

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
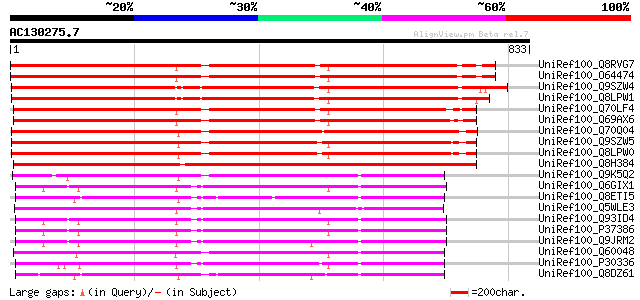
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RVG7 Putative heavy metal transporter [Arabidopsis t... 779 0.0
UniRef100_O64474 Potential cadmium/zinc-transporting ATPase 2 [A... 779 0.0
UniRef100_Q9SZW4 Potential cadmium/zinc-transporting ATPase 3 [A... 766 0.0
UniRef100_Q8LPW1 Putative heavy-metal transporter [Arabidopsis t... 765 0.0
UniRef100_Q70LF4 Putative heavy metal transporting P-type ATPase... 761 0.0
UniRef100_Q69AX6 P1B-type heavy metal transporting ATPase [Thlas... 756 0.0
UniRef100_Q70Q04 Putative cadmium/zinc-transporting ATPase 3 [Ar... 742 0.0
UniRef100_Q9SZW5 Potential cadmium/zinc-transporting ATPase 4 [A... 741 0.0
UniRef100_Q8LPW0 Putative heavy metal transporter HMA3 [Arabidop... 735 0.0
UniRef100_Q8H384 Putative potential cadmium/zinc-transporting AT... 638 0.0
UniRef100_Q9K5Q2 Cadmium-transporting ATPase [Bacillus halodurans] 343 1e-92
UniRef100_Q6GIX1 Probable cadmium-transporting ATPase [Staphyloc... 341 5e-92
UniRef100_Q8ETI5 Cadmium-transporting ATPase [Oceanobacillus ihe... 340 1e-91
UniRef100_Q5WLE3 Cadmium-transporting ATPase [Bacillus clausii] 335 3e-90
UniRef100_Q93ID4 Cadmium resistance protein B [Staphylococcus au... 334 6e-90
UniRef100_P37386 Probable cadmium-transporting ATPase [Staphyloc... 334 6e-90
UniRef100_Q9JRM2 CadA protein [Xanthomonas maltophilia] 334 7e-90
UniRef100_Q60048 Probable cadmium-transporting ATPase [Listeria ... 334 7e-90
UniRef100_P30336 Probable cadmium-transporting ATPase [Bacillus ... 333 1e-89
UniRef100_Q8DZ61 Cation-transporting ATPase, E1-E2 family [Strep... 332 2e-89
>UniRef100_Q8RVG7 Putative heavy metal transporter [Arabidopsis thaliana]
Length = 1172
Score = 779 bits (2012), Expect = 0.0
Identities = 414/796 (52%), Positives = 546/796 (68%), Gaps = 49/796 (6%)
Query: 2 VENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIA 61
V+ +++S F+VLG+CC +E ++E ILK L GVK SVIVP+RTV VVHD LLIS +IA
Sbjct: 13 VKKLQKSYFDVLGICCTSEVPIIENILKSLDGVKEYSVIVPSRTVIVVHDSLLISPFQIA 72
Query: 62 DALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGF 121
ALN ARLEA+ R GET+ + K P + GLLL LSFLK++Y PL WLA+ +VA G
Sbjct: 73 KALNEARLEANVRVNGETSFKNKWPSPFAVVSGLLLLLSFLKFVYSPLRWLAVAAVAAGI 132
Query: 122 PKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKAL 181
+L +A ASI+ ++INILV++ V T A+QDF + ++FLF+I+ WLETRA++KA
Sbjct: 133 YPILAKAFASIKRPRIDINILVIITVIATLAMQDFMEAAAVVFLFTISDWLETRASYKAT 192
Query: 182 VAMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKML 241
M SL ++APQ AI+AETGE V+V++VK++T++AVKAG+ IP+DGIVV+G CEVDEK L
Sbjct: 193 SVMQSLMSLAPQKAIIAETGEEVEVDEVKVDTVVAVKAGETIPIDGIVVDGNCEVDEKTL 252
Query: 242 TGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNV 296
TGE+FPV K+ DS VWAGTIN+N Y + D ++ K + + S + +
Sbjct: 253 TGEAFPVPKQRDSTVWAGTINLNGYICVKTTSLAGDCVVAKMAKLVEEAQSSKTKSQRLI 312
Query: 297 KTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALI 356
C + + +I ++ +C+A+VP + V +++ WFHLALVVL+SGCPC LI
Sbjct: 313 DKCSQYYTPAII----------LVSACVAIVPVIMKVHNLKHWFHLALVVLVSGCPCGLI 362
Query: 357 LSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDD 415
LSTPVA FCALTKAA SGLL+K DYL+TLS+IK VAFDKTGTITRGEF V DF S D
Sbjct: 363 LSTPVATFCALTKAATSGLLIKSADYLDTLSKIKIVAFDKTGTITRGEFIVIDFKSLSRD 422
Query: 416 INNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRD 475
IN +LLYW+SS+ESKSSHP+A +VDYA+ S++P PE VE++QNFPGEGI+G IDG D
Sbjct: 423 INLRSLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRPEEVEDYQNFPGEGIYGKIDGND 482
Query: 476 IYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGAL 529
I+IGNK++ RA C E+ TK E L G F+L DACRSG
Sbjct: 483 IFIGNKKIASRAGCS------TVPEIEVDTKGGKTVGYVYVGERLAGFFNLSDACRSGVS 536
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+AM ELK LG+++ MLTGD+ AM+ Q QL N +D+VH +LLP +K+++I+ FKKEGP
Sbjct: 537 QAMAELKSLGIKTAMLTGDNQAAAMHAQEQLGNVLDVVHGDLLPEDKSRIIQEFKKEGPT 596
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
AM+GDG+NDAPALATADIGISMGISGSALA +T N ILMSNDIR++P+A++LAR+ RK+
Sbjct: 597 AMVGDGVNDAPALATADIGISMGISGSALATQTGNIILMSNDIRRIPQAVKLARRARRKV 656
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTK 709
VENV +S+ K ILALA AG+PL+W AVL DVGTCLLVI NSML+L+E K ++ +
Sbjct: 657 VENVCLSIILKAGILALAFAGHPLIWAAVLVDVGTCLLVIFNSMLLLREKKKIGNKKCYR 716
Query: 710 GSK---YGKFLEDKTKPLLNKQSNIDEEKGLLS---GEECGKDCCKNATHYVETSKERND 763
S G+ LE + +D E GLL+ +C CC ++N
Sbjct: 717 ASTSKLNGRKLEG------DDDYVVDLEAGLLTKSGNGQCKSSCC---------GDKKNQ 761
Query: 764 ESCGVSKLSLLNGNDH 779
E+ + K S +DH
Sbjct: 762 ENVVMMKPSSKTSSDH 777
>UniRef100_O64474 Potential cadmium/zinc-transporting ATPase 2 [Arabidopsis thaliana]
Length = 1172
Score = 779 bits (2012), Expect = 0.0
Identities = 414/796 (52%), Positives = 546/796 (68%), Gaps = 49/796 (6%)
Query: 2 VENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIA 61
V+ +++S F+VLG+CC +E ++E ILK L GVK SVIVP+RTV VVHD LLIS +IA
Sbjct: 13 VKKLQKSYFDVLGICCTSEVPIIENILKSLDGVKEYSVIVPSRTVIVVHDSLLISPFQIA 72
Query: 62 DALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGF 121
ALN ARLEA+ R GET+ + K P + GLLL LSFLK++Y PL WLA+ +VA G
Sbjct: 73 KALNEARLEANVRVNGETSFKNKWPSPFAVVSGLLLLLSFLKFVYSPLRWLAVAAVAAGI 132
Query: 122 PKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKAL 181
+L +A ASI+ ++INILV++ V T A+QDF + ++FLF+I+ WLETRA++KA
Sbjct: 133 YPILAKAFASIKRPRIDINILVIITVIATLAMQDFMEAAAVVFLFTISDWLETRASYKAT 192
Query: 182 VAMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKML 241
M SL ++APQ AI+AETGE V+V++VK++T++AVKAG+ IP+DGIVV+G CEVDEK L
Sbjct: 193 SVMQSLMSLAPQKAIIAETGEEVEVDEVKVDTVVAVKAGETIPIDGIVVDGNCEVDEKTL 252
Query: 242 TGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNV 296
TGE+FPV K+ DS VWAGTIN+N Y + D ++ K + + S + +
Sbjct: 253 TGEAFPVPKQRDSTVWAGTINLNGYICVKTTSLAGDCVVAKMAKLVEEAQSSKTKSQRLI 312
Query: 297 KTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALI 356
C + + +I ++ +C+A+VP + V +++ WFHLALVVL+SGCPC LI
Sbjct: 313 DKCSQYYTPAII----------LVSACVAIVPVIMKVHNLKHWFHLALVVLVSGCPCGLI 362
Query: 357 LSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDD 415
LSTPVA FCALTKAA SGLL+K DYL+TLS+IK VAFDKTGTITRGEF V DF S D
Sbjct: 363 LSTPVATFCALTKAATSGLLIKSADYLDTLSKIKIVAFDKTGTITRGEFIVIDFKSLSRD 422
Query: 416 INNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRD 475
IN +LLYW+SS+ESKSSHP+A +VDYA+ S++P PE VE++QNFPGEGI+G IDG D
Sbjct: 423 INLRSLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRPEEVEDYQNFPGEGIYGKIDGND 482
Query: 476 IYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGAL 529
I+IGNK++ RA C E+ TK E L G F+L DACRSG
Sbjct: 483 IFIGNKKIASRAGCS------TVPEIEVDTKGGKTVGYVYVGERLAGFFNLSDACRSGVS 536
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+AM ELK LG+++ MLTGD+ AM+ Q QL N +D+VH +LLP +K+++I+ FKKEGP
Sbjct: 537 QAMAELKSLGIKTAMLTGDNQAAAMHAQEQLGNVLDVVHGDLLPEDKSRIIQEFKKEGPT 596
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
AM+GDG+NDAPALATADIGISMGISGSALA +T N ILMSNDIR++P+A++LAR+ RK+
Sbjct: 597 AMVGDGVNDAPALATADIGISMGISGSALATQTGNIILMSNDIRRIPQAVKLARRARRKV 656
