

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126787.14 - phase: 0
(370 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
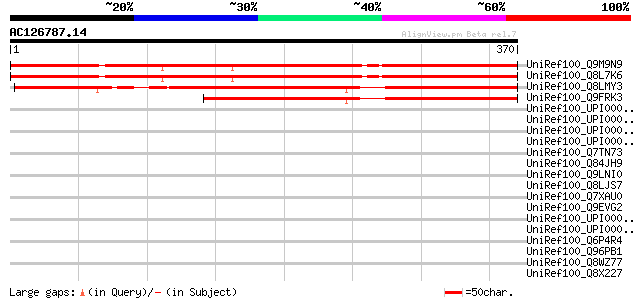
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M9N9 T17B22.10 protein [Arabidopsis thaliana] 422 e-116
UniRef100_Q8L7K6 Hypothetical protein At3g03210 [Arabidopsis tha... 419 e-116
UniRef100_Q8LMY3 Hypothetical protein OSJNBa0025B05.3 [Oryza sat... 333 6e-90
UniRef100_Q9FRK3 Hypothetical protein OSJNBb0064P21.16 [Oryza sa... 221 3e-56
UniRef100_UPI0000348884 UPI0000348884 UniRef100 entry 47 0.001
UniRef100_UPI0000365663 UPI0000365663 UniRef100 entry 45 0.003
UniRef100_UPI00003AB9C2 UPI00003AB9C2 UniRef100 entry 43 0.016
UniRef100_UPI000023EF23 UPI000023EF23 UniRef100 entry 42 0.027
UniRef100_Q7TN73 O-acetyltransferase [Mus musculus] 39 0.23
UniRef100_Q84JH9 Hypothetical protein At1g01430 [Arabidopsis tha... 39 0.23
UniRef100_Q9LNI0 F6F3.23 protein [Arabidopsis thaliana] 39 0.23
UniRef100_Q8LJS7 Homeodomain protein GhHOX2 [Gossypium hirsutum] 39 0.30
UniRef100_Q7XAU0 Homeodomain protein BNLGHi6863 [Gossypium hirsu... 39 0.30
UniRef100_Q9EVG2 3-isopropylmalate dehydratase large subunit [Bu... 38 0.51
UniRef100_UPI0000365664 UPI0000365664 UniRef100 entry 37 0.88
UniRef100_UPI000036DE1E UPI000036DE1E UniRef100 entry 37 0.88
UniRef100_Q6P4R4 O-acetyltransferase [Homo sapiens] 37 0.88
UniRef100_Q96PB1 C7ORF12 [Homo sapiens] 37 0.88
UniRef100_Q8WZ77 O-acetyltransferase [Homo sapiens] 37 0.88
UniRef100_Q8X227 O-acetyltransferase [Cryptococcus neoformans va... 37 0.88
>UniRef100_Q9M9N9 T17B22.10 protein [Arabidopsis thaliana]
Length = 368
Score = 422 bits (1084), Expect = e-116
Identities = 217/376 (57%), Positives = 274/376 (72%), Gaps = 14/376 (3%)
Query: 1 MFGAVQLGLFAACVVLFVPMGMAGWHLSRNKVLFFSGALFITLAVGVHLTPYFPSVSDFM 60
M GA+ LG+ AAC VLFVPM MAGWHLSRNK+LFFSGALFI+LAV VHLTPYFPSVSD +
Sbjct: 1 MLGAIHLGVLAACFVLFVPMAMAGWHLSRNKMLFFSGALFISLAVCVHLTPYFPSVSDIV 60
Query: 61 SSVKSSNDDVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNN-SLNYDKF---WSWSRL 116
+SV S VV + R C++ +++I W+V P + + RNN S D F W W +
Sbjct: 61 ASVSS----VVVYDHRISCINEVNQIVWDVKPVPNPESVRRNNGSTKLDYFVKNWDWMKS 116
Query: 117 NSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVL--DSERVESVRSFLF 174
V CEFQ+L + DV DLLNGSWV++ GDSQAR LSLL+LVL DS+ ++SVR LF
Sbjct: 117 RKVLSCEFQKLDKFDVSDLLNGSWVVVAGDSQARFVALSLLNLVLGSDSKAMDSVRGDLF 176
Query: 175 KRHSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNA 234
+RHSDY I + E+G+KLDF+WAPY +L DLV+ +KK YPDV++MG+GLWHMLH+ NA
Sbjct: 177 RRHSDYSIVVKEIGMKLDFVWAPYEKDLDDLVVSYKKMKKYPDVVIMGTGLWHMLHVNNA 236
Query: 235 SDYGGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEK 294
SD+G L L + V S++P++PK ++ GSVS RS HLFW+GMP LIN MLNT EK
Sbjct: 237 SDFGFRLRQLSSHVESLVPLTPK---EQEGGGSVSG-RSVHLFWIGMPVLINGMLNTDEK 292
