALPTGSSGHVNSTPYMGSSSASHSSTASLEA 484
Query: 503 AMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVSESGNLAASINTPHDGSDIQTIMSRIQ 562
+ S + G +N+ G S R R SR S +S NT H+G+D Q +++RI
Sbjct: 485 GVASLEIPGGVNLYGVSARGDSRDRILQSRFAGSGLAEGRSSRNTQHEGADAQPVVNRIP 544
Query: 563 SELATSVAAAAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEMGAHFSPC 622
SELA+S+AAA ELPCTVKLRVWSHDIK+PCS L +D+CRL I HAVLCSEMGAHFSPC
Sbjct: 545 SELASSIAAA---ELPCTVKLRVWSHDIKDPCSILKSDKCRLTIHHAVLCSEMGAHFSPC 601
Query: 623 GRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYSLEEATFG 682
GR+LAACVAC++PH E DP LQT V Q+SG+ATSPTRHP++AHQVMYELR+YSLE+ +FG
Sbjct: 602 GRYLAACVACVIPHAETDPSLQTLVQQDSGLATSPTRHPVTAHQVMYELRVYSLEKESFG 661
Query: 683 LVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRV 742
VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIV DGETT +TVLE+YRV
Sbjct: 662 SVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVSDGETTSHFFTVLEIYRV 721
Query: 743 SDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEE 802
SDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N T P+ P+E
Sbjct: 722 SDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFRYNTAAASNLTAPNSSPDE 781
Query: 803 TI 804
+
Sbjct: 782 NL 783
>UniRef100_Q93XZ2 Hypothetical protein At5g43930; MRH10.2 [Arabidopsis thaliana]
Length = 726
Score = 742 bits (1916), Expect = 0.0
Identities = 421/782 (53%), Positives = 506/782 (63%), Gaps = 104/782 (13%)
Query: 28 NVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILRVEA 87
+VFN+L +RE+SP+ K V R + G WY + S T + M T + VEA
Sbjct: 36 SVFNMLVQREMSPKAKFVPRK-RWGKSRWYTDS-SCGTNNESM----KETGQSLTSWVEA 89
Query: 88 DSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMR 147
+SL+HLSAKYCPL PRSTIAAAFS DG+ LASTHGDHTVKIIDCETG CLKVL GH R
Sbjct: 90 ESLQHLSAKYCPLGAPPRSTIAAAFSTDGRTLASTHGDHTVKIIDCETGNCLKVLTGHRR 149
Query: 148 TPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
TPWVVRFHP H +I+ASGSLD EVRLW+ TSECI SH FYRPIASIAFHA+GE++AVAS
Sbjct: 150 TPWVVRFHPHHSEIVASGSLDLEVRLWNTTTSECIRSHLFYRPIASIAFHAEGELLAVAS 209
Query: 208 GHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMTEA 267
GHKL++WHY+++GE S SP VLKTRRSLRAVHFHPH AP LLTAEVN++DS DSSM+ A
Sbjct: 210 GHKLHMWHYNRRGEGS-SPTVVLKTRRSLRAVHFHPHGAPLLLTAEVNEIDSLDSSMSRA 268
Query: 268 TSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHDAG 327
TS+GYL+YPPPA+ T+ +
Sbjct: 269 TSMGYLRYPPPAILFTSTESNQ-------------------------------------- 290
Query: 328 SGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTAFSNGMGIG 387
+S A+ + T+S T S + +P NS SN
Sbjct: 291 -------TSLAAENENRTSSPPLPLATSSGPSGPNSVPGNSP-----------SNIFLTR 332
Query: 388 IGNLT---MDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGH--PF 442
G+ T +DGM+ DE +P RN G+ Q +N PE G
Sbjct: 333 AGDRTSPAVDGMDVDEAQPVG----RN------------GIPSQVSNRSDFPELGQIRQL 376
Query: 443 VSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADV 502
RD WELPFLQGW+M Q ++ TG S ++ S T++ +
Sbjct: 377 FHFRDRVSWELPFLQGWLMAQGHGVANPVVTPTGSSNHGISAPSS---------TASLEA 427
Query: 503 AMPSSAMSGSINIPGSSVRSGLRSHFSH---SRTPVSESGNLAASINTPHDGSDIQTIMS 559
