

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.5 - phase: 0
(312 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
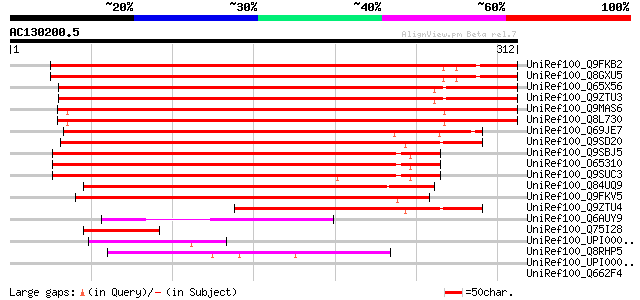
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FKB2 Gb|AAC72543.1 [Arabidopsis thaliana] 385 e-106
UniRef100_Q8GXU5 Hypothetical protein At5g48850/K24G6_19 [Arabid... 385 e-106
UniRef100_Q65X56 Hypothetical protein OJ1123_F01.11 [Oryza sativa] 371 e-101
UniRef100_Q9ZTU3 Hypothetical protein [Oryza sativa] 369 e-101
UniRef100_Q9MAS6 F13M7.24 protein [Arabidopsis thaliana] 325 1e-87
UniRef100_Q8L730 Hypothetical protein At1g04770 [Arabidopsis tha... 325 1e-87
UniRef100_Q69JE7 Putative pollenless3 [Oryza sativa] 288 2e-76
UniRef100_Q9SD20 MS5-like protein [Arabidopsis thaliana] 275 9e-73
UniRef100_Q9SBJ5 Male sterility MS5 [Arabidopsis thaliana] 253 4e-66
UniRef100_O65310 Pollenless3 [Arabidopsis thaliana] 253 4e-66
UniRef100_Q9SUC3 Hypothetical protein T13K14.60 [Arabidopsis tha... 243 5e-63
UniRef100_Q84UQ9 Putative pollenless3 [Oryza sativa] 223 7e-57
UniRef100_Q9FKV5 Male sterility MS5; pollenless3 [Arabidopsis th... 213 5e-54
UniRef100_Q9ZTU4 MS5-like protein [Arabidopsis thaliana] 122 2e-26
UniRef100_Q6AUY9 Hypothetical protein OSJNBa0004G03.21 [Oryza sa... 82 2e-14
UniRef100_Q75I28 Hypothetical protein OSJNBa0091E13.29 [Oryza sa... 70 9e-11
UniRef100_UPI00002867B7 UPI00002867B7 UniRef100 entry 54 5e-06
UniRef100_Q8RHP5 Tetratricopeptide repeat family protein [Fusoba... 48 4e-04
UniRef100_UPI00002E7317 UPI00002E7317 UniRef100 entry 43 0.012
UniRef100_Q662F4 Surface-located membrane protein 1 [Borrelia ga... 43 0.012
>UniRef100_Q9FKB2 Gb|AAC72543.1 [Arabidopsis thaliana]
Length = 326
Score = 385 bits (989), Expect = e-106
Identities = 190/292 (65%), Positives = 247/292 (84%), Gaps = 7/292 (2%)
Query: 26 KSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
KS+ K ++L+HVIHKVP GDTPYV+AKHAQL++K+PE+AIV+FWKAIN GD+VDSALKD
Sbjct: 37 KSNLMKDDELFHVIHKVPCGDTPYVRAKHAQLIEKNPEMAIVWFWKAINTGDRVDSALKD 96
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MAVVMKQLDR+EEAIEAIKSFR C+K+SQ+SLDNVL+DLYKKCGR+EEQ+ELLKRKLR
Sbjct: 97 MAVVMKQLDRSEEAIEAIKSFRPRCSKNSQDSLDNVLIDLYKKCGRMEEQVELLKRKLRQ 156
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV+++QE +RLLGNLGWAYMQ+ Y+ AE V++KAQM+
Sbjct: 157 IYQGEAFNGKPTKTARSHGKKFQVTVQQEISRLLGNLGWAYMQQAKYLSAEAVYRKAQMV 216
Query: 206 DADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELSANLPQ 265
+ DANK+ NLA+CL++Q R+EE L+L+ VL+ ++ G+D+ ++R RAEELL EL ++LP+
Sbjct: 217 EPDANKSCNLAMCLIKQGRFEEGRLVLDDVLEYRVLGADDCRTRQRAEELLSELESSLPR 276
Query: 266 ---PKFMDDLG--LDDDLLKGIDGLLNVWSPTRSRRLPIFEEISSFRDQLAC 312
