

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
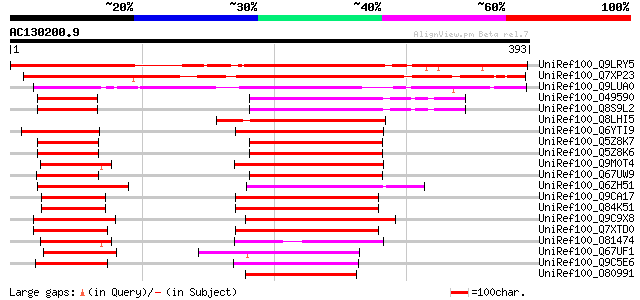
Query= AC130200.9 - phase: 0
(393 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LRY5 Gb|AAF16548.1 [Arabidopsis thaliana] 381 e-104
UniRef100_Q7XP23 OSJNBa0027H09.16 protein [Oryza sativa] 319 7e-86
UniRef100_Q9LUA0 Gb|AAF16548.1 [Arabidopsis thaliana] 198 2e-49
UniRef100_O49590 Hypothetical protein AT4g31410 [Arabidopsis tha... 139 1e-31
UniRef100_Q8S9L2 AT4g31410/F8F16_230 [Arabidopsis thaliana] 139 1e-31
UniRef100_Q8LHI5 Hypothetical protein P0684A03.102-1 [Oryza sativa] 132 2e-29
UniRef100_Q6YTI9 Hypothetical protein P0020D05.10 [Oryza sativa] 127 7e-28
UniRef100_Q5Z8K7 Hypothetical protein P0550B04.19-1 [Oryza sativa] 124 5e-27
UniRef100_Q5Z8K6 Hypothetical protein P0550B04.19-2 [Oryza sativa] 124 5e-27
UniRef100_Q9M0T4 Hypothetical protein AT4g08460 [Arabidopsis tha... 124 6e-27
UniRef100_Q67UW9 Hypothetical protein OSJNBa0050G13.17 [Oryza sa... 122 2e-26
UniRef100_Q6ZH51 Hypothetical protein OJ1079_F11.33-1 [Oryza sat... 120 7e-26
UniRef100_Q9CA17 Hypothetical protein T32E8.10 [Arabidopsis thal... 119 2e-25
UniRef100_Q84K51 Hypothetical protein At1g77770 [Arabidopsis tha... 119 2e-25
UniRef100_Q9C9X8 Hypothetical protein T23K23.1 [Arabidopsis thal... 119 2e-25
UniRef100_Q7XTD0 OSJNBa0064H22.8 protein [Oryza sativa] 107 8e-22
UniRef100_O81474 T15F16.1 protein [Arabidopsis thaliana] 99 2e-19
UniRef100_Q67UF1 Hypothetical protein OJ1163_C07.30 [Oryza sativa] 92 3e-17
UniRef100_Q9C5E6 Hypothetical protein At1g15430 [Arabidopsis tha... 83 1e-14
UniRef100_O80991 Hypothetical protein At2g26050 [Arabidopsis tha... 79 3e-13
>UniRef100_Q9LRY5 Gb|AAF16548.1 [Arabidopsis thaliana]
Length = 354
Score = 381 bits (979), Expect = e-104
Identities = 209/402 (51%), Positives = 261/402 (63%), Gaps = 60/402 (14%)
Query: 1 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
MAG KR+L +SD+HALH+ELDEVSCP+CMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS
Sbjct: 1 MAGVKRKLSTESDVHALHKELDEVSCPVCMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
Query: 61 NCLDRFKKLRDNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTL 120
NCLDRFKKL S +P +++L + +++ + ++
Sbjct: 61 NCLDRFKKLHSESANDPTPEANLASREHNNESLYE------------------------- 95
Query: 121 LSVGIPQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSL 180
G A S H + + G + DSE+L+ R +EEE++ SED ++L
Sbjct: 96 ----------HGTASRSSFHRESGNRG--SSWDSESLRRRRRVEEEVE----SEDI-TNL 138
Query: 181 HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVD 240
CPLCRGTVLGW+VVEE R YL++K RSCSR+SCSF G+Y +LRRHARR HPT+RPSD D
Sbjct: 139 KCPLCRGTVLGWKVVEEVRTYLDHKNRSCSRESCSFTGNYQDLRRHARRTHPTTRPSDTD 198
