

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.4 - phase: 0
(359 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
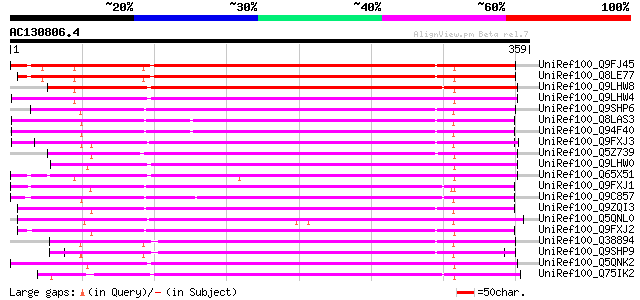
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJ45 GDSL-motif lipase/hydrolase-like protein [Arabi... 306 4e-82
UniRef100_Q8LE77 GDSL-motif lipase/hydrolase-like protein [Arabi... 301 2e-80
UniRef100_Q9LHW8 Putative esterase [Oryza sativa] 293 4e-78
UniRef100_Q9LHW4 Putative esterase [Oryza sativa] 285 1e-75
UniRef100_Q9SHP6 F1K23.16 [Arabidopsis thaliana] 280 4e-74
UniRef100_Q8LAS3 Lipase, putative [Arabidopsis thaliana] 276 8e-73
UniRef100_Q94F40 At1g28600/F1K23_6 [Arabidopsis thaliana] 274 2e-72
UniRef100_Q9FXJ3 F1K23.17 [Arabidopsis thaliana] 274 2e-72
UniRef100_Q5Z739 Putative lipase [Oryza sativa] 272 9e-72
UniRef100_Q9LHW0 Putative esterase [Oryza sativa] 270 3e-71
UniRef100_Q65X51 Hypothetical protein OJ1123_F01.16 [Oryza sativa] 265 1e-69
UniRef100_Q9FXJ1 F1K23.19 [Arabidopsis thaliana] 265 1e-69
UniRef100_Q9C857 Hypothetical protein T8E3.19 [Arabidopsis thali... 261 2e-68
UniRef100_Q9ZQI3 Putative lipase [Arabidopsis thaliana] 261 3e-68
UniRef100_Q5QNL0 Putative esterase [Oryza sativa] 259 6e-68
UniRef100_Q9FXJ2 F1K23.18 [Arabidopsis thaliana] 259 8e-68
UniRef100_Q38894 Lipase [Arabidopsis thaliana] 257 3e-67
UniRef100_Q9SHP9 F1K23.13 [Arabidopsis thaliana] 257 3e-67
UniRef100_Q5QNK2 Putative esterase [Oryza sativa] 257 4e-67
UniRef100_Q75IK2 Hypothetical protein OSJNBb0016G07.10 [Oryza sa... 256 7e-67
>UniRef100_Q9FJ45 GDSL-motif lipase/hydrolase-like protein [Arabidopsis thaliana]
Length = 372
Score = 306 bits (785), Expect = 4e-82
Identities = 169/375 (45%), Positives = 228/375 (60%), Gaps = 30/375 (8%)
Query: 1 MNVSTLFLITFTCGFLQNVVS--NANPLSYEAFFNFGDSISDTGN---AASIFLPMPNPI 55
M ++ LF++ F+ FL +V S L YE+ FNFGDS+SDTGN + + P +
Sbjct: 1 MRINMLFIVAFS--FLVSVRSLPMRPTLKYESIFNFGDSLSDTGNFLLSGDVDSPNIGRL 58
Query: 56 PYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAY-----ENKSIDQDIKKGVNFAFAG 110
PYG ++F +GR S+GRLIIDFIAEA GLP++P Y N S+D K+G NFA AG
Sbjct: 59 PYGQTFFNRSTGRCSDGRLIIDFIAEASGLPYIPPYLQSLRTNDSVD--FKRGANFAVAG 116
Query: 111 ATVLNVEYYVKNGLPLPD-TNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIG 169
AT ++ GL + TN +L IQL WFK +KP LCK+K +C YF+KSLF+VGEIG
Sbjct: 117 ATANEFSFFKNRGLSVTLLTNKTLDIQLDWFKKLKPSLCKTKPECEQYFRKSLFLVGEIG 176
Query: 170 GNDI-MKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTL 228
GND + ++ ++VPF++ I + T+ LIEEGA+ L+VPGN P+GCSAA+
Sbjct: 177 GNDYNYPLLAFRSFKHAMDLVPFVINKIMDVTSALIEEGAMTLIVPGNLPIGCSAALLER 236
Query: 229 VNSNKKEDYDEFG-CLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQ 287
N N YD C + NNL + N +LK + LR+K+P KIIY DYY+ A + +
Sbjct: 237 FNDNSGWLYDSRNQCYMPLNNLAKLHNDKLKKGLAALRKKYPYAKIIYADYYSSAMQFFN 296
Query: 288 TPQQYGFDKDAIFKACCGG------------CGSLIATVCSDPSKRINWDGPHFTEAAYK 335
+P +YGF ++ KACCGG CG +T C DPS NWDG H TEAAY+
Sbjct: 297 SPSKYGF-TGSVLKACCGGGDGRYNVQPNVRCGEKGSTTCEDPSTYANWDGIHLTEAAYR 355
Query: 336 LIAKGLVEGPFSNPS 350
IA GL+ G F+ P+
Sbjct: 356 HIATGLISGRFTMPT 370
>UniRef100_Q8LE77 GDSL-motif lipase/hydrolase-like protein [Arabidopsis thaliana]
Length = 368
Score = 301 bits (771), Expect = 2e-80
Identities = 167/370 (45%), Positives = 224/370 (60%), Gaps = 30/370 (8%)
Query: 6 LFLITFTCGFLQNVVS--NANPLSYEAFFNFGDSISDTGN---AASIFLPMPNPIPYGSS 60
LF++ F+ FL +V S L YE+ FNFGDS+SDTGN + + P +PYG +
Sbjct: 2 LFIVAFS--FLVSVRSLPMRPTLKYESIFNFGDSLSDTGNFLLSGDVDSPNIGRLPYGQT 59
Query: 61 YFKHPSGRMSNGRLIIDFIAEAYGLPFLPAY-----ENKSIDQDIKKGVNFAFAGATVLN 115
+F +GR S+GRLIIDFIAEA GLP++P Y N S+D K+G NFA AGAT
Sbjct: 60 FFNRSTGRCSDGRLIIDFIAEASGLPYIPPYLQSLRTNDSVD--FKRGANFAVAGATANE 117
Query: 116 VEYYVKNGLPLPD-TNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI- 173
++ GL + TN +L IQL WFK +KP LCK+K +C YF+KSLF+VGEI GND
Sbjct: 118 FSFFKNRGLSVTLLTNKTLDIQLDWFKKLKPSLCKTKPECERYFRKSLFLVGEISGNDYN 177
Query: 174 MKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNK 233
+ ++ ++VPF++ I + T+ LIEEGA+ L+VPGN P+GCSAA+ N N
Sbjct: 178 YPLLAFRSFKHAMDLVPFVINKIMDVTSALIEEGAMTLIVPGNLPIGCSAALLERFNDNS 237
Query: 234 KEDYDEFG-CLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQY 292
YD C + NNL + N +LK + LR+K+P KIIY DYY+ A + + +P +Y
Sbjct: 238 GWLYDSRNQCYMPLNNLAKLHNDKLKKGLAALRKKYPYAKIIYADYYSSAMQFFNSPSKY 297
Query: 293 GFDKDAIFKACCGG------------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKG 340
GF ++ KACCGG CG +T C DPS NWDG H TEAAY+ IA G