+ D LG LDDD + G++ + + + +S+RLPIFE+ISSFR+ L C
Sbjct: 277 MRDAEMEDVLGNILDDDFVLGLEEMTS--TSFKSKRLPIFEQISSFRNTLVC 326
>UniRef100_Q8GXU5 Hypothetical protein At5g48850/K24G6_19 [Arabidopsis thaliana]
Length = 306
Score = 385 bits (989), Expect = e-106
Identities = 190/292 (65%), Positives = 247/292 (84%), Gaps = 7/292 (2%)
Query: 26 KSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
KS+ K ++L+HVIHKVP GDTPYV+AKHAQL++K+PE+AIV+FWKAIN GD+VDSALKD
Sbjct: 17 KSNLMKDDELFHVIHKVPCGDTPYVRAKHAQLIEKNPEMAIVWFWKAINTGDRVDSALKD 76
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MAVVMKQLDR+EEAIEAIKSFR C+K+SQ+SLDNVL+DLYKKCGR+EEQ+ELLKRKLR
Sbjct: 77 MAVVMKQLDRSEEAIEAIKSFRPRCSKNSQDSLDNVLIDLYKKCGRMEEQVELLKRKLRQ 136
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV+++QE +RLLGNLGWAYMQ+ Y+ AE V++KAQM+
Sbjct: 137 IYQGEAFNGKPTKTARSHGKKFQVTVQQEISRLLGNLGWAYMQQAKYLSAEAVYRKAQMV 196
Query: 206 DADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELSANLPQ 265
+ DANK+ NLA+CL++Q R+EE L+L+ VL+ ++ G+D+ ++R RAEELL EL ++LP+
Sbjct: 197 EPDANKSCNLAMCLIKQGRFEEGRLVLDDVLEYRVLGADDCRTRQRAEELLSELESSLPR 256
Query: 266 ---PKFMDDLG--LDDDLLKGIDGLLNVWSPTRSRRLPIFEEISSFRDQLAC 312
+ D LG LDDD + G++ + + + +S+RLPIFE+ISSFR+ L C
Sbjct: 257 MRDAEMEDVLGNILDDDFVLGLEEMTS--TSFKSKRLPIFEQISSFRNTLVC 306
>UniRef100_Q65X56 Hypothetical protein OJ1123_F01.11 [Oryza sativa]
Length = 299
Score = 371 bits (952), Expect = e-101
Identities = 180/284 (63%), Positives = 239/284 (83%), Gaps = 3/284 (1%)
Query: 31 KKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVM 90
+K+DL+HV+HKVP GD+PYV+AKH QLVDKDPE AIV+FWKAIN+ DKVDSALKDMAVVM
Sbjct: 17 EKKDLFHVVHKVPAGDSPYVRAKHLQLVDKDPETAIVWFWKAINSRDKVDSALKDMAVVM 76
Query: 91 KQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGE 150
KQ DRA+EAIEAI+SFR LC++ +QESLDN+L+DLYKKCG+V+EQI+LLK+KL++IY GE
Sbjct: 77 KQQDRAKEAIEAIRSFRHLCSRQAQESLDNLLIDLYKKCGKVDEQIDLLKQKLKMIYLGE 136
Query: 151 AFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADAN 210
AFNG+ TKTARSHGKKFQVSI+QET+R+LGNLGWAYMQ++NY AE+V++KAQ I+ DAN
Sbjct: 137 AFNGKATKTARSHGKKFQVSIQQETSRILGNLGWAYMQQSNYSAAELVYRKAQSIEPDAN 196
Query: 211 KALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELS--ANLPQPKF 268
+A NL LCL++QSR++EA +L V+ ++ GS++ K RA++LL EL ++ P
Sbjct: 197 RACNLGLCLIKQSRHDEARQVLHDVVLRRISGSEDDKVVARAKQLLHELEPVTHVTSPN- 255
Query: 269 MDDLGLDDDLLKGIDGLLNVWSPTRSRRLPIFEEISSFRDQLAC 312
L + +++++ +D +LN W+P RSRRLP+FEEI++ RDQ+AC
Sbjct: 256 NAGLSVSEEIMERLDLVLNEWTPFRSRRLPVFEEIATLRDQIAC 299
>UniRef100_Q9ZTU3 Hypothetical protein [Oryza sativa]
Length = 321
Score = 369 bits (947), Expect = e-101
Identities = 179/284 (63%), Positives = 239/284 (84%), Gaps = 3/284 (1%)
