

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.4 - phase: 0
(362 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
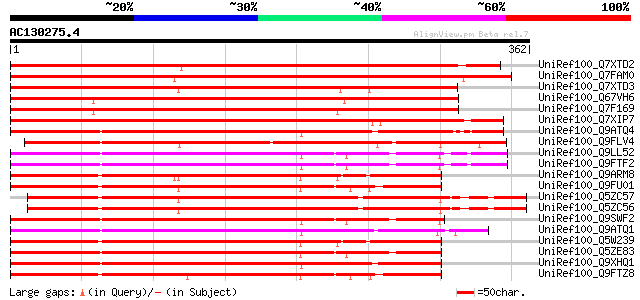
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XTD2 OSJNBb0022F16.13 protein [Oryza sativa] 412 e-113
UniRef100_Q7FAM0 OSJNBa0071I13.6 protein [Oryza sativa] 392 e-108
UniRef100_Q7XTD3 OSJNBb0022F16.12 protein [Oryza sativa] 389 e-107
UniRef100_Q67VH6 S-receptor kinase PK3-like [Oryza sativa] 369 e-101
UniRef100_Q7F169 S-receptor kinase PK3-like protein [Oryza sativa] 363 3e-99
UniRef100_Q7XIP7 Receptor-like kinase TAK33-like protein [Oryza ... 349 6e-95
UniRef100_Q9ATQ4 TAK33 [Triticum aestivum] 264 3e-69
UniRef100_Q9FLV4 Receptor-like protein kinase [Arabidopsis thali... 263 6e-69
UniRef100_Q9LL52 Receptor-like kinase [Oryza sativa] 261 3e-68
UniRef100_Q9FTF2 Putative rust resistance kinase Lr10 [Oryza sat... 261 3e-68
UniRef100_Q9ARM8 Putative rust resistance kinase Lr10 [Oryza sat... 260 4e-68
UniRef100_Q9FU01 Putative rust resistance kinase Lr10 [Oryza sat... 259 1e-67
UniRef100_Q5ZC57 Putative receptor serine/threonine kinase PR5K ... 258 1e-67
UniRef100_Q5ZC56 Putative receptor serine/threonine kinase PR5K ... 258 1e-67
UniRef100_Q9SWF2 Receptor kinase [Oryza sativa] 258 2e-67
UniRef100_Q9ATQ1 TAK14 [Triticum aestivum] 258 2e-67
UniRef100_Q5W239 Ser/Thr receptor-like kinase precursor [Zea mays] 258 2e-67
UniRef100_Q5ZE83 Putative rust resistance kinase Lr10 [Oryza sat... 258 2e-67
UniRef100_Q9XHQ1 Receptor-like kinase ARK1AS [Hordeum vulgare] 257 4e-67
UniRef100_Q9FTZ8 Putative rust resistance kinase Lr10 [Oryza sat... 256 5e-67
>UniRef100_Q7XTD2 OSJNBb0022F16.13 protein [Oryza sativa]
Length = 419
Score = 412 bits (1058), Expect = e-113
Identities = 200/345 (57%), Positives = 251/345 (71%), Gaps = 8/345 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
+D+FL+D+ REKP RFT + LR T +Y+ LG+GGFG VY+G F G VAVK+L +
Sbjct: 67 VDRFLDDILREKPARFTPENLREFTGDYAERLGAGGFGVVYRGRFPGGVQVAVKILHRTL 126
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-- 118
+++ +EQFMAEV T GR +H NLVRLYGFCF+ ALVYEY+ NGSLDR LF
Sbjct: 127 DRRAEEQFMAEVATAGRTYHINLVRLYGFCFDATTKALVYEYLENGSLDRVLFDAAAAAA 186
Query: 119 LGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRE 178
L ++ LH I +GTARG+ YLHEECQHRIIHYDIKPGN+LL ++ PKVADFGLAK C+R+
Sbjct: 187 LEFDTLHGIVVGTARGVRYLHEECQHRIIHYDIKPGNVLLAGDYAPKVADFGLAKLCSRD 246
Query: 179 NTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKN-TESQEW 237
NTH+TMTG RGTPGYAAPELW+P P+THKCDVYSFGML+FEI+GRRRNL + ESQEW
Sbjct: 247 NTHLTMTGARGTPGYAAPELWLPLPVTHKCDVYSFGMLVFEILGRRRNLDTQRPAESQEW 306
Query: 238 FPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+P W W++ D G GE M GI K+ E AERM KVALWC+QY+PE RP MS VV+MLE
Sbjct: 307 YPRWAWQRFDQGRFGEVMAASGIRSKDGEKAERMCKVALWCIQYQPEARPSMSSVVRMLE 366
Query: 298 GSLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSS 342
G +I + NPF ++ T ++ S++ VS+ + +S
Sbjct: 367 GEEQIARPVNPFAYMA-----TMDAISSSSSGGGGVSTATSASAS 406
>UniRef100_Q7FAM0 OSJNBa0071I13.6 protein [Oryza sativa]
Length = 430
Score = 392 bits (1007), Expect = e-108
Identities = 198/360 (55%), Positives = 253/360 (70%), Gaps = 10/360 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
++KFLN++ EKP+RFT +QL T NYS+ LGSGGFG VY+G G VAVKVL+ S
Sbjct: 69 VEKFLNEILSEKPMRFTSEQLAACTGNYSSELGSGGFGVVYRGELPKGLQVAVKVLKVSM 128
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF-----HE 115
NKK+ E FMAE+GTIGR +H +LVRLYGFCF+ + ALVYE++ NGSL++YL+
Sbjct: 129 NKKVQEAFMAEIGTIGRTYHVHLVRLYGFCFDADTKALVYEFLENGSLEKYLYGGGGEDR 188