A+ + +N+ S R G + S SRT + E +S NT H GSD Q +++
Sbjct: 428 AVALLEIPSGVNLHAVSRRGGAQEQTSQPQFSRTGLPEG---VSSRNTQH-GSDAQPVVN 483
Query: 560 RIQSELATSVAA----AAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEM 615
R+QSELATS+AA AAA ELPCTVKLR+WSHDIK+P + L +DRC IPHAVLCSEM
Sbjct: 484 RVQSELATSIAASAAAAAAAELPCTVKLRMWSHDIKDPYAQLKSDRCLFTIPHAVLCSEM 543
Query: 616 GAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYS 675
GAH+SPCGR+LAACVAC+ PH E DPGLQT Q+SG+ATSPTRHP++AHQV+YELR+YS
Sbjct: 544 GAHYSPCGRYLAACVACVFPHGEIDPGLQTQAQQDSGLATSPTRHPVTAHQVIYELRVYS 603
Query: 676 LEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYT 735
L++ +FG VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLL+SIV DGETT +T
Sbjct: 604 LQKESFGSVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLRSIVSDGETTSHFFT 663
Query: 736 VLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTG 795
VLE+YRVSDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N TG
Sbjct: 664 VLEIYRVSDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFQYNTAATSNFTG 723
Query: 796 PS 797
P+
Sbjct: 724 PN 725
>UniRef100_Q9FND3 Similarity to unknown protein [Arabidopsis thaliana]
Length = 729
Score = 740 bits (1911), Expect = 0.0
Identities = 421/782 (53%), Positives = 508/782 (64%), Gaps = 101/782 (12%)
Query: 28 NVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILRVEA 87
+VFN+L +RE+SP+ K V R + G WY + S T + M + + + L VEA
Sbjct: 36 SVFNMLVQREMSPKAKFVPRK-RWGKSRWYTDS-SCGTNNESMKETGQSLTSWYTL-VEA 92
Query: 88 DSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMR 147
+SL+HLSAKYCPL PRSTIAAAFS DG+ LASTHGDHTVKIIDCETG CLKVL GH R
Sbjct: 93 ESLQHLSAKYCPLGAPPRSTIAAAFSTDGRTLASTHGDHTVKIIDCETGNCLKVLTGHRR 152
Query: 148 TPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
TPWVVRFHP H +I+ASGSLD EVRLW+ TSECI SH FYRPIASIAFHA+GE++AVAS
Sbjct: 153 TPWVVRFHPHHSEIVASGSLDLEVRLWNTTTSECIRSHLFYRPIASIAFHAEGELLAVAS 212
Query: 208 GHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMTEA 267
GHKL++WHY+++GE S SP VLKTRRSLRAVHFHPH AP LLTAEVN++DS DSSM+ A
Sbjct: 213 GHKLHMWHYNRRGEGS-SPTVVLKTRRSLRAVHFHPHGAPLLLTAEVNEIDSLDSSMSRA 271
Query: 268 TSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHDAG 327
TS+GYL+YPPPA+ T+ +
Sbjct: 272 TSMGYLRYPPPAILFTSTESNQ-------------------------------------- 293
Query: 328 SGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTAFSNGMGIG 387
+S A+ + T+S T S + +P NS SN
Sbjct: 294 -------TSLAAENENRTSSPPLPLATSSGPSGPNSVPGNSP-----------SNIFLTR 335
Query: 388 IGNLT---MDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGH--PF 442
G+ T +DGM+ DE +P RN G+ Q +N PE G
Sbjct: 336 AGDRTSPAVDGMDVDEAQPVG----RN------------GIPSQVSNRSDFPELGQIRQL 379
Query: 443 VSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADV 502
RD WELPFLQGW+M Q ++ TG S ++ S T++ +
Sbjct: 380 FHFRDRVSWELPFLQGWLMAQGHGVANPVVTPTGSSNHGISAPSS---------TASLEA 430
Query: 503 AMPSSAMSGSINIPGSSVRSGLRSHFSH---SRTPVSESGNLAASINTPHDGSDIQTIMS 559
A+ + +N+ S R G + S SRT + E +S NT H GSD Q +++
Sbjct: 431 AVALLEIPSGVNLHAVSRRGGAQEQTSQPQFSRTGLPEG---VSSRNTQH-GSDAQPVVN 486