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTK 709
VENV +S+ K ILALA AG+PL+W AVL DVGTCLLVI NSML+L+E K ++ +
Sbjct: 657 VENVCLSIILKAGILALAFAGHPLIWAAVLVDVGTCLLVIFNSMLLLREKKKIGNKKCYR 716
Query: 710 GSK---YGKFLEDKTKPLLNKQSNIDEEKGLLS---GEECGKDCCKNATHYVETSKERND 763
S G+ LE + +D E GLL+ +C CC ++N
Sbjct: 717 ASTSKLNGRKLEG------DDDYVVDLEAGLLTKSGNGQCKSSCC---------GDKKNQ 761
Query: 764 ESCGVSKLSLLNGNDH 779
E+ + K S +DH
Sbjct: 762 ENVVMMKPSSKTSSDH 777
>UniRef100_Q9SZW4 Potential cadmium/zinc-transporting ATPase 3 [Arabidopsis thaliana]
Length = 951
Score = 766 bits (1979), Expect = 0.0
Identities = 420/829 (50%), Positives = 555/829 (66%), Gaps = 47/829 (5%)
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ + +S F+VLG+CC +E L+E IL + GVK SVIVP+RTV VVHD L++S+ +I
Sbjct: 4 KKMTKSYFDVLGICCTSEVPLIENILNSMDGVKEFSVIVPSRTVIVVHDTLILSQFQIVK 63
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN A+LEA+ R GETN + K P + G+LL LSF KY+Y P WLA+ +V G
Sbjct: 64 ALNQAQLEANVRVTGETNFKNKWPSPFAVVSGILLLLSFFKYLYSPFRWLAVAAVVAGIY 123
Query: 123 KVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALV 182
+L +A+AS+ ++INILV++ V T +QD+T+ V++FLF+IA+WL++RA++KA
Sbjct: 124 PILAKAVASLARFRIDINILVVVTVGATIGMQDYTEAAVVVFLFTIAEWLQSRASYKASA 183
Query: 183 AMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLT 242
M SL ++APQ A++AETGE V+V+++K NT++AVKAG+ IP+DG+VV+G CEVDEK LT
Sbjct: 184 VMQSLMSLAPQKAVIAETGEEVEVDELKTNTVIAVKAGETIPIDGVVVDGNCEVDEKTLT 243
Query: 243 GESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RS 302
GE+FPV K DS VWAGTIN+N Y I +T C + K +N KT +
Sbjct: 244 GEAFPVPKLKDSTVWAGTINLNGY--ITVNTTAL-AEDCVVAKMAKLVEEAQNSKTETQR 300
Query: 303 F*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVA 362
F K +K + ++ C +P AL V +++ W HLALVVL+S CPC LILSTPVA
Sbjct: 301 FIDK--CSKYYTPAIILISICFVAIPFALKVHNLKHWVHLALVVLVSACPCGLILSTPVA 358
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNETL 421
FCALTKAA SGLL+KG DYLETL++IK VAFDKTGTITRGEF V DF S +DI+ ++L
Sbjct: 359 TFCALTKAATSGLLIKGADYLETLAKIKIVAFDKTGTITRGEFIVMDFQSLSEDISLQSL 418
Query: 422 LYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNK 481
LYW+SS ESKSSHP+A A+VDYAR S++P PE VE++QNFPGEGI+G IDG+++YIGNK
Sbjct: 419 LYWVSSTESKSSHPMAAAVVDYARSVSVEPKPEAVEDYQNFPGEGIYGKIDGKEVYIGNK 478
Query: 482 RVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAMEEL 535
R+ RA C + ++ TK ETL GVF+L DACRSG +AM+EL
Sbjct: 479 RIASRAGC------LSVPDIDVDTKGGKTIGYVYVGETLAGVFNLSDACRSGVAQAMKEL 532
Query: 536 KLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK-EGPIAMIGD 594
K LG++ MLTGD+ AM+ Q QL NA+DIV AELLP +K+++I+ K+ EGP AM+GD
Sbjct: 533 KSLGIKIAMLTGDNHAAAMHAQEQLGNAMDIVRAELLPEDKSEIIKQLKREEGPTAMVGD 592
Query: 595 GINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVI 654
G+NDAPALATADIGISMG+SGSALA ET N ILMSNDIR++P+AI+LA++ RK+VENV+
Sbjct: 593 GLNDAPALATADIGISMGVSGSALATETGNIILMSNDIRRIPQAIKLAKRAKRKVVENVV 652
Query: 655 ISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYG 714
IS+ K AILALA AG+PL+W AVL DVGTCLLVILNSML+L + HK + + S
Sbjct: 653 ISITMKGAILALAFAGHPLIWAAVLADVGTCLLVILNSMLLLSDKHKTGNKCYRESSSSS 712
Query: 715 KFLEDKTKPLLNKQSNIDEEKGLL---SGEECGKDCCKNATH-----------------Y 754
+ +K L + D E GLL S + C CC T
Sbjct: 713 VLIAEK----LEGDAAGDMEAGLLPKISDKHCKPGCCGTKTQEKAMKPAKASSDHSHSGC 768
Query: 755 VETSKERN----DESCGVSKLSLLNGNDHGMFMFVEVHVVKHCVMIQDS 799
ET ++ N +SC + L +G+D G +H V +Q S
Sbjct: 769 CETKQKDNVTVVKKSCCAEPVDLGHGHDSGCCGDKSQQPHQHEVQVQQS 817
>UniRef100_Q8LPW1 Putative heavy-metal transporter [Arabidopsis thaliana]
Length = 951
Score = 765 bits (1975), Expect = 0.0
Identities = 410/784 (52%), Positives = 542/784 (68%), Gaps = 31/784 (3%)
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ + +S F+VLG+CC +E L+E IL + GVK SVIVP+RTV VVHD L++S+ +I
Sbjct: 4 KKMTKSYFDVLGICCTSEVPLIENILNSMDGVKEFSVIVPSRTVIVVHDTLILSQFQIVK 63
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN A+LEA+ R GETN + K P + G+LL LSF KY+Y P WLA+ +V G
Sbjct: 64 ALNQAQLEANVRVTGETNFKNKWPSPFAVVSGILLLLSFFKYLYSPFRWLAVAAVVAGIY 123
Query: 123 KVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALV 182
+L +A+AS+ ++INILV++ V T +QD+T+ V++FLF+IA+WL++RA++KA
Sbjct: 124 PILAKAVASLARFRIDINILVVVTVGATIGMQDYTEAAVVVFLFTIAEWLQSRASYKASA 183
Query: 183 AMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLT 242
M SL ++APQ A++AETGE V+V+++K NT++AVKAG+ IP+DG+VV+G CEVDEK LT
Sbjct: 184 VMQSLMSLAPQKAVIAETGEEVEVDELKTNTVIAVKAGETIPIDGVVVDGNCEVDEKTLT 243
Query: 243 GESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RS 302
GE+FPV K DS VWAGTIN+N Y I +T C + K +N KT +
Sbjct: 244 GEAFPVPKLKDSTVWAGTINLNGY--ITVNTTAL-AEDCVVAKMAKLVEEAQNSKTETQR 300
Query: 303 F*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVA 362
F K +K + ++ C +P AL V +++ W HLALVVL+S CPC LILSTPVA
Sbjct: 301 FIDK--CSKYYTPAIILISICFVAIPFALKVHNLKHWVHLALVVLVSACPCGLILSTPVA 358
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNETL 421
FCALTKAA SGLL+KG DYLETL++IK VAFDKTGTITRGEF V DF S +DI+ ++L
Sbjct: 359 TFCALTKAATSGLLIKGADYLETLAKIKIVAFDKTGTITRGEFIVMDFQSLSEDISLQSL 418
Query: 422 LYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNK 481
LYW+SS ESKSSHP+A A+VDYAR S++P PE VE++QN PGEGI+G IDG+++YIGNK
Sbjct: 419 LYWVSSTESKSSHPMAAAVVDYARSVSVEPKPEAVEDYQNLPGEGIYGKIDGKEVYIGNK 478
Query: 482 RVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAMEEL 535
R+ RA C + ++ TK ETL GVF+L DACRSG +AM+EL
Sbjct: 479 RIASRAGC------LSVPDIDVDTKGGKTIGYVYVGETLAGVFNLSDACRSGVAQAMKEL 532
Query: 536 KLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK-EGPIAMIGD 594
K LG++ MLTGD+ AM+ Q QL NA+DIV AELLP +K+++I+ K+ EGP AM+GD
Sbjct: 533 KSLGIKIAMLTGDNHAAAMHAQEQLGNAMDIVRAELLPEDKSEIIKQLKREEGPTAMVGD 592
Query: 595 GINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVI 654
G+NDAPALATADIGISMG+SGSALA ET N ILMSNDIR++P+AI+LA++ RK+VENV+
Sbjct: 593 GLNDAPALATADIGISMGVSGSALATETGNIILMSNDIRRIPQAIKLAKRAKRKVVENVV 652
Query: 655 ISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYG 714
IS+ K AILALA AG+PL+W AVL DVGTCLLVILNSML+L + HK + + S
Sbjct: 653 ISITMKGAILALAFAGHPLIWAAVLADVGTCLLVILNSMLLLSDKHKTGNKCYRESSSSS 712
Query: 715 KFLEDKTKPLLNKQSNIDEEKGLL---SGEECGKDCC-----KNATHYVETSKERNDESC 766
+ +K L + D E GLL S + C CC + A +TS + + C
Sbjct: 713 VLIAEK----LEGDAAGDMEAGLLPKISDKHCKPGCCGTKTQEKAMKPAKTSSDHSHSGC 768
Query: 767 GVSK 770
+K
Sbjct: 769 CETK 772
>UniRef100_Q70LF4 Putative heavy metal transporting P-type ATPase [Thlaspi
caerulescens]
Length = 1186
Score = 761 bits (1966), Expect = 0.0
Identities = 401/755 (53%), Positives = 523/755 (69%), Gaps = 39/755 (5%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN 65
++S F+VLG+CC +E L+E ILK L GVK +VIVP+RTV VVHD LLIS +IA ALN
Sbjct: 21 QKSYFDVLGICCTSEIPLIENILKSLDGVKEYTVIVPSRTVIVVHDSLLISPFQIAKALN 80
Query: 66 TARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPKVL 125
ARLEA+ + GET+ + K P + G+ L LSFLK++YPPL WLA+ VA G +L
Sbjct: 81 QARLEANVKVNGETSFKNKWPSPFAVVSGIFLLLSFLKFVYPPLRWLAVVGVAAGIYPIL 140
Query: 126 FRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMS 185
+A+ASIR L ++INILV++ V T A+QD+ + ++FLF+IA WLETR ++KA M