Sbjct: 298 GF-TGSVLKACCGGGDGRYNVQPNVRCGEKGSTTCEDPSTYANWDGIHLTEAAYRHIATG 356
Query: 341 LVEGPFSNPS 350
L+ G F+ P+
Sbjct: 357 LISGRFTMPT 366
>UniRef100_Q9LHW8 Putative esterase [Oryza sativa]
Length = 374
Score = 293 bits (751), Expect = 4e-78
Identities = 148/341 (43%), Positives = 215/341 (62%), Gaps = 19/341 (5%)
Query: 27 SYEAFFNFGDSISDTGN---AASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+Y + F+FGDS++DTGN ++ + + PYG +YF P+GR S+GRL++DF+A+A+
Sbjct: 36 NYTSMFSFGDSLTDTGNLVVSSPLSFSIVGKYPYGMTYFHRPTGRCSDGRLVVDFLAQAF 95
Query: 84 GLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPD-TNNSLSIQLGWFKN 142
GLP L Y ++ +D+ +GVNFA GAT ++ ++ + G TN SLS+QLGWF+
Sbjct: 96 GLPLLQPYLSRG--EDVTRGVNFAVGGATAMDPPFFEEIGASDKLWTNLSLSVQLGWFEQ 153
Query: 143 IKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITNTTN 201
+KP LC S +DC +F KSLF+VGEIGGND K K++ + + VP + A+ + T
Sbjct: 154 LKPSLCSSPKDCKEFFSKSLFLVGEIGGNDYNYAFFKGKSLDDAKSYVPTVAGAVADATE 213
Query: 202 VLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQLKNSI 261
LI+ GAV LVVPGN P+GCS+A TL S+ + DYD GCL YN+ ++ N L++ +
Sbjct: 214 RLIKAGAVHLVVPGNLPIGCSSAYLTLHPSSNRSDYDSTGCLKTYNDFAQHHNAVLQDKL 273
Query: 262 ETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCGG-----------CGSL 310
LR+ +PE +I+Y DYY A Q P+Q+GF A+ + CCGG CG
Sbjct: 274 RLLRRSYPEARIMYADYYGAAMSFAQNPKQFGFRHGAL-RTCCGGGGPYNFNPKASCGVR 332
Query: 311 IATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSL 351
++VC+DPS NWDG H TEA Y IA ++ GP+++P L
Sbjct: 333 GSSVCTDPSAYANWDGVHLTEAGYHAIANSILNGPYTSPRL 373
>UniRef100_Q9LHW4 Putative esterase [Oryza sativa]
Length = 397
Score = 285 bits (729), Expect = 1e-75
Identities = 154/370 (41%), Positives = 213/370 (56%), Gaps = 22/370 (5%)
Query: 2 NVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGN----AASIFLPMP-NPIP 56
N TL L+ F G + ++ ++ F+FG+S +DTGN AA +F +P N +P
Sbjct: 7 NEMTLLLLLFLLGCTHYGHAGSDRPKIDSIFSFGNSYADTGNFVKLAAPVFPGIPFNNLP 66
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GR SNGRL +DFIAE G+P L Y +S QD G NFA GAT L++
Sbjct: 67 YGETFFGHPTGRASNGRLNVDFIAEGLGVPLLAPYHGES--QDFSHGANFAVVGATALDL 124
Query: 117 EYYVKNGLP-LPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGND-IM 174
++ KN + +P N SLS+Q+ WF+ +KP LC + + C YF++SLF +GEIGGND +
Sbjct: 125 AFFQKNNITSVPPFNTSLSVQVEWFQKLKPTLCSTTQGCKDYFERSLFFMGEIGGNDYVF 184
Query: 175 KHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKK 234
+ KTV E VP +V+AI+ +I+EGA +VVPG P GC + TL S
Sbjct: 185 LYAAGKTVDEAMSYVPKVVQAISAGVEAVIKEGARYVVVPGQLPTGCLPIILTLYASPAA 244
Query: 235 EDYDE-FGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYG 293
DYD GCL +N L Y N L ++ LR KHP V I++ DYY + Q P ++G
Sbjct: 245 ADYDAGTGCLWRFNALARYHNAVLFAAVSLLRAKHPSVAIVFADYYRPVIKFVQNPDEFG 304
Query: 294 FDKDAIFKACCGG------------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGL 341
F + + +ACCGG CG A C DP INWDG H TEAAY +A G
Sbjct: 305 FSESSKLRACCGGGGGAYNYDVAAACGFPGAAACPDPDAAINWDGIHLTEAAYGQVAAGW 364
Query: 342 VEGPFSNPSL 351
+ GP+++P +
Sbjct: 365 LRGPYAHPPI 374
>UniRef100_Q9SHP6 F1K23.16 [Arabidopsis thaliana]
Length = 383
Score = 280 bits (716), Expect = 4e-74
Identities = 154/352 (43%), Positives = 205/352 (57%), Gaps = 18/352 (5%)
Query: 15 FLQNVVSNANPLSYEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMS 70
F+ V S + E+ +FGDSI+DTGN + LP+ +PYG ++F HP+GR
Sbjct: 16 FVTIVSSQTQCRNLESIISFGDSITDTGNLVGLSDRNHLPVTAFLPYGETFFHHPTGRSC 75
Query: 71 NGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTN 130
NGR+IIDFIAE GLP +P + S + + +KGVNFA AGAT L K G+ P +N
Sbjct: 76 NGRIIIDFIAEFLGLPHVPPFYG-SKNGNFEKGVNFAVAGATALETSILEKRGIYYPHSN 134
Query: 131 NSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVP 190
SL IQL FK P LC S DC + I+GEIGGND E++E+VP
Sbjct: 135 ISLGIQLKTFKESLPNLCGSPTDCRDMIGNAFIIMGEIGGNDFNFAFFVNKTSEVKELVP 194
Query: 191 FMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNL 249
++ I++ L++ G +VPGNFP+GCSA TL ++ KE+YD GCL N+
Sbjct: 195 LVITKISSAIVELVDMGGRTFLVPGNFPLGCSATYLTLYQTSNKEEYDPLTGCLTWLNDF 254
Query: 250 IEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG---- 305
EY+N +L+ + L + +P V IIY DY+N RLYQ P ++GF D ACCG
Sbjct: 255 SEYYNEKLQAELNRLSKLYPHVNIIYGDYFNALLRLYQEPSKFGF-MDRPLPACCGLGGP 313
Query: 306 -------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CGS+ CSDPSK +NWDG H TEAAYK IA GL++GP++ PS
Sbjct: 314 YNFTLSKKCGSVGVKYCSDPSKYVNWDGVHMTEAAYKWIADGLLKGPYTIPS 365
>UniRef100_Q8LAS3 Lipase, putative [Arabidopsis thaliana]
Length = 393
Score = 276 bits (705), Expect = 8e-73