Query: 31 KKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVM 90
+K+DL+HV+HKVP G++PYV+AKH QLVDKDPE AIV+FWKAIN+ DKVDSALKDMAVVM
Sbjct: 39 EKKDLFHVVHKVPAGNSPYVRAKHLQLVDKDPETAIVWFWKAINSRDKVDSALKDMAVVM 98
Query: 91 KQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGE 150
KQ DRA+EAIEAI+SFR LC++ +QESLDN+L+DLYKKCG+V+EQI+LLK+KL++IY GE
Sbjct: 99 KQQDRAKEAIEAIRSFRHLCSRQAQESLDNLLIDLYKKCGKVDEQIDLLKQKLKMIYLGE 158
Query: 151 AFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADAN 210
AFNG+ TKTARSHGKKFQVSI+QET+R+LGNLGWAYMQ++NY AE+V++KAQ I+ DAN
Sbjct: 159 AFNGKATKTARSHGKKFQVSIQQETSRILGNLGWAYMQQSNYSAAELVYRKAQSIEPDAN 218
Query: 211 KALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELS--ANLPQPKF 268
+A NL LCL++QSR++EA +L V+ ++ GS++ K RA++LL EL ++ P
Sbjct: 219 RACNLGLCLIKQSRHDEARQVLHDVVLRRISGSEDDKVVARAKQLLHELEPVTHVTSPN- 277
Query: 269 MDDLGLDDDLLKGIDGLLNVWSPTRSRRLPIFEEISSFRDQLAC 312
L + +++++ +D +LN W+P RSRRLP+FEEI++ RDQ+AC
Sbjct: 278 NAGLSVSEEIMERLDLVLNEWTPFRSRRLPVFEEIATLRDQIAC 321
>UniRef100_Q9MAS6 F13M7.24 protein [Arabidopsis thaliana]
Length = 364
Score = 325 bits (833), Expect = 1e-87
Identities = 168/294 (57%), Positives = 223/294 (75%), Gaps = 11/294 (3%)
Query: 30 GKKED----LYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
G+++D Y+V+HK+P+GD+PYV+AKH QLV+KD E AI FW AI A D+VDSALKD
Sbjct: 71 GERQDSSAAAYNVVHKLPHGDSPYVRAKHVQLVEKDAEAAIELFWIAIKARDRVDSALKD 130
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MA++MKQ +RAEEAI+AI+SFR LC++ +QESLDNVL+DLYKKCGR+EEQ+ELLK+KL +
Sbjct: 131 MALLMKQQNRAEEAIDAIQSFRDLCSRQAQESLDNVLIDLYKKCGRIEEQVELLKQKLWM 190
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV++++ET+R+LGNLGWAYMQ +Y AE V++KAQ+I
Sbjct: 191 IYQGEAFNGKPTKTARSHGKKFQVTVEKETSRILGNLGWAYMQLMDYTAAEAVYRKAQLI 250
Query: 206 DADANKALNLALCLMRQSRYEEAYLIL-EQVLQGKLPGSDEIKSRNRAEELLVELSANLP 264
+ DANKA NL CL++Q +++EA IL VL GS + + R +ELL EL
Sbjct: 251 EPDANKACNLCTCLIKQGKHDEARSILFRDVLMENKEGSGDPRLMARVQELLSELKPQEE 310
Query: 265 QP----KFMDDLGLDD-DLLKGIDGLLNVW-SPTRSRRLPIFEEISSFRDQLAC 312
+ ++G+D+ +++G+D + W P R+RRLPIFEEI RDQLAC
Sbjct: 311 EAAASVSVECEVGIDEIAVVEGLDEFVKEWRRPYRTRRLPIFEEILPLRDQLAC 364
>UniRef100_Q8L730 Hypothetical protein At1g04770 [Arabidopsis thaliana]
Length = 303
Score = 325 bits (833), Expect = 1e-87
Identities = 168/294 (57%), Positives = 223/294 (75%), Gaps = 11/294 (3%)
Query: 30 GKKED----LYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
G+++D Y+V+HK+P+GD+PYV+AKH QLV+KD E AI FW AI A D+VDSALKD
Sbjct: 10 GERQDSSAAAYNVVHKLPHGDSPYVRAKHVQLVEKDAEAAIELFWIAIKARDRVDSALKD 69
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MA++MKQ +RAEEAI+AI+SFR LC++ +QESLDNVL+DLYKKCGR+EEQ+ELLK+KL +
Sbjct: 70 MALLMKQQNRAEEAIDAIQSFRDLCSRQAQESLDNVLIDLYKKCGRIEEQVELLKQKLWM 129