Sbjct: 141 AKAVASIRRLRVDINILVIITVAATLAMQDYMEAAAVVFLFTIADWLETRTSYKANSVMQ 200
Query: 186 SLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGES 245
SL ++APQ A++AETGE V+V++V++NTI+AVKAG+ IP+DGIVV+G CEVDEK LTGE+
Sbjct: 201 SLMSLAPQKAVIAETGEEVEVDEVQLNTIIAVKAGETIPIDGIVVDGNCEVDEKTLTGEA 260
Query: 246 FPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC* 300
FPV K+ DS V AGT+N+N Y + D ++ K + G + S + + C
Sbjct: 261 FPVPKQRDSTVLAGTMNLNGYISVNTTALASDCVVAKMAKLVEEAQGSKTKSQRLIDKCS 320
Query: 301 RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTP 360
+ + +I ++ + A+VPA + V ++ WFHLALVVL+S CP LILSTP
Sbjct: 321 QYYTPAII----------IISAGFAIVPAIMKVHNLNHWFHLALVVLVSACPSGLILSTP 370
Query: 361 VAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNE 419
VA FCALTKAA SGLL+K DYL+TLS+IK AFDKTGTITRGEF V +F S DI+
Sbjct: 371 VATFCALTKAATSGLLIKSADYLDTLSKIKIAAFDKTGTITRGEFIVIEFKSLSRDISLR 430
Query: 420 TLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIG 479
+LLYW+SS+ESKSSHP+A +VDYA+ S++P E VE++QNFPGEGI+G IDG ++YIG
Sbjct: 431 SLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRSEEVEDYQNFPGEGIYGKIDGNNVYIG 490
Query: 480 NKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAME 533
NKR+ RA C E+ TKK E L GVF+L DACRSG +AM+
Sbjct: 491 NKRIASRAGCS------TVPEIEVDTKKGKTVGYVYVGERLAGVFNLSDACRSGVAQAMK 544
Query: 534 ELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIAMIG 593
ELK LG+++ MLTGD+ AM Q QL NA+D+VH ELLP +K+K+I+ FKKEGP M+G
Sbjct: 545 ELKDLGIKTAMLTGDNQDSAMQAQEQLGNALDVVHGELLPEDKSKIIQEFKKEGPTCMVG 604
Query: 594 DGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENV 653
DG+NDAPALA ADIGISMGISGSAL +T + ILMSNDIR++P+AI+LAR+ RK+++NV
Sbjct: 605 DGVNDAPALANADIGISMGISGSALTTQTGHIILMSNDIRRIPQAIKLARRAQRKVLQNV 664
Query: 654 IISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKY 713
IS+ K IL LA AG+PL+W AVLTDVGTCL+VILNSML+L+E K SK
Sbjct: 665 FISITLKVGILVLAFAGHPLIWAAVLTDVGTCLIVILNSMLLLREKDK---------SKI 715
Query: 714 GKFLEDKTKPLLNKQSNIDEEKGLLSGEECGKDCC 748
K K + +D E GLLS +C CC
Sbjct: 716 KKCYRKKVEG--GDDQGLDLEAGLLSKSQCNSGCC 748
>UniRef100_Q69AX6 P1B-type heavy metal transporting ATPase [Thlaspi caerulescens]
Length = 1186
Score = 756 bits (1951), Expect = 0.0
Identities = 398/755 (52%), Positives = 524/755 (68%), Gaps = 39/755 (5%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN 65
++S F+VLG+CC +E L+E ILK L GVK +VIVP+RTV VVHD LLI +IA ALN
Sbjct: 21 QKSYFDVLGICCTSEIPLIENILKSLDGVKEYTVIVPSRTVIVVHDSLLIPPFQIAKALN 80
Query: 66 TARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPKVL 125
ARLEA+ + GET+ + K P + G+ L SFLK++YPPL WLA+ VA G +L
Sbjct: 81 QARLEANVKVNGETSFKNKWPSPFAVVSGIFLLPSFLKFVYPPLRWLAVVGVAAGIYPIL 140
Query: 126 FRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMS 185
+A+ASIR L ++INIL+++ V T A+QD+ + ++FLF+ A WLETRA++KA M
Sbjct: 141 AKAVASIRRLRVDINILIIITVAATLAMQDYMEAAAVVFLFTTADWLETRASYKANSVMQ 200
Query: 186 SLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGES 245
SL ++APQ A++AETGE V+V++V++NTI+AVKAG+ IP+DGIVV+G CEVDEK LTGE+
Sbjct: 201 SLMSLAPQKAVIAETGEEVEVDEVQLNTIIAVKAGETIPIDGIVVDGNCEVDEKTLTGEA 260
Query: 246 FPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC* 300
FPV K+ DS V AGT+N+N Y + D ++ K + G + S + + C
Sbjct: 261 FPVPKQRDSTVLAGTMNLNGYISVNTTALASDCVVAKMAKLVEEAQGSKTKSQRLIDKCS 320
Query: 301 RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTP 360
+ + +I ++ + A+VPA + V ++ WFHLALVVL+S CPC LILSTP
Sbjct: 321 QYYTPAII----------IISAGFAIVPAIMKVHNLNHWFHLALVVLVSACPCGLILSTP 370
Query: 361 VAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNE 419
VA FCALTKAA SGLL+K +L+TLS+IK AFDKTGTITRGEF V +F S DI+
Sbjct: 371 VATFCALTKAATSGLLIKSAGHLDTLSKIKIAAFDKTGTITRGEFIVIEFKSLSRDISLR 430
Query: 420 TLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIG 479
+LLYW+SS+ESKSSHP+A +VDYA+ S++P E VE++QNFPGEGI+G IDG ++YIG
Sbjct: 431 SLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRSEEVEDYQNFPGEGIYGKIDGNNVYIG 490
Query: 480 NKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAME 533
NKR+ RA C E+ TKK E L GVF+L DACRSG +AM+
Sbjct: 491 NKRIASRAGCS------TVPEIEVDTKKGKTVGYVYVGERLAGVFNLSDACRSGVAQAMK 544
Query: 534 ELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIAMIG 593
LK LG+++ MLTGD+ AM Q QL NA+D+VH ELLP +K+K+I+ FKKEGP M+G
Sbjct: 545 GLKDLGIKTAMLTGDNQDSAMQAQEQLGNALDVVHGELLPEDKSKIIQEFKKEGPTCMVG 604
Query: 594 DGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENV 653
DG+NDAPALA ADIGISMGISGSALA +T + ILMSNDIR++P+AI+LAR+ RK+++NV
Sbjct: 605 DGVNDAPALANADIGISMGISGSALATQTGHIILMSNDIRRIPQAIKLARRAQRKVLQNV 664
Query: 654 IISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKY 713
IIS+ K IL LA AG+PL+W AVLTDVGTCL+VILNSML+L+E K + ++ Y
Sbjct: 665 IISITLKVGILVLAFAGHPLIWAAVLTDVGTCLIVILNSMLLLREKDKSKIKKC-----Y 719
Query: 714 GKFLEDKTKPLLNKQSNIDEEKGLLSGEECGKDCC 748
K LE +D E GLLS +C CC
Sbjct: 720 RKKLEGV------DDQGLDLEAGLLSKSQCNSGCC 748
>UniRef100_Q70Q04 Putative cadmium/zinc-transporting ATPase 3 [Arabidopsis halleri
subsp. halleri]
Length = 757
Score = 742 bits (1916), Expect = 0.0
Identities = 392/758 (51%), Positives = 524/758 (68%), Gaps = 29/758 (3%)
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+N++ S F+V+G+CC +E ++V +L+PL GVK SVIVP+RTV VVHD LIS +I
Sbjct: 10 KNLQTSYFDVVGICCTSEVSIVGDVLRPLDGVKEFSVIVPSRTVIVVHDTFLISPLQIVK 69
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN ARLEAS RP GET+ + + P + G+ LALSF KY Y L WLA+ +V G
Sbjct: 70 ALNQARLEASVRPYGETSLKSQWPSPFAILSGVFLALSFFKYFYSLLEWLAVVAVVAGIF 129
Query: 123 KVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALV 182
+L +A+AS+ L+IN L +AV T +Q+FT+ I+FLFS+A WLE+ A HKA
Sbjct: 130 PILAKAVASVTRFRLDINALTFIAVIATLCMQNFTEAATIVFLFSVADWLESSAAHKAST 189
Query: 183 AMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLT 242
MSSL ++AP+ A++AETG VDV++V++NTI++VKAG++IP+DG+VV+G C+VDEK LT
Sbjct: 190 VMSSLMSLAPRKAVIAETGHEVDVDEVRINTIVSVKAGESIPIDGVVVDGSCDVDEKTLT 249
Query: 243 GESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGKNVK 297
GESFPV+K+ DS V A TIN+N Y K+ D ++ K + + + + +
Sbjct: 250 GESFPVSKQRDSTVLAATINLNGYIKVKTTALARDCVVAKMTKLVEEAQKSQTKTQRFID 309
Query: 298 TC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALIL 357
C R + V+ VL +C AV+P L + D+ WFHLALVVL+SGCPC LIL
Sbjct: 310 KCSRYYTPAVV----------VLAACFAVIPVLLKLQDLSHWFHLALVVLVSGCPCGLIL 359
Query: 358 STPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDI 416
STP+A FCALTKAA+SG L+K GD LETL++IK VAFDKTGTIT+ EF V+DF S +I
Sbjct: 360 STPIATFCALTKAAMSGFLIKTGDCLETLAKIKIVAFDKTGTITKAEFMVSDFRSLSHNI 419
Query: 417 NNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDI 476
N LLYW+SSIESKSSHP+A AL+DYAR S++P P+ VENFQNFPGEG++G IDG+DI
Sbjct: 420 NLHNLLYWVSSIESKSSHPMAAALIDYARSVSVEPKPDLVENFQNFPGEGVYGRIDGQDI 479
Query: 477 YIGNKRVGVRAICKR-DNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSGALEAMEEL 535
YIGNKR+ RA C + E +R + + L G F+L+D+CR G +A++EL
Sbjct: 480 YIGNKRIAQRAGCLTVPDMEANMKRGK-TIGYIYIGAKLSGSFNLIDSCRYGVAQALKEL 538
Query: 536 KLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIAMIGDG 595
K LG+++ MLTGD+ A+ Q QL+NA+DIVH+ELLP +KA++I+ FK +GP M+GDG
