

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.11 - phase: 0
(275 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
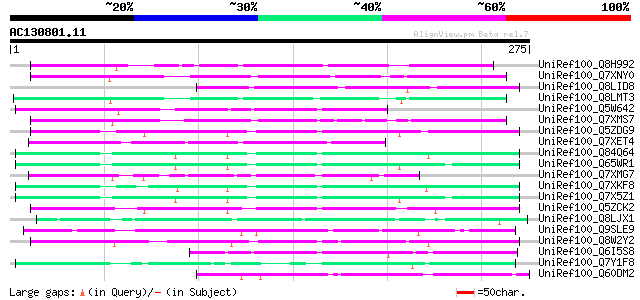
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 104 2e-21
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 100 4e-20
UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [... 86 1e-15
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 85 2e-15
UniRef100_Q5W642 Retrotransposon protein, putative, unclassified... 81 3e-14
UniRef100_Q7XMS7 OSJNBa0029L02.10 protein [Oryza sativa] 80 8e-14
UniRef100_Q5ZDG9 Hypothetical protein P0480E02.9 [Oryza sativa] 79 1e-13
UniRef100_Q7XET4 Contains similarity to non-LTR reverse transcri... 77 5e-13
UniRef100_Q84Q64 Hypothetical protein OSJNBa0071M09.16 [Oryza sa... 75 1e-12
UniRef100_Q65WR1 Putative polyprotein [Oryza sativa] 75 1e-12
UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa] 75 2e-12
UniRef100_Q7XKF8 OSJNBb0065J09.11 protein [Oryza sativa] 75 2e-12
UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa] 75 2e-12
UniRef100_Q5ZCK2 Rust resistance protein Rp1-dp8-like [Oryza sat... 74 3e-12
UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor] 72 1e-11
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 72 2e-11
UniRef100_Q8W2Y2 Putative reverse transcriptase [Oryza sativa] 72 2e-11
UniRef100_Q6I5S8 Putative polyprotein [Oryza sativa] 72 2e-11
UniRef100_Q7Y1F8 Putative reverse transcriptase [Oryza sativa] 69 1e-10
UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa] 69 1e-10
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 104 bits (260), Expect = 2e-21
Identities = 78/253 (30%), Positives = 116/253 (45%), Gaps = 38/253 (15%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPL--------RLSRGGEG 63
+GDGS T FW W G L QF LF + D TV +L RLS +
Sbjct: 926 IGDGSRTSFWHDRWTGNMTLAAQFESLFSHSTDDLATVEQILSFGIYDILTPRLSSAAQN 985
Query: 64 WSWRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGNS 123
L+E +T+L + SL T+ SD+ L N +G + + Y++ + G
Sbjct: 986 -------------DLQELQTILQNFSL-TNESDTR--LNNRLGNLTTKTIYDLRSPPG-- 1027
Query: 124 ITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSD 183
IW ++PLK+ +FAW L+RDRLST+ NL + I+Q TA C + ET+D
Sbjct: 1028 FPSPNWKFIWDCRMPLKIKLFAWLLVRDRLSTKLNLLKKKIVQ--TATCDICSTTDETAD 1085
Query: 184 HLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTT 243
HL +CPF W+ L + I N + L A Q ++L
Sbjct: 1086 HLSFNCPFAISFWQALHI----------QPIINETKYLSQLKAPATIPTRHFQSFFMLCF 1135
Query: 244 WVIWSERNNRLFK 256
WV+W+ R++ +F+
Sbjct: 1136 WVLWNHRHDVVFR 1148
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 100 bits (249), Expect = 4e-20
Identities = 72/261 (27%), Positives = 112/261 (42%), Gaps = 32/261 (12%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVAN--------MLPLRLSRGGEG 63
VG+G TLFW W GG PL QFP L+ + + K T+A+ M R G +