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV++++ET+R+LGNLGWAYMQ +Y AE V++KAQ+I
Sbjct: 130 IYQGEAFNGKPTKTARSHGKKFQVTVEKETSRILGNLGWAYMQLMDYTAAEAVYRKAQLI 189
Query: 206 DADANKALNLALCLMRQSRYEEAYLIL-EQVLQGKLPGSDEIKSRNRAEELLVELSANLP 264
+ DANKA NL CL++Q +++EA IL VL GS + + R +ELL EL
Sbjct: 190 EPDANKACNLCTCLIKQGKHDEARSILFRDVLMENKEGSGDPRLMARVQELLSELKPQEE 249
Query: 265 QP----KFMDDLGLDD-DLLKGIDGLLNVW-SPTRSRRLPIFEEISSFRDQLAC 312
+ ++G+D+ +++G+D + W P R+RRLPIFEEI RDQLAC
Sbjct: 250 EAAASVSVECEVGIDEIAVVEGLDEFVKEWRRPYRTRRLPIFEEILPLRDQLAC 303
>UniRef100_Q69JE7 Putative pollenless3 [Oryza sativa]
Length = 515
Score = 288 bits (736), Expect = 2e-76
Identities = 152/269 (56%), Positives = 189/269 (69%), Gaps = 13/269 (4%)
Query: 34 DLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQL 93
D +HV HKVP GDTPYV+AK QLVDKDPE AI FW AINAGD+VDSALKDMA+VMKQ
Sbjct: 52 DSFHVAHKVPVGDTPYVRAKRVQLVDKDPEKAIALFWAAINAGDRVDSALKDMAIVMKQQ 111
Query: 94 DRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFN 153
+RAEEAIEAIKS R C+ +QESLDN+LLDLYK+CGR+++QI LLK KL+LI+QG AFN
Sbjct: 112 NRAEEAIEAIKSLRSRCSDQAQESLDNILLDLYKRCGRLDDQISLLKHKLQLIHQGHAFN 171
Query: 154 GRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKAL 213
G+ TKTARS G+KFQV+++QE RLLGNLGWA MQK NY AE +++A +I D NK
Sbjct: 172 GKRTKTARSQGRKFQVTLEQEATRLLGNLGWALMQKENYTEAEGAYRRALLIGPDNNKMC 231
Query: 214 NLALCLMRQSRYEEAYLILEQV----LQGKLPGSDEIKSRNRAEELLVELSANL------ 263
NL +CLM+Q R EA +L+QV + G +K+ RA+E+L +L A L
Sbjct: 232 NLGICLMKQGRVLEAKDVLKQVRPAGVDGLRGADSHLKAYERAQEMLRDLEAKLVGRRLP 291
Query: 264 -PQPKFMDDLGLDDDLLKGIDGLLNVWSP 291
+ +D L D LL G ++W P
Sbjct: 292 RAGDQLVDKSWLFDALLLGSSS--SIWQP 318
>UniRef100_Q9SD20 MS5-like protein [Arabidopsis thaliana]
Length = 430
Score = 275 bits (704), Expect = 9e-73
Identities = 144/264 (54%), Positives = 188/264 (70%), Gaps = 5/264 (1%)
Query: 32 KEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMK 91
+ + +H IHKVP GD+PYV+AK+ QLV+KDPE AI FWKAINAGD+VDSALKDMA+VMK
Sbjct: 26 QSESFHAIHKVPVGDSPYVRAKNVQLVEKDPERAIPLFWKAINAGDRVDSALKDMAIVMK 85
Query: 92 QLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEA 151
Q +RAEEAIEAIKS R C+ +QESLDN+LLDLYK+CGR+++QI LLK KL LI +G A
Sbjct: 86 QQNRAEEAIEAIKSLRVRCSDQAQESLDNILLDLYKRCGRLDDQIGLLKHKLFLIQKGLA 145
Query: 152 FNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANK 211
FNG+ TKTARS GKKFQVS++QE RLLGNLGWA MQ+ N++ AE +++A I D NK
Sbjct: 146 FNGKRTKTARSQGKKFQVSVEQEATRLLGNLGWALMQRDNFVEAEDAYRRALSIAPDNNK 205
Query: 212 ALNLALCLMRQSRYEEAYLILEQVLQGKLPG----SDEIKSRNRAEELLVELSANLPQPK 267
NL +CLM+Q R +EA L +V + G +K+ RA+++L +L + + + +
Sbjct: 206 MCNLGICLMKQGRIDEAKETLRRVKPAVVDGPRGVDSHLKAYERAQQMLNDLGSEMMR-R 264