Query: 241 PTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNG 300
P+RE+AW++ E QREYGDIVSAI+SA+PGAVVVGDYV+ENGD G RE NG
Sbjct: 199 PSRERAWRRLENQREYGDIVSAIRSAMPGAVVVGDYVIENGDRF-----AGERETGNGGS 253
Query: 301 NVPWLTTTTILFQM---MDNTIEIVR----EPRARSSNGWSRHRRSSDRRRYLWGENLLG 353
+ L TT +LFQM +DN R+ S W HRRSS R YLWGENLLG
Sbjct: 254 D---LWTTLLLFQMIGSLDNGGSSASGSGGGSRSHRSRAWRNHRRSSSDRPYLWGENLLG 310
Query: 354 LQD---NEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNEDH 392
LQD N +E+ R+ N+ ++ VPRRRRR R RS+ +H
Sbjct: 311 LQDERNNNDDEEFRLQNDAGGASTPVPRRRRRFGRPRSSGNH 352
>UniRef100_Q7XP23 OSJNBa0027H09.16 protein [Oryza sativa]
Length = 371
Score = 319 bits (818), Expect = 7e-86
Identities = 180/386 (46%), Positives = 244/386 (62%), Gaps = 37/386 (9%)
Query: 11 DSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLR 70
D+++ ALH+E D+ CPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK++
Sbjct: 15 DANIAALHKEWDDALCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKMK 74
Query: 71 DNSKENPNLQSSLINTNNSSGN-----SFDINFTMQSDMHDVDELLLYQNEINTLLSVGI 125
+ + + QSS + + SS N FD +Q+ + + E+ +++ I + +
Sbjct: 75 VDHNDGSSQQSSSLPRDISSQNVPQRSHFDPTGEIQTGISESHEIFNHRDAIQSSAGLSG 134
Query: 126 PQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSK-SSLHCPL 184
QG + +++ + T +++ + + +E SSE ++ + L CPL
Sbjct: 135 QQGE------------NSYNQDLDLTLEAQQRESSSTVE-------SSELTRLNQLACPL 175
Query: 185 CRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTRE 244
CRGTV GW++++EAR YL+ K RSCSR++C+F+G+Y ELRRHARRVHPT+RP+DVDP+R
Sbjct: 176 CRGTVKGWKIIKEAREYLDEKSRSCSRETCAFSGNYRELRRHARRVHPTTRPADVDPSRR 235
Query: 245 QAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPW 304
+AW + E QREYGDI+SAI+SA+PGAVV GDYV+E GD GG +G+
Sbjct: 236 RAWHRLEHQREYGDILSAIRSAMPGAVVFGDYVVEGGDMFSPDQEGGMPNEPSGS----- 290
Query: 305 LTTTTILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRRRYLWGENLLGLQDNEVEEDLR 364
L TT LF M+ ++ + SS G R RRRYLWGENLLGLQ + +ED
Sbjct: 291 LLTTFFLFHMISSSPMRSGDEIRGSSRGLRR-----QRRRYLWGENLLGLQYEDEDEDDE 345
Query: 365 IFNELVEDASHVPRRRRRLNRTRSNE 390
N L ED PR RRR R+ S E
Sbjct: 346 EEN-LDEDVQR-PRSRRRFVRSXSEE 369
>UniRef100_Q9LUA0 Gb|AAF16548.1 [Arabidopsis thaliana]
Length = 372
Score = 198 bits (504), Expect = 2e-49
Identities = 124/382 (32%), Positives = 192/382 (49%), Gaps = 60/382 (15%)
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPN 78
+E +E CP+CM+HPHN +LL+CSS++ GCR Y+CDTS+RHSNC D+F+K SKE P+
Sbjct: 36 KEWEEARCPVCMEHPHNGILLICSSYENGCRPYMCDTSHRHSNCFDQFRKA---SKEKPS 92
Query: 79 LQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPS 138
L SL+ S ++ + SD V+ L +EI + +G + + ++