Sbjct: 731 VGNGEKTLFWEDSWLGGKPLAIQFPSLYGIVITKRITIADLNRKGIDCMKFRRDLHGDKL 790
Query: 64 WSWRRWLWAWEEI-VLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGN 122
WR+ + +WE + ++E C+ D W + G ++VR Y L
Sbjct: 791 RDWRKIVNSWEGLNLVENCK-------------DKLWWTLSKDGKFTVRSFYRALKLQQT 837
Query: 123 SITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETS 182
S + IW +VPLK+ IF W ++++ T+ NL RG + D C +ET
Sbjct: 838 SFPNKK---IWKFRVPLKIRIFIWFFTKNKILTKDNLLKRGWRKGDNK--CQFCDKVETV 892
Query: 183 DHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLT 242
HLF CP +W I C + V ++S + + + K ++ + +
Sbjct: 893 QHLFFDCPLARLIWN----IIACALNV-KPVLSRQDLFGSWIQSMDKFTKNLVIVGIAAV 947
Query: 243 TWVIWSERNNRLFKNVVTEAP 263
W IW RN F+ + P
Sbjct: 948 LWSIWKCRNKACFERKLPNDP 968
>UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [Oryza sativa]
Length = 210
Score = 85.9 bits (211), Expect = 1e-15
Identities = 55/176 (31%), Positives = 78/176 (44%), Gaps = 13/176 (7%)
Query: 100 WLPNPIGGYSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNL 159
W P P G +SV AY++ S IW P +V F W R R T NL
Sbjct: 25 WKPGPDGAFSVSSAYDMFFMARQSYPFG--QHIWQTNAPSRVRFFFWLAARGRCQTADNL 82
Query: 160 ASRGILQLDT-ALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGV----DSNII 214
A +G D+ ALC+ E HLF++C F +WE +R WI + D ++I
Sbjct: 83 AKKGWPHEDSCALCMR---EQEDCHHLFVTCEFSGRVWELMRAWISVDFPIPGQIDCSLI 139
Query: 215 SNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
F C F R + +L W+IW ERN R+F+ + +L++ IK
Sbjct: 140 DWWFEARRC---FRTGYREIFDAVLMLVCWLIWKERNARIFEQRLRSPEQLVEDIK 192
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 84.7 bits (208), Expect = 2e-15
Identities = 74/273 (27%), Positives = 106/273 (38%), Gaps = 39/273 (14%)
Query: 3 WFGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANM-------LPL 55
W KVG+G T FW W G PL+ Q+P ++ + DK TV+ M + L
Sbjct: 454 WMDLGSSYKVGNGKATNFWSDVWIGETPLKTQYPNIYRMCADKEKTVSQMCLEGDWYIEL 513
Query: 56 RLSRGGEGWS-WRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAY 114
R S G + W EI L+E R D IW G Y + Y
Sbjct: 514 RRSLGERDLNEWNDLHNTPREIHLKEER-------------DCIIWKLTKNGFYKAKTLY 560
Query: 115 EVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVT 174
+ L+ G + D + +W +PLKV I W +++ R+ L ++ +
Sbjct: 561 QALSFGG--VKDKVMQDLWRSSIPLKVKILFWLMLKGRIQAAGQLKK---MKWSGSPNCK 615
Query: 175 GCGSIETSDHLFLSCPFFAELWEQLRLWIGCC----VGVDSNIISNHFVQFTCLSGFAKA 230
CG E DHL C +W CC G D NI S+ L+
Sbjct: 616 LCGQTEDVDHLMFRCHISQFMW--------CCYRDAFGWD-NIPSSREDLLEKLNIQKDG 666
Query: 231 RRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAP 263
++S L TW IW RN+ +F +T P
Sbjct: 667 QKSVLLACLAAGTWAIWLMRNDWVFNGKLTSDP 699
>UniRef100_Q5W642 Retrotransposon protein, putative, unclassified [Oryza sativa]
Length = 1090
Score = 80.9 bits (198), Expect = 3e-14