Query: 116 TKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
K L + LH+IA+GTA+GI YLHEECQ RI+HYDIKP NILL +F PKVADFGLA+
Sbjct: 189 GKKLEWRTLHDIAVGTAKGIRYLHEECQQRIVHYDIKPANILLTADFTPKVADFGLARLG 248
Query: 176 NRENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAI-KNTES 234
RENTH+++TGGRGTPGYAAPELWM P T KCDVYSFGM+LFE++GRRRN + ES
Sbjct: 249 ERENTHMSLTGGRGTPGYAAPELWMALPATEKCDVYSFGMVLFEVLGRRRNYDLAAQAES 308
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
QEWFP WVW + + G + + GI E+++ AE M KVALWCVQ++P RP MS VV+
Sbjct: 309 QEWFPKWVWDRYEQGDMECVVSAAGIGEEDRAKAEMMCKVALWCVQFQPSARPKMSSVVR 368
Query: 295 MLEGSLEIPKTFNPFQHLIDG----TEFTTHSVQESNTYTTSVSSVMVSDSSIVCATPIM 350
MLEG + I NPF +++ G + T+ S S+ TT+ SS + +T +M
Sbjct: 369 MLEGEMAIVPPVNPFHYVMSGGSGSSTLTSSSTNLSSGGTTTGSSEVAVSLPAKKSTDVM 428
>UniRef100_Q7XTD3 OSJNBb0022F16.12 protein [Oryza sativa]
Length = 431
Score = 389 bits (1000), Expect = e-107
Identities = 190/327 (58%), Positives = 237/327 (72%), Gaps = 15/327 (4%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
M F+ ++ E+P+RF+ +QLR T +Y++ +GSGGFG VY+G+F +G VAVKVL +
Sbjct: 77 MSHFIEGLQNERPVRFSARQLRAFTKSYAHKVGSGGFGVVYRGVFPSGAPVAVKVLNSTL 136
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---- 116
K+ +EQFMAEVGTIGR +H NLVRLYGFCF+ ++ ALVYEYM GSLDRYLF +
Sbjct: 137 GKRAEEQFMAEVGTIGRTYHINLVRLYGFCFDADVKALVYEYMEKGSLDRYLFDSSPSPA 196
Query: 117 -KVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
+ +G+EKLHEIA+GTA+ + YLHEEC RIIHYDIKP N+LL PKV+DFGLAK C
Sbjct: 197 AERIGFEKLHEIAVGTAKAVRYLHEECAQRIIHYDIKPENVLLGAGMAPKVSDFGLAKLC 256
Query: 176 NRENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAI------ 229
+RE+TH+T+TG RGTPGYAAPELWMP P+THKCDVYS+GMLLFE++GRRRNL +
Sbjct: 257 DREDTHLTITGARGTPGYAAPELWMPLPVTHKCDVYSYGMLLFEMLGRRRNLELGAGAGA 316
Query: 230 KNTESQEWFPIWVWKKKDAG----LLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPEL 285
SQEW+P WVW + +AG +L A + +E AER+ VALWCVQYRPE
Sbjct: 317 HGHGSQEWYPRWVWHRFEAGETEAVLARATAAAAGGGREREKAERVCMVALWCVQYRPED 376
Query: 286 RPIMSVVVKMLEGSLEIPKTFNPFQHL 312
RP M VV+MLEG I NPF HL
Sbjct: 377 RPSMGNVVRMLEGEDHIAAPRNPFAHL 403
>UniRef100_Q67VH6 S-receptor kinase PK3-like [Oryza sativa]
Length = 444
Score = 369 bits (947), Expect = e-101
Identities = 183/321 (57%), Positives = 231/321 (71%), Gaps = 8/321 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLR--- 57
+++FL ++ EKPIRFT QQL T+NYS LG+GGFGTVYKG+ NG VAVK L
Sbjct: 83 VERFLKEIAGEKPIRFTAQQLAGFTNNYSARLGAGGFGTVYKGMLPNGLTVAVKRLHVGG 142
Query: 58 -GSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET 116
G EQFMAEVG++GRIHH NLVRL+GFCF+ ++ ALVYEYM NG+LD YLF +
Sbjct: 143 HGDGWSTSQEQFMAEVGSVGRIHHINLVRLFGFCFDADVRALVYEYMDNGALDAYLFDRS 202
Query: 117 KVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCN 176
+ + IA+G ARG+ YLHEECQH+I+HYDIKPGN+LLD PKVADFGLA+ +
Sbjct: 203 RAVAVATRRAIAVGVARGLRYLHEECQHKIVHYDIKPGNVLLDGGLTPKVADFGLARLAS 262
Query: 177 RENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNT--ES 234
R +TH++++G RGTPGYAAPE+WM +T KCDVYSFG+LLFEI+ RRRNL
Sbjct: 263 RGDTHVSVSGMRGTPGYAAPEMWMQAGVTEKCDVYSFGVLLFEIVRRRRNLDDGGAPGSQ 322
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVC-GIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVV 293
Q+WFP+ W K +AG L EA+ C ++++ +E ERM KVA WCVQ +PE RP MS VV
Sbjct: 323 QQWFPMLAWSKHEAGHLAEAIEGCDAMDKQERETVERMCKVAFWCVQQQPEARPPMSAVV 382
Query: 294 KMLEGSLEI-PKTFNPFQHLI 313
+MLEG ++I NPFQHL+
Sbjct: 383 RMLEGEVDIDAPPVNPFQHLV 403
>UniRef100_Q7F169 S-receptor kinase PK3-like protein [Oryza sativa]
Length = 444
Score = 363 bits (933), Expect = 3e-99