Query: 560 RIQSELATSVAA----AAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEM 615
R+QSELATS+AA AAA ELPCTVKLR+WSHDIK+P + L +DRC IPHAVLCSEM
Sbjct: 487 RVQSELATSIAASAAAAAAAELPCTVKLRMWSHDIKDPYAQLKSDRCLFTIPHAVLCSEM 546
Query: 616 GAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYS 675
GAH+SPCGR+LAACVAC+ PH E DPGLQT Q+SG+ATSPTRHP++AHQV+YELR+YS
Sbjct: 547 GAHYSPCGRYLAACVACVFPHGEIDPGLQTQAQQDSGLATSPTRHPVTAHQVIYELRVYS 606
Query: 676 LEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYT 735
L++ +FG VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLL+SIV DGETT +T
Sbjct: 607 LQKESFGSVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLRSIVSDGETTSHFFT 666
Query: 736 VLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTG 795
VLE+YRVSDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N TG
Sbjct: 667 VLEIYRVSDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFQYNTAATSNFTG 726
Query: 796 PS 797
P+
Sbjct: 727 PN 728
>UniRef100_Q6K884 Transducin-like [Oryza sativa]
Length = 786
Score = 716 bits (1849), Expect = 0.0
Identities = 419/827 (50%), Positives = 495/827 (59%), Gaps = 126/827 (15%)
Query: 28 NVFNLLARREISPRTKHVAR---------------------DAKRGLLSWYLNYFSL*TI 66
N+F+LLA+REISPRTKH A+ DAK + SW
Sbjct: 32 NIFDLLAQREISPRTKHQAKKLWSKSPGNDADSNELRYAATDAKHDIYSW---------- 81
Query: 67 STKM*AS*ITTSLKMILRVEADSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDH 126
E+ SL H SAKYCPLLP PRSTIAAAFSPDGK LASTHGDH
Sbjct: 82 ------------------AESQSLHHWSAKYCPLLPPPRSTIAAAFSPDGKTLASTHGDH 123
Query: 127 TVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHH 186
TVKIID +TG+CLKVL GH RTPWVVR+HPL P ILASGSLDQEVRLWDA TS+CI S
Sbjct: 124 TVKIIDYQTGKCLKVLSGHRRTPWVVRYHPLLPDILASGSLDQEVRLWDAKTSDCIGSQD 183
Query: 187 FYRPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAA 246
F+RPIASIAFHA+GEI+AVASGHKLYIW+Y+K+ EA+ +P +L+TRRSLRAVHFHPH A
Sbjct: 184 FHRPIASIAFHARGEILAVASGHKLYIWNYNKRDEAA-APTIILRTRRSLRAVHFHPHGA 242
Query: 247 PYLLTAEVNDLDSSDSSMTEATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFL 306
PYLLTAEVN+LDS+DS +T ATS GY Y P AVF N++ N S P L
Sbjct: 243 PYLLTAEVNNLDSADSPLTLATSSGYSNY-PSAVFFANINSR---NCPHHEANSSSPCLL 298
Query: 307 VPPYTVDESRAELQHASHDAGSGRIQIESSAVAQFQ-ADTNSTEQHDTTVSPMDTVSEIP 365
P Y D+ L + S + S++AQ A +Q D V+PMD P
Sbjct: 299 WPAYLRDDGSLCLIRNDLVSSSTNVHQRPSSLAQNPLASDVENQQPDQLVTPMDVCPGEP 358
Query: 366 TNSQAGTEYPAHTAFSNGMGIGIGNLTMDGMETDETRPAEGSQHRNPTDASSLNGM--LH 423
+ S A A M I G + + + T E S R+ SL+ +
Sbjct: 359 STSHAS----ASGLSGVEMQIDRGQPSSRLLGSSSTSNHESSTARDDVQMPSLSNSVPIP 414
Query: 424 GLSRQTANHGVHPEDGHPFVSS----------------------RDPSGWELPFLQGWMM 461
S+ + + G H + F +S D WELPFLQGW M
Sbjct: 415 ATSQPSEHDGRHGMPMNSFTTSSGLDVHMILRNSEGGNHHHDLFSDSRSWELPFLQGWFM 474
Query: 462 GQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADVAMPSSAMSGSINIPGSSVR 521
Q+ HTG S SI I S R
Sbjct: 475 AQN---------HTGA--------------------------------SPSIPIDVGSSR 493
Query: 522 SGLRSHFSHSRTPVSESGNLAASINTPHDGSDIQTIMSRIQSELATSVAAAAATELPCTV 581
R H S S G ++ + D +++ + SEL TS+ AA A ELPCTV