Identities = 153/365 (41%), Positives = 213/365 (57%), Gaps = 20/365 (5%)
Query: 2 NVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIPY 57
++ +L + F+ F+ V S ++++ +FGDSI+DTGN + LP+ PY
Sbjct: 3 SLDSLVIFLFSTLFVTIVSSETPCRNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPY 62
Query: 58 GSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVE 117
G ++F HP+GR +GR+I+DFIAE GLP++P Y S +++ KGVNFA AGAT L
Sbjct: 63 GETFFHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFG-SKNRNFDKGVNFAVAGATALKSS 121
Query: 118 YYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKH 176
+ K G+ P TN SL +QL FK P LC S DC +L ++GEIGGND
Sbjct: 122 FLKKRGIQ-PHTNVSLRVQLKSFKKSLPNLCGSPSDCRDMIGNALILMGEIGGNDYNFPF 180
Query: 177 MKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKED 236
K V E+ E+VPF++ +I++T LI G +VPG FP+GCS TL ++ K++
Sbjct: 181 FNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKTSNKDE 240
Query: 237 YDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFD 295
YD GCL N EY + +LK + LR+ +P V IIY DYYN R+++ P ++GF
Sbjct: 241 YDPTTGCLKWLNKFGEYHSEKLKAELNRLRKLYPHVNIIYADYYNSLLRIFKEPAKFGF- 299
Query: 296 KDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEG 344
D F ACCG CGS+ C DPSK + WDG H TEAAYK IA G++ G
Sbjct: 300 MDRPFPACCGIGGPYNFNFTRKCGSVGVKSCKDPSKYVGWDGVHMTEAAYKWIADGILNG 359
Query: 345 PFSNP 349
P++NP
Sbjct: 360 PYANP 364
>UniRef100_Q94F40 At1g28600/F1K23_6 [Arabidopsis thaliana]
Length = 393
Score = 274 bits (701), Expect = 2e-72
Identities = 152/365 (41%), Positives = 213/365 (57%), Gaps = 20/365 (5%)
Query: 2 NVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIPY 57
++ +L + F+ F+ V S ++++ +FGDSI+DTGN + LP+ PY
Sbjct: 3 SLDSLVIFLFSTLFVTIVSSETPCPNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPY 62
Query: 58 GSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVE 117
G ++F HP+GR +GR+I+DFIAE GLP++P Y S +++ KGVNFA AGAT L
Sbjct: 63 GETFFHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFG-SKNRNFDKGVNFAVAGATALKSS 121
Query: 118 YYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKH 176
+ K G+ P TN SL +QL FK P LC S DC +L ++GEIGGND
Sbjct: 122 FLKKRGIQ-PHTNVSLGVQLKSFKKSLPNLCGSPSDCRDMIGNALILMGEIGGNDYNFPF 180
Query: 177 MKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKED 236
K V E+ E+VPF++ +I++T LI G +VPG FP+GCS TL ++ K++
Sbjct: 181 FNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKTSNKDE 240
Query: 237 YD-EFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFD 295
YD GCL N EY + +LK + LR+ +P V IIY DYYN R+++ P ++GF
Sbjct: 241 YDPSTGCLKWLNKFGEYHSEKLKVELNRLRKLYPHVNIIYADYYNSLLRIFKEPAKFGF- 299
Query: 296 KDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEG 344
+ F ACCG CGS+ C DPSK + WDG H TEAAYK IA G++ G
Sbjct: 300 MERPFPACCGIGGPYNFNFTRKCGSVGVKSCKDPSKYVGWDGVHMTEAAYKWIADGILNG 359
Query: 345 PFSNP 349
P++NP
Sbjct: 360 PYANP 364
>UniRef100_Q9FXJ3 F1K23.17 [Arabidopsis thaliana]
Length = 823
Score = 274 bits (701), Expect = 2e-72
Identities = 152/365 (41%), Positives = 213/365 (57%), Gaps = 20/365 (5%)
Query: 2 NVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIPY 57
++ +L + F+ F+ V S ++++ +FGDSI+DTGN + LP+ PY
Sbjct: 3 SLDSLVIFLFSTLFVTIVSSETPCPNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPY 62
Query: 58 GSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVE 117
G ++F HP+GR +GR+I+DFIAE GLP++P Y S +++ KGVNFA AGAT L
Sbjct: 63 GETFFHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFG-SKNRNFDKGVNFAVAGATALKSS 121
Query: 118 YYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKH 176
+ K G+ P TN SL +QL FK P LC S DC +L ++GEIGGND
Sbjct: 122 FLKKRGIQ-PHTNVSLGVQLKSFKKSLPNLCGSPSDCRDMIGNALILMGEIGGNDYNFPF 180
Query: 177 MKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKED 236
K V E+ E+VPF++ +I++T LI G +VPG FP+GCS TL ++ K++
Sbjct: 181 FNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKTSNKDE 240
Query: 237 YD-EFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFD 295
YD GCL N EY + +LK + LR+ +P V IIY DYYN R+++ P ++GF
Sbjct: 241 YDPSTGCLKWLNKFGEYHSEKLKVELNRLRKLYPHVNIIYADYYNSLLRIFKEPAKFGF- 299
Query: 296 KDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEG 344
+ F ACCG CGS+ C DPSK + WDG H TEAAYK IA G++ G
Sbjct: 300 MERPFPACCGIGGPYNFNFTRKCGSVGVKSCKDPSKYVGWDGVHMTEAAYKWIADGILNG 359
Query: 345 PFSNP 349
P++NP
Sbjct: 360 PYANP 364
Score = 268 bits (685), Expect = 2e-70
Identities = 149/352 (42%), Positives = 208/352 (58%), Gaps = 19/352 (5%)
Query: 18 NVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSSYFKHPSGRMSNGR 73
+V S ++++ +FGDSI+DTGN + P P PYG ++F HP+GR S+GR
Sbjct: 444 SVNSQTQCRNFKSIISFGDSIADTGNLLGLSDPNDLPASAFPPYGETFFHHPTGRYSDGR 503
Query: 74 LIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSL 133
LIIDFIAE G P +P + + + KKGVNFA AGAT L + + G+ TN SL
Sbjct: 504 LIIDFIAEFLGFPLVPPFYGCQ-NANFKKGVNFAVAGATALEPSFLEERGIHSTITNVSL 562
Query: 134 SIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFM 192
S+QL F P LC S DC + +L ++GEIGGND + K V E+ E+VPF+
Sbjct: 563 SVQLRSFTESLPNLCGSPSDCRDMIENALILMGEIGGNDYNFALFQRKPVKEVEELVPFV 622
Query: 193 VEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIE 251
+ I++ L+ G +VPGNFP+G SA+ TL ++ KE+YD GCL N+ E
Sbjct: 623 IATISSAITELVCMGGRTFLVPGNFPIGYSASYLTLYKTSNKEEYDPLTGCLKWLNDFSE 682
Query: 252 YFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------ 305
Y+N QL+ + LR+ +P V IIY DYYN RL+Q P ++GF + ACCG
Sbjct: 683 YYNKQLQEELNGLRKLYPHVNIIYADYYNALLRLFQEPAKFGF-MNRPLPACCGVGGSYN 741
Query: 306 -----GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSLK 352
CGS+ C DPS+ +N+DG H TEAAY+LI++GL++GP++ P K
Sbjct: 742 FNFSRRCGSVGVEYCDDPSQYVNYDGIHMTEAAYRLISEGLLKGPYAIPPFK 793
>UniRef100_Q5Z739 Putative lipase [Oryza sativa]
Length = 395
Score = 272 bits (696), Expect = 9e-72
Identities = 144/344 (41%), Positives = 206/344 (59%), Gaps = 23/344 (6%)
Query: 27 SYEAFFNFGDSISDTGNAASIFLPMPNPI---PYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
SYEA F+FGDS+SD GN + +P PYG ++F P+GR SNGRL++DF+AE +
Sbjct: 54 SYEAIFSFGDSLSDAGNLIADGIPKSLTTARAPYGMTFFGRPTGRCSNGRLVVDFLAEHF 113
Query: 84 GLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNN-SLSIQLGWFKN 142
GLP PA +K+ D KG NFA GAT L ++ ++G+ N S++ Q+GW ++
Sbjct: 114 GLPLPPA--SKAHGADFSKGANFAITGATALEYSFFKQHGIDQRIWNTGSINTQIGWLQD 171
Query: 143 IKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVI-ELREIVPFMVEAITNTTN 201
+KP LCKS ++C YF KSLF+VGE GGND + E++ VP + +AI N
Sbjct: 172 MKPSLCKSDQECKDYFGKSLFVVGEFGGNDYNAPLFSGVAFSEVKTYVPLVAKAIANGVE 231
Query: 202 VLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYD-EFGCLIAYNNLIEYFNGQLKNS 260
LIE GA +L+VPG P+GC TL N++ K DY+ GCL YN L + N +LK
Sbjct: 232 KLIELGAKDLLVPGVLPIGCFPLYLTLYNTSSKADYNARTGCLRRYNRLAFHHNRELKQQ 291
Query: 261 IETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCGG-------------C 307
++ L++K+PE KI+Y DY+ A + +P +GF + +ACCG C
Sbjct: 292 LDELQKKYPETKIMYGDYFKAAMQFVVSPGNFGF--SSTMQACCGAGGQGNYNFNLKKKC 349
Query: 308 GSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSL 351
G A+VCS+PS ++WDG H TEAAY+ +A G + GP++ P +
Sbjct: 350 GEEGASVCSNPSSYVSWDGIHMTEAAYRYVANGWLNGPYAEPPI 393
>UniRef100_Q9LHW0 Putative esterase [Oryza sativa]
Length = 382
Score = 270 bits (691), Expect = 3e-71
Identities = 142/342 (41%), Positives = 196/342 (56%), Gaps = 21/342 (6%)
Query: 29 EAFFNFGDSISDTGNAASIFLPMP-----NPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
++ F+FG+S SDTGN + P+ N +PYG ++F HP+GR S+GRL +DFIAE +
Sbjct: 37 DSIFSFGNSYSDTGNFVKLAAPVIPVIAFNNLPYGETFFGHPTGRASDGRLNVDFIAEDF 96
Query: 84 GLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLP-LPDTNNSLSIQLGWFKN 142
G+P LP Y +S ++ G NFA GAT L++ ++ KN + +P N SLS+Q+ WF
Sbjct: 97 GVPLLPPYLGES--KNFSHGANFAVVGATALDLAFFQKNNITSVPPFNTSLSVQVEWFHK 154
Query: 143 IKPLLCKSKEDCNIYFKKSLFIVGEIGGND-IMKHMKHKTVIELREIVPFMVEAITNTTN 201
+KP LC + + C YF++SLF +GE GGND + KTV E VP +V I+
Sbjct: 155 LKPTLCSTTQGCRDYFERSLFFMGEFGGNDYVFLLAAGKTVDEAMSYVPKVVGVISAGVE 214
Query: 202 VLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDE-FGCLIAYNNLIEYFNGQLKNS 260
+IEEGA +VVPG P GC + TL S DY+ GCL +N L Y N L +
Sbjct: 215 AVIEEGARYVVVPGQLPTGCLPIILTLYASANATDYESGAGCLRRFNELARYHNAALFAA 274
Query: 261 IETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCGG-----------CGS 309
+ LR KHP I++ DYY + P+ +GF + + +ACCGG CG
Sbjct: 275 VSLLRGKHPSAAIVFADYYQPVIEFVRMPENFGFSRSSRLRACCGGGGRYNYNATAACGL 334
Query: 310 LIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSL 351
AT C DP+ INWDG H TEAAY IA G + GP++ P +
Sbjct: 335 AGATACPDPAASINWDGVHLTEAAYGRIAAGWLRGPYAQPPI 376
>UniRef100_Q65X51 Hypothetical protein OJ1123_F01.16 [Oryza sativa]
Length = 371
Score = 265 bits (678), Expect = 1e-69
Identities = 155/370 (41%), Positives = 209/370 (55%), Gaps = 24/370 (6%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGN---AASIFLPMPNP-IP 56
M + +F I F F + VS+ + + + F+ GDS DTGN AS +P+ N +P
Sbjct: 1 MELKLVFPIAFL--FCLSRVSSTSQF-FTSMFSLGDSYIDTGNFVIMASPVVPVWNDKLP 57
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GRMS+GR+IIDFIAE +GLPFLPA S + GVNFA GA V