Identities = 59/202 (29%), Positives = 87/202 (42%), Gaps = 16/202 (7%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLR-----LS 58
F + R +GDG + W W GG PL+ P LF + K +V+ L L
Sbjct: 802 FAAATRVTLGDGRKAILWESNWIGGQPLKSIAPNLFKHSKRKNRSVSEALTNNAWIGDLR 861
Query: 59 RGGEGWSWRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLT 118
+G + ++ W+ + E + L D+ W+ G YSV Y V
Sbjct: 862 QGITTQLLQEFVQVWQVLYNAE-------IELHEGEEDTIQWIGTASGSYSVSSTYNV-Q 913
Query: 119 ATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGS 178
G + ++SA DLIW P K FAW L +R+ T L RG + C +
Sbjct: 914 FMGRTKSNSA-DLIWKTWAPGKHKFFAWLLAHNRIWTADRLQCRG--WPNNYFCQLCFRN 970
Query: 179 IETSDHLFLSCPFFAELWEQLR 200
+ET+ HLF CPF ++W + R
Sbjct: 971 LETAQHLFKECPFVQQIWNEAR 992
>UniRef100_Q7XMS7 OSJNBa0029L02.10 protein [Oryza sativa]
Length = 389
Score = 79.7 bits (195), Expect = 8e-14
Identities = 66/257 (25%), Positives = 111/257 (42%), Gaps = 28/257 (10%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANML-----PLRLSRGGEGWSW 66
VG+G +TLFW W GG PL EQF L+++ + K TVA++ ++ R G
Sbjct: 15 VGNGRNTLFWEDNWIGGKPLAEQFRDLYNITLTKKITVADVKQVGIDSIKFRRDVLGNKL 74
Query: 67 RRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGNSITD 126
+ +W ++VLE+ D+ W G ++V+ + L
Sbjct: 75 LKIKQSWSDLVLED------------GTRDTLRWNLTKDGKFTVKSFHTALKMQQVIFPH 122
Query: 127 SAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLF 186
IW +VPLK+ IF W ++++ T+ NL RG + A C C ET LF
Sbjct: 123 KK---IWGLKVPLKIKIFIWMFTKNKILTKDNLFKRG-WRKGKANC-QFCDQTETLQPLF 177
Query: 187 LSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVI 246
CP LW ++ C + ++S +++ + + ++ K + + + W I
Sbjct: 178 Y-CPLARLLWNIVK----CALNINS-VLNRQDLFGSWMNKIDKFTKKLVVVGLAAIIWTI 231
Query: 247 WSERNNRLFKNVVTEAP 263
W RN F+ + P
Sbjct: 232 WKFRNKACFEKKLPADP 248
>UniRef100_Q5ZDG9 Hypothetical protein P0480E02.9 [Oryza sativa]
Length = 385
Score = 79.0 bits (193), Expect = 1e-13
Identities = 70/274 (25%), Positives = 115/274 (41%), Gaps = 32/274 (11%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWL- 70
VGDG+ FW W G L++ P ++ + K +T L + W + L
Sbjct: 113 VGDGTKVSFWDSGWLQGRRLKDVAPLVYAASKKKTST--------LQQASLSDQWMQDLD 164
Query: 71 ------WAWEEI-VLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAY--EVLTATG 121
W+ E I L E +++ ++ L D W G Y+ AY ++L T
Sbjct: 165 LPENTGWSIELIDQLIEVWSVVQNLHLIEHEEDKITWKLTSHGEYTTTSAYKAQLLGTTA 224
Query: 122 NSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIET 181
+ + LIW P K FAW ++++R+ T + LA+RG + ++C ET
Sbjct: 225 TNFNN----LIWKPWAPHKCKTFAWLIIQNRVWTSNRLATRG--WPNGSICPLCRHRQET 278
Query: 182 SDHLFLSCPFFAELWEQLRLWIGC-----CVGVDSNIISNHFVQFTCLSGFAKARRSFLQ 236
+ HL C + +W + W C + ++ +S + + A + L+
Sbjct: 279 AIHLLAECRYTRRIWGAIAEWTACEQLNPSLWHQASTVSEWW---EATANTKDAPKKALR 335
Query: 237 LIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+ LL W IW+ERN R F+ LL KIK
Sbjct: 336 TLTLLVAWEIWNERNRRTFQQKELSMGSLLAKIK 369