Sbjct: 93 L--SLLREEEESNEPTEME-DVDSDSTAVNLLGEAASEITVVDLSDGERGEEEVEEEEEE 149
Query: 139 RHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEA 198
+++ +EGI+ T + + ++ L CPLCRG + W VV+ A
Sbjct: 150 VVVEEEEEGIVTTEEDQE-----------------KNKPQKLTCPLCRGHIKEWVVVKAA 192
Query: 199 RNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGD 258
R ++N+K RSCS ++C F+G Y +LR+HAR +HP RPS+ DP R+++W++ ERQ + GD
Sbjct: 193 RCFMNSKHRSCSCETCDFSGSYSDLRKHARLLHPGVRPSEADPERQRSWRRLERQSDLGD 252
Query: 259 IVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPWLTTTTILFQMMDNT 318
++S +QS+ GG E SN +G +L T L +
Sbjct: 253 LLSTLQSSF-----------------------GGDEISNDDG---FLFADTRLLTVYFLI 286
Query: 319 IEIVREPRARSSNGWS---------RHRRSSDRRRYLWGENLLGLQDNEVEEDLRIFNEL 369
E S+ WS RR S R LWGE+ G ++ N+
Sbjct: 287 RVFRPESSGSRSSSWSGTSRARTHTSGRRRSSRPASLWGESYEGNTGTSPRDEEN--NQS 344
Query: 370 VEDASHVPRRRRRLNRTRSNED 391
++ RRRR RT ++D
Sbjct: 345 SDEQVSGTRRRRSRRRTVIDDD 366
>UniRef100_O49590 Hypothetical protein AT4g31410 [Arabidopsis thaliana]
Length = 380
Score = 139 bits (351), Expect = 1e-31
Identities = 76/164 (46%), Positives = 95/164 (57%), Gaps = 10/164 (6%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V GW VVEEAR L+ KKR C + C F G YLELR+HA+ HP SRPS++DP
Sbjct: 183 CPLCRGEVTGWLVVEEARLRLDEKKRCCEEERCRFMGTYLELRKHAQSEHPDSRPSEIDP 242
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGN 301
R+ W+ F++ E D++S I S +P VV+GDYV+E GD G E N
Sbjct: 243 ARKLDWENFQQSSEIIDVLSTIHSEVPRGVVLGDYVIEYGDD----DTGDEFEDVPNNEG 298
Query: 302 VPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRRRY 345
W T+ IL+QM DN +R R R + S RR S R Y
Sbjct: 299 NWW--TSCILYQMFDN----IRNARNRRRSRMSESRRGSRRSSY 336
Score = 75.9 bits (185), Expect = 2e-12
Identities = 28/45 (62%), Positives = 38/45 (84%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRF 66
D+++CPIC+D PHN VLL CSS+ GCR+++C+T + HSNCLDRF
Sbjct: 111 DDLTCPICLDFPHNGVLLQCSSYGNGCRAFVCNTDHLHSNCLDRF 155
>UniRef100_Q8S9L2 AT4g31410/F8F16_230 [Arabidopsis thaliana]
Length = 308
Score = 139 bits (351), Expect = 1e-31
Identities = 76/164 (46%), Positives = 95/164 (57%), Gaps = 10/164 (6%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V GW VVEEAR L+ KKR C + C F G YLELR+HA+ HP SRPS++DP
Sbjct: 102 CPLCRGEVTGWLVVEEARLRLDEKKRCCEEERCRFMGTYLELRKHAQSEHPDSRPSEIDP 161
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGN 301
R+ W+ F++ E D++S I S +P VV+GDYV+E GD G E N
Sbjct: 162 ARKLDWENFQQSSEIIDVLSTIHSEVPRGVVLGDYVIEYGDD----DTGDEFEDVPNNEG 217
Query: 302 VPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRRRY 345
W T+ IL+QM DN +R R R + S RR S R Y
Sbjct: 218 NWW--TSCILYQMFDN----IRNARNRRRSRMSESRRGSRRSSY 255
Score = 75.9 bits (185), Expect = 2e-12