Identities = 181/321 (56%), Positives = 230/321 (71%), Gaps = 8/321 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLR--- 57
+++FL ++ EKPIRFT QQL T+NYS LG+GGFGTVYKG+ NG VAVK L
Sbjct: 83 VERFLKEIAGEKPIRFTAQQLAGFTNNYSARLGAGGFGTVYKGMLPNGLTVAVKRLHVGG 142
Query: 58 -GSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET 116
G EQFMAEVG++GRIHH NLVRL+GFCF+ ++ ALVYEYM NG+LD YLF +
Sbjct: 143 HGDGWSTSQEQFMAEVGSVGRIHHINLVRLFGFCFDADVRALVYEYMDNGALDAYLFDRS 202
Query: 117 KVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCN 176
+ + IA+G ARG+ YLHEECQH+I+HYDIKPGN+LLD PKVADFGLA+ +
Sbjct: 203 RAVPVATRRAIAVGVARGLRYLHEECQHKIVHYDIKPGNVLLDGGLTPKVADFGLARLAS 262
Query: 177 RENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNL--AIKNTES 234
R +TH++++G RGTPGYAAPE+WM +T KCDVYSFG+ LFEI+ RRRNL +
Sbjct: 263 RGDTHVSVSGMRGTPGYAAPEMWMQAGVTEKCDVYSFGVHLFEIVRRRRNLDDGGEPGSQ 322
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVC-GIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVV 293
+WFP+ W K +AG L EA+ C ++++ +E ERM KVA WCVQ +PE RP MS VV
Sbjct: 323 HQWFPMLAWSKHEAGHLAEAIEGCDAMDKQERETVERMCKVAFWCVQQQPEARPPMSAVV 382
Query: 294 KMLEGSLEI-PKTFNPFQHLI 313
+MLEG ++I NPFQHL+
Sbjct: 383 RMLEGEVDIDAPPVNPFQHLV 403
>UniRef100_Q7XIP7 Receptor-like kinase TAK33-like protein [Oryza sativa]
Length = 447
Score = 349 bits (896), Expect = 6e-95
Identities = 188/370 (50%), Positives = 237/370 (63%), Gaps = 30/370 (8%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
+++FL +M EKPIRFT +QL T YS LG+G FGTVY G NG VAVKVLRG
Sbjct: 72 VERFLWEMAHEKPIRFTPRQLAGFTRGYSARLGAGVFGTVYGGALPNGLAVAVKVLRGGM 131
Query: 61 NKK-IDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
+++ +EQFMAEVGTIGR HH NLVRL+GFC++ + ALVYEYMGNG+LD YLF ++ +
Sbjct: 132 DRRRSEEQFMAEVGTIGRTHHINLVRLFGFCYDAAVRALVYEYMGNGALDAYLFDLSRDV 191
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
G IAIG ARG+ YLHEEC+H+I+HYDIKPGN+LLD PKVADFGLA+ NR +
Sbjct: 192 GVPARRAIAIGVARGLRYLHEECEHKIVHYDIKPGNVLLDGGMTPKVADFGLARLVNRGD 251
Query: 180 THITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFP 239
TH++++G RGTPGYAAPE M +T KCDVYSFGMLL +I+GRRRN ESQ+W+P
Sbjct: 252 THVSVSGMRGTPGYAAPETLMQSGVTEKCDVYSFGMLLLKIVGRRRNFDEAAPESQQWWP 311
Query: 240 IWVWKKKDAGLL----------------------GEAMIV---CGIEEKNKEIAERMIKV 274
+ W + + G L GEA++ E + KE RM +V
Sbjct: 312 MEAWARYERGELMMVDDAAAAINHPSDEICSGSDGEAVVTVAEADDERRCKEAVVRMYQV 371
Query: 275 ALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVS 334
A WCVQ RPE RP M VVKMLEG +++ NPF HL+ V TT+ S
Sbjct: 372 AFWCVQQRPEARPPMGAVVKMLEGEMDVAPPVNPFLHLMAAPA----PVPNPWATTTASS 427
Query: 335 SVMVSDSSIV 344
VS++ +V
Sbjct: 428 GNAVSENVVV 437
>UniRef100_Q9ATQ4 TAK33 [Triticum aestivum]
Length = 708
Score = 264 bits (675), Expect = 3e-69
Identities = 155/349 (44%), Positives = 215/349 (61%), Gaps = 14/349 (4%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIF-SNGTMVAVKVLRGS 59
++KFL + P R++ + T ++ + LG GG+G+VYKG+ G VAVK+L G+
Sbjct: 363 VEKFLRIQQMIGPARYSYTDIVAVTSHFRDKLGQGGYGSVYKGVLLPGGVHVAVKMLEGN 422
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
SN E F++EV TIGRIHH N+VRL GFC E L ALVYEYM NGSLD+Y+F K
Sbjct: 423 SNCN-GEDFISEVSTIGRIHHVNVVRLVGFCSEEMLRALVYEYMPNGSLDKYIFSTEKSF 481
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
+EKL+EIA+G ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R +
Sbjct: 482 SWEKLNEIALGIARGINYLHQGCDMQILHFDIKPHNILLDSNFVPKVADFGLAKLYPRGD 541
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNL-AIKNTESQE 236
+ + ++ RGT GY APE+ I+ K DVYSFGMLL E+ G RRN + SQ
Sbjct: 542 SFVPLSAMRGTVGYIAPEMISRSFGVISSKSDVYSFGMLLLEMAGGRRNADPNMGSSSQA 601
