

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
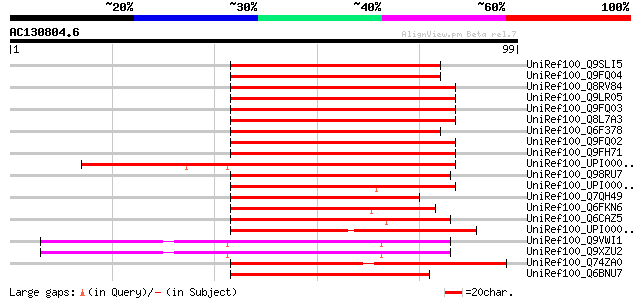
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SLI5 F20D21.30 protein [Arabidopsis thaliana] 66 2e-10
UniRef100_Q9FQ04 XRN4 [Arabidopsis thaliana] 66 2e-10
UniRef100_Q8RV84 Putative 5-3 exoribonuclease [Oryza sativa] 63 2e-09
UniRef100_Q9LR05 F10A5.15 [Arabidopsis thaliana] 62 3e-09
UniRef100_Q9FQ03 XRN3 [Arabidopsis thaliana] 62 3e-09
UniRef100_Q8L7A3 Hypothetical protein At1g75660 [Arabidopsis tha... 62 3e-09
UniRef100_Q6F378 Putative 5'-3' exonuclease [Oryza sativa] 62 3e-09
UniRef100_Q9FQ02 XRN2 [Arabidopsis thaliana] 61 6e-09
UniRef100_Q9FH71 5'-3' exoribonuclease 2 [Arabidopsis thaliana] 61 6e-09
UniRef100_UPI00002D8B13 UPI00002D8B13 UniRef100 entry 52 4e-06
UniRef100_Q98RU7 Very similar to mouse Dhm1 and Dhm2 [Guillardia... 50 1e-05
UniRef100_UPI000004DE22 UPI000004DE22 UniRef100 entry 49 3e-05
UniRef100_Q7QH49 ENSANGP00000004085 [Anopheles gambiae str. PEST] 49 3e-05
UniRef100_Q6FKN6 Similar to sp|Q02792 Saccharomyces cerevisiae Y... 49 3e-05
UniRef100_Q6CAZ5 Yarrowia lipolytica chromosome C of strain CLIB... 49 3e-05
UniRef100_UPI00002A0B92 UPI00002A0B92 UniRef100 entry 49 4e-05
UniRef100_Q9VWI1 CG3291-PA [Drosophila melanogaster] 49 4e-05
UniRef100_Q9XZU2 Pacman protein [Drosophila melanogaster] 49 4e-05
UniRef100_Q74ZA0 AGR303Wp [Ashbya gossypii] 49 4e-05
UniRef100_Q6BNU7 Similar to sp|Q02792 Saccharomyces cerevisiae Y... 49 4e-05
>UniRef100_Q9SLI5 F20D21.30 protein [Arabidopsis thaliana]
Length = 965
Score = 66.2 bits (160), Expect = 2e-10
Identities = 32/41 (78%), Positives = 35/41 (85%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
IDGV PRAKMN+QR+RRFR AKD EAEEERLRK+FEME
Sbjct: 103 IDGVAPRAKMNQQRSRRFRAAKDAAEAEAEEERLRKDFEME 143
>UniRef100_Q9FQ04 XRN4 [Arabidopsis thaliana]
Length = 947
Score = 66.2 bits (160), Expect = 2e-10
Identities = 32/41 (78%), Positives = 35/41 (85%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
IDGV PRAKMN+QR+RRFR AKD EAEEERLRK+FEME
Sbjct: 103 IDGVAPRAKMNQQRSRRFRAAKDAAEAEAEEERLRKDFEME 143
>UniRef100_Q8RV84 Putative 5-3 exoribonuclease [Oryza sativa]
Length = 1064
Score = 62.8 bits (151), Expect = 2e-09
Identities = 31/44 (70%), Positives = 34/44 (76%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR AKD AEEERLR+EFE E K
Sbjct: 102 IDGVAPRAKMNQQRSRRFRAAKDAADAAAEEERLREEFEREGRK 145
>UniRef100_Q9LR05 F10A5.15 [Arabidopsis thaliana]
Length = 1037
Score = 62.4 bits (150), Expect = 3e-09
Identities = 30/44 (68%), Positives = 35/44 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR+AKD AEEERLR+EFE E +
Sbjct: 102 IDGVAPRAKMNQQRSRRFRSAKDASDAAAEEERLREEFEREGRR 145
>UniRef100_Q9FQ03 XRN3 [Arabidopsis thaliana]
Length = 1020
Score = 62.4 bits (150), Expect = 3e-09
Identities = 30/44 (68%), Positives = 35/44 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR+AKD AEEERLR+EFE E +
Sbjct: 102 IDGVAPRAKMNQQRSRRFRSAKDASDAAAEEERLREEFEREGRR 145
>UniRef100_Q8L7A3 Hypothetical protein At1g75660 [Arabidopsis thaliana]