Query: 268 FMDDLGLDDDLLKGIDGLLNVWSP 291
DD L I G ++W P
Sbjct: 265 GGDDKVEQRRLFDAIFGSSSIWQP 288
>UniRef100_Q9SBJ5 Male sterility MS5 [Arabidopsis thaliana]
Length = 434
Score = 253 bits (647), Expect = 4e-66
Identities = 130/245 (53%), Positives = 173/245 (70%), Gaps = 8/245 (3%)
Query: 27 SSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDM 86
SS ++ D +H++HKVP GD+PYV+AKHAQL+DKDP AI FW AINAGD+VDSALKDM
Sbjct: 42 SSSSERRDPFHIVHKVPSGDSPYVRAKHAQLIDKDPNRAISLFWTAINAGDRVDSALKDM 101
Query: 87 AVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
AVVMKQL R++E IEAIKSFR LC+ SQ+S+DN+LL+LYKK GR+EE+ LL+ KL+ +
Sbjct: 102 AVVMKQLGRSDEGIEAIKSFRYLCSFESQDSIDNLLLELYKKSGRIEEEAVLLEHKLQTL 161
Query: 147 YQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMID 206
QG F GR ++ R GK ++I+QE AR+LGNLGW ++Q NY +AE +++A ++
Sbjct: 162 EQGMGFGGRVSRAKRVQGKHVIMTIEQEKARILGNLGWVHLQLHNYGIAEQHYRRALGLE 221
Query: 207 ADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDE------IKSRNRAEELLVELS 260
D NK NLA+CLMR SR EA +L+ V P E KS +RA E+L E+
Sbjct: 222 RDKNKLCNLAICLMRMSRIPEAKSLLDDVRDS--PAESECGDEPFAKSYDRAVEMLAEIE 279
Query: 261 ANLPQ 265
+ P+
Sbjct: 280 SKKPE 284
>UniRef100_O65310 Pollenless3 [Arabidopsis thaliana]
Length = 434
Score = 253 bits (647), Expect = 4e-66
Identities = 130/245 (53%), Positives = 173/245 (70%), Gaps = 8/245 (3%)
Query: 27 SSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDM 86
SS ++ D +H++HKVP GD+PYV+AKHAQL+DKDP AI FW AINAGD+VDSALKDM
Sbjct: 42 SSSSERRDPFHIVHKVPSGDSPYVRAKHAQLIDKDPNRAISLFWTAINAGDRVDSALKDM 101
Query: 87 AVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
AVVMKQL R++E IEAIKSFR LC+ SQ+S+DN+LL+LYKK GR+EE+ LL+ KL+ +
Sbjct: 102 AVVMKQLGRSDEGIEAIKSFRYLCSFESQDSIDNLLLELYKKSGRIEEEAVLLEHKLQTL 161
Query: 147 YQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMID 206
QG F GR ++ R GK ++I+QE AR+LGNLGW ++Q NY +AE +++A ++
Sbjct: 162 EQGMGFGGRVSRAKRVQGKHVIMTIEQEKARILGNLGWVHLQLHNYGIAEQHYRRALGLE 221
Query: 207 ADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDE------IKSRNRAEELLVELS 260
D NK NLA+CLMR SR EA +L+ V P E KS +RA E+L E+
Sbjct: 222 RDKNKLCNLAICLMRMSRIPEAKSLLDDVRDS--PAESECGDEPFAKSYDRAVEMLAEIE 279
Query: 261 ANLPQ 265
+ P+
Sbjct: 280 SKKPE 284
>UniRef100_Q9SUC3 Hypothetical protein T13K14.60 [Arabidopsis thaliana]
Length = 450
Score = 243 bits (620), Expect = 5e-63
Identities = 130/261 (49%), Positives = 173/261 (65%), Gaps = 24/261 (9%)
Query: 27 SSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDM 86
SS ++ D +H++HKVP GD+PYV+AKHAQL+DKDP AI FW AINAGD+VDSALKDM
Sbjct: 42 SSSSERRDPFHIVHKVPSGDSPYVRAKHAQLIDKDPNRAISLFWTAINAGDRVDSALKDM 101