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
++P WV+ + GE + + E+ +++ V LWC+Q R RP MS V+++L
Sbjct: 602 YYPSWVYDRLTQEEAGE---ISDVAADMHELEKKLCVVGLWCIQMRSRDRPTMSEVIEIL 658
Query: 297 EGSLE-IPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIV 344
E ++ + PF D E H V++S +T+ +S+V + S+V
Sbjct: 659 EAGVDGLQMPSRPF--FCD--EGHIH-VEDSYQFTSELSAVSEEELSVV 702
>UniRef100_Q9FLV4 Receptor-like protein kinase [Arabidopsis thaliana]
Length = 872
Score = 263 bits (672), Expect = 6e-69
Identities = 158/355 (44%), Positives = 217/355 (60%), Gaps = 23/355 (6%)
Query: 11 EKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMA 70
+ P+ FT + L+ T+N+S LLGSGGFGTVYKG + T+VAVK L + + + +F+
Sbjct: 515 DSPVSFTYRDLQNCTNNFSQLLGSGGFGTVYKGTVAGETLVAVKRLDRALSHG-EREFIT 573
Query: 71 EVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK---VLGYEKLHEI 127
EV TIG +HH NLVRL G+C E + LVYEYM NGSLD+++F + +L + EI
Sbjct: 574 EVNTIGSMHHMNLVRLCGYCSEDSHRLLVYEYMINGSLDKWIFSSEQTANLLDWRTRFEI 633
Query: 128 AIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGG 187
A+ TA+GIAY HE+C++RIIH DIKP NILLD NF PKV+DFGLAK RE++H+ +T
Sbjct: 634 AVATAQGIAYFHEQCRNRIIHCDIKPENILLDDNFCPKVSDFGLAKMMGREHSHV-VTMI 692
Query: 188 RGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKD 247
RGT GY APE PIT K DVYS+GMLL EI+G RRNL + ++P W +K+
Sbjct: 693 RGTRGYLAPEWVSNRPITVKADVYSYGMLLLEIVGGRRNLDMSYDAEDFFYPGWAYKELT 752
Query: 248 AGLLGEAM--IVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEI 302
G +A+ + G+ E+ + + + +KVA WC+Q +RP M VVK+LEG+ + +
Sbjct: 753 NGTSLKAVDKRLQGVAEEEEVV--KALKVAFWCIQDEVSMRPSMGEVVKLLEGTSDEINL 810
Query: 303 PKTFNPFQHLI-DGTEFTTHSVQES----------NTYTTSVSSVMVSDSSIVCA 346
P LI +G E +++ NT TTS S S S C+
Sbjct: 811 PPMPQTILELIEEGLEDVYRAMRREFNNQLSSLTVNTITTSQSYRSSSRSHATCS 865
>UniRef100_Q9LL52 Receptor-like kinase [Oryza sativa]
Length = 633
Score = 261 bits (666), Expect = 3e-68
Identities = 155/356 (43%), Positives = 215/356 (59%), Gaps = 21/356 (5%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGS 59
++KFL + P R+ L T ++ + LG GG+G+VYKG+ +G + VAVK+L G+
Sbjct: 289 VEKFLQMQQVLGPTRYAYTDLTAVTSHFRDKLGQGGYGSVYKGVLLSGDVHVAVKMLNGT 348
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
S E+F++EV TIGRIHH N+VRL GFC E ALVYEYM GSLD+Y+F +
Sbjct: 349 STYD-GEEFISEVSTIGRIHHVNVVRLVGFCSEELRRALVYEYMPQGSLDKYIFSSERSF 407
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIAIG ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R
Sbjct: 408 SWDKLNEIAIGIARGINYLHQGCDMQILHFDIKPHNILLDDNFVPKVADFGLAKLYPRNK 467
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTE---S 234
+ ++ RGT GY APE+ I+ KCDVYSFGMLL E+ G RRN A NT S
Sbjct: 468 SFVSDRALRGTVGYIAPEMVSRSFGVISSKCDVYSFGMLLLEMAGGRRN-ADPNTNPNAS 526
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
Q ++P WV+ + +GE + E K ++ V LWC+Q + RP MS ++
Sbjct: 527 QSYYPSWVYGQLTGEQVGETSGAADMHELQK----KLCLVGLWCIQMKSHDRPTMSETIE 582
Query: 295 MLEG---SLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIVCAT 347
MLEG +L++P P DG +V +S +++ ++++ D +I A+
Sbjct: 583 MLEGDVNALQVP----PRPFFCDGDFMP--NVMDSYLHSSELTAISEDDGAIEFAS 632
>UniRef100_Q9FTF2 Putative rust resistance kinase Lr10 [Oryza sativa]
Length = 672
Score = 261 bits (666), Expect = 3e-68
Identities = 155/356 (43%), Positives = 215/356 (59%), Gaps = 21/356 (5%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGS 59
++KFL + P R+ L T ++ + LG GG+G+VYKG+ +G + VAVK+L G+
Sbjct: 328 VEKFLQMQQVLGPTRYAYTDLTAVTSHFRDKLGQGGYGSVYKGVLLSGDVHVAVKMLNGA 387
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