Identities = 28/45 (62%), Positives = 38/45 (84%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRF 66
D+++CPIC+D PHN VLL CSS+ GCR+++C+T + HSNCLDRF
Sbjct: 30 DDLTCPICLDFPHNGVLLQCSSYGNGCRAFVCNTDHLHSNCLDRF 74
>UniRef100_Q8LHI5 Hypothetical protein P0684A03.102-1 [Oryza sativa]
Length = 335
Score = 132 bits (331), Expect = 2e-29
Identities = 62/128 (48%), Positives = 85/128 (65%), Gaps = 4/128 (3%)
Query: 157 LQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSF 216
++D ++EE D SS D K CPLCRG+V GW E R YLN K R+CS DSC F
Sbjct: 62 VEDSILMEECHDAMQSSADLK----CPLCRGSVSGWIPAGEVRKYLNEKLRTCSHDSCKF 117
Query: 217 AGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDY 276
G Y +LR HAR H ++P+ VD +R++ W + ER++E GD++SAI+S PGA++VGDY
Sbjct: 118 VGTYEQLREHARTAHLLAKPAHVDLSRKRTWDRLEREQEVGDVISAIRSQNPGAIIVGDY 177
Query: 277 VLENGDGI 284
V+E D +
Sbjct: 178 VIETRDAM 185
>UniRef100_Q6YTI9 Hypothetical protein P0020D05.10 [Oryza sativa]
Length = 362
Score = 127 bits (318), Expect = 7e-28
Identities = 57/112 (50%), Positives = 72/112 (63%)
Query: 172 SSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVH 231
S + L CPLCRG V GW VVE AR YLN KKR+C D CSF G Y EL +H H
Sbjct: 125 SKQQCAMELACPLCRGDVKGWTVVEPARQYLNRKKRACMHDGCSFIGSYKELCKHVNSKH 184
Query: 232 PTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDG 283
P+++P +VDP W++FE +RE D +S I+S PGAV++GDYV+E G
Sbjct: 185 PSAKPREVDPAHADEWKKFECERERQDAISTIRSMTPGAVIMGDYVVEFNGG 236
Score = 82.0 bits (201), Expect = 3e-14
Identities = 32/59 (54%), Positives = 46/59 (77%)
Query: 10 NDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK 68
ND + + +E SCP+C++HPH+AVLLLC+SH KGCR Y+C T+++HSNCL+ FK+
Sbjct: 38 NDIALVSEKKEWKGASCPVCLEHPHDAVLLLCTSHHKGCRPYMCGTNHQHSNCLEHFKE 96
>UniRef100_Q5Z8K7 Hypothetical protein P0550B04.19-1 [Oryza sativa]
Length = 316
Score = 124 bits (311), Expect = 5e-27
Identities = 50/101 (49%), Positives = 73/101 (71%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V+GW V+ EAR +LN KKR C D CSF G++ EL++H ++ HP SRPS++DP
Sbjct: 109 CPLCRGDVIGWIVIGEARLHLNQKKRCCEEDCCSFVGNFNELQKHTQQKHPDSRPSEIDP 168
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGD 282
R+ W+ F++ + D++S I + +P +V+GDYV+E GD
Sbjct: 169 ARQVDWENFQQSSDIVDVLSTIHAQVPNGIVLGDYVIEYGD 209
Score = 77.8 bits (190), Expect = 5e-13
Identities = 29/46 (63%), Positives = 40/46 (86%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
++V+CPIC+D+PHNAVLL C+S++KGCR ++CDT SNCL+RFK
Sbjct: 29 EDVTCPICLDYPHNAVLLRCTSYEKGCRPFVCDTDQTRSNCLERFK 74
>UniRef100_Q5Z8K6 Hypothetical protein P0550B04.19-2 [Oryza sativa]
Length = 317
Score = 124 bits (311), Expect = 5e-27