Sbjct: 494 GSNRHHASRRHVVGSLRGVGSSLLGPQIDEAEVHAASLGVGSELTTSLLAAGAAELPCTV 553
Query: 582 KLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADP 641
KLR+W HDIK+PC L + CRL I HAVLCSEMGAHFSPCGRFL ACVAC+LP E D
Sbjct: 554 KLRIWRHDIKDPCVTLEPEACRLTISHAVLCSEMGAHFSPCGRFLVACVACLLPQTEGDR 613
Query: 642 GLQTPVHQES-GIATSPTRHPISAHQVMYELRIYSLEEATFGLVLASRAIRAAHCLTSIQ 700
G Q PV +S G TSPTRHP+ +H+V+YELR+YSLEEATFG VL SRAIRAAHCLTSIQ
Sbjct: 614 GSQLPVQYDSAGAGTSPTRHPLPSHRVIYELRVYSLEEATFGKVLTSRAIRAAHCLTSIQ 673
Query: 701 FSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNV 760
FSPTSEHILLAYGRRH S+L+SIV+DGET +P+YT+LEVYRVSDMELVRVLPSAEDEVNV
Sbjct: 674 FSPTSEHILLAYGRRHNSMLRSIVMDGETGIPVYTILEVYRVSDMELVRVLPSAEDEVNV 733
Query: 761 ACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEETIVGV 807
ACFHP PGGGLVYGTKEGKLRIL ++GA + TG + F EE ++ +
Sbjct: 734 ACFHPSPGGGLVYGTKEGKLRILQHNGA-DITSTGLNCFIEENMLEI 779
>UniRef100_Q69TB2 Transducin family protein-like [Oryza sativa]
Length = 796
Score = 665 bits (1716), Expect = 0.0
Identities = 391/787 (49%), Positives = 479/787 (60%), Gaps = 55/787 (6%)
Query: 28 NVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILR-VE 86
N+++LLA+REISPRTKH A++ G L A +T + + E
Sbjct: 37 NIYHLLAQREISPRTKHQAKNIWSGSPECATGSIEL--------AFWVTDARHDLFSWAE 88
Query: 87 ADSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHM 146
+ SL SA+YC LLPAPRSTIAAAFSPDG+ LASTHGDHTVKIIDC+TG+CLKVL+GH
Sbjct: 89 SQSLHRWSARYCLLLPAPRSTIAAAFSPDGRTLASTHGDHTVKIIDCQTGKCLKVLLGHW 148
Query: 147 RTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVA 206
RTPWVVR+HPLHP I+ASGSLD EV LWDA TS CI SH FY+ IASIAFHAKGE++AVA
Sbjct: 149 RTPWVVRYHPLHPDIVASGSLDNEVCLWDAKTSRCIGSHFFYKQIASIAFHAKGELLAVA 208
Query: 207 SGHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMTE 266
SGHKL+IW Y+K+ EAS P+ +L+TRRSLRAV FHP+ APYLLTAEVN+LDS+DS +T
Sbjct: 209 SGHKLFIWDYNKRDEASDPPM-ILRTRRSLRAVQFHPNGAPYLLTAEVNNLDSADSELTH 267
Query: 267 ATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHDA 326
ATS GY P F + S P + P Y D+ L +
Sbjct: 268 ATSSGYSNSPSAVFFAI----MNSACCPYSESRFSSPCLIWPAYVRDDGSICLLRNDWVS 323
Query: 327 GSGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPA--------HT 378
GS +Q Q + T+Q V+PMD P + E A HT
Sbjct: 324 GSSDVQ---------QPSDSETQQAGHMVTPMDVCPGEPGVNNYDDEDSASLSNRIEMHT 374
Query: 379 -AFSNGMGIGIGNLTMD----------GMETDETRPAEGSQHRNPTDAS---SLNGMLHG 424
++ N + D + +D P + R S L +
Sbjct: 375 PSWQNSSRFHNSSAATDLHRIDIRQVSDLSSDTPNPEMPAHSRIDVPNSMPMDLFAFSNT 434
Query: 425 LSRQTANHGVHPEDGHPFVSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAP 484
+ Q V H + S WELPFLQGW+M Q++ GL + LP+ V D P
Sbjct: 435 IDVQMFLRDVEAGHHHNNYTGGSHS-WELPFLQGWLMAQNRTGLRATLPNNEVIGDL--P 491
Query: 485 QISSSVMANTLPTSNADVAMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVSESGNLAAS 544
++ N + S+ + S SI I S+R GL H S G S
Sbjct: 492 IGGTAGTDNVMNESSNMYSFERVGPSSSIPITTDSLR-GLSKH---RHMLASVPGGAGTS 547
Query: 545 INTPHDG-SDIQTIMSRIQSELATSVAAAAATELPCTVKLRVWSHDIKNPCSPLNADRCR 603
+ +G + + + + SE ATS+ A TELPCTVKLR+W H+I NPC+ L + C