Query: 87 AVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
AVVMKQL R++E IEAIKSFR LC+ SQ+S+DN+LL+LYKK GR+EE+ LL+ KL+ +
Sbjct: 102 AVVMKQLGRSDEGIEAIKSFRYLCSFESQDSIDNLLLELYKKSGRIEEEAVLLEHKLQTL 161
Query: 147 YQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFK------ 200
QG F GR ++ R GK ++I+QE AR+LGNLGW ++Q NY +AE ++
Sbjct: 162 EQGMGFGGRVSRAKRVQGKHVIMTIEQEKARILGNLGWVHLQLHNYGIAEQHYRFGFVTK 221
Query: 201 ----------KAQMIDADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDE----- 245
+A ++ D NK NLA+CLMR SR EA +L+ V P E
Sbjct: 222 IPNIDYCLVMRALGLERDKNKLCNLAICLMRMSRIPEAKSLLDDVRDS--PAESECGDEP 279
Query: 246 -IKSRNRAEELLVELSANLPQ 265
KS +RA E+L E+ + P+
Sbjct: 280 FAKSYDRAVEMLAEIESKKPE 300
>UniRef100_Q84UQ9 Putative pollenless3 [Oryza sativa]
Length = 815
Score = 223 bits (567), Expect = 7e-57
Identities = 111/216 (51%), Positives = 155/216 (71%), Gaps = 1/216 (0%)
Query: 46 DTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKS 105
D+PYV+AK AQ+++KDP A+ FW AIN+GD+++SALKDMA V+KQ +RAEEAIEAI+S
Sbjct: 85 DSPYVRAKQAQVIEKDPNKAVPLFWAAINSGDRIESALKDMATVLKQANRAEEAIEAIRS 144
Query: 106 FRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGK 165
FR C +QESLDN+LLDLYKKCGR +EQIE+L KLR++ + A TK ++SHG+
Sbjct: 145 FRDRCPNEAQESLDNILLDLYKKCGRTKEQIEMLTLKLRIVDEELASGRWKTKLSKSHGR 204
Query: 166 KFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKALNLALCLMRQSRY 225
+S++ E ARLLGNL WA+MQ NY AE+++++A I+AD NK NLA+CL++ +
Sbjct: 205 VVYLSLRDEKARLLGNLAWAHMQSENYDEAEMLYRQALAIEADYNKECNLAICLIKTGKV 264
Query: 226 EEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELSA 261
EA +L Q + ++S RA E+L+EL +
Sbjct: 265 AEAKYLL-QSIPDNCSDESHVRSLARAREMLMELES 299
>UniRef100_Q9FKV5 Male sterility MS5; pollenless3 [Arabidopsis thaliana]
Length = 469
Score = 213 bits (542), Expect = 5e-54
Identities = 115/221 (52%), Positives = 148/221 (66%), Gaps = 3/221 (1%)
Query: 41 KVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAI 100
+V GD+PYV+AKHAQLV KDP AI FW AINAGD+VDSALKDM VV+KQL+R +E I
Sbjct: 49 RVRTGDSPYVRAKHAQLVSKDPNRAISLFWAAINAGDRVDSALKDMVVVLKQLNRFDEGI 108
Query: 101 EAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTA 160
EAIKSFR LC SQ+S+DN+LL+LY K GR+ E ELL+ KLR + Q + + GR
Sbjct: 109 EAIKSFRYLCPFESQDSIDNLLLELYMKSGRITEVAELLEHKLRTLEQDKHYGGRIKIAK 168
Query: 161 RSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKALNLALCLM 220
RSH ++ +I+QE AR+LGNL W ++Q NY +AE ++ A ++ D NK NLA+CL+
Sbjct: 169 RSHEEQNNKTIEQEKARILGNLAWVHLQLHNYGIAEQYYRNALSLEPDNNKLCNLAICLI 228
Query: 221 RQSRYEEAYLILEQVLQ---GKLPGSDEIKSRNRAEELLVE 258
R R EA +LE V Q + KS RA E+L E
Sbjct: 229 RMERTHEAKSLLEDVKQSLGNQWKNEPFCKSFERATEMLAE 269
>UniRef100_Q9ZTU4 MS5-like protein [Arabidopsis thaliana]