Identities = 50/101 (49%), Positives = 73/101 (71%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V+GW V+ EAR +LN KKR C D CSF G++ EL++H ++ HP SRPS++DP
Sbjct: 109 CPLCRGDVIGWIVIGEARLHLNQKKRCCEEDCCSFVGNFNELQKHTQQKHPDSRPSEIDP 168
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGD 282
R+ W+ F++ + D++S I + +P +V+GDYV+E GD
Sbjct: 169 ARQVDWENFQQSSDIVDVLSTIHAQVPNGIVLGDYVIEYGD 209
Score = 77.8 bits (190), Expect = 5e-13
Identities = 29/46 (63%), Positives = 40/46 (86%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
++V+CPIC+D+PHNAVLL C+S++KGCR ++CDT SNCL+RFK
Sbjct: 29 EDVTCPICLDYPHNAVLLRCTSYEKGCRPFVCDTDQTRSNCLERFK 74
>UniRef100_Q9M0T4 Hypothetical protein AT4g08460 [Arabidopsis thaliana]
Length = 274
Score = 124 bits (310), Expect = 6e-27
Identities = 53/113 (46%), Positives = 73/113 (63%)
Query: 171 NSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRV 230
+ D L CPLCRG V GW VVE+ R YLN+KKRSC D C F G Y +L++H +
Sbjct: 96 DEKSDKPPELLCPLCRGQVKGWTVVEKERKYLNSKKRSCMNDECLFYGSYRQLKKHVKEN 155
Query: 231 HPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDG 283
HP ++P +DP E W++ E +RE D++S + S+ PGA+V GDYV+E +G
Sbjct: 156 HPRAKPRAIDPVLEAKWKKLEVERERSDVISTVMSSTPGAMVFGDYVIEPYNG 208
Score = 72.0 bits (175), Expect = 3e-11
Identities = 29/56 (51%), Positives = 42/56 (74%), Gaps = 2/56 (3%)
Query: 24 VSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK--LRDNSKENP 77
V+CP+C++ PHN+V+LLCSS+ KGCR Y+C T R SNCL+++KK +D + P
Sbjct: 47 VTCPVCLEVPHNSVVLLCSSYHKGCRPYMCATGNRFSNCLEQYKKAYAKDEKSDKP 102
>UniRef100_Q67UW9 Hypothetical protein OSJNBa0050G13.17 [Oryza sativa]
Length = 316
Score = 122 bits (305), Expect = 2e-26
Identities = 48/101 (47%), Positives = 72/101 (70%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V+GW V++EAR +LN KKR C CS+ G++ EL++H ++ HP SRPS++DP
Sbjct: 104 CPLCRGDVIGWVVIDEARLHLNQKKRCCEESCCSYVGNFHELQKHTQQKHPNSRPSEIDP 163
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGD 282
R W+ F++ + D++S I + +P +V+GDYV+E GD
Sbjct: 164 ARRVDWENFQQSSDIIDVLSTIHAQVPNGIVLGDYVIEYGD 204
Score = 77.4 bits (189), Expect = 6e-13
Identities = 29/47 (61%), Positives = 40/47 (84%)
Query: 21 LDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
+++++CPIC+D PHNAVLL C+S++KGCR +ICDT SNCL+RFK
Sbjct: 23 MEDITCPICLDFPHNAVLLRCTSYEKGCRPFICDTDQSRSNCLERFK 69
>UniRef100_Q6ZH51 Hypothetical protein OJ1079_F11.33-1 [Oryza sativa]
Length = 345
Score = 120 bits (301), Expect = 7e-26
Identities = 61/135 (45%), Positives = 81/135 (59%), Gaps = 1/135 (0%)
Query: 180 LHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDV 239
L CPLCRG V GW +VE AR+YLN K+R+C +D CSF G Y ELR+H + HP ++P +V
Sbjct: 130 LACPLCRGKVKGWTIVEPARSYLNGKRRTCMQDGCSFVGTYKELRKHVKSEHPLAKPREV 189