Sbjct: 539 KSLGIKTAMLTGDNRDAALSTQEQLENALDIVHSELLPQDKARIIDEFKIQGPTMMVGDG 598
Query: 596 INDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVII 655
+NDAPALA ADIG+SMGISGSALA ET + ILMSNDIRK+P+ +RLA+++ +K++ENV++
Sbjct: 599 LNDAPALAKADIGLSMGISGSALATETGDIILMSNDIRKIPKGMRLAKRSHKKVIENVVL 658
Query: 656 SVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYGK 715
SV K AI+ LA GYPLVW AVL D GTCLLVILNSM++L++ + S K
Sbjct: 659 SVSIKGAIMVLAFVGYPLVWAAVLADAGTCLLVILNSMMLLRDEREAVSTCYRASSSPVK 718
Query: 716 FLEDKTKPLLNKQSNIDEEKGLL--SGEECGKDCCKNA 751
ED+ + D E GLL S E K CC +
Sbjct: 719 LEEDEAE---------DLEVGLLQKSEETNKKSCCSGS 747
>UniRef100_Q9SZW5 Potential cadmium/zinc-transporting ATPase 4 [Arabidopsis thaliana]
Length = 760
Score = 741 bits (1913), Expect = 0.0
Identities = 398/764 (52%), Positives = 524/764 (68%), Gaps = 46/764 (6%)
Query: 4 NIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADA 63
N++ S F+V+G+CC++E ++V +L+ + GVK SVIVP+RTV VVHD LIS +I A
Sbjct: 11 NLQTSYFDVVGICCSSEVSIVGNVLRQVDGVKEFSVIVPSRTVIVVHDTFLISPLQIVKA 70
Query: 64 LNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPK 123
LN ARLEAS RP GET+ + + P + G+LL LSF KY Y PL WLA+ +V G
Sbjct: 71 LNQARLEASVRPYGETSLKSQWPSPFAIVSGVLLVLSFFKYFYSPLEWLAIVAVVAGVFP 130
Query: 124 VLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVA 183
+L +A+AS+ L+IN L L+AV T +QDFT+ I+FLFS+A WLE+ A HKA +
Sbjct: 131 ILAKAVASVTRFRLDINALTLIAVIATLCMQDFTEAATIVFLFSVADWLESSAAHKASIV 190
Query: 184 MSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTG 243
MSSL ++AP+ A++A+TG VDV++V +NT+++VKAG++IP+DG+VV+G C+VDEK LTG
Sbjct: 191 MSSLMSLAPRKAVIADTGLEVDVDEVGINTVVSVKAGESIPIDGVVVDGSCDVDEKTLTG 250
Query: 244 ESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGKNVKT 298
ESFPV+K+ +S V A TIN+N Y K+ D ++ K + + + + +
Sbjct: 251 ESFPVSKQRESTVMAATINLNGYIKVKTTALARDCVVAKMTKLVEEAQKSQTKTQRFIDK 310
Query: 299 C*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILS 358
C R + V+ V +C AV+P L V D+ WFHLALVVL+SGCPC LILS
Sbjct: 311 CSRYYTPAVV----------VSAACFAVIPVLLKVQDLSHWFHLALVVLVSGCPCGLILS 360
Query: 359 TPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDIN 417
TPVA FCALTKAA SG L+K GD LETL++IK VAFDKTGTIT+ EF V+DF S IN
Sbjct: 361 TPVATFCALTKAATSGFLIKTGDCLETLAKIKIVAFDKTGTITKAEFMVSDFRSLSPSIN 420
Query: 418 NETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIY 477
LLYW+SSIE KSSHP+A AL+DYAR S++P P+ VENFQNFPGEG++G IDG+DIY
Sbjct: 421 LHKLLYWVSSIECKSSHPMAAALIDYARSVSVEPKPDIVENFQNFPGEGVYGRIDGQDIY 480
Query: 478 IGNKRVGVRAICKRDNCEVQFQRPEI-STKKNN-------CEETLVGVFSLVDACRSGAL 529
IGNKR+ RA C DN P+I +T K L G F+L+D CR G
Sbjct: 481 IGNKRIAQRAGCLTDNV------PDIEATMKRGKTIGYIYMGAKLTGSFNLLDGCRYGVA 534
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+A++ELK LG+++ MLTGD+ AM Q QL+NA+DIVH+ELLP +KA++I++FK +GP
Sbjct: 535 QALKELKSLGIQTAMLTGDNQDAAMSTQEQLENALDIVHSELLPQDKARIIDDFKIQGPT 594
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
M+GDG+NDAPALA ADIGISMGISGSALA ET + ILMSNDIRK+P+ +RLA+++ +K+
Sbjct: 595 MMVGDGLNDAPALAKADIGISMGISGSALATETGDIILMSNDIRKIPKGMRLAKRSHKKV 654
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEK---RE 706
+ENV++SV K AI+ L GYPLVW AVL D GTCLLVILNSM++L++ + R
Sbjct: 655 IENVVLSVSIKGAIMVLGFVGYPLVWAAVLADAGTCLLVILNSMMLLRDEREAVSTCYRA 714
Query: 707 STKGSKYGKFLEDKTKPLLNKQSNIDEEKGLL--SGEECGKDCC 748
ST S K ED+ + D E GLL S E K CC
Sbjct: 715 ST--SSPVKLEEDEVE---------DLEVGLLQKSEETSKKSCC 747
>UniRef100_Q8LPW0 Putative heavy metal transporter HMA3 [Arabidopsis thaliana]
Length = 760
Score = 735 bits (1897), Expect = 0.0
Identities = 396/764 (51%), Positives = 522/764 (67%), Gaps = 46/764 (6%)
Query: 4 NIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADA 63
N++ S F+V+G+CC++E ++V +L+ + GVK SVIVP+RTV VVHD LIS +I A
Sbjct: 11 NLQTSYFDVVGICCSSEVSIVGNVLRQVDGVKEFSVIVPSRTVIVVHDTFLISPLQIVKA 70
Query: 64 LNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPK 123
LN ARLEAS RP GET+ + + P + G+LL LSF KY Y PL WLA+ +V G
Sbjct: 71 LNQARLEASVRPYGETSLKSQWPSPFAIVSGVLLVLSFFKYFYSPLEWLAIVAVVAGVFP 130
Query: 124 VLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVA 183
+L +A+AS+ L+IN L L+AV T +QDFT+ I+FLFS+A WLE+ A HKA +
Sbjct: 131 ILAKAVASVTRFRLDINALTLIAVIATLCMQDFTEAATIVFLFSVADWLESSAAHKASIV 190
Query: 184 MSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTG 243
MSSL ++AP+ A++A+TG VDV++V +NT+++VKAG++IP+DG+VV+G C+VDEK LTG
Sbjct: 191 MSSLMSLAPRKAVIADTGLEVDVDEVGINTVVSVKAGESIPIDGVVVDGSCDVDEKTLTG 250
Query: 244 ESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGKNVKT 298
ESFPV+K+ +S V A TIN+N Y K+ D ++ K + + + + +
Sbjct: 251 ESFPVSKQRESTVMAATINLNGYIKVKTTALARDCVVAKMTKLVEEAQKSQTKTQRFIDK 310
Query: 299 C*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILS 358
C R + V+ V +C AV+P L V D+ WFHLALVVL+SGCPC LILS
Sbjct: 311 CSRYYTPAVV----------VSAACFAVIPVLLKVQDLSHWFHLALVVLVSGCPCGLILS 360
Query: 359 TPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDIN 417
TPVA FCALTKAA SG L+K GD LETL++IK VAFDKTGTIT+ EF V+DF S IN
Sbjct: 361 TPVATFCALTKAATSGFLIKTGDCLETLAKIKIVAFDKTGTITKAEFMVSDFRSLSPSIN 420
Query: 418 NETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIY 477
LL W+SSIE KSSHP+A AL+DYA S++P P+ VENFQNFPGEG++G IDG+DIY
Sbjct: 421 LHKLLNWVSSIECKSSHPMAAALIDYAISVSVEPKPDIVENFQNFPGEGVYGRIDGQDIY 480
Query: 478 IGNKRVGVRAICKRDNCEVQFQRPEI-STKKNN-------CEETLVGVFSLVDACRSGAL 529
IGNKR+ RA C DN P+I +T K L G F+L+D CR G
Sbjct: 481 IGNKRIAQRAGCLTDNV------PDIEATMKRGKTIGYIYMGAKLTGSFNLLDGCRYGVA 534
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+A++ELK LG+++ MLTGD+ AM Q QL+NA+DIVH+ELLP +KA++I++FK +GP
Sbjct: 535 QALKELKSLGIQTAMLTGDNQDAAMSTQEQLENALDIVHSELLPQDKARIIDDFKIQGPT 594
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
M+GDG+NDAPALA ADIGISMGISGSALA ET + ILMSNDIRK+P+ +RLA+++ +K+
Sbjct: 595 MMVGDGLNDAPALAKADIGISMGISGSALATETGDIILMSNDIRKIPKGMRLAKRSHKKV 654
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEK---RE 706
+ENV++SV K AI+ L GYPLVW AVL D GTCLLVILNSM++L++ + R
Sbjct: 655 IENVVLSVSIKGAIMVLGFVGYPLVWAAVLADAGTCLLVILNSMILLRDEREAVSTCYRS 714
Query: 707 STKGSKYGKFLEDKTKPLLNKQSNIDEEKGLL--SGEECGKDCC 748
ST S K ED+ + D E GLL S E K CC
Sbjct: 715 ST--SSPVKLEEDEVE---------DLEVGLLQKSEETSKKSCC 747
>UniRef100_Q8H384 Putative potential cadmium/zinc-transporting ATPase 4 [Oryza
sativa]
Length = 1004
Score = 638 bits (1646), Expect = 0.0
Identities = 354/749 (47%), Positives = 484/749 (64%), Gaps = 13/749 (1%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN 65
K++ +VLG+CC+ E LVER+L PL GV+ VSV+V +RTV V HD ES I ALN
Sbjct: 42 KKTYLDVLGVCCSAEVALVERLLAPLDGVRVVSVVVASRTVVVEHDPAAAPESAIVKALN 101
Query: 66 TARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPKVL 125
A LEAS R G + + P +A G+LL SF ++++PPL LA+ +V G P ++
Sbjct: 102 KAGLEASVRAYGSSGVVSRWPSPYIVASGVLLTASFFEWLFPPLQCLAVAAVVAGAPPMV 161
Query: 126 FRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMS 185
R A+ L+L+IN+L+L+AV G L D+T+ G I+FLF+ A+WLET A KA MS
Sbjct: 162 RRGFAAASRLSLDINVLMLIAVAGALCLGDYTEAGAIVFLFTTAEWLETLACTKASAGMS 221
Query: 186 SLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGES 245