Length = 763
Score = 62.4 bits (150), Expect = 3e-09
Identities = 30/44 (68%), Positives = 35/44 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR+AKD AEEERLR+EFE E +
Sbjct: 102 IDGVAPRAKMNQQRSRRFRSAKDASDAAAEEERLREEFEREGRR 145
>UniRef100_Q6F378 Putative 5'-3' exonuclease [Oryza sativa]
Length = 988
Score = 62.4 bits (150), Expect = 3e-09
Identities = 31/41 (75%), Positives = 33/41 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
IDGV PRAKMN+QR+RRFR AKD AEEERLRKEFE E
Sbjct: 103 IDGVAPRAKMNQQRSRRFRAAKDAADAAAEEERLRKEFEAE 143
>UniRef100_Q9FQ02 XRN2 [Arabidopsis thaliana]
Length = 1012
Score = 61.2 bits (147), Expect = 6e-09
Identities = 30/44 (68%), Positives = 33/44 (74%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR RRFR AKD AEEE+LR+EFE E K
Sbjct: 104 IDGVAPRAKMNQQRARRFRAAKDAAEAAAEEEQLREEFEREGKK 147
>UniRef100_Q9FH71 5'-3' exoribonuclease 2 [Arabidopsis thaliana]
Length = 746
Score = 61.2 bits (147), Expect = 6e-09
Identities = 30/44 (68%), Positives = 33/44 (74%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR RRFR AKD AEEE+LR+EFE E K
Sbjct: 104 IDGVAPRAKMNQQRARRFRAAKDAAEAAAEEEQLREEFEREGKK 147
>UniRef100_UPI00002D8B13 UPI00002D8B13 UniRef100 entry
Length = 495
Score = 51.6 bits (122), Expect = 4e-06
Identities = 31/76 (40%), Positives = 48/76 (62%), Gaps = 3/76 (3%)
Query: 15 ANTLKQKIHETLHAKFNRW--LIAPPPTL-PDIDGVTPRAKMNKQRTRRFRTAKDNEMRE 71
A T ++++ E + +R +I P L IDGV PRAKMN+QR+RRFR+A++ + +
Sbjct: 14 APTTEEEVFECIFDYIDRLFLMIRPRKVLYMAIDGVAPRAKMNQQRSRRFRSAQEAKEKA 73
Query: 72 AEEERLRKEFEMEANK 87
EEE+LR++ E K
Sbjct: 74 IEEEKLREKLIAEGVK 89
>UniRef100_Q98RU7 Very similar to mouse Dhm1 and Dhm2 [Guillardia theta]
Length = 542
Score = 50.4 bits (119), Expect = 1e-05
Identities = 23/43 (53%), Positives = 30/43 (69%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEAN 86
IDGV PRAKMN+QR+RRFR KDNE + + R R +++N
Sbjct: 81 IDGVAPRAKMNQQRSRRFRAIKDNEKKSDFDHRNRLNLTLDSN 123
>UniRef100_UPI000004DE22 UPI000004DE22 UniRef100 entry
Length = 968
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/50 (52%), Positives = 32/50 (64%), Gaps = 6/50 (12%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMR------EAEEERLRKEFEMEANK 87
+DGV PRAKMN+QR RRFR AKD E++ E +E LR E +A K
Sbjct: 97 VDGVAPRAKMNQQRARRFRAAKDAELKAKQLEIEVQERELRGEIINDAIK 146
>UniRef100_Q7QH49 ENSANGP00000004085 [Anopheles gambiae str. PEST]
Length = 902
Score = 48.9 bits (115), Expect = 3e-05
Identities = 22/37 (59%), Positives = 29/37 (77%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
IDGV PRAKMN+QR+RRFR +K+ + AE R+R+E
Sbjct: 102 IDGVAPRAKMNQQRSRRFRASKETAEKAAEMARIREE 138
>UniRef100_Q6FKN6 Similar to sp|Q02792 Saccharomyces cerevisiae YOR048c RAT1 5 -3
exoribonuclease [Candida glabrata]
Length = 1018
Score = 48.9 bits (115), Expect = 3e-05
Identities = 23/41 (56%), Positives = 34/41 (82%), Gaps = 1/41 (2%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEM-REAEEERLRKEFEM 83
+DGV PRAKMN+QR+RRFR+A+D E+ EA EE +R++ ++
Sbjct: 101 VDGVAPRAKMNQQRSRRFRSARDAEIENEAREEIMRQKEQL 141