Length = 307
Score = 122 bits (305), Expect = 2e-26
Identities = 69/157 (43%), Positives = 96/157 (60%), Gaps = 5/157 (3%)
Query: 139 LKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVV 198
LK KL LI +G AFNG+ TKTARS GKKFQVS++QE RLLGNLGWA MQ+ N++ AE
Sbjct: 1 LKHKLFLIQKGLAFNGKRTKTARSQGKKFQVSVEQEATRLLGNLGWALMQRDNFVEAEDA 60
Query: 199 FKKAQMIDADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPG----SDEIKSRNRAEE 254
+++A I D NK NL +CLM+Q R +EA L +V + G +K+ RA++
Sbjct: 61 YRRALSIAPDNNKMCNLGICLMKQGRIDEAKETLRRVKPAVVDGPRGVDSHLKAYERAQQ 120
Query: 255 LLVELSANLPQPKFMDDLGLDDDLLKGIDGLLNVWSP 291
+L +L + + + + DD L I G ++W P
Sbjct: 121 MLNDLGSEMMR-RGGDDKVEQRRLFDAIFGSSSIWQP 156
>UniRef100_Q6AUY9 Hypothetical protein OSJNBa0004G03.21 [Oryza sativa]
Length = 274
Score = 82.0 bits (201), Expect = 2e-14
Identities = 48/143 (33%), Positives = 71/143 (49%), Gaps = 39/143 (27%)
Query: 57 LVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQE 116
+++KDP A+ FW AIN+GD+++SALK
Sbjct: 166 VIEKDPNKAVPLFWAAINSGDRIESALK-------------------------------- 193
Query: 117 SLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETA 176
D+ KC R +EQIE+L KL + + A TK ++SHG+ +S++ E A
Sbjct: 194 -------DMATKCDRTKEQIEMLTLKLIFVDEELASGRWKTKLSKSHGRVVYLSLRDEKA 246
Query: 177 RLLGNLGWAYMQKTNYMMAEVVF 199
LLGNL WA+MQ NY AE+++
Sbjct: 247 WLLGNLAWAHMQSENYDGAEMLY 269
>UniRef100_Q75I28 Hypothetical protein OSJNBa0091E13.29 [Oryza sativa]
Length = 160
Score = 69.7 bits (169), Expect = 9e-11
Identities = 29/47 (61%), Positives = 40/47 (84%)
Query: 46 DTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQ 92
D+PYV+AK AQ+++KDP A+ FW AIN+GD+++SALKDMA V+KQ
Sbjct: 112 DSPYVRAKQAQVIEKDPNKAVPLFWAAINSGDRIESALKDMATVLKQ 158
>UniRef100_UPI00002867B7 UPI00002867B7 UniRef100 entry
Length = 174
Score = 53.9 bits (128), Expect = 5e-06
Identities = 35/88 (39%), Positives = 48/88 (53%), Gaps = 3/88 (3%)
Query: 49 YVKAKHAQLVDKDPEVAIVYFWKAINA-GDKVDSALKDMAVVMKQLDRAEEAIEAIKSFR 107
Y +AKHAQL KD A + I + G+ SALKD+ ++KQ+ EA I +R
Sbjct: 86 YARAKHAQLTLKDLAQARDLMVQEIESKGESSQSALKDLVCILKQMGSHAEATATIVRYR 145
Query: 108 GLC--NKHSQESLDNVLLDLYKKCGRVE 133
+ QESLDN+LLDL+K +E
Sbjct: 146 AAWPDDDRMQESLDNMLLDLFKHARNLE 173
>UniRef100_Q8RHP5 Tetratricopeptide repeat family protein [Fusobacterium nucleatum]
Length = 345
Score = 47.8 bits (112), Expect = 4e-04
Identities = 53/189 (28%), Positives = 79/189 (41%), Gaps = 15/189 (7%)
Query: 61 DPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDN 120
D E + Y +A G + +M + +L+RAEE +E +K L + E+++
Sbjct: 89 DYEQGLKYLLEAEKLGRDDEWLNTEMGQCLGRLERAEEGLERLKKSLKLIETEAPENINE 148
Query: 121 VLL---DLYKKCGRVEEQIELLK---------RKLRLIYQGEAFNGRTTKTARSHGKKFQ 168