SL M P A++A TGE V V DV++ ++AV+AG+ +P+DG+VV+G+ EVDE+ LTGES
Sbjct: 222 SLMGMLPVKAVIATTGEVVSVRDVRVGDVVAVRAGEIVPVDGVVVDGQSEVDERSLTGES 281
Query: 246 FPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*Q 305
FPV K+ S VWAGT+N + Y + +N +V K + + RS Q
Sbjct: 282 FPVPKQPHSEVWAGTMNFDGYIAVRTTAL------AENSTVAKMERLVEAAQNS-RSKTQ 334
Query: 306 KVIST--KIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAI 363
++I + K + V+ + +A++PA L +E W+ LALV+L+S CPCAL+LSTPVA
Sbjct: 335 RLIDSCAKYYTPAVVVVAAGVALIPALLGADGLEQWWKLALVMLVSACPCALVLSTPVAS 394
Query: 364 FCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDD--INNETL 421
FCA+ +AA G+ +KGGD LE+L I+ VAFDKTGTITRGEFS+ F D + + L
Sbjct: 395 FCAMLRAARMGIFIKGGDVLESLGEIRAVAFDKTGTITRGEFSIDSFHLVGDHKVEMDHL 454
Query: 422 LYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNK 481
LYWI+SIESKSSHP+A ALV+YA+ SI+P PENV +F+ +PGEGI+G I G+ IYIGN+
Sbjct: 455 LYWIASIESKSSHPMAAALVEYAQSKSIQPNPENVGDFRIYPGEGIYGEIHGKHIYIGNR 514
Query: 482 RVGVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSGALEAMEELKLLGVR 541
R RA + E+ +S C+ L GVFSL D CR+GA EA+ EL LG++
Sbjct: 515 RTLARASSPQSTQEMGEMIKGVSIGYVICDGELAGVFSLSDDCRTGAAEAIRELGSLGIK 574
Query: 542 SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFK-KEGPIAMIGDGINDAP 600
SVMLTGDSS A + Q QL ++ +H+ELLP +K +L+ K + GP M+GDG+NDA
Sbjct: 575 SVMLTGDSSAAATHAQGQLGGVMEELHSELLPEDKVRLVSGLKARFGPTMMVGDGMNDAA 634
Query: 601 ALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFK 660
ALA AD+G+SMGISGSA A ETS+A LMS+D+ +VPEA+RL R R + NV SV K
Sbjct: 635 ALAAADVGVSMGISGSAAAMETSHATLMSSDVLRVPEAVRLGRCARRTIAVNVAGSVAVK 694
Query: 661 CAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYGKFLEDK 720
A+LALA A P++W AVL DVGTCLLV+LNSM +L+E K +E + L +
Sbjct: 695 AAVLALAAAWRPVLWAAVLADVGTCLLVVLNSMTLLREEWKGGAKEDGACRATARSLVMR 754
Query: 721 TKPLLNKQSNIDEEKGLLSGEEC-GKDCC 748
++ + Q+ + G E+ G CC
Sbjct: 755 SQLAADSQAPNAADAGAAGREQTNGCRCC 783
>UniRef100_Q9K5Q2 Cadmium-transporting ATPase [Bacillus halodurans]
Length = 707
Score = 343 bits (880), Expect = 1e-92
Identities = 228/712 (32%), Positives = 376/712 (52%), Gaps = 36/712 (5%)
Query: 5 IKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADAL 64
+K++ + V G CA A E+ +K LHGV V A +TV D + + + A A
Sbjct: 9 VKKTTYRVQGFSCANCANTFEKNVKDLHGVVEAQVNFGASKLTVYGDAT-VEDLERAGAF 67
Query: 65 NTARLEASFRPQGETNNEKKCPDILT----MACGLLLALSFLKYIYPPLG--W---LALG 115
++ RP+ E E K P + LLL + + Y W L
Sbjct: 68 EHLKV----RPENEKAIETKEPFWKAYGHVIVSALLLLVGVVSYFMKGERHLWTVSLFAS 123
Query: 116 SVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETR 175
S+ IG + ++ L ++ L+ +AV G A + ++ +G V++ LF+I++ LE
Sbjct: 124 SIVIGGYSLFKAGFYNLIRLRFDMKTLMTVAVIGAALIGEWLEGAVVVILFAISEALERF 183
Query: 176 ATHKALVAMSSLTNMAPQMAIVAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEGK 233
+ +A ++ SL +MAP+ A+V GE + V +++ I+ +K G + +DGIV G+
Sbjct: 184 SMDRARQSIRSLVDMAPKEALVRRNGEELVIPVEAIEVGDIMLIKPGQKLAMDGIVRSGR 243
Query: 234 CEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGK 288
++E +TGES PV K V+AGT+N + ++ + DT +I + G+
Sbjct: 244 SSMNEAAITGESVPVEKMVGDDVFAGTLNEEGFLEVEVTKRVEDTTLAKIIHLVEEAQGE 303
Query: 289 RHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLL 348
R S V + + ++ V+ + +A VP E W + L VL+
Sbjct: 304 RAPSQAFVDMFAKYYTPAIM----------VIAALVATVPPLFFAGSWETWIYQGLAVLV 353
Query: 349 SGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVT 408
GCPCAL++STPV+I A+ AA G+L+KGG YLE +R++ VAFDKTGT+T+G+ +VT
Sbjct: 354 VGCPCALVISTPVSIVTAIGNAARHGILVKGGIYLEEAARLRAVAFDKTGTLTKGKPAVT 413
Query: 409 DFSAGD-DINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGI 467
D+ D D + LL ++++E +S HP+A A++ A + V F + G+GI
Sbjct: 414 DYQVYDADTDGRELLAAVAALEYRSQHPLAAAVMKKADDEQLSYTDMEVSEFSSITGKGI 473
Query: 468 FGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSG 527
GT++G I +G + + + Q + E+ ++ + ++ D R
Sbjct: 474 AGTVNGIAINVGKPALFGELPPAIEADVARLQAAGKTIMLIGAEQKVLALLAVADEVREV 533
Query: 528 ALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKE 586
+ A+E L LG+ ++VMLTGD+ A + Q+ + V AELLP +K ++ K+E
Sbjct: 534 SRTAVERLHALGITKTVMLTGDNRMTARAIGKQV--GVAEVEAELLPEDKLGFVKGLKEE 591
Query: 587 -GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKT 645
G +AM+GDG+NDAPALA + +G++MG +G+ A ET++ LM +D+RK+P IRL+RK
Sbjct: 592 HGRVAMVGDGVNDAPALAASSVGVAMGGAGTDTALETADVALMGDDLRKLPFLIRLSRKA 651
Query: 646 TRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+ EN+ S+G K L L G+ +W+A+L D+G LLV LN + +L+
Sbjct: 652 LLIIKENIAFSLGIKLVALFLVAPGWLTLWIAILADMGATLLVTLNGLRLLR 703
>UniRef100_Q6GIX1 Probable cadmium-transporting ATPase [Staphylococcus aureus]
Length = 726
Score = 341 bits (875), Expect = 5e-92
Identities = 228/725 (31%), Positives = 381/725 (52%), Gaps = 46/725 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K L GV+ V A + V + L +
Sbjct: 15 YRVEGFSCANCAGKFEKNVKQLAGVQDAKVNFGASKIDVYGNASVQELEKAGAFENLKVF 74
Query: 57 ESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LGW 111
K+A++ A E + P+ E K L A LL+A +L +
Sbjct: 75 PEKLANSSMQAVKEDTKAPKEEKIPFYKKHSTLLFAT-LLIAFGYLSHFVNGEDNLVTSM 133
Query: 112 LALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQW 171
L +GS+ IG + ++ ++ L+ +AV G A + ++ + +++ LF+I++
Sbjct: 134 LFVGSIVIGGYSLFKVGFQNLIRFDFDMKTLMTVAVIGAAIIGEWAEASIVVVLFAISEA 193
Query: 172 LETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIV 229
LE + +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI+
Sbjct: 194 LERFSMDRARQSIRSLMDIAPKEALVMRNGQEIMIHVDDIAVGDIMIVKPGEKIAMDGII 253
Query: 230 VEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNY 284
+ G V++ +TGES PV K D V+AGT+N K + DT +I
Sbjct: 254 INGVSAVNQAAITGESVPVAKTVDDEVFAGTLNEEGLLEVKITKYVEDTTISKIIHLVEE 313
Query: 285 SVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLAL 344
+ G+R + ++F K K + V+ + +AVVP + W + L
Sbjct: 314 AQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQGL 363
Query: 345 VVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGE 404
VL+ GCPCAL++STP++I A+ AA G+L+KGG YLE L IK +AFDKTGT+T+G
Sbjct: 364 AVLVVGCPCALVISTPISIVSAIGNAAKKGVLIKGGVYLEELGAIKAIAFDKTGTLTKGV 423
Query: 405 FSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFP 463
VTDF D + + L I+++E +S HP+A A++ A +I VE+F +
Sbjct: 424 PVVTDFKVLNDQVEEKELFSIITALEYRSQHPLASAIMKKAEQDNITYSDVRVEDFTSIT 483
Query: 464 GEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVF 518
G GI G IDG YIG+ R+ + E + + + + ++T++GV
Sbjct: 484 GRGIQGNIDGTTYYIGSPRLFKELNVSDFSLEFENKVKVLQNQGKTAMIIGTDQTILGVI 543
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + +++L LG++ ++MLTGD+ A + + + + + +EL+P +K
Sbjct: 544 AVADEVRETSKNVIQKLHQLGIKQTIMLTGDNQGTAEAIGAHV--GVSDIQSELMPQDKL 601
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ K E G +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K+P
Sbjct: 602 DYIKKMKAEHGNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSKLP 661
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+ ++
Sbjct: 662 FAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLRLM 721
Query: 697 QENHK 701
+ K
Sbjct: 722 RVKDK 726
>UniRef100_Q8ETI5 Cadmium-transporting ATPase [Oceanobacillus iheyensis]
Length = 711