>UniRef100_Q7XET4 Contains similarity to non-LTR reverse transcriptase [Oryza sativa]
Length = 379
Score = 77.0 bits (188), Expect = 5e-13
Identities = 57/189 (30%), Positives = 84/189 (44%), Gaps = 10/189 (5%)
Query: 11 KVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWL 70
+VGDG+ T FW W L F L+ A++K TV N++ L S L
Sbjct: 153 QVGDGARTSFWHDCWISKPALANVFMPLYTHALNKDDTVRNVMQRGLVS-----SLVPRL 207
Query: 71 WAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVD 130
+ L+ + L +L + D + + + YE G I D
Sbjct: 208 TVTANLELQRLQAALQSFTLSNE-PDKRTYGTKAAPNVTCKQFYEARLLPG--IVSPLWD 264
Query: 131 LIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCP 190
IW + PLKV FAW L++DRLST+ NL + I+ D +C G+ ET+ HL CP
Sbjct: 265 AIWKCKAPLKVKFFAWLLVKDRLSTKKNLHKKTIVPND--ICDICNGATETASHLCFFCP 322
Query: 191 FFAELWEQL 199
F W+++
Sbjct: 323 FAKSFWDKI 331
>UniRef100_Q84Q64 Hypothetical protein OSJNBa0071M09.16 [Oryza sativa]
Length = 1143
Score = 75.5 bits (184), Expect = 1e-12
Identities = 69/277 (24%), Positives = 111/277 (39%), Gaps = 22/277 (7%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEG 63
F S + VGDG+ T FW W G ++ P +++++ ++ + LR + +
Sbjct: 857 FAASTKITVGDGNTTRFWDSAWINGQRPKDLMPLVYEISKNRKKS------LRQGKEDDS 910
Query: 64 WSWRRWLWAWEEIVLEECRTLLL------DVSLFTDISDSWIWLPNPIGGYSVRGAY--E 115
W L A I + L+ +V L ++ D +W G Y+ AY +
Sbjct: 911 WVHDLTLDAGSSITINLLDQLVRLWEAVRNVHLDSEEPDQIVWKFTSSGHYTASSAYHAQ 970
Query: 116 VLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTG 175
L A + LIW P K AW ++++R+ T LA+RG + C
Sbjct: 971 CLGAPNTNFNS----LIWKVWAPGKCKFHAWLIIQNRVWTSDRLATRG--WRNNGHCPLC 1024
Query: 176 CGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQ--FTCLSGFAKARRS 233
ET+ HL C + +W + W G + V + L+ +
Sbjct: 1025 RCDTETALHLVAMCRYTKRIWRLVATWAGYQQFEPAQWEEARSVHQWWESLANTPGIPKK 1084
Query: 234 FLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
L+ + LL W IW ERN R+F + LL KIK
Sbjct: 1085 GLRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIK 1121
>UniRef100_Q65WR1 Putative polyprotein [Oryza sativa]
Length = 1023
Score = 75.5 bits (184), Expect = 1e-12
Identities = 69/280 (24%), Positives = 111/280 (39%), Gaps = 28/280 (10%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEG 63
F S + VGDG+ T FW W G ++ P +++++ ++ + LR + +
Sbjct: 737 FAASTKITVGDGNTTRFWDSAWINGRRPKDLMPLVYEISKNRKKS------LRQGKEDDS 790
Query: 64 WSWRRWLWAWEEIVLEECRTLLL------DVSLFTDISDSWIWLPNPIGGYSVRGAY--E 115
W L A I + L+ +V L ++ D +W G Y+ AY +
Sbjct: 791 WVHDLTLDAGSSITINLLDQLVRLWEAVRNVHLDSEEPDQIVWKFTSSGHYTASSAYHAQ 850
Query: 116 VLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTG 175
L A + LIW P K AW ++++R+ T LA+RG + C
Sbjct: 851 CLGAPNTNFNS----LIWKVWAPGKCKFHAWLIIQNRVWTSDRLATRG--WRNNGHCPLC 904
Query: 176 CGSIETSDHLFLSCPFFAELWEQLRLWIGC-----CVGVDSNIISNHFVQFTCLSGFAKA 230
ET+ HL C + +W + W G ++ + + G K