+ ++ G +E E LK RK IY FN +K K F+
Sbjct: 149 KIFINSEIGYLYGFLENSEEALKYFHIAKDLGRKDDWIYMHLWFNLERSKGKEEALKYFE 208
Query: 169 VSIKQE--TARLLGNLGWAYMQK-TNYMMAEVVFKKAQMIDADANKALNLALCLMRQSRY 225
K + A + +LG YM NY AE VFKKA + D N + L RY
Sbjct: 209 NEAKTDDKNAIVWASLGQIYMNFFRNYEEAEKVFKKAFGLSGDGLYLYNRGMVLRILGRY 268
Query: 226 EEAYLILEQ 234
+EA +L Q
Sbjct: 269 KEAVEVLLQ 277
>UniRef100_UPI00002E7317 UPI00002E7317 UniRef100 entry
Length = 455
Score = 42.7 bits (99), Expect = 0.012
Identities = 45/173 (26%), Positives = 76/173 (43%), Gaps = 11/173 (6%)
Query: 65 AIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLD 124
AI + KAIN A ++ + +++ +EAIE K L N + E+L+N L
Sbjct: 66 AIENYKKAININPNYAQAYNNLGIAFHKINNLDEAIENYKKALKLKNNFA-EALNN-LGS 123
Query: 125 LYKKCGRVEEQIELLKRKLRLIYQ-GEAFNGRTT--KTARSHGK-----KFQVSIKQETA 176
K + +E I ++ ++L EAFNG + + R + K K + I +
Sbjct: 124 ALKDLKKYKESISYFEKSIQLKENYAEAFNGLASVYEEVRDNEKAIENYKIALKINPNFS 183
Query: 177 RLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKAL-NLALCLMRQSRYEEA 228
NLG Y + + + ++KA ++ + KA NL L RY +A
Sbjct: 184 EAYNNLGSLYSNLVMFDKSLICYEKAIEVNPNNEKAYNNLGNLLNDLGRYNDA 236
>UniRef100_Q662F4 Surface-located membrane protein 1 [Borrelia garinii]
Length = 906
Score = 42.7 bits (99), Expect = 0.012
Identities = 49/227 (21%), Positives = 101/227 (43%), Gaps = 18/227 (7%)
Query: 46 DTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKS 105
DT Y + A+ + D + A F A N K++ ALK +V L +++ E +
Sbjct: 630 DTAYYQKGIAEEKNGDMQQAFTSFKNAYNIDKKLNYALK-AGIVSNNLGNFKKSEEYLSF 688
Query: 106 FRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL-------IYQGEAFNGRTTK 158
F K ++ ++ N+ + ++ ++EE +E++ + L L +Y + N +
Sbjct: 689 FNDNAKKPNEIAIYNLSIAKFEN-NKLEESLEIINKALNLNPEKSEYLYLKASINLKNEN 747
Query: 159 TARSHGKKFQVSIKQ-ETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKAL-NLA 216
+ V K E NL AY + N A+ + ++I+ + AL NL
Sbjct: 748 YQNAISLYSSVIEKNPENTSAYINLAKAYEKLGN--KAQAISTLEKIINKNNKLALNNLG 805
Query: 217 LCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELSANL 263
+ +Q Y++A I E+ ++ + +I+++ L+E++ N+
Sbjct: 806 ILYKKQKNYQKAIEIFEKAIK-----NSDIEAKYNLATTLIEINDNI 847
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 485,122,781
Number of Sequences: 2790947
Number of extensions: 19382462
Number of successful extensions: 56909
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 56690
Number of HSP's gapped (non-prelim): 351
length of query: 312
length of database: 848,049,833
effective HSP length: 127
effective length of query: 185
effective length of database: 493,599,564
effective search space: 91315919340
effective search space used: 91315919340
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC130200.5