Query: 240 DPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGN 299
DP EQ W+ E +RE D +S I + + A+V GDYVL+ D L E N N
Sbjct: 190 DPILEQKWRLLEIERERQDALSTITATMGRAIVFGDYVLDLEDE-DDLDDVESDEDDNAN 248
Query: 300 GNVPWLTTTTILFQM 314
G+ T ++F M
Sbjct: 249 GHGTDNTRRMLMFLM 263
Score = 90.5 bits (223), Expect = 7e-17
Identities = 38/69 (55%), Positives = 50/69 (72%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPNLQS 81
++ +C +CM++PHNAVLLLCSSHDKGCR Y+C TS+RHSNCLD+FKK L +
Sbjct: 48 EDANCSVCMEYPHNAVLLLCSSHDKGCRPYMCGTSHRHSNCLDQFKKAYTKGALLEELPA 107
Query: 82 SLINTNNSS 90
+ + TN S
Sbjct: 108 NTVGTNLDS 116
>UniRef100_Q9CA17 Hypothetical protein T32E8.10 [Arabidopsis thaliana]
Length = 264
Score = 119 bits (297), Expect = 2e-25
Identities = 49/108 (45%), Positives = 76/108 (70%)
Query: 172 SSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVH 231
+ + L CPLCRG V GW VV++AR + N+K+R+C +D+CSF G++ +L++H + H
Sbjct: 77 NENSGQPELLCPLCRGQVKGWTVVKDARMHFNSKRRTCMQDNCSFLGNFRKLKKHMKEKH 136
Query: 232 PTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
P + P +DP E W++ ER+R+ D++S I S+ PGAVV+GDYV+E
Sbjct: 137 PHACPRAIDPALETKWKRLERERDRRDVISTIMSSTPGAVVLGDYVIE 184
Score = 73.6 bits (179), Expect = 9e-12
Identities = 29/48 (60%), Positives = 39/48 (80%)
Query: 25 SCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDN 72
+CP+C++ PHNAVLLLCSS+ KGCR Y+C TS R +NCLD+++K N
Sbjct: 30 TCPVCLESPHNAVLLLCSSYHKGCRPYMCATSSRFANCLDQYRKSYGN 77
>UniRef100_Q84K51 Hypothetical protein At1g77770 [Arabidopsis thaliana]
Length = 265
Score = 119 bits (297), Expect = 2e-25
Identities = 49/108 (45%), Positives = 76/108 (70%)
Query: 172 SSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVH 231
+ + L CPLCRG V GW VV++AR + N+K+R+C +D+CSF G++ +L++H + H
Sbjct: 77 NENSGQPELLCPLCRGQVKGWTVVKDARMHFNSKRRTCMQDNCSFLGNFRKLKKHMKEKH 136
Query: 232 PTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
P + P +DP E W++ ER+R+ D++S I S+ PGAVV+GDYV+E
Sbjct: 137 PHACPRAIDPALETKWKRLERERDRRDVISTIMSSTPGAVVLGDYVIE 184
Score = 73.6 bits (179), Expect = 9e-12
Identities = 29/48 (60%), Positives = 39/48 (80%)
Query: 25 SCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDN 72
+CP+C++ PHNAVLLLCSS+ KGCR Y+C TS R +NCLD+++K N
Sbjct: 30 TCPVCLESPHNAVLLLCSSYHKGCRPYMCATSSRFANCLDQYRKSYGN 77
>UniRef100_Q9C9X8 Hypothetical protein T23K23.1 [Arabidopsis thaliana]
Length = 334
Score = 119 bits (297), Expect = 2e-25
Identities = 52/114 (45%), Positives = 78/114 (67%)
Query: 179 SLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSD 238
+L CPLCRG V GW +V+ AR++LN KKR C +++C +AG + ELR+H + HP+++P +
Sbjct: 117 NLTCPLCRGQVKGWTIVQPARDFLNLKKRICMQENCVYAGTFKELRKHMKVDHPSAKPRE 176