Sbjct: 905 RCDTETALHLVAMCRYTKRIWRLVATWAGYQQLEPAQWEEARSVHQWWESLANTPGIPKK 964
Query: 231 RRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
L+ + LL W IW ERN R+F + LL KIK
Sbjct: 965 G---LRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIK 1001
>UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa]
Length = 593
Score = 74.7 bits (182), Expect = 2e-12
Identities = 68/218 (31%), Positives = 98/218 (44%), Gaps = 32/218 (14%)
Query: 11 KVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANML---PLRLSRGGEGWSWR 67
K+ DG FW W G L + +P L++L +K VA +L PL +S +R
Sbjct: 368 KLQDGLQIRFWEDCWLGNQSLEKIYPSLYNLVRNKNVVVAKVLERVPLNVS-------FR 420
Query: 68 RW-----LWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGN 122
R L AW EIV ++D++L D + W + G ++V Y+ L GN
Sbjct: 421 RAIIGDNLKAWHEIVAN-----VVDINLRGQ-KDRFTWDASKNGTFTVNSMYKKLMF-GN 473
Query: 123 SITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETS 182
I + IW ++PLK+ IF W L + T+ NLA R + +L C S ET
Sbjct: 474 VIPRQS--FIWKLRLPLKIKIFLWYLKEGVILTKDNLAKR---RWKGSLRCCFCRSHETI 528
Query: 183 DHLFLSCP---FFAELWEQLRLWIGCCVGVDSNIISNH 217
+LF CP F W RLW D N++ N+
Sbjct: 529 QYLFFDCPMVIFRGTHW--ARLWSLLLKEEDGNMLKNY 564
>UniRef100_Q7XKF8 OSJNBb0065J09.11 protein [Oryza sativa]
Length = 436
Score = 74.7 bits (182), Expect = 2e-12
Identities = 68/285 (23%), Positives = 114/285 (39%), Gaps = 38/285 (13%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEG 63
F S + VGDG+ T FW W G ++ P ++ ++ ++ + L +G E
Sbjct: 150 FAASTKITVGDGNMTRFWDSTWIDGRRPKDLMPLVYAISKNRKKS--------LHQGKED 201
Query: 64 WSWRRWLWAWEEIVLEECRTLLLD--------------VSLFTDISDSWIWLPNPIGGYS 109
+W ++++ + T+ ++ V L ++ D +W G Y+
Sbjct: 202 DTWV------DDLIFDAGSTITVNLVEQLVRLWEAVRNVQLVSEEPDQIVWKFTGNGHYT 255
Query: 110 VRGAY--EVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQL 167
AY + L A ++ LIW P K + W ++++R+ T LA RG
Sbjct: 256 ASSAYHAQCLEAPSTNLNS----LIWKAWAPGKCKFYVWLIIQNRVWTSDRLAIRG--WQ 309
Query: 168 DTALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFV--QFTCLS 225
+ C ET HL +C + +W + W+G S V + L+
Sbjct: 310 NNGHCPLCRCEAETGLHLVATCRYTKRIWHHVAGWVGYHQLEPSQWEEAQSVCQWWESLA 369
Query: 226 GFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+ L+ + LL W IW ERN R+F N LL KIK
Sbjct: 370 NTPNIPKKGLRSLILLVVWEIWKERNRRIFDNKEMAVGLLLAKIK 414
>UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa]
Length = 1189
Score = 74.7 bits (182), Expect = 2e-12
Identities = 69/280 (24%), Positives = 111/280 (39%), Gaps = 28/280 (10%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEG 63
F S + VGDG+ T FW W G ++ P +++++ ++ + LR + +
Sbjct: 903 FAASTKITVGDGNTTRFWDSAWINGRRPKDLMPLVYEISKNRKKS------LRQGKEDDS 956
Query: 64 WSWRRWLWAWEEIVLEECRTLLL------DVSLFTDISDSWIWLPNPIGGYSVRGAY--E 115
W L A I + L+ +V L ++ D +W G Y+ AY +
Sbjct: 957 WVHDLTLDAGLSITINLLDQLVRLWEAVRNVHLDSEEPDQIVWKFTSSGHYTASSAYHAQ 1016