Score = 340 bits (871), Expect = 1e-91
Identities = 224/710 (31%), Positives = 385/710 (53%), Gaps = 40/710 (5%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL 69
+ + G+ C A E ++ L GV+ V A V V + + K N
Sbjct: 15 YRIQGLSCTNCAAKFENNVRKLPGVENAQVNFGASKVYVEGNTTIQDLEKAGSFENLKIT 74
Query: 70 EASFRPQGETNNEKKCPDILTM-ACGLLLALSFL-------KYIYPPLGWLALGSVAIGF 121
R +T + K D + + ++ +S + + I P +G+ A ++ IG
Sbjct: 75 NDRERKSEKTISFWKQKDNIRLYIAAFIIVISLIASYSLNEQSIIPTIGFGA--AILIGG 132
Query: 122 PKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKAL 181
+ + + ++ ++N L+ +AV G A + ++ +G ++ LF+I++ LE + KA
Sbjct: 133 YSLFMKGLKNLIQFQFDMNTLMTIAVIGAALIGEWLEGAAVVILFAISEALERFSMDKAR 192
Query: 182 VAMSSLTNMAPQMAIVAETG--ERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEK 239
++ SL ++AP A++ + +DVND+++N I+ VK G I +DGIV++G V++
Sbjct: 193 QSIQSLMDIAPNEALIRRDNIEQTIDVNDIQINDIMIVKPGQKIAMDGIVIKGSSSVNQS 252
Query: 240 MLTGESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGK 294
+TGES PV + +D V+AGT N + +I + DT +I + +R S
Sbjct: 253 AITGESIPVARMTDDEVFAGTFNEDGLLEIRVTKKVEDTTISKIIHLVEEAQAERAPS-- 310
Query: 295 NVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCA 354
++F K K + ++ +A+VP+A++ D W + L +L+ GCPCA
Sbjct: 311 ------QAFVDKF--AKYYTPVIMLIALMVAIVPSAIT-GDWSTWIYQGLAILVVGCPCA 361
Query: 355 LILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSA-- 412
L++STPVAI A+ AA +G+L+KGG YLE ++++AFDKTGT+T G VTD +
Sbjct: 362 LVISTPVAIVTAIGNAAKNGVLIKGGVYLEETGSLQSIAFDKTGTLTEGVPHVTDIVSYQ 421
Query: 413 GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTID 472
G++ +N +L I++IE+ S HP+A A++ A +I +ENF++ G+G+ TID
Sbjct: 422 GEEGSNLSL---IAAIENGSQHPLAAAIIRKANEDNITFKNVEMENFKSITGKGVQATID 478
Query: 473 GRDIYIGNKRVGVRAICKRDNCEVQ-FQRPEISTKKN---NCEETLVGVFSLVDACRSGA 528
+D Y+G+ D VQ + ++ K + ++ ++++ D R+ +
Sbjct: 479 NKDYYVGSPDFFKELQTSIDPTIVQQIEDMQLEGKTVIILGTHQDILSLYAIADKVRAAS 538
Query: 529 LEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENF-KKEG 587
+ +L LG+ +VMLTGD+ A + Q+ + V + LLP +K K I+ +
Sbjct: 539 KHVVTKLNKLGMSTVMLTGDNQNTASAIGKQV--GVTNVKSNLLPEDKLKYIKELTNNDQ 596
Query: 588 PIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTR 647
AM+GDGIND+PALA + +GI+MG +G+ A ET++ +LMS+D+ K+P + L+RKT R
Sbjct: 597 KTAMVGDGINDSPALAASTVGIAMGGAGTDTALETADIVLMSDDLNKLPYTMNLSRKTLR 656
Query: 648 KLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+ EN+I S+G K A L L I G+ +WLA+ D+G L+V LNS+ +L+
Sbjct: 657 IIKENIIFSLGIKLAALLLVIPGWLTLWLAIFADIGATLIVTLNSLRLLR 706
>UniRef100_Q5WLE3 Cadmium-transporting ATPase [Bacillus clausii]
Length = 709
Score = 335 bits (860), Expect = 3e-90
Identities = 230/712 (32%), Positives = 381/712 (53%), Gaps = 36/712 (5%)
Query: 8 SNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLL--ISESKIADALN 65
S + V G CA A E+ +K L GVK V A +TV + + + E+ + L
Sbjct: 7 SVYRVDGFTCANCAGKFEKNVKDLPGVKDAKVNFGASKLTVYGEATIEALEEAGAFEQLK 66
Query: 66 TARLEASFRPQGETNNE--KKCPDILTMACGLLLALSF-LKYIYPPLGWLALGSVAI--- 119
+ + +P T KK IL L+ F KY +G + L AI
Sbjct: 67 VTKEPSRRQPFQYTQIPFYKKHRSILLSVLLLVFGYVFQFKYGESNIGTVGLFLTAILVG 126
Query: 120 GFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHK 179
GFP +L I ++ L ++ L+ LAV G A + ++ + V++ LF+I++ LE + +
Sbjct: 127 GFP-LLKTGIRNVARLEFDMKTLMTLAVIGGAIIGEWGEVAVVVILFAISEALERFSMDR 185
Query: 180 ALVAMSSLTNMAPQMAIVAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVD 237
A ++ SL ++AP A++ G+ V V D+ + I+ VK G I +DGIVV G V+
Sbjct: 186 ARKSIQSLMDIAPNEALIKRNGQELTVHVGDIVVGDIMIVKPGQKIAMDGIVVHGHSSVN 245
Query: 238 EKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSS 292
+ +TGES PV K D V+A T+N K++ DT +I + G+R +
Sbjct: 246 QAAITGESVPVEKAVDDEVFASTLNEEGLLEIEVTKLVEDTTISKIIHLVEEAQGERAPA 305
Query: 293 GKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCP 352
++F K K + V+ + +A++P + + W + L VL+ GCP
Sbjct: 306 --------QAFVDKF--AKYYTPIIMVVAALVAIIPPLFAGGNWSTWVYQGLAVLVVGCP 355
Query: 353 CALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSA 412
CAL++STP++I A+ AA G+L+KGG YLE ++ +K +AFDKTGT+T+G VTDF+
Sbjct: 356 CALVISTPISIVSAIGNAAKKGVLVKGGVYLEEMASLKAIAFDKTGTLTKGIPVVTDFAL 415
Query: 413 -GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTI 471
++ L ++++E++S HP+A A+V A + VEN G+GI GT+
Sbjct: 416 LNPQLDQRQLFAIVAALENRSQHPLASAIVAKAEDEQLPYHDYMVENVTALTGKGITGTV 475
Query: 472 DGRDIYIGNKRVGVRAICKRDNCEV-----QFQRPEISTKKNNCEETLVGVFSLVDACRS 526
+G+ Y+G+ ++ + RD Q+ + E L+ + ++ D R
Sbjct: 476 NGKTYYVGSPKLFQERLPTRDITPFAQNIKSLQQQGKTAMLVGTETELLAIVAVADEVRP 535
Query: 527 GALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK 585
+ E + +L G+ ++VMLTGD+ Q A + Q DI AEL+P +K + I+ +++
Sbjct: 536 SSKEVIRKLHEAGIAKTVMLTGDNKQTAHAI-GQAVGVADI-QAELMPEDKLRFIKQWQE 593
Query: 586 E-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK 644
G +AM+GDG+NDAPALA A +GI+MG +G+ A ET++ LM +D++K+P ++L+RK
Sbjct: 594 NHGHVAMVGDGVNDAPALAAATVGIAMGGAGTDTALETADIALMGDDLKKLPFTVKLSRK 653
Query: 645 TTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
T + N+ ++ K A L L + G+ +W+A+L+D+G LLV LNS+ ++
Sbjct: 654 TLNIIKANIAFAIAIKLAALLLVVPGWLTLWIAILSDMGATLLVALNSIRLI 705
>UniRef100_Q93ID4 Cadmium resistance protein B [Staphylococcus aureus]
Length = 804
Score = 334 bits (857), Expect = 6e-90
Identities = 226/725 (31%), Positives = 378/725 (51%), Gaps = 46/725 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K L GV+ V A + V + L +
Sbjct: 93 YRVEGFSCANCAGKFEKNVKQLAGVQDAKVNFGASKIDVYGNTSVEELEKAGAFENLKVI 152
Query: 57 ESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LGW 111
K+A+ A E + P+ E K L A LL+A +L +
Sbjct: 153 PEKLANPSIQAVKEDTKAPKEEKIPFYKKHSTLLFAT-LLIAFGYLSHFVNGEDNLVTSM 211
Query: 112 LALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQW 171
L + S+ IG + ++ ++ L+ +AV G A + ++ + +++ LF+I++
Sbjct: 212 LFVSSIVIGGYSLFKVGFQNLIRFDFDMKTLMTVAVIGAAIIGEWAEASIVVILFAISEA 271
Query: 172 LETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIV 229
LE + +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI+
Sbjct: 272 LERFSMDRARQSIRSLMDIAPKEALVRRNGQEIMIHVDDIAVGDIMIVKPGEKIAMDGII 331
Query: 230 VEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNY 284
+ G V++ +TGES PV K D V+AGT+N K + DT +I
Sbjct: 332 INGVSAVNQAAITGESVPVAKTVDDEVFAGTLNEEGLLEVKITKYVEDTTISKIIHLVEE 391
Query: 285 SVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLAL 344
+ G+R + ++F K K + V+ + +AVVP + W + L
Sbjct: 392 AQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQGL 441
Query: 345 VVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGE 404
VL+ GCPCAL+++TP++I A+ AA G+L+KGG YLE L IK +AFDKTGT+T+G
Sbjct: 442 AVLVVGCPCALVITTPISIVSAIGNAAKKGVLIKGGVYLEELGAIKAIAFDKTGTLTKGV 501
Query: 405 FSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFP 463
VTDF D + + L I+++E +S HP+A A++ A +I V++F +
Sbjct: 502 PVVTDFKVLNDQVEEKELFSIITALEYRSQHPLASAIMKKAEQDNITYSDVRVKDFTSIT 561
Query: 464 GEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVF 518
G GI G IDG YIG+ R+ + E + + + + ++T++GV
Sbjct: 562 GRGIQGNIDGTTYYIGSPRLFKELNVSDFSLEFENKVKVLQNQGKTAMIIGTDQTILGVI 621
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + + +L LG++ ++MLTGD+ A + + + + + +ELLP +K