Query: 239 VDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGG 292
VDP EQ W++ E + + D++S I+S +PG VV GDYV+E + G GG
Sbjct: 177 VDPDVEQNWRRLEIEHDRDDVMSTIRSTMPGTVVYGDYVIERNNANGSDSDEGG 230
Score = 84.7 bits (208), Expect = 4e-15
Identities = 36/62 (58%), Positives = 47/62 (75%)
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPN 78
R+ + V C +CM+ PHNAVLLLCSSHDKGCR Y+C TS+R+SNCLD++KK K + +
Sbjct: 48 RDWENVICSVCMECPHNAVLLLCSSHDKGCRPYMCGTSFRYSNCLDQYKKASAKLKTSGH 107
Query: 79 LQ 80
Q
Sbjct: 108 QQ 109
>UniRef100_Q7XTD0 OSJNBa0064H22.8 protein [Oryza sativa]
Length = 267
Score = 107 bits (266), Expect = 8e-22
Identities = 44/108 (40%), Positives = 67/108 (61%)
Query: 172 SSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVH 231
S + + L CP+CRG V GW VVE AR +LN K+R+C + CSF G Y +LR H R H
Sbjct: 113 SKKPQEMELVCPICRGDVKGWTVVEPARRFLNRKRRTCMHEGCSFGGSYRKLRNHVRSNH 172
Query: 232 PTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
P+S P ++D W++ E +++ D +S I + PG+ ++GDY ++
Sbjct: 173 PSSNPREIDSASLAEWKELEYEKDRQDAISIITALNPGSTIMGDYFID 220
Score = 79.3 bits (194), Expect = 2e-13
Identities = 31/56 (55%), Positives = 41/56 (72%)
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSK 74
++ +C IC++HPH AVLLLCSSH KGCR Y+CDT+ +HSNCL++FK K
Sbjct: 45 KDWKRATCSICLEHPHKAVLLLCSSHSKGCRPYMCDTNRQHSNCLEQFKNAYSRGK 100
>UniRef100_O81474 T15F16.1 protein [Arabidopsis thaliana]
Length = 260
Score = 99.0 bits (245), Expect = 2e-19
Identities = 46/113 (40%), Positives = 66/113 (57%), Gaps = 14/113 (12%)
Query: 171 NSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRV 230
+ D L CPLCRG V GW VVE+ R YLN+KKR +L++H +
Sbjct: 96 DEKSDKPPELLCPLCRGQVKGWTVVEKERKYLNSKKR--------------QLKKHVKEN 141
Query: 231 HPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDG 283
HP ++P +DP E W++ E +RE D++S + S+ PGA+V GDYV+E +G
Sbjct: 142 HPRAKPRAIDPVLEAKWKKLEVERERSDVISTVMSSTPGAMVFGDYVIEPYNG 194
Score = 72.0 bits (175), Expect = 3e-11
Identities = 29/56 (51%), Positives = 42/56 (74%), Gaps = 2/56 (3%)
Query: 24 VSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK--LRDNSKENP 77
V+CP+C++ PHN+V+LLCSS+ KGCR Y+C T R SNCL+++KK +D + P
Sbjct: 47 VTCPVCLEVPHNSVVLLCSSYHKGCRPYMCATGNRFSNCLEQYKKAYAKDEKSDKP 102
>UniRef100_Q67UF1 Hypothetical protein OJ1163_C07.30 [Oryza sativa]
Length = 231
Score = 92.0 bits (227), Expect = 3e-17
Identities = 47/126 (37%), Positives = 67/126 (52%), Gaps = 4/126 (3%)
Query: 144 HDEGILETADSENLQDRAVLEEELDVDNSSEDSKS----SLHCPLCRGTVLGWEVVEEAR 199
HD+G + N L++ +D SS+D + L CPLCRG V G+ +VE AR
Sbjct: 67 HDKGCRPYICATNYHHSNCLDQLIDSRRSSKDCEDLDSIELTCPLCRGEVKGYTLVEPAR 126