Sbjct: 548 LQGAQNGEAHVNVVSLGVGSEFATSLFAGDGTELPCTVKLRIWRHNIDNPCAVLAPEACC 607
Query: 604 LIIPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGL-QTPVHQES-GIATSPTRHP 661
L I HAVLCSEMG HFSPCGRFL ACVAC+LP E + Q+PV +S G TSPTRHP
Sbjct: 608 LTISHAVLCSEMGTHFSPCGRFLVACVACLLPQTEVGEHVSQSPVQYDSTGAGTSPTRHP 667
Query: 662 ISAHQVMYELRIYSLEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLK 721
+ + +V+YELR+YSLEE TFG VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRH SLL+
Sbjct: 668 LPSRRVIYELRVYSLEEETFGTVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHNSLLR 727
Query: 722 SIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLR 781
I +DG+TT+P+YTVLEVYRVSDMELVRV+PSAEDEVNVACFHP PG GLVYGTKEGKLR
Sbjct: 728 GIFMDGKTTIPVYTVLEVYRVSDMELVRVIPSAEDEVNVACFHPSPGAGLVYGTKEGKLR 787
Query: 782 ILHYDGA 788
I+ ++ A
Sbjct: 788 IIQHNVA 794
>UniRef100_UPI000034CC12 UPI000034CC12 UniRef100 entry
Length = 198
Score = 189 bits (481), Expect = 2e-46
Identities = 95/186 (51%), Positives = 122/186 (65%), Gaps = 25/186 (13%)
Query: 603 RLIIPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPI 662
+L++P VLCSEMGAHFSPCGR++A C AC D
Sbjct: 2 QLVLPQTVLCSEMGAHFSPCGRYIATCQACKPESSRQDV--------------------- 40
Query: 663 SAHQVMYELRIYSLEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKS 722
Q + E+R YS++ FG VL++RAI+AAHCLTSIQFSPTS H+LLAYGRRH SLL
Sbjct: 41 ---QYVCEVRTYSIDAHQFGQVLSARAIKAAHCLTSIQFSPTSSHVLLAYGRRHPSLLL- 96
Query: 723 IVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRI 782
++ D E+ ++T+LE++ M L RV+PSAEDEVN ACFHP PG G+ YGTKEG+LR+
Sbjct: 97 LMADAESFSQVHTILEIFDAKSMRLARVIPSAEDEVNAACFHPIPGQGIAYGTKEGRLRV 156
Query: 783 LHYDGA 788
L +D +
Sbjct: 157 LLHDNS 162
>UniRef100_UPI00003ADD70 UPI00003ADD70 UniRef100 entry
Length = 1247
Score = 123 bits (309), Expect = 2e-26
Identities = 72/205 (35%), Positives = 106/205 (51%), Gaps = 18/205 (8%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD ++ASTH +H + I + +TG+C+ L+GH RTPW V FH
Sbjct: 51 KKVELPDSPRSTFLLAFSPDRTLMASTHVNHNIYITEVKTGKCVHSLVGHRRTPWCVTFH 110
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 111 PTIPGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 170
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVNDLDSSDSSMTEATSIGY 272
+ ++ P V+KT + R V F P YLLTA VN + S S+
Sbjct: 171 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVNPSNQQQSDEEVEISVDS 224
Query: 273 LQYPPPAVFVTNVHPTEHVTLSSEP 297
+ P H + L S+P
Sbjct: 225 TEMP---------HYRQRAILPSQP 240
>UniRef100_UPI00003ADD73 UPI00003ADD73 UniRef100 entry
Length = 739
Score = 121 bits (303), Expect = 9e-26
Identities = 66/174 (37%), Positives = 96/174 (54%), Gaps = 9/174 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD ++ASTH +H + I + +TG+C+ L+GH RTPW V FH
Sbjct: 45 KKVELPDSPRSTFLLAFSPDRTLMASTHVNHNIYITEVKTGKCVHSLVGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTIPGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVNDLDSSDSSMTE 266
+ ++ P V+KT + R V F P YLLTA VN + E
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVNPSNQQSDEEVE 212
>UniRef100_UPI00003ADD72 UPI00003ADD72 UniRef100 entry