Sbjct: 58 YGMTFFDHPTGRMSDGRVIIDFIAEEFGLPFLPASLANS--SSVSHGVNFAVGGAPATGV 115
Query: 117 EYYVKNGL-PLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIY--FKKSLFIVGEIGGNDI 173
EY+ N + P NNSL +QLGWF+ +KP +C S ++ N F K+LFIVGE G ND
Sbjct: 116 EYFENNNIVPFKLLNNSLDVQLGWFEELKPSICNSTDETNGLNCFGKTLFIVGEFGVNDY 175
Query: 174 -MKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSN 232
M K E+ VP +V+ IT LI +GA +VVPGN P GC+ A+ T S
Sbjct: 176 NFMWMAGKPKQEVDSYVPQVVKKITTAVERLITQGAAYVVVPGNPPTGCAPALLTSRMSP 235
Query: 233 KKEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQY 292
K DYD GCL N+++E N L+ ++ LR K+P KII D+YN R+ Q P +
Sbjct: 236 NKTDYDGLGCLRFINDVVERHNTMLRAALGVLRGKYPHAKIILADFYNPIIRVLQNPSHF 295
Query: 293 GFDKDAIFKACCGG-----------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGL 341
G D + KACCG C C DPS ++WDG H+TEA IA+G
Sbjct: 296 GVAADGVLKACCGTGGAYNWNASAICAMPGVVACQDPSAAVSWDGVHYTEAINSYIAQGW 355
Query: 342 VEGPFSNPSL 351
+ GP+++P +
Sbjct: 356 LHGPYADPPI 365
>UniRef100_Q9FXJ1 F1K23.19 [Arabidopsis thaliana]
Length = 389
Score = 265 bits (678), Expect = 1e-69
Identities = 151/366 (41%), Positives = 213/366 (57%), Gaps = 20/366 (5%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNP----IP 56
M + + FLI T L V S ++++ +FGDSI+DTGN ++ P P +P
Sbjct: 6 MKLVSFFLILSTF-CLTTVNSEPQCHNFKSIISFGDSIADTGNLLALSDPTNLPKVAFLP 64
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GR SNGRLIIDFIAE G P +P + S + + +KGVNFA GAT L
Sbjct: 65 YGETFFHHPTGRFSNGRLIIDFIAEFLGFPLVPPFYG-SQNANFEKGVNFAVGGATALER 123
Query: 117 EYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MK 175
+ + G+ P TN SL++QL FK P LC S DC + SL ++GEIGGND
Sbjct: 124 SFLEERGIHFPYTNVSLAVQLSSFKESLPNLCVSPSDCRDMIENSLILMGEIGGNDYNYA 183
Query: 176 HMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKE 235
K + E++E+VP ++E I++ LI G +VPG FP+GCS A +L ++ E
Sbjct: 184 FFVGKNIEEIKELVPLVIETISSAITELIGMGGKTFLVPGEFPLGCSVAYLSLYQTSNIE 243
Query: 236 DYDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGF 294
+YD GCL N EY + QL+ + L++ +P V IIY DYYN RL Q P ++GF
Sbjct: 244 EYDPLTGCLKWLNKFSEYHDEQLQAELNRLQKLYPHVNIIYADYYNTLLRLAQEPAKFGF 303
Query: 295 DKDAIFKACC--GG---------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
+ ACC GG G+ + C DPSK ++WDG H TEAAY+L+A+G+++
Sbjct: 304 ISRPL-PACCALGGPFNFTLGRKRGTQVPECCDDPSKYVSWDGVHMTEAAYRLMAEGILK 362
Query: 344 GPFSNP 349
GP++ P
Sbjct: 363 GPYAIP 368
>UniRef100_Q9C857 Hypothetical protein T8E3.19 [Arabidopsis thaliana]
Length = 394
Score = 261 bits (668), Expect = 2e-68
Identities = 153/366 (41%), Positives = 209/366 (56%), Gaps = 23/366 (6%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIP 56
M + +LFL T F+ V S + ++E+ +FGDSI+DTGN + LPM P
Sbjct: 10 MKLGSLFLSTL---FVSIVSSESQCRNFESIISFGDSIADTGNLLGLSDHNNLPMSAFPP 66
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GR S+GRLIIDFIAE GLP++P Y S + + +KGVNFA A AT L
Sbjct: 67 YGETFFHHPTGRFSDGRLIIDFIAEFLGLPYVPPYFG-STNGNFEKGVNFAVASATALES 125
Query: 117 EYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MK 175
+ + G P N SL +QL FK P LC DC +L ++GEIG ND
Sbjct: 126 SFLEEKGYHCPH-NFSLGVQLKIFKQSLPNLCGLPSDCRDMIGNALILMGEIGANDYNFP 184
Query: 176 HMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKE 235
+ + + E++E+VP ++ I++ LI G +VPG FP+GCS A TL ++ E
Sbjct: 185 FFQLRPLDEVKELVPLVISTISSAITELIGMGGRTFLVPGGFPLGCSVAFLTLHQTSNME 244
Query: 236 DYDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGF 294
+YD GCL N EY + QL+ + LR+ +P V IIY DYYN + RL + P +YGF
Sbjct: 245 EYDPLTGCLKWLNKFGEYHSEQLQEELNRLRKLNPHVNIIYADYYNASLRLGREPSKYGF 304
Query: 295 DKDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
+ ACCG CGS+ CSDPSK + WDG H TEAA+K +A GLV+
Sbjct: 305 INRHL-SACCGVGGPYNFNLSRSCGSVGVEACSDPSKYVAWDGLHMTEAAHKSMADGLVK 363
Query: 344 GPFSNP 349
GP++ P
Sbjct: 364 GPYAIP 369
>UniRef100_Q9ZQI3 Putative lipase [Arabidopsis thaliana]
Length = 394
Score = 261 bits (666), Expect = 3e-68
Identities = 151/363 (41%), Positives = 204/363 (55%), Gaps = 21/363 (5%)
Query: 6 LFLITFTCGFLQNVV-SNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSS 60
+ L F FL VV S ++++ +FGDSI+DTGN + P P PYG +
Sbjct: 8 MLLSFFISTFLITVVTSQTRCRNFKSIISFGDSITDTGNLLGLSSPNDLPESAFPPYGET 67
Query: 61 YFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYV 120
+F HPSGR S+GRLIIDFIAE G+P +P + S + + +KGVNFA GAT L
Sbjct: 68 FFHHPSGRFSDGRLIIDFIAEFLGIPHVPPFYG-SKNGNFEKGVNFAVGGATALECSVLE 126
Query: 121 KNGLPLPDTNNSLSIQLGWFKNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMK 178