Query: 200 NYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDI 259
LN KRSC +D CS+ G Y EL +H R+ HP+ +P VDP W++ + D+
Sbjct: 127 EQLNQNKRSCMQDGCSYMGSYGELCKHVRKKHPSVKPHSVDPVHTYRWRRLLFRSSLQDM 186
Query: 260 VSAIQS 265
+ A S
Sbjct: 187 ICATSS 192
Score = 84.7 bits (208), Expect = 4e-15
Identities = 36/56 (64%), Positives = 44/56 (78%)
Query: 26 CPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPNLQS 81
CP+C++ PHNAVLLLCSSHDKGCR YIC T+Y HSNCLD+ R +SK+ +L S
Sbjct: 49 CPVCLECPHNAVLLLCSSHDKGCRPYICATNYHHSNCLDQLIDSRRSSKDCEDLDS 104
>UniRef100_Q9C5E6 Hypothetical protein At1g15430 [Arabidopsis thaliana]
Length = 259
Score = 83.2 bits (204), Expect = 1e-14
Identities = 37/95 (38%), Positives = 56/95 (58%)
Query: 170 DNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARR 229
++SS S +LHCP CRG V G AR ++N + R C+ D C F+G Y +L+ H +
Sbjct: 97 NSSSRCSGKTLHCPYCRGEVQGTMKSTCARRFMNARPRCCTVDKCDFSGTYAQLKNHLKT 156
Query: 230 VHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQ 264
HP P +DP + W+Q ER+ EY ++++A Q
Sbjct: 157 EHPGFTPPKLDPWEQHMWEQLEREAEYIEMLNARQ 191
Score = 76.3 bits (186), Expect = 1e-12
Identities = 31/55 (56%), Positives = 42/55 (76%)
Query: 20 ELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSK 74
E ++V C ICM+ PHNAVLL CSS KGCR+Y+CDTS RHSNC ++++ +S+
Sbjct: 47 EWEDVRCVICMEPPHNAVLLQCSSFSKGCRAYMCDTSARHSNCFKQYRRSNSSSR 101
>UniRef100_O80991 Hypothetical protein At2g26050 [Arabidopsis thaliana]
Length = 221
Score = 78.6 bits (192), Expect = 3e-13
Identities = 36/85 (42%), Positives = 52/85 (60%), Gaps = 1/85 (1%)
Query: 179 SLHCPLCRGTVLGW-EVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPS 237
+LHCPLCRG V +V AR ++N K RSCS + C F+G + +L +H + H P
Sbjct: 71 TLHCPLCRGEVSETTKVTSTARRFMNAKPRSCSVEDCKFSGTFSQLTKHLKTEHRGIVPP 130
Query: 238 DVDPTREQAWQQFERQREYGDIVSA 262
VDP R+Q W+ ER EY ++++A
Sbjct: 131 KVDPLRQQRWEMMERHSEYVELMTA 155
Score = 38.9 bits (89), Expect = 0.25
Identities = 13/23 (56%), Positives = 18/23 (77%)
Query: 46 KGCRSYICDTSYRHSNCLDRFKK 68
+GC Y+CDTS RHSNC +F++
Sbjct: 38 RGCHPYMCDTSVRHSNCFKQFRR 60
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 706,804,556
Number of Sequences: 2790947
Number of extensions: 31701412
Number of successful extensions: 104014
Number of sequences better than 10.0: 101
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 103784
Number of HSP's gapped (non-prelim): 274
length of query: 393
length of database: 848,049,833
effective HSP length: 129
effective length of query: 264
effective length of database: 488,017,670
effective search space: 128836664880
effective search space used: 128836664880
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC130200.9