Length = 1204
Score = 121 bits (303), Expect = 9e-26
Identities = 66/174 (37%), Positives = 96/174 (54%), Gaps = 9/174 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD ++ASTH +H + I + +TG+C+ L+GH RTPW V FH
Sbjct: 45 KKVELPDSPRSTFLLAFSPDRTLMASTHVNHNIYITEVKTGKCVHSLVGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTIPGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVNDLDSSDSSMTE 266
+ ++ P V+KT + R V F P YLLTA VN + E
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVNPSNQQSDEEVE 212
>UniRef100_UPI00003ADD71 UPI00003ADD71 UniRef100 entry
Length = 1228
Score = 121 bits (303), Expect = 9e-26
Identities = 66/174 (37%), Positives = 96/174 (54%), Gaps = 9/174 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD ++ASTH +H + I + +TG+C+ L+GH RTPW V FH
Sbjct: 50 KKVELPDSPRSTFLLAFSPDRTLMASTHVNHNIYITEVKTGKCVHSLVGHRRTPWCVTFH 109
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 110 PTIPGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 169
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVNDLDSSDSSMTE 266
+ ++ P V+KT + R V F P YLLTA VN + E
Sbjct: 170 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVNPSNQQSDEEVE 217
>UniRef100_UPI000024A8F2 UPI000024A8F2 UniRef100 entry
Length = 594
Score = 120 bits (301), Expect = 2e-25
Identities = 62/156 (39%), Positives = 94/156 (59%), Gaps = 9/156 (5%)
Query: 103 APRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKIL 162
+PRST AFSPD ++ASTH +H + I + ++G+C+ L+GH RTPW + FHP+ P ++
Sbjct: 47 SPRSTFLLAFSPDRSLMASTHVNHNIYITEVKSGKCVHSLVGHRRTPWCLTFHPIIPGLI 106
Query: 163 ASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEA 222
ASG LD EVR+WD + IAS+AFH +++ +A+ +++++W + +K
Sbjct: 107 ASGCLDGEVRIWDLHGGSESWLTESNSAIASLAFHPTAQLLLIATNNEVHLWDWSRK--- 163
Query: 223 SYSPIFVLKT---RRSLRAVHFHPHAAPYLLTAEVN 255
P V+KT +R V F P YLLTA VN
Sbjct: 164 --EPFTVVKTASETERVRLVRFDP-LGHYLLTAIVN 196
>UniRef100_UPI00002490D0 UPI00002490D0 UniRef100 entry
Length = 775
Score = 120 bits (301), Expect = 2e-25
Identities = 62/156 (39%), Positives = 94/156 (59%), Gaps = 9/156 (5%)
Query: 103 APRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKIL 162
+PRST AFSPD ++ASTH +H + I + ++G+C+ L+GH RTPW + FHP+ P ++
Sbjct: 47 SPRSTFLLAFSPDRSLMASTHVNHNIYITEVKSGKCVHSLVGHRRTPWCLTFHPIIPGLI 106
Query: 163 ASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEA 222
ASG LD EVR+WD + IAS+AFH +++ +A+ +++++W + +K
Sbjct: 107 ASGCLDGEVRIWDLHGGSESWLTESNSAIASLAFHPTAQLLLIATNNEVHLWDWSRK--- 163
Query: 223 SYSPIFVLKT---RRSLRAVHFHPHAAPYLLTAEVN 255
P V+KT +R V F P YLLTA VN
Sbjct: 164 --EPFTVVKTASETERVRLVRFDP-LGHYLLTAIVN 196
>UniRef100_UPI00002478E6 UPI00002478E6 UniRef100 entry
Length = 720
Score = 120 bits (301), Expect = 2e-25
Identities = 62/156 (39%), Positives = 94/156 (59%), Gaps = 9/156 (5%)
Query: 103 APRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKIL 162
+PRST AFSPD ++ASTH +H + I + ++G+C+ L+GH RTPW + FHP+ P ++
Sbjct: 47 SPRSTFLLAFSPDRSLMASTHVNHNIYITEVKSGKCVHSLVGHRRTPWCLTFHPIIPGLI 106