Query: 116 VLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTG 175
L A + LIW P K AW ++++R+ T LA+RG + C
Sbjct: 1017 CLGAPNTNFNS----LIWKVWAPGKCKFHAWLIIQNRVWTSDRLATRG--WRNNGHCPLC 1070
Query: 176 CGSIETSDHLFLSCPFFAELWEQLRLWIGC-----CVGVDSNIISNHFVQFTCLSGFAKA 230
ET+ HL C + +W + W G ++ + + G K
Sbjct: 1071 RCDTETAFHLVAMCRYTKRIWRLVATWAGYQQLEPAQWEEARSVHQWWESLANTPGIPKK 1130
Query: 231 RRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
L+ + LL W IW ERN R+F + LL KIK
Sbjct: 1131 G---LRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIK 1167
>UniRef100_Q5ZCK2 Rust resistance protein Rp1-dp8-like [Oryza sativa]
Length = 1374
Score = 74.3 bits (181), Expect = 3e-12
Identities = 69/279 (24%), Positives = 114/279 (40%), Gaps = 45/279 (16%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWL- 70
+G+G+ FW W +++ P ++ ++ K +T+ + L +WL
Sbjct: 1116 IGNGATVSFWESAWWQERRMKDVAPLVYAVSKKKNSTLQHALQSD-----------QWLL 1164
Query: 71 ---------WAWEEI-VLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAY--EVLT 118
W E I L T + + L D W G Y+ AY ++L
Sbjct: 1165 DLGFPADVGWTTELIDQLVTVWTAIQTIELTEHEDDQISWKLTSHGQYTATSAYNAQLLG 1224
Query: 119 ATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGS 178
T N+ LIW K FAW ++++R+ T L++RG + ++C
Sbjct: 1225 TTANN-------LIWKPWALRKCKTFAWLVIQNRVWTSDRLSTRG--WPNGSICPLCRRV 1275
Query: 179 IETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCL-------SGFAKAR 231
ET+ HL C + +W L W C +NI NH+ Q + + + +A
Sbjct: 1276 QETALHLLAGCRYTKRIWATLAEWTAC-----NNIHPNHWWQVSSVLEWWDQTTSVKEAP 1330
Query: 232 RSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+ L+ + L W IW+ERN R F++ LL KIK
Sbjct: 1331 KKALRSLTFLVVWEIWNERNRRTFQHKELPPSGLLTKIK 1369
>UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 72.4 bits (176), Expect = 1e-11
Identities = 73/263 (27%), Positives = 107/263 (39%), Gaps = 24/263 (9%)
Query: 15 GSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWE 74
G +FW W G LR + P LF K+ + A +L G E ++ R
Sbjct: 1737 GESIVFWQDKW-GMQKLRLEMPELFSFVQHKFLSAAEVL------GQEDFT-RLLQLPIS 1788
Query: 75 EIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVDLIWH 134
E + + L + + W +S AY L G+ IW
Sbjct: 1789 ETAFHQLQQLTTRIDNLERTQEQDQWKYKWGNNFSATKAYSELM--GHQQIHEVHKWIWK 1846
Query: 135 HQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGC-GSIETSDHLFLSCPFFA 193
K +F W L++DRLST N+ R +QL + CV S ET+ HLFL C +
Sbjct: 1847 SFCQPKHKVFFWLLIKDRLST-CNILKRKNMQLQSYNCVLCLQNSEETTSHLFLHCAYAR 1905
Query: 194 ELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNR 253
+ W+ L + I D I++HF GF F + +L W IWS RNN
Sbjct: 1906 QCWQLLDMDIP--PNSDFPEITDHFKH-----GF---NNQFFMVTTILLCWAIWSTRNNL 1955
Query: 254 LFKNV--VTEAPRLLDKIKLLSL 274
+F+ + E + + K +LL L
Sbjct: 1956 IFRGIQPTIEGTKEIFKRELLLL 1978
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 72.0 bits (175), Expect = 2e-11