+ G +N SL QL FK P LC S DC + + ++GEIGGND
Sbjct: 127 EKGTHCSQSNISLGNQLKSFKESLPYLCGSSSPDCRDMIENAFILIGEIGGNDYNFPLFD 186
Query: 179 HKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYD 238
K + E++E+VP ++ I++ + L++ GA +VPGNFP+GCS A TL + KE+Y+
Sbjct: 187 RKNIEEVKELVPLVITTISSAISELVDMGARTFLVPGNFPLGCSVAYLTLYETPNKEEYN 246
Query: 239 EF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKD 297
GCL N+ Y N QL+ ++ LR +P V IIY DYYN RL Q P ++G D
Sbjct: 247 PLTGCLTWLNDFSVYHNEQLQAELKRLRNLYPHVNIIYGDYYNTLLRLMQEPSKFGL-MD 305
Query: 298 AIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPF 346
ACCG CGS CSDPSK +NWDG H TEAAYK I++G++ GP+
Sbjct: 306 RPLPACCGLGGPYNFTFSIKCGSKGVEYCSDPSKYVNWDGIHMTEAAYKWISEGVLTGPY 365
Query: 347 SNP 349
+ P
Sbjct: 366 AIP 368
>UniRef100_Q5QNL0 Putative esterase [Oryza sativa]
Length = 409
Score = 259 bits (663), Expect = 6e-68
Identities = 147/395 (37%), Positives = 216/395 (54%), Gaps = 46/395 (11%)
Query: 6 LFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLP----MP-NPIPYGSS 60
L L+ F C ++ F+FG+S +DTGN + P MP N +PYG +
Sbjct: 14 LLLLLFGCLHYAQANPGHRRPKIDSVFSFGNSFADTGNFVELAAPLLPIMPFNNLPYGET 73
Query: 61 YFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYV 120
+F HP+GR +NGR+I+DFIA+ + +PF+P + + Q+ G NFA GA+ L++ +++
Sbjct: 74 FFGHPTGRATNGRIIMDFIADEFHVPFVPPFLGQG-RQNFTHGANFAVVGASALDLAFFL 132
Query: 121 KNGLP-LPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKH 179
KN + +P N SLS+QL WF+ +KP LC++ ++C YFK+SLF +GE GGND + +
Sbjct: 133 KNNITNVPPLNISLSVQLEWFQKLKPTLCQTAQECREYFKRSLFFMGEFGGNDYVFILAA 192
Query: 180 -KTVIELREIVPFMVEAIT----------------------NTTNVLIE----EGAVELV 212
KT+ EL VP +V+AI+ T N++I+ EGA +V
Sbjct: 193 GKTLEELVPYVPKVVQAISAGIEAAVKFSLTIYTELTLPLSRTNNIVIQAVIKEGARYVV 252
Query: 213 VPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVK 272
VPG P GC + TL S + DYD GCL N L Y N L ++ LR ++P VK
Sbjct: 253 VPGELPNGCVPIILTLYASKSRGDYDARGCLKKQNALARYHNSALFEAVSRLRHRYPWVK 312
Query: 273 IIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------------GCGSLIATVCSDPSK 320
I+Y DYY + P ++GF+ + +ACCG CG A C DP+
Sbjct: 313 IVYADYYKPVIDFIKKPARFGFNGSSTLRACCGAGGGPYNYDATAACGLPGAAACPDPAA 372
Query: 321 RINWDGPHFTEAAYKLIAKGLVEGPFSNPSLKSPL 355
I+WDG H TEAAY I+ G + GP+++P + S L
Sbjct: 373 FISWDGIHLTEAAYARISAGWLHGPYAHPPILSAL 407
>UniRef100_Q9FXJ2 F1K23.18 [Arabidopsis thaliana]
Length = 390
Score = 259 bits (662), Expect = 8e-68
Identities = 146/361 (40%), Positives = 201/361 (55%), Gaps = 22/361 (6%)
Query: 6 LFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSSY 61
+FL TF + NV S +++ +FGDSI+DTGN + P P PYG ++
Sbjct: 16 IFLSTFV---VTNVSSETKCREFKSIISFGDSIADTGNLLGLSDPKDLPHMAFPPYGENF 72
Query: 62 FKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVK 121
F HP+GR SNGRLIIDFIAE GLP +P + S + + +KGVNFA GAT L +
Sbjct: 73 FHHPTGRFSNGRLIIDFIAEFLGLPLVPPFYG-SHNANFEKGVNFAVGGATALERSFLED 131
Query: 122 NGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHK 180
G+ P TN SL +QL FK P +C S DC + +L ++GEIGGND K
Sbjct: 132 RGIHFPYTNVSLGVQLNSFKESLPSICGSPSDCRDMIENALILMGEIGGNDYNYAFFVDK 191
Query: 181 TVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF 240
+ E++E++P ++ I++ LI G +VPG FP+GCS T ++ E+YD
Sbjct: 192 GIEEIKELMPLVITTISSAITELIGMGGRTFLVPGEFPVGCSVLYLTSHQTSNMEEYDPL 251
Query: 241 -GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAI 299
GCL N E QL+ + L++ +P V IIY DYYN LYQ P ++GF +
Sbjct: 252 TGCLKWLNKFGENHGEQLRAELNRLQKLYPHVNIIYADYYNALFHLYQEPAKFGF-MNRP 310
Query: 300 FKACCGG-----------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSN 348
ACCG CG+ I C DPSK + WDG H TEAAY+L+A+G++ GP++
Sbjct: 311 LSACCGAGGPYNYTVGRKCGTDIVESCDDPSKYVAWDGVHMTEAAYRLMAEGILNGPYAI 370
Query: 349 P 349
P
Sbjct: 371 P 371
>UniRef100_Q38894 Lipase [Arabidopsis thaliana]
Length = 384
Score = 257 bits (657), Expect = 3e-67
Identities = 147/344 (42%), Positives = 202/344 (57%), Gaps = 26/344 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+++ +FGDSI+DTGN + LP +PYG S+F PSGR SNGRLIIDFIAE
Sbjct: 33 FKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGESFFHPPSGRASNGRLIIDFIAEFL 92
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P Y +N S +Q G+NFA GAT L+ + + G+ TN SLS+QL F
Sbjct: 93 GLPYVPPYFGSQNVSFEQ----GINFAVYGATALDRAFLLGKGIESDFTNVSLSVQLDTF 148
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIGGND + K++ E++E+VP +V+AI++
Sbjct: 149 KQILPNLCASSTRDCKEMLGDSLILMGEIGGNDYNYPFFEGKSINEIKELVPLIVKAISS 208
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQL 257