Query: 163 ASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEA 222
ASG LD EVR+WD + IAS+AFH +++ +A+ +++++W + +K
Sbjct: 107 ASGCLDGEVRIWDLHGGSESWLTESNSAIASLAFHPTAQLLLIATNNEVHLWDWSRK--- 163
Query: 223 SYSPIFVLKT---RRSLRAVHFHPHAAPYLLTAEVN 255
P V+KT +R V F P YLLTA VN
Sbjct: 164 --EPFTVVKTASETERVRLVRFDP-LGHYLLTAIVN 196
>UniRef100_UPI000021ECEF UPI000021ECEF UniRef100 entry
Length = 781
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 45 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 201
>UniRef100_UPI000021D4D3 UPI000021D4D3 UniRef100 entry
Length = 752
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 45 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 201
>UniRef100_Q8BYW8 Mus musculus 16 days neonate thymus cDNA, RIKEN full-length
enriched library, clone:A130023A14 product:ISCHEMIA
RELATED FACTOR NYW-1 homolog [Mus musculus]
Length = 752
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 45 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 201
>UniRef100_Q7TQK6 Hypothetical protein D030051N19Rik [Mus musculus]
Length = 813
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 45 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 201
>UniRef100_Q9C0C7 KIAA1736 protein [Homo sapiens]
Length = 1244
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 51 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 110
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 111 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 170
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 171 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 207
>UniRef100_Q86XD6 Hypothetical protein FLJ20294 [Homo sapiens]
Length = 1208
Score = 119 bits (297), Expect = 5e-25
Identities = 66/163 (40%), Positives = 93/163 (56%), Gaps = 9/163 (5%)
Query: 96 KYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFH 155
K L +PRST AFSPD +LASTH +H + I + +TG+C+ LIGH RTPW V FH
Sbjct: 45 KRVELPDSPRSTFLLAFSPDRTLLASTHVNHNIYITEVKTGKCVHSLIGHRRTPWCVTFH 104
Query: 156 PLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWH 215
P ++ASG LD EVR+WD + IAS+AFH +++ +A+ ++++ W
Sbjct: 105 PTISGLIASGCLDGEVRIWDLHGGSESWFTDSNNAIASLAFHPTAQLLLIATANEIHFWD 164
Query: 216 YDKKGEASYSPIFVLKTRRSL---RAVHFHPHAAPYLLTAEVN 255
+ ++ P V+KT + R V F P YLLTA VN
Sbjct: 165 WSRR-----EPFAVVKTASEMERVRLVRFDP-LGHYLLTAIVN 201
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,327,905,089
Number of Sequences: 2790947
Number of extensions: 55444703
Number of successful extensions: 154411
Number of sequences better than 10.0: 2968
Number of HSP's better than 10.0 without gapping: 800
Number of HSP's successfully gapped in prelim test: 2187
Number of HSP's that attempted gapping in prelim test: 141944
Number of HSP's gapped (non-prelim): 10127
length of query: 808
length of database: 848,049,833
effective HSP length: 136
effective length of query: 672
effective length of database: 468,481,041
effective search space: 314819259552
effective search space used: 314819259552
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC133341.5