Identities = 70/275 (25%), Positives = 113/275 (40%), Gaps = 33/275 (12%)
Query: 8 VRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANML-PLRLSRGGEGWSW 66
++R +G+G T W W R P ++ + V++++ PL
Sbjct: 874 LKRLIGNGEQTNVWIDKWLFDGHSRR--PMNLHSLMNIHMKVSHLIDPLT---------- 921
Query: 67 RRW-LWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATG--NS 123
R W L E+ E+ L++ DS+ W G Y+V+ YE + N
Sbjct: 922 RNWNLKKLTELFHEKDVQLIMHQRPLISSEDSYCWAGTNNGLYTVKSGYERSSRETFKNL 981
Query: 124 ITDSAV--------DLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTG 175
++ V D +W + K+ +F W+ ++ L+ L SRGI D C+
Sbjct: 982 FKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTADG--CLFC 1039
Query: 176 CGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIIS--NHFVQFTCLSGFAKARRS 233
IET +HL CPF ++W L L G ++I S NH +Q + G + R+
Sbjct: 1040 KEEIETINHLLFQCPFARQVW-ALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPRHMRT 1098
Query: 234 FLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDK 268
WLL W IW RN LF+ + ++ K
Sbjct: 1099 VSP--WLL--WEIWKNRNKTLFQGTGLTSSEIVAK 1129
>UniRef100_Q8W2Y2 Putative reverse transcriptase [Oryza sativa]
Length = 581
Score = 71.6 bits (174), Expect = 2e-11
Identities = 70/271 (25%), Positives = 117/271 (42%), Gaps = 29/271 (10%)
Query: 12 VGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLP-------LRLSRGGEGW 64
VGDG FW W + P +F + K TV + L + +SR E
Sbjct: 304 VGDGKIARFWGSKWLDNQTPQLIAPDIFVASRRKNRTVHDALAENKWIRDIDVSRIFEAN 363
Query: 65 SWRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEV--LTATGN 122
++++ W ++ D+ + DS W G YS + AY + L A +
Sbjct: 364 HLQQYIRLWR---------MIRDMRPLRENHDSIRWNLTANGQYSAKSAYRLQFLGAAKS 414
Query: 123 SITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETS 182
+ S IW P K F+W + ++R+ T L RG + +C C +++ S
Sbjct: 415 TFYQS----IWKAWAPPKCKFFSWLVTQNRVWTADRLEKRG--WQNQKICPL-CYNVDES 467
Query: 183 DHLFLS-CPFFAELWEQLRLWIGCCVGVDSNIISNHFVQ--FTCLSGFAKARRSFLQLIW 239
L L+ C +W +++ W G + +D N S H V+ +T ++ L+ +
Sbjct: 468 ALLLLAKCKLTKRIWAEMQRWTGTDLQMD-NSESCHSVEQWWTLVTESPNTPCKPLRTLV 526
Query: 240 LLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+L +W IW ERN R+F+N + +L KIK
Sbjct: 527 ILISWEIWCERNARIFRNNASTPSSILAKIK 557
>UniRef100_Q6I5S8 Putative polyprotein [Oryza sativa]
Length = 1419
Score = 71.6 bits (174), Expect = 2e-11
Identities = 53/178 (29%), Positives = 86/178 (47%), Gaps = 11/178 (6%)
Query: 96 DSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLST 155
D+ +W + G Y+ + AY V+ G + +DS +L+W P K FAW L ++R+ T
Sbjct: 253 DTILWRWSASGLYTAKSAY-VMQFMGKTFSDST-ELVWETWAPGKHKFFAWLLAQNRIWT 310
Query: 156 RSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELWEQL----RLWIGCCVGVDS 211
+L RG + C ++ET HLF+ CPF ++W + R V D+
Sbjct: 311 ADHLQCRG--WPNNYFCQLCFRNLETVQHLFMECPFTQQVWRDIGRKFRPQRFLHVTTDN 368