LI+ G +VPG FP GCSAA TL + ++D D GC N E+ N QL
Sbjct: 209 AIVDLIDLGGKTFLVPGGFPTGCSAAYLTLFQTVAEKDQDPLTGCYPLLNEFGEHHNEQL 268
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K ++ L++ +P V IIY DY+N R YQ P +YGF K+ ACCG
Sbjct: 269 KTELKRLQKFYPHVNIIYADYHNSLYRFYQEPAKYGF-KNKPLAACCGVGGKYNFTIGKE 327
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CG C +PS+ +NWDG H TEAAY+ + +G++ GP++ P+
Sbjct: 328 CGYEGVNYCQNPSEYVNWDGYHLTEAAYQKMTEGILNGPYATPA 371
>UniRef100_Q9SHP9 F1K23.13 [Arabidopsis thaliana]
Length = 1411
Score = 257 bits (657), Expect = 3e-67
Identities = 147/344 (42%), Positives = 202/344 (57%), Gaps = 26/344 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+++ +FGDSI+DTGN + LP +PYG S+F PSGR SNGRLIIDFIAE
Sbjct: 33 FKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGESFFHPPSGRASNGRLIIDFIAEFL 92
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P Y +N S +Q G+NFA GAT L+ + + G+ TN SLS+QL F
Sbjct: 93 GLPYVPPYFGSQNVSFEQ----GINFAVYGATALDRAFLLGKGIESDFTNVSLSVQLDTF 148
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIGGND + K++ E++E+VP +V+AI++
Sbjct: 149 KQILPNLCASSTRDCKEMLGDSLILMGEIGGNDYNYPFFEGKSINEIKELVPLIVKAISS 208
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQL 257
LI+ G +VPG FP GCSAA TL + ++D D GC N E+ N QL
Sbjct: 209 AIVDLIDLGGKTFLVPGGFPTGCSAAYLTLFQTVAEKDQDPLTGCYPLLNEFGEHHNEQL 268
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K ++ L++ +P V IIY DY+N R YQ P +YGF K+ ACCG
Sbjct: 269 KTELKRLQKFYPHVNIIYADYHNSLYRFYQEPAKYGF-KNKPLAACCGVGGKYNFTIGKE 327
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CG C +PS+ +NWDG H TEAAY+ + +G++ GP++ P+
Sbjct: 328 CGYEGVNYCQNPSEYVNWDGYHLTEAAYQKMTEGILNGPYATPA 371
Score = 255 bits (651), Expect = 2e-66
Identities = 145/344 (42%), Positives = 205/344 (59%), Gaps = 26/344 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+++ +FGDSI+DTGN + LP +PYG S+F PSGR S+GRLIIDFIAE
Sbjct: 1054 FKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGESFFHPPSGRYSDGRLIIDFIAEFL 1113
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P+Y +N S DQ G+NFA GAT L+ + V G+ TN SLS+QL F
Sbjct: 1114 GLPYVPSYFGSQNVSFDQ----GINFAVYGATALDRVFLVGKGIESDFTNVSLSVQLNIF 1169
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIG ND + K++ E++++VP +++AI++
Sbjct: 1170 KQILPNLCTSSSRDCREMLGDSLILMGEIGVNDYNYPFFEGKSINEIKQLVPLVIKAISS 1229
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQL 257
LI+ G +VPGNFP+GC A TL + +ED+D F GC+ N EY N QL
Sbjct: 1230 AIVDLIDLGGKTFLVPGNFPLGCYPAYLTLFQTAAEEDHDPFTGCIPRLNEFGEYHNEQL 1289
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K ++ L++ + V IIY DYYN RLYQ P +YGF K+ ACCG
Sbjct: 1290 KTELKRLQELYDHVNIIYADYYNSLFRLYQEPVKYGF-KNRPLAACCGVGGQYNFTIGKE 1348
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CG + C +PS+ +NWDG H TEA ++ +A+ ++ G +++P+
Sbjct: 1349 CGHRGVSCCQNPSEYVNWDGYHLTEATHQKMAQVILNGTYASPA 1392
Score = 250 bits (639), Expect = 4e-65
Identities = 144/335 (42%), Positives = 197/335 (57%), Gaps = 25/335 (7%)
Query: 28 YEAFFNFGDSISDTGNAASIF----LPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
Y++ +FGDSI+DTGN + LP +PYG S+F PSGR S+GRL+IDFIAE
Sbjct: 683 YKSIISFGDSIADTGNYVHLSNVNNLPQAAFLPYGESFFHPPSGRYSDGRLVIDFIAEFL 742
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P Y +N S +Q G+NFA GAT L+ + VK G+ TN SLS+QL F
Sbjct: 743 GLPYVPPYFGSQNVSFNQ----GINFAVYGATALDRAFLVKQGIKSDFTNISLSVQLNTF 798
Query: 141 KNIKPLLCKSK-EDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIGGND + K++ E++E+VP +++AI++
Sbjct: 799 KQILPNLCASSTRDCREMLGDSLILMGEIGGNDYNYPFFEGKSINEIKELVPLIIKAISS 858
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQLK 258
LI+ G +VPGNFP+GCS A TL + E GC+ N E+ N QLK
Sbjct: 859 AIVDLIDLGGKTFLVPGNFPIGCSTAYLTLFQTATVEHDPFTGCIPWLNKFGEHHNEQLK 918
Query: 259 NSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------GC 307
++ L++ +P V IIY DYYN L+Q P +YGF K+ ACCG C
Sbjct: 919 IELKQLQKLYPHVNIIYADYYNSLYGLFQEPAKYGF-KNRPLAACCGVGGQYNFTIGKEC 977
Query: 308 GSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLV 342
G + C +PS+ +NWDG H TEA Y+ +A+GL+
Sbjct: 978 GENGVSYCQNPSEYVNWDGYHLTEATYQKMAQGLL 1012
Score = 218 bits (554), Expect = 3e-55
Identities = 135/332 (40%), Positives = 175/332 (52%), Gaps = 62/332 (18%)
Query: 39 SDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAY--- 91
SDTGN + LP PYG S+F PSGR S+GRLIIDFIAE GLP++P Y
Sbjct: 379 SDTGNILHLSDVNHLPQTAFFPYGESFFHPPSGRASDGRLIIDFIAEFLGLPYVPPYFGS 438
Query: 92 ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLC-KS 150