S E+F++EV TIGRIHH N+VRL GFC E ALVYEYM GSLD+Y+F +
Sbjct: 388 STYD-GEEFISEVSTIGRIHHVNVVRLVGFCSEELRRALVYEYMPQGSLDKYIFSSERSF 446
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIAIG ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R
Sbjct: 447 SWDKLNEIAIGIARGINYLHQGCDMQILHFDIKPHNILLDDNFVPKVADFGLAKLYPRNK 506
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTE---S 234
+ ++ RGT GY APE+ I+ KCDVYSFGMLL E+ G RRN A NT S
Sbjct: 507 SFVSDRALRGTVGYIAPEMVSRSFGVISSKCDVYSFGMLLLEMAGGRRN-ADPNTNPNAS 565
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
Q ++P WV+ + +GE + E K ++ V LWC+Q + RP MS ++
Sbjct: 566 QSYYPSWVYGQLTGEQVGETSGAADMHELQK----KLCLVGLWCIQMKSHDRPTMSETIE 621
Query: 295 MLEG---SLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIVCAT 347
MLEG +L++P P DG +V +S +++ ++++ D +I A+
Sbjct: 622 MLEGDVNALQVP----PRPFFCDGDFMP--NVMDSYLHSSELTAISEDDGAIEFAS 671
>UniRef100_Q9ARM8 Putative rust resistance kinase Lr10 [Oryza sativa]
Length = 630
Score = 260 bits (665), Expect = 4e-68
Identities = 141/310 (45%), Positives = 200/310 (64%), Gaps = 14/310 (4%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
++ FL KP R+T Q++ T + +G GGFGTVYKG NG VAVK+L +
Sbjct: 308 VEMFLRTYGTSKPTRYTFSQVKKITRRFKEKVGQGGFGTVYKGKLLNGVPVAVKMLENPT 367
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF---HET- 116
E F+ EV TIGRIHH N++ L GFC E AL+YE+M N SL++Y+F H T
Sbjct: 368 GD--GEDFITEVATIGRIHHANIIHLLGFCSEGTRRALIYEFMPNESLEKYIFLHDHNTP 425
Query: 117 -KVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
++L K+ +IA+G ARG+ YLH+ C RI+H+DIKP NILLD NF PK++DFGLAK C
Sbjct: 426 QELLSPNKMLDIALGIARGMEYLHQGCNQRILHFDIKPHNILLDYNFSPKISDFGLAKLC 485
Query: 176 NRENTHITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNL--AIKN 231
R+ + +TMT RGT GY APEL+ I++K DVYSFGML+ E++ RR+ +IKN
Sbjct: 486 PRDQSIVTMTKARGTMGYIAPELYSRNFGEISYKSDVYSFGMLVLEMVSGRRSWDPSIKN 545
Query: 232 TESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSV 291
+++ +FP W+++K G E ++ + E+ K++ ++ VALWC+Q+ P RP M+
Sbjct: 546 -QNEVYFPEWIYEKVITG--QEFVLSREMTEEEKQMVRQLALVALWCIQWNPRNRPSMTK 602
Query: 292 VVKMLEGSLE 301
VV M+ G L+
Sbjct: 603 VVNMITGRLQ 612
>UniRef100_Q9FU01 Putative rust resistance kinase Lr10 [Oryza sativa]
Length = 635
Score = 259 bits (661), Expect = 1e-67
Identities = 143/313 (45%), Positives = 202/313 (63%), Gaps = 20/313 (6%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
++ FL KP R+T +++ + +G GGFG+VY+G NG VAVK+L S
Sbjct: 313 VEMFLKTYGTSKPTRYTFSEVKKIARRFKVKVGQGGFGSVYRGELPNGVPVAVKMLENSE 372
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF-HET--- 116
+ ++F+ EV TIGRIHH N+VRL GFC E AL+YEYM N SL++Y+F H++
Sbjct: 373 GE--GDEFINEVATIGRIHHANIVRLLGFCSEGTRRALIYEYMPNDSLEKYVFSHDSDTS 430
Query: 117 -KVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
+VL K+ +IAIG ARG+ YLH+ C RI+H+DIKP NILLD NF PK++DFGLAK C
Sbjct: 431 QEVLVPSKMLDIAIGIARGMEYLHQGCNQRILHFDIKPNNILLDYNFSPKISDFGLAKLC 490
Query: 176 NRENTHITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE 233
R+ + +T+T RGT GY APEL+ I++K DVYSFGML+ E++ RRN + + E
Sbjct: 491 ARDQSIVTLTAARGTMGYIAPELYSRNFGEISYKSDVYSFGMLVLEMVSGRRN-SDPSVE 549
Query: 234 SQE--WFPIWVWKKKDAG---LLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPI 288
SQ +FP W++++ ++G LG M ++ KE ++ VALWC+Q+ P+ RP
Sbjct: 550 SQNVVYFPEWIYEQVNSGQDLALGREM-----TQEEKETVRQLAIVALWCIQWNPKNRPS 604
Query: 289 MSVVVKMLEGSLE 301
M+ VV ML G L+
Sbjct: 605 MTKVVNMLTGRLQ 617
>UniRef100_Q5ZC57 Putative receptor serine/threonine kinase PR5K [Oryza sativa]
Length = 674
Score = 258 bits (660), Expect = 1e-67
Identities = 149/356 (41%), Positives = 219/356 (60%), Gaps = 23/356 (6%)