Query: 212 NIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKI 269
++ N + + T K ++ L+ L T W IW ERN+R+F+ + L KI
Sbjct: 369 TLL-NWWREQT--DEAEKLQQKGLRSAVLTTLWEIWMERNDRIFRGKESTPSALASKI 423
>UniRef100_Q7Y1F8 Putative reverse transcriptase [Oryza sativa]
Length = 1014
Score = 69.3 bits (168), Expect = 1e-10
Identities = 71/271 (26%), Positives = 107/271 (39%), Gaps = 49/271 (18%)
Query: 4 FGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEG 63
F S +GDG LFW W + L+ P LF A K TVA L
Sbjct: 780 FAASTSVTIGDGRKALFWESPWACSSTLKAMAPNLFHHARKKKRTVAEAL---------- 829
Query: 64 WSWRRWLWAWEEIVLEECRTLLLDV-SLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGN 122
+W+ ++I L+ D +F I S I L I ++R + T +G
Sbjct: 830 -QDNKWI---DDIRYNLTTVLVTDFFKVFEIIWTSGITLTQDIED-NIRWKW---TTSGV 881
Query: 123 SITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETS 182
T SA +IW DRL R + C ++ET+
Sbjct: 882 YTTKSAYQIIW---------------TADRLQQRE--------WPNNYFCQLCFRNLETA 918
Query: 183 DHLFLSCPFFAELWEQLRLWIGCCVGVDSN-----IISNHFVQFTCLSGFAKARRSFLQL 237
HLF CP ++W+ + +G N I+ + + Q +S K +R +
Sbjct: 919 HHLFYECPITRKIWDAIATRLGMPQLRPQNWSKTTILEDRWTQ--TVSTMPKEKRKGAKS 976
Query: 238 IWLLTTWVIWSERNNRLFKNVVTEAPRLLDK 268
I +L +W IW ERNN +FKN+ T +++K
Sbjct: 977 ILILLSWEIWKERNNMVFKNIETATQGIVEK 1007
>UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa]
Length = 855
Score = 68.9 bits (167), Expect = 1e-10
Identities = 54/191 (28%), Positives = 82/191 (42%), Gaps = 25/191 (13%)
Query: 100 WLPNPIGGYSVRGAYEVLTATG-------------NSITDSAVDL--IWHHQVPLKVSIF 144
W +G YSVR AY ++ + + + +S D +W P K+ I
Sbjct: 492 WPHEKLGLYSVRSAYNMVRSEAFFADQSNSGRGMASRLLESQKDWKGLWKINAPGKMKII 551
Query: 145 AWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIG 204
WR + L+T L R I D CV C +T +H+FL CPF A++WE+++
Sbjct: 552 LWRAAHECLATGFQLPRRHIPSTDG--CVF-CNRDDTVEHVFLFCPFAAQIWEEIKRKCA 608
Query: 205 CCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPR 264
+G + H++ F K S + +T W IW RNN N R
Sbjct: 609 VKLGRNGFSTMRHWI-----FDFLKRGSSHTNTLLAVTFWHIWEARNNTKNNNGTVHPQR 663
Query: 265 LLDKIKLLSLV 275
++ IK+LS V
Sbjct: 664 VV--IKILSYV 672
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.140 0.483
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 498,020,370
Number of Sequences: 2790947
Number of extensions: 20052034
Number of successful extensions: 52316
Number of sequences better than 10.0: 214
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 140
Number of HSP's that attempted gapping in prelim test: 51909
Number of HSP's gapped (non-prelim): 251
length of query: 275
length of database: 848,049,833
effective HSP length: 125
effective length of query: 150
effective length of database: 499,181,458
effective search space: 74877218700
effective search space used: 74877218700
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC130801.11