+N S +Q G+NFA GAT L+ Y+V G+ TN SL +QL FK I P LC S
Sbjct: 439 QNVSFEQ----GINFAVYGATALDRAYFVAKGIESDFTNVSLGVQLDIFKQILPNLCASS 494
Query: 151 KEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVE 210
DC SL ++GEIGG KT +
Sbjct: 495 SRDCREMLGDSLILMGEIGGG--------KTFL--------------------------- 519
Query: 211 LVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHP 269
VPG FP GCSAA T + +EDYD GC+ N L E+ N QLK ++ L++ +P
Sbjct: 520 --VPGGFPAGCSAACLTQYQNATEEDYDPLTGCIPRLNELGEHDNEQLKTELKRLQKLYP 577
Query: 270 EVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------GCGSLIATVCSDP 318
+V IIY DY+N R YQ P +YGF K+ ACCG CG + C +P
Sbjct: 578 DVNIIYADYHNSLYRFYQEPAKYGF-KNKPLAACCGVGGKYNFTIGKECGYEGVSYCQNP 636
Query: 319 SKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
S+ +NWDG H TEAAY+ +A+G++ GP++ P+
Sbjct: 637 SEYVNWDGYHLTEAAYQKMAEGILNGPYATPA 668
>UniRef100_Q5QNK2 Putative esterase [Oryza sativa]
Length = 386
Score = 257 bits (656), Expect = 4e-67
Identities = 145/372 (38%), Positives = 201/372 (53%), Gaps = 23/372 (6%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMP-----NPI 55
M L L+ G +N ++ F+FG+S SDTGN + P+ + +
Sbjct: 9 MTFLLLLLLQLLIGCTHYAQANPGHHMIDSIFSFGNSYSDTGNFVKLAAPLLPVIPLDNL 68
Query: 56 PYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLN 115
PYG ++F HP+GR SNGRLIIDFIA +G+PFLP Y + Q+ G NFA GAT L+
Sbjct: 69 PYGETFFGHPAGRASNGRLIIDFIAGHFGVPFLPPYLGQV--QNFSHGANFAVVGATALD 126
Query: 116 VEYYVKNGLP-LPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGND-I 173
+ ++ KN + +P N+SLS+QL WF ++P LC + C YF++SLF +GE GGND +
Sbjct: 127 LAFFQKNNITNVPPFNSSLSVQLEWFHKLRPTLCSKTQGCKHYFERSLFFMGEFGGNDYV 186
Query: 174 MKHMKHKTVIELRE-IVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSN 232
KTV E+ VP ++ AI+ +IEEGA +VVPG P GC + T S
Sbjct: 187 FLLAAGKTVDEVMSCYVPKVIGAISAGVEAVIEEGARYVVVPGQQPTGCLPVVLTPYASP 246
Query: 233 KKEDYDE-FGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQ 291
DYD GCL +N L Y N L ++ LR+K+P I++ DYY+ Q P
Sbjct: 247 NATDYDAGTGCLWRFNELARYHNAALLAAVSLLRRKYPSATIVFADYYDPVIEFMQKPDD 306
Query: 292 YGFDKDAIFKACCGG------------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAK 339
+ F + +ACCGG CG +VC P+ INWDG H TEAAY IA
Sbjct: 307 FAFSDSSKLRACCGGGGGPYNYNATVACGLPGTSVCPTPNTSINWDGIHLTEAAYARIAA 366
Query: 340 GLVEGPFSNPSL 351
+ GP ++P +
Sbjct: 367 CWLHGPHAHPPI 378
>UniRef100_Q75IK2 Hypothetical protein OSJNBb0016G07.10 [Oryza sativa]
Length = 370
Score = 256 bits (654), Expect = 7e-67
Identities = 139/351 (39%), Positives = 204/351 (57%), Gaps = 25/351 (7%)
Query: 20 VSNANPLS--YEAFFNFGDSISDTGNAASIFL-PMPNPIPYGSSYFKHPSGRMSNGRLII 76
V+ A+PL Y A F+FGDS SDTGN I +PN F P R SNGRL+I
Sbjct: 20 VAMADPLPSYYNAIFSFGDSFSDTGNFVIINSGKLPN-----MPKFPPPYARCSNGRLVI 74
Query: 77 DFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGL-PLPDTNNSLSI 135
DF+AEA+GLP LP NK + +G NFA GAT L+++Y+ N + +P N S+++
Sbjct: 75 DFLAEAFGLPLLPPSANKGTN--FSQGANFAVMGATALDLKYFKDNNVWSIPPFNTSMNV 132
Query: 136 QLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIE-LREIVPFMVE 194
QL WF +K +C S ++C +F K+LF+ GE GGND K + +E ++ +VP +V
Sbjct: 133 QLQWFDEVKQTICSSPQECREFFSKALFVFGEFGGNDYSFAWKAEWSLEKVKTMVPSVVA 192
Query: 195 AITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYD-EFGCLIAYNNLIEYF 253
++ L++EGA +VVPGN P GC T+ + + +YD GCL YN++ Y
Sbjct: 193 SMAGGIERLLDEGARHVVVPGNLPAGCIPITLTMYATEDRSEYDPRTGCLKKYNSVALYH 252
Query: 254 NGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCGG------- 306
N L+ +++ L+++HP+ +I+Y DYY + +TP YG+ + A+ +ACCGG
Sbjct: 253 NAMLRIALDQLQRRHPDSRIVYADYYTPYIQFARTPHLYGYKRGAL-RACCGGGGPYNYN 311
Query: 307 ----CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSLKS 353
CG AT C DP ++WDG H TEA Y+ IA + GP+++P L S
Sbjct: 312 MSASCGLPGATTCEDPDAHVSWDGIHLTEAPYRFIANTWIRGPYAHPPLAS 362
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.140 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 660,615,044
Number of Sequences: 2790947
Number of extensions: 30392719
Number of successful extensions: 71226
Number of sequences better than 10.0: 338
Number of HSP's better than 10.0 without gapping: 201
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 69961
Number of HSP's gapped (non-prelim): 378
length of query: 359
length of database: 848,049,833
effective HSP length: 128
effective length of query: 231
effective length of database: 490,808,617
effective search space: 113376790527
effective search space used: 113376790527
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC130806.4