Query: 13 PIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
P R+ ++ T + LG GG+G V+KG +G +VAVK L S E+F+ EV
Sbjct: 322 PTRYKYSEVTKITSFLNYKLGEGGYGVVFKGRLQDGRLVAVKFLHDSKGN--GEEFVNEV 379
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET--KVLGYEKLHEIAIG 130
+IGR H N+V L+GFC E + AL+YEYM NGSLD Y++ E ++LG+EKL+ IAIG
Sbjct: 380 MSIGRTSHINIVSLFGFCLEGSKRALLYEYMPNGSLDDYIYSENPKEILGWEKLYGIAIG 439
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
ARG+ YLH C RIIH+DIKP NILLD++F PK+ADFGLAK C + + ++MTG RGT
Sbjct: 440 IARGLEYLHHSCNTRIIHFDIKPQNILLDQDFCPKIADFGLAKLCRTKESKLSMTGARGT 499
Query: 191 PGYAAPE-LWMPFPI-THKCDVYSFGMLLFEIIGRRRNL-AIKNTESQEWFPIWVWKKKD 247
G+ APE ++ F I + K DVYS+GM+L E++G R+N ++ S+++FP W++ D
Sbjct: 500 IGFIAPEVIYRSFGIVSTKSDVYSYGMMLLEMVGGRKNAKSMVENSSEKYFPDWIY---D 556
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEIPK 304
L + + C + + ++IA++M + LWCVQ P RP ++ V+ M E S LE+P
Sbjct: 557 HFALDDGLQACEVTSEVEQIAKKMTLIGLWCVQVLPMHRPTITQVLDMFERSLDELEMPP 616
Query: 305 TFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIVCATPIMRKYEIELASS 360
N F L++ H QE NT +TS + ++ + + ++R E L +S
Sbjct: 617 KQN-FSELLE------HPAQEINTESTSST---INTKAAQALSEVLRVEETSLVNS 662
>UniRef100_Q5ZC56 Putative receptor serine/threonine kinase PR5K [Oryza sativa]
Length = 419
Score = 258 bits (660), Expect = 1e-67
Identities = 149/356 (41%), Positives = 219/356 (60%), Gaps = 23/356 (6%)
Query: 13 PIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
P R+ ++ T + LG GG+G V+KG +G +VAVK L S E+F+ EV
Sbjct: 67 PTRYKYSEVTKITSFLNYKLGEGGYGVVFKGRLQDGRLVAVKFLHDSKGN--GEEFVNEV 124
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET--KVLGYEKLHEIAIG 130
+IGR H N+V L+GFC E + AL+YEYM NGSLD Y++ E ++LG+EKL+ IAIG
Sbjct: 125 MSIGRTSHINIVSLFGFCLEGSKRALLYEYMPNGSLDDYIYSENPKEILGWEKLYGIAIG 184
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
ARG+ YLH C RIIH+DIKP NILLD++F PK+ADFGLAK C + + ++MTG RGT
Sbjct: 185 IARGLEYLHHSCNTRIIHFDIKPQNILLDQDFCPKIADFGLAKLCRTKESKLSMTGARGT 244
Query: 191 PGYAAPE-LWMPFPI-THKCDVYSFGMLLFEIIGRRRNL-AIKNTESQEWFPIWVWKKKD 247
G+ APE ++ F I + K DVYS+GM+L E++G R+N ++ S+++FP W++ D
Sbjct: 245 IGFIAPEVIYRSFGIVSTKSDVYSYGMMLLEMVGGRKNAKSMVENSSEKYFPDWIY---D 301
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEIPK 304
L + + C + + ++IA++M + LWCVQ P RP ++ V+ M E S LE+P
Sbjct: 302 HFALDDGLQACEVTSEVEQIAKKMTLIGLWCVQVLPMHRPTITQVLDMFERSLDELEMPP 361
Query: 305 TFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIVCATPIMRKYEIELASS 360
N F L++ H QE NT +TS + ++ + + ++R E L +S
Sbjct: 362 KQN-FSELLE------HPAQEINTESTSST---INTKAAQALSEVLRVEETSLVNS 407
>UniRef100_Q9SWF2 Receptor kinase [Oryza sativa]
Length = 608
Score = 258 bits (659), Expect = 2e-67
Identities = 147/312 (47%), Positives = 196/312 (62%), Gaps = 15/312 (4%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGS 59
++KFL + P R+ L T ++ + LG GG+G+VYKG+ +G + VAVK+L G+
Sbjct: 290 VEKFLQMQQVLGPTRYAYTDLTAVTSHFRDKLGQGGYGSVYKGVLLSGDVHVAVKMLNGT 349
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
S E+F++EV TIGRIHH N+VRL GFC E ALVYEYM GSLD+Y+F +
Sbjct: 350 STYD-GEEFISEVSTIGRIHHVNVVRLVGFCSEELRRALVYEYMPQGSLDKYIFSSERSF 408
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIAIG ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R
Sbjct: 409 SWDKLNEIAIGIARGINYLHQGCDMQILHFDIKPHNILLDDNFVPKVADFGLAKLYPRNK 468
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTE---S 234
+ ++ RGT GY APE+ I+ KCDVYSFGMLL E+ G RRN A NT S
Sbjct: 469 SFVSDRALRGTVGYIAPEMVSRSFGVISSKCDVYSFGMLLLEMAGGRRN-ADPNTNPNAS 527