>UniRef100_Q6CAZ5 Yarrowia lipolytica chromosome C of strain CLIB99 of Yarrowia
lipolytica [Yarrowia lipolytica]
Length = 1526
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/46 (56%), Positives = 33/46 (71%), Gaps = 3/46 (6%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREA---EEERLRKEFEMEAN 86
IDGV PRAKMN+QR+RRFRTA D E +A EE + KE + ++N
Sbjct: 84 IDGVAPRAKMNQQRSRRFRTALDAEKAKAKALEEGTILKEDQFDSN 129
>UniRef100_UPI00002A0B92 UPI00002A0B92 UniRef100 entry
Length = 245
Score = 48.5 bits (114), Expect = 4e-05
Identities = 27/48 (56%), Positives = 32/48 (66%), Gaps = 1/48 (2%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLN 91
IDGV PRAKMN+QR+RRFR+A + +E EE R R E E E F N
Sbjct: 31 IDGVAPRAKMNQQRSRRFRSASE-AAKEREEARRRGEPEPEGEPFDSN 77
>UniRef100_Q9VWI1 CG3291-PA [Drosophila melanogaster]
Length = 1612
Score = 48.5 bits (114), Expect = 4e-05
Identities = 32/84 (38%), Positives = 46/84 (54%), Gaps = 6/84 (7%)
Query: 7 NKFACHLLANTLKQKIHETLHAKFNRWLIAPPPTL-PDIDGVTPRAKMNKQRTRRFRTAK 65
N HL + Q+I + F +LI P +DGV PRAKMN+QR+RRFRTA+
Sbjct: 49 NNIHFHLEEEQIFQEIFNYVDKLF--YLIKPQRLFFLSVDGVAPRAKMNQQRSRRFRTAR 106
Query: 66 DNEMRE---AEEERLRKEFEMEAN 86
+ E +E A+ LR+ ++N
Sbjct: 107 EAEQQEAKAAQRGELREHERFDSN 130
>UniRef100_Q9XZU2 Pacman protein [Drosophila melanogaster]
Length = 1613
Score = 48.5 bits (114), Expect = 4e-05
Identities = 32/84 (38%), Positives = 46/84 (54%), Gaps = 6/84 (7%)
Query: 7 NKFACHLLANTLKQKIHETLHAKFNRWLIAPPPTL-PDIDGVTPRAKMNKQRTRRFRTAK 65
N HL + Q+I + F +LI P +DGV PRAKMN+QR+RRFRTA+
Sbjct: 49 NNIHFHLEEEQIFQEIFNYVDKLF--YLIKPQRLFFLSVDGVAPRAKMNQQRSRRFRTAR 106
Query: 66 DNEMRE---AEEERLRKEFEMEAN 86
+ E +E A+ LR+ ++N
Sbjct: 107 EAEQQEAKAAQRGELREHERFDSN 130
>UniRef100_Q74ZA0 AGR303Wp [Ashbya gossypii]
Length = 945
Score = 48.5 bits (114), Expect = 4e-05
Identities = 26/54 (48%), Positives = 37/54 (68%), Gaps = 2/54 (3%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVKCQ 97
+DGV PRAKMN+QR+RRFR+A+D ++ A EE+ R E EA ++ VK +
Sbjct: 101 VDGVAPRAKMNQQRSRRFRSARDAKL--ANEEKARVLAEREAYGEMIDDAVKAK 152
>UniRef100_Q6BNU7 Similar to sp|Q02792 Saccharomyces cerevisiae YOR048c RAT1 5'-3'
exoribonuclease [Debaryomyces hansenii]
Length = 1003
Score = 48.5 bits (114), Expect = 4e-05
Identities = 22/39 (56%), Positives = 31/39 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFE 82
+DGV PRAKMN+QR+RRFR+A+D ++ E+ER +E E
Sbjct: 98 VDGVAPRAKMNQQRSRRFRSAQDAKIAHEEKERQIRERE 136
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 168,298,587
Number of Sequences: 2790947
Number of extensions: 6068754
Number of successful extensions: 27338
Number of sequences better than 10.0: 144
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 27102
Number of HSP's gapped (non-prelim): 236
length of query: 99
length of database: 848,049,833
effective HSP length: 75
effective length of query: 24
effective length of database: 638,728,808
effective search space: 15329491392
effective search space used: 15329491392
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC130804.6