Sbjct: 622 AVADEVRETSKNVILKLHQLGIKQTIMLTGDNQGTAEAIGAHV--GVSDIQSELLPQDKL 679
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ K E G +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K+P
Sbjct: 680 DYIKKMKAEHGNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSKLP 739
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+ ++
Sbjct: 740 FAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLRLM 799
Query: 697 QENHK 701
+ K
Sbjct: 800 RVKDK 804
>UniRef100_P37386 Probable cadmium-transporting ATPase [Staphylococcus aureus]
Length = 804
Score = 334 bits (857), Expect = 6e-90
Identities = 226/725 (31%), Positives = 378/725 (51%), Gaps = 46/725 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K L GV+ V A + V + L +
Sbjct: 93 YRVEGFSCANCAGKFEKNVKQLAGVQDAKVNFGASKIDVYGNASVEELEKAGAFENLKVI 152
Query: 57 ESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LGW 111
K+A+ A E + P+ E K L A LL+A +L +
Sbjct: 153 PEKLANPSIQAVKEDTKAPKEEKIPFYKKHSTLLFAT-LLIAFGYLSHFVNGEDNLVTSM 211
Query: 112 LALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQW 171
L + S+ IG + ++ ++ L+ +AV G A + ++ + +++ LF+I++
Sbjct: 212 LFVSSIVIGGYSLFKVGFQNLIRFDFDMKTLMTVAVIGAAIIGEWAEASIVVILFAISEA 271
Query: 172 LETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIV 229
LE + +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI+
Sbjct: 272 LERFSMDRARQSIRSLMDIAPKEALVRRNGQEIMIHVDDIAVGDIMIVKPGEKIAMDGII 331
Query: 230 VEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNY 284
+ G V++ +TGES PV K D V+AGT+N K + DT +I
Sbjct: 332 INGVSAVNQAAITGESVPVAKTVDDEVFAGTLNEEGLLEVKITKYVEDTTISKIIHLVEE 391
Query: 285 SVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLAL 344
+ G+R + ++F K K + V+ + +AVVP + W + L
Sbjct: 392 AQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQGL 441
Query: 345 VVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGE 404
VL+ GCPCAL+++TP++I A+ AA G+L+KGG YLE L IK +AFDKTGT+T+G
Sbjct: 442 AVLVVGCPCALVITTPISIVSAIGNAAKKGVLIKGGVYLEELGAIKAIAFDKTGTLTKGV 501
Query: 405 FSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFP 463
VTDF D + + L I+++E +S HP+A A++ A +I V++F +
Sbjct: 502 PVVTDFKVLNDQVEEKELFSIITALEYRSQHPLASAIMKKAEQDNITYSDVRVKDFTSIT 561
Query: 464 GEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVF 518
G GI G IDG YIG+ R+ + E + + + + ++T++GV
Sbjct: 562 GRGIQGNIDGTTYYIGSPRLFKELNVSDFSLEFENKVKVLQNQGKTAMIIGTDQTILGVI 621
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + + +L LG++ ++MLTGD+ A + + + + + +ELLP +K
Sbjct: 622 AVADEVRETSKNVILKLHQLGIKQTIMLTGDNQGTAEAIGAHV--GVSDIQSELLPQDKL 679
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ K E G +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K+P
Sbjct: 680 DYIKKMKAEHGNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSKLP 739
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+ ++
Sbjct: 740 FAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLRLM 799
Query: 697 QENHK 701
+ K
Sbjct: 800 RVKDK 804
>UniRef100_Q9JRM2 CadA protein [Xanthomonas maltophilia]
Length = 727
Score = 334 bits (856), Expect = 7e-90
Identities = 226/727 (31%), Positives = 382/727 (52%), Gaps = 50/727 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K + GV+ V A + V + L +S
Sbjct: 16 YRVQGFTCANCAGKFEKNVKKIPGVQDAKVNFGASKIDVYGNASVEELEKAGAFENLKVS 75
Query: 57 ESKIAD-ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LG 110
K+A+ + + + + +T KK +L LL+A +L +
Sbjct: 76 PEKLANQTIQRVKDDTKAHKEEKTPFYKKHSTLLFAT--LLIAFGYLSHFVNGEDNLVTS 133
Query: 111 WLALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQ 170
L +GS+ IG + ++ ++ L+ +AV G + + + +++ LF+I++
Sbjct: 134 MLFVGSIVIGGYSLFKVGFPNLIRFDFDMKTLITVAVIGATIIGKWAEASIVVILFAISE 193
Query: 171 WLETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGI 228
LE + +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI
Sbjct: 194 ALERFSMDRARQSIRSLMDIAPKEALVRRNGQEIIIHVDDIAVGDIMIVKPGEKIAMDGI 253
Query: 229 VVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKN 283
+V G V+E +TGES PV+K D V+AGT+N K + DT +I
Sbjct: 254 IVNGLSVVNEAAITGESVPVSKAVDDEVFAGTLNEEGLIEVKITKYVEDTTIAKIIHLVE 313
Query: 284 YSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLA 343
+ G+R + ++F K K + V+ + +AVVP + W +
Sbjct: 314 EAQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQG 363
Query: 344 LVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRG 403
L VL+ GCPCAL++STP++I A+ AA G+L+KGG YLE L IKTVAFDKTGT+T+G
Sbjct: 364 LAVLVVGCPCALVISTPISIVSAIGNAAKKGVLVKGGVYLEKLGDIKTVAFDKTGTLTKG 423
Query: 404 EFSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNF 462
VTDF D + + L I+++E +S HP+A A++ A +I VE F +
Sbjct: 424 VPVVTDFEVLNDQVEEKELFSTITALEYRSQHPLASAIMKKAEQDNIPYSNVQVEEFTSI 483
Query: 463 PGEGIFGTIDGRDIYIGNKR------VGVRAICKRDNCEVQFQRPEISTKKNNCEETLVG 516
G GI G ++G YIG+ + V ++ +N ++ Q + E+T++G
Sbjct: 484 TGRGIKGIVNGTTYYIGSPKLFKELNVSDFSLGFENNVKI-LQNQGKTAMIIGTEKTILG 542
Query: 517 VFSLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHE 575
V ++ D R + +++L LG++ ++MLTGD+ A + + + + + +EL+P +
Sbjct: 543 VIAVADEVRETSKNVIQKLHQLGIKQTIMLTGDNQGTANAIGTHV--GVSDIQSELMPQD 600
Query: 576 KAKLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRK 634
K I+ + E +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K
Sbjct: 601 KLDYIKKMQSEYDNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSK 660
Query: 635 VPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSML 694
+P A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+
Sbjct: 661 LPFAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLR 720
Query: 695 ILQENHK 701
+++ K
Sbjct: 721 LMRVKDK 727
>UniRef100_Q60048 Probable cadmium-transporting ATPase [Listeria monocytogenes]
Length = 711
Score = 334 bits (856), Expect = 7e-90
Identities = 216/718 (30%), Positives = 381/718 (52%), Gaps = 38/718 (5%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLL--ISESKIADA 63
+++ + V G+ C A ER +K + GV V A +TV + + + ++ +
Sbjct: 3 EKTVYRVDGLSCTNCAAKFERNVKEIEGVTEAIVNFGASKITVTGEASIQQVEQAGAFEH 62
Query: 64 LNTARLEASFR-PQGETNNEKKC-PDILTMACGLLLALSFLKYI-----YPPLGWLALGS 116
L + SF P+ T+++ + + GL +A+ + I + L + +
Sbjct: 63 LKIIPEKESFTDPEHFTDHQSFIRKNWRLLLSGLFIAVGYASQIMNGEDFYLTNALFIFA 122
Query: 117 VAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRA 176
+ IG + ++ + L+ +A+ G A + ++ +G +++ LF++++ LE +
Sbjct: 123 IFIGGYSLFKEGFKNLLKFEFTMETLMTIAIIGAAFIGEWAEGSIVVILFAVSEALERYS 182
Query: 177 THKALVAMSSLTNMAPQMAIVAETG--ERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKC 234
KA ++ SL ++AP+ A+V +G V V+D+++ I+ +K G I +DG VV+G
Sbjct: 183 MDKARQSIRSLMDIAPKEALVRRSGTDRMVHVDDIQIGDIMIIKPGQKIAMDGHVVKGYS 242
Query: 235 EVDEKMLTGESFPVTKESDSLVWAGTINMN-----AYPKIIFDTYDFRIYKCKNYSVGKR 289
V++ +TGES PV K D V+AGT+N A K + DT +I + G+R
Sbjct: 243 AVNQAAITGESIPVEKNIDDSVFAGTLNEEGLLEVAVTKRVEDTTISKIIHLVEEAQGER 302
Query: 290 HSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLS 349
+ V T + + +I V+ + IA VP L + E W + L VL+
Sbjct: 303 APAQAFVDTFAKYYTPAII----------VIAALIATVPPLLFGGNWETWVYQGLSVLVV 352
Query: 350 GCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTD 409