Query: 235 QEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
Q ++P WV+ + +GE + E K ++ V LWC+Q + RP MS ++
Sbjct: 528 QSYYPSWVYGQLTGEQVGETSGAADMHELQK----KLCLVGLWCIQMKSHDRPTMSETIE 583
Query: 295 MLEG---SLEIP 303
MLEG +L++P
Sbjct: 584 MLEGDVNALQVP 595
>UniRef100_Q9ATQ1 TAK14 [Triticum aestivum]
Length = 689
Score = 258 bits (659), Expect = 2e-67
Identities = 152/347 (43%), Positives = 209/347 (59%), Gaps = 18/347 (5%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGS 59
++KFL + P R++ + T ++ + LG GG+GTVYKG+ G + VAVK+L G+
Sbjct: 342 VEKFLRIQQMIGPTRYSYTDIVAITSHFRDKLGQGGYGTVYKGVLLPGDVHVAVKMLEGN 401
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
SN E F++EV TIGRIHH N+VRL GFC E ALVYEYM +GSLD+Y+F K
Sbjct: 402 SNCN-GEDFISEVSTIGRIHHVNVVRLVGFCSEEMRRALVYEYMPHGSLDKYIFSAEKSF 460
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIA+G ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R +
Sbjct: 461 SWDKLNEIALGIARGINYLHQGCDMQILHFDIKPHNILLDSNFVPKVADFGLAKLYPRGD 520
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNL-AIKNTESQE 236
+ + ++ RGT GY APE+ I+ K DVYSFGMLL E+ G RRN + SQ
Sbjct: 521 SFVPLSAMRGTVGYIAPEMISRSFGVISSKSDVYSFGMLLLEMAGGRRNADPNAGSSSQA 580
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
++P WV+ + GE + + E+ +++ V LWC+Q R RP M V+++L
Sbjct: 581 YYPSWVYDQLTREEAGEE--ISPVAADMHELEKKLCVVGLWCIQMRSSDRPTMGEVIEIL 638
Query: 297 E---GSLEIPKTFNPF------QHLIDGTEFTTHSVQESNTYTTSVS 334
E G L++P PF H+ D FT+ S ++VS
Sbjct: 639 EAGAGGLQMPS--RPFFCDEGHIHVEDSYHFTSELTAVSEEELSAVS 683
>UniRef100_Q5W239 Ser/Thr receptor-like kinase precursor [Zea mays]
Length = 607
Score = 258 bits (659), Expect = 2e-67
Identities = 138/305 (45%), Positives = 196/305 (64%), Gaps = 9/305 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
++ FL KP R+T +++ + +G GGFGTVYKG NG VAVK+L S+
Sbjct: 298 VEMFLKTYGTSKPTRYTFSEVKKIARRFKEKVGQGGFGTVYKGQLPNGVPVAVKMLENST 357
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLG 120
+ E F+ EV TIG+IHH N+VRL GFC E AL+YE+M N SL RY+F ++L
Sbjct: 358 GE--GEDFINEVATIGQIHHANIVRLLGFCSEGTRRALIYEFMPNESLGRYIFLPQELLV 415
Query: 121 YEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENT 180
EK+ +IA G ARG+ YLH+ C RI+H+DIKP NILLD +F PK++DFGLAK C R+ +
Sbjct: 416 PEKMLDIATGIARGMEYLHQGCNQRILHFDIKPHNILLDYSFNPKISDFGLAKLCARDQS 475
Query: 181 HITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNL--AIKNTESQE 236
+T+T RGT GY APE++ P +++K DVYSFGML+ E++ RRN I+N ++
Sbjct: 476 IVTLTAARGTMGYIAPEIYSPNFGGVSYKSDVYSFGMLVLEMVSGRRNSDPGIEN-QNGV 534
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
+ P WV+++ G + + I ++ KE ++ VALWC+Q+ P+ RP M+ VV ML
Sbjct: 535 YLPEWVYERVVTG--QDLTLSKKIADQEKETVRQLAIVALWCIQWNPKNRPSMTKVVNML 592
Query: 297 EGSLE 301
G L+
Sbjct: 593 TGRLQ 597
>UniRef100_Q5ZE83 Putative rust resistance kinase Lr10 [Oryza sativa]
Length = 677
Score = 258 bits (658), Expect = 2e-67
Identities = 145/306 (47%), Positives = 196/306 (63%), Gaps = 11/306 (3%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGS 59
++KFL + P R++ + T +Y + LG GG+G+VYKG+ G + VA+K+L+G
Sbjct: 333 VEKFLQLQQMLTPTRYSYTDIIAITSHYRDKLGQGGYGSVYKGVLLPGDVRVAIKMLKGD 392
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
+N K E+F++EV TIGRIHH N+VRL GFC E ALVYEYM GSLD+Y+F K
Sbjct: 393 ANCK-GEEFISEVSTIGRIHHVNVVRLVGFCSEEIRRALVYEYMPQGSLDKYIFSSEKSF 451
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIA+G ARGI YLH C +I+H+DIKP NILLD NF PKVADFGLAK R+
Sbjct: 452 SWDKLNEIALGIARGINYLHHGCDMQILHFDIKPHNILLDNNFVPKVADFGLAKLYPRDK 511
Query: 180 THITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE--SQ 235
+ + ++ RGT GY APE+ I+ K DVYSFGMLL E+ G RRN A N E SQ
Sbjct: 512 SFVPVSAARGTVGYIAPEMISRGFGAISSKSDVYSFGMLLLEMAGGRRN-ADPNAENSSQ 570
Query: 236 EWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKM 295
++P V+++ GE + E K ++ V LWC+Q R RP+MS V++M
Sbjct: 571 AYYPSRVYRQLTRQETGEITAAADMHELEK----KLCIVGLWCIQMRSCDRPMMSEVIEM 626
Query: 296 LEGSLE 301
LEG ++
Sbjct: 627 LEGGVD 632
>UniRef100_Q9XHQ1 Receptor-like kinase ARK1AS [Hordeum vulgare]
Length = 669
Score = 257 bits (656), Expect = 4e-67
Identities = 141/305 (46%), Positives = 193/305 (63%), Gaps = 8/305 (2%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIF-SNGTMVAVKVLRGS 59
++KFL + P R+ + T ++ + LG GG+G+VYKG+ G VAVK+L G+
Sbjct: 349 VEKFLRVQQLIGPTRYAYTDIIAITRHFRDNLGQGGYGSVYKGVLLPGGVHVAVKMLEGN 408
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL 119
SN E F++EV TIGRIHH N+VRL GFC E +ALVYEYM NGSLD+Y+F K
Sbjct: 409 SNCN-GEDFISEVSTIGRIHHVNIVRLVGFCSEEMRMALVYEYMPNGSLDKYIFSAEKSF 467
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++KL+EIA+G ARGI YLH+ C +I+H+DIKP NILLD NF PKVADFGLAK R +
Sbjct: 468 SWDKLNEIALGVARGINYLHQGCDMQILHFDIKPHNILLDSNFVPKVADFGLAKLYPRGD 527
Query: 180 THITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNL-AIKNTESQE 236
+ + ++ RGT GY APE+ I+ K DVYSFGMLL E+ G RRN + SQ
Sbjct: 528 SFVPLSAMRGTVGYIAPEMISRSFGVISSKSDVYSFGMLLLEMAGGRRNADPNAGSSSQA 587
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
++P WV+ + +GE + + E+ + + V LWC+Q R RP MS V+++L
Sbjct: 588 YYPSWVYDQLTREEVGE---ISPVAADMHELEKNLCVVGLWCIQMRSRDRPTMSEVIEIL 644
Query: 297 EGSLE 301
E ++
Sbjct: 645 EAGVD 649
>UniRef100_Q9FTZ8 Putative rust resistance kinase Lr10 [Oryza sativa]
Length = 636
Score = 256 bits (655), Expect = 5e-67
Identities = 142/313 (45%), Positives = 197/313 (62%), Gaps = 20/313 (6%)
Query: 1 MDKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSS 60
++ FL KP R+T +++ + + +G GGFG+VY+G NG VAVK+L S
Sbjct: 305 VEMFLKTYGTSKPTRYTFSEVKKISRRFKVKVGQGGFGSVYRGELPNGVPVAVKMLENSE 364
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLG 120
+ ++F+ EV TIGRIHH N+VRL GFC E AL+YEYM N SL++Y+F +
Sbjct: 365 GE--GDEFINEVATIGRIHHANIVRLLGFCSEGTRRALIYEYMPNDSLEKYIFSQDSDTS 422
Query: 121 YE-----KLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
E K+ +IA+G ARG+ YLH+ C RI+H+DIKP NILLD NF PK++DFGLAK C
Sbjct: 423 QELLVPSKMLDIALGIARGMEYLHQGCNQRILHFDIKPNNILLDYNFSPKISDFGLAKLC 482
Query: 176 NRENTHITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE 233
R+ + IT+T RGT GY APEL+ I++K DVYSFGML+ E++ RRN + + E
Sbjct: 483 ARDQSIITLTAARGTMGYIAPELYSRNFGEISYKSDVYSFGMLVLEMVSGRRN-SDPSVE 541
Query: 234 SQE--WFPIWVWKKKDAGL---LGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPI 288
SQ +FP W++++ G LG M E+ K I ++ VALWC+Q+ P+ RP
Sbjct: 542 SQNVVYFPEWIYEQVTIGQDLELGREM-----TEEEKAIMRQLAIVALWCIQWNPKNRPS 596
Query: 289 MSVVVKMLEGSLE 301
M+ VV ML G L+
Sbjct: 597 MTKVVNMLTGRLQ 609
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 632,281,702
Number of Sequences: 2790947
Number of extensions: 27860342
Number of successful extensions: 101002
Number of sequences better than 10.0: 18190
Number of HSP's better than 10.0 without gapping: 9212
Number of HSP's successfully gapped in prelim test: 8980
Number of HSP's that attempted gapping in prelim test: 64527
Number of HSP's gapped (non-prelim): 20319
length of query: 362
length of database: 848,049,833
effective HSP length: 128
effective length of query: 234
effective length of database: 490,808,617
effective search space: 114849216378
effective search space used: 114849216378
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC130275.4