GCPCAL++STPVAI A+ AA +G+L+KGG YLE + +K +AFDKTGT+T+G VTD
Sbjct: 353 GCPCALVVSTPVAIVTAIGNAAKNGVLVKGGVYLEEIGGLKAIAFDKTGTLTKGVPVVTD 412
Query: 410 F---SAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEG 466
+ + +I + ++++E S HP+A A++ Y + NV +F + G+G
Sbjct: 413 YIELTEATNIQHNKNYIIMAALEQLSQHPLASAIIKYGETREMDLTSINVNDFTSITGKG 472
Query: 467 IFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVFSLV 521
I GT+DG Y+G+ + + + + Q ++ K + L+ + ++
Sbjct: 473 IRGTVDGNTYYVGSPVLFKELLASQFTDSIHRQVSDLQLKGKTAMLFGTNQKLISIVAVA 532
Query: 522 DACRSGALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLI 580
D RS + ++ L LG+ +++MLTGD+ A + Q+ + + EL+P +K I
Sbjct: 533 DEVRSSSQHVIKRLHELGIEKTIMLTGDNQATAQAIGQQV--GVSEIEGELMPQDKLDYI 590
Query: 581 ENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAI 639
+ K G +AM+GDGINDAPALA A +GI+MG +G+ A ET++ LM +D++K+P +
Sbjct: 591 KQLKINFGKVAMVGDGINDAPALAAATVGIAMGGAGTDTAIETADVALMGDDLQKLPFTV 650
Query: 640 RLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+L+RKT + + +N+ S+ K L L I G+ +W+A++ D+G LLV LN + +++
Sbjct: 651 KLSRKTLQIIKQNITFSLVIKLIALLLVIPGWLTLWIAIMADMGATLLVTLNGLRLMK 708
>UniRef100_P30336 Probable cadmium-transporting ATPase [Bacillus pseudofirmus]
Length = 723
Score = 333 bits (854), Expect = 1e-89
Identities = 223/721 (30%), Positives = 384/721 (52%), Gaps = 49/721 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL 69
+ V G CA A E+ +K L GV+ V A + V + I E + A A ++
Sbjct: 16 YRVQGFTCANCAGKFEKNVKQLSGVEDAKVNFGASKIAVYGNAT-IEELEKAGAFENLKV 74
Query: 70 --EASFRPQG----ETNNEKKCP----DILTMACGLLLALSFLK-YIYPPLG----WLAL 114
E S R E E K P + LL+ +L Y+ L L
Sbjct: 75 TPEKSARQASQEVKEDTKEDKVPFYKKHSTLLYASLLITFGYLSSYVNGEENIVTTLLFL 134
Query: 115 GSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLET 174
S+ IG + + ++ ++ L+ +AV G A + ++ + +++ LF+I++ LE
Sbjct: 135 ASMFIGGLSLFKVGLQNLLRFEFDMKTLMTVAVIGGAIIGEWAEVAIVVILFAISEALER 194
Query: 175 RATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIVVEG 232
+ +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G I +DG+VV G
Sbjct: 195 FSMDRARQSIRSLMDIAPKEALVKRNGQEIMIHVDDIAVGDIMIVKPGQKIAMDGVVVSG 254
Query: 233 KCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVG 287
V++ +TGES PV K D+ V+AGT+N K++ DT +I + G
Sbjct: 255 YSAVNQTAITGESVPVEKTVDNEVFAGTLNEEGLLEVEITKLVEDTTISKIIHLVEEAQG 314
Query: 288 KRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVL 347
+R S ++F K K + ++ + +A+VP E W + L VL
Sbjct: 315 ERAPS--------QAFVDKF--AKYYTPIIMIIATLVAIVPPLFFDGSWETWIYQGLAVL 364
Query: 348 LSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
+ GCPCAL++STP++I A+ AA G+L+KGG YLE + +K +AFDKTGT+T+G +V
Sbjct: 365 VVGCPCALVISTPISIVSAIGNAAKKGVLVKGGVYLEEMGALKAIAFDKTGTLTKGVPAV 424
Query: 408 TDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEG 466
TD++ IN + LL I+++E +S HP+A A++ A +I VE+F + G+G
Sbjct: 425 TDYNVLNKQINEKELLSIITALEYRSQHPLASAIMKKAEEENITYSDVQVEDFSSITGKG 484
Query: 467 IFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN--------CEETLVGVF 518
I G ++G YIG+ ++ + + +++ ++T +N E+ ++ V
Sbjct: 485 IKGIVNGTTYYIGSPKLFKELLTNDFDKDLE---QNVTTLQNQGKTAMIIGTEKEILAVI 541
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + E +++L LG++ ++MLTGD+ A + Q+ + + AEL+P +K
Sbjct: 542 AVADEVRESSKEILQKLHQLGIKKTIMLTGDNKGTANAIGGQV--GVSDIEAELMPQDKL 599
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ + E G +AM+GDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+RK+P
Sbjct: 600 DFIKQLRSEYGNVAMVGDGVNDAPALAASTVGIAMGGAGTDTALETADVALMGDDLRKLP 659
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
++L+RKT + N+ ++ K L I G+ +W+A+L+D+G LLV LN + ++
Sbjct: 660 STVKLSRKTLNIIKANITFAIAIKFIASLLVIPGWLTLWIAILSDMGATLLVALNGLRLM 719
Query: 697 Q 697
+
Sbjct: 720 R 720
>UniRef100_Q8DZ61 Cation-transporting ATPase, E1-E2 family [Streptococcus agalactiae]
Length = 709
Score = 332 bits (852), Expect = 2e-89
Identities = 220/711 (30%), Positives = 380/711 (52%), Gaps = 43/711 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL 69
+ V G C A + E +K L GV+ V A V V I E + A A ++
Sbjct: 16 YRVQGFTCTNCAAIFENNVKELPGVQDAKVNFGASKV-YVKGTTTIEELEKAGAFENLKI 74
Query: 70 --EASFRPQGETNNEKKCPDILTMACGLLLALSFL-------KYIYPPLGWLALGSVAIG 120
E R GE ++K +I LLL +S+ +++ P +G+ A S+ IG
Sbjct: 75 RDEKEQRVGGEPFWKQK-ENIKVYISALLLVVSWFLGEQYGEEHVLPTIGYAA--SILIG 131
Query: 121 FPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKA 180
+ + + ++R L ++N L+ +A+ G A + ++ +G ++ LF+I++ LE + KA
Sbjct: 132 GYSLFIKGLKNLRRLNFDMNTLMTIAIIGAAIIGEWGEGATVVILFAISEALERYSMDKA 191
Query: 181 LVAMSSLTNMAPQMAIVAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDE 238
++ SL ++AP+ A++ E + V+++++ I+ VK G + +DGIVV+G +++
Sbjct: 192 RQSIESLMDIAPKEALIRRGNEEMMIHVDEIQVGDIMIVKPGQKLAMDGIVVKGTSTLNQ 251
Query: 239 KMLTGESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSG 293
+TGES PVTK ++ V+AGT+N ++ + DT +I + +R S
Sbjct: 252 AAITGESVPVTKITNDEVFAGTLNEEGLLEVKVTKRVEDTTLSKIIHLVEEAQAERAPSQ 311
Query: 294 KNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPC 353
V + + ++ +L IAVVP D W + L VL+ GCPC
Sbjct: 312 AFVDKFAKYYTPAIV----------ILALLIAVVPPLFG-GDWSQWIYQGLAVLVVGCPC 360
Query: 354 ALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAG 413
AL++STPVA+ A+ AA +G+L+KGG +LE +K +AFDKTGT+T+G +VTD
Sbjct: 361 ALVVSTPVAVVTAIGNAAKNGVLIKGGIHLEAAGHLKAIAFDKTGTLTKGIPAVTDIVTY 420
Query: 414 DDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDG 473
NE L+ S+IE S HP+A A++ A + +K VE+FQ+ G+G+ I+
Sbjct: 421 GRNENE-LITITSAIEKGSQHPLASAIMRKAEENGLKFNEVTVEDFQSITGKGVKAKINN 479
Query: 474 RDIYIGNKRV------GVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSG 527
Y+G++ + + + K ++Q Q + E+ ++ ++ D R
Sbjct: 480 EMYYVGSQNLFEELHGSISSDKKEKIADMQTQGKTVMVL--GTEKEILSFIAVADEMRES 537
Query: 528 ALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFK-KE 586
+ E + +L +G+ +VMLTGD+ + A + Q+ + + A+LLP +K I+ + K
Sbjct: 538 SKEVIGKLNNMGIETVMLTGDNQRTATAIGKQV--GVSDIKADLLPEDKLNFIKELREKH 595
Query: 587 GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTT 646
+ M+GDG+NDAPALA + +G++MG +G+ A ET++ LMS+D+ K+P I+L+RK
Sbjct: 596 QSVGMVGDGVNDAPALAASTVGVAMGGAGTDTALETADIALMSDDLSKLPYTIKLSRKAL 655
Query: 647 RKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+ +N+ S+ K L L + G+ +W+A+ D+G LLV LNS+ +L+
Sbjct: 656 AIIKQNITFSLAIKLVALLLVMPGWLTLWIAIFADMGATLLVTLNSLRLLK 706
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,278,070,003
Number of Sequences: 2790947
Number of extensions: 51604799
Number of successful extensions: 152495
Number of sequences better than 10.0: 2661
Number of HSP's better than 10.0 without gapping: 2351
Number of HSP's successfully gapped in prelim test: 310
Number of HSP's that attempted gapping in prelim test: 140811
Number of HSP's gapped (non-prelim): 7764
length of query: 833
length of database: 848,049,833
effective HSP length: 136
effective length of query: 697
effective length of database: 468,481,041
effective search space: 326531285577
effective search space used: 326531285577
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 79 (35.0 bits)
Medicago: description of AC130275.7