Query: 295 RVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAE 354
+ KM+D + EY+R + +S ILR+ GGP LLDI S + NCG +CT DGMHYD AVY+A
Sbjct: 293 KEKMSDTVWHEYDRSLGESKILRQMGGPLILLDIQSFTWNCGPQCTLDGMHYDSAVYDAA 352
Query: 355 LHIMFNALLIESHQKL 370
+H+M NALLIESHQ L
Sbjct: 353 VHVMLNALLIESHQSL 368
>UniRef100_Q8L7K6 Hypothetical protein At3g03210 [Arabidopsis thaliana]
Length = 368
Score = 419 bits (1076), Expect = e-116
Identities = 216/376 (57%), Positives = 273/376 (72%), Gaps = 14/376 (3%)
Query: 1 MFGAVQLGLFAACVVLFVPMGMAGWHLSRNKVLFFSGALFITLAVGVHLTPYFPSVSDFM 60
M GA+ LG+ AAC VLFVPM MAGWHLSRNK+LFFSGALFI+LAV VHLTPYFPSVSD +
Sbjct: 1 MLGAIHLGVLAACFVLFVPMAMAGWHLSRNKMLFFSGALFISLAVCVHLTPYFPSVSDIV 60
Query: 61 SSVKSSNDDVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNN-SLNYDKF---WSWSRL 116
+SV S VV + R C++ +++I W+V P + + RNN S D F W W +
Sbjct: 61 ASVSS----VVVYDHRISCINEVNQIVWDVKPVPNPESVRRNNGSTKLDYFVKNWDWMKS 116
Query: 117 NSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVL--DSERVESVRSFLF 174
V CEFQ+L + DV DLLNGSWV++ GDSQAR LSLL+LVL DS+ ++SVR LF
Sbjct: 117 RKVLSCEFQKLDKFDVSDLLNGSWVVVAGDSQARFVALSLLNLVLGSDSKAMDSVRGDLF 176
Query: 175 KRHSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNA 234
+RHSDY I + E+G+KLDF+WAPY +L DLV+ +KK YPDV++M +GLWHMLH+ NA
Sbjct: 177 RRHSDYSIVVKEIGMKLDFVWAPYEKDLDDLVVSYKKMKKYPDVVIMRTGLWHMLHVNNA 236
Query: 235 SDYGGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEK 294
SD+G L L + V S++P++PK ++ GSVS RS HLFW+GMP LIN MLNT EK
Sbjct: 237 SDFGFRLRQLSSHVESLVPLTPK---EQEGGGSVSG-RSVHLFWIGMPVLINGMLNTDEK 292
Query: 295 RVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAE 354
+ KM+D + EY+R + +S ILR+ GGP LLDI S + NCG +CT DGMHYD AVY+A
Sbjct: 293 KEKMSDTVWHEYDRSLGESKILRQMGGPLILLDIQSFTWNCGPQCTLDGMHYDSAVYDAA 352
Query: 355 LHIMFNALLIESHQKL 370
+H+M NALLIESHQ L
Sbjct: 353 VHVMLNALLIESHQSL 368
>UniRef100_Q8LMY3 Hypothetical protein OSJNBa0025B05.3 [Oryza sativa]
Length = 351
Score = 333 bits (853), Expect = 6e-90
Identities = 179/378 (47%), Positives = 229/378 (60%), Gaps = 42/378 (11%)
Query: 4 AVQLGLFAACVVLFVPMGMAGWHLSRNKVLFFSGALFITLAVGVHLTPYFPSVSDFMSS- 62
A Q G+ AACVVLFVPMG+AGWHLSRNKVLFFSGALF++LAVGVHL+PY PS+ + +
Sbjct: 5 AAQAGVVAACVVLFVPMGLAGWHLSRNKVLFFSGALFVSLAVGVHLSPYLPSLPHLLLAA 64
Query: 63 ------VKSSNDDVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNNSLNYDKFWSWSRL 116
+ SS+ CV LLH ++W + W+W
Sbjct: 65 SFHPHPISSSSSSSAASSS---CVPLLHRVSWADA----------GGESGVGRAWAWPP- 110
Query: 117 NSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKR 176
+ S C RL R D LLNGSWVM+ GDSQAR+ L+LL L+LD + LF+R
Sbjct: 111 SLASTCGLARLSRDDASLLLNGSWVMVAGDSQARLLVLALLRLLLDPAAAAAAEPELFRR 170
Query: 177 HSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMR-FKKKHLYPDVLVMGSGLWHMLHITNAS 235
HSDY + G+ +DF+WAP+ +NLT L+ + PDVLV+GSGLWHMLH+T+A+
Sbjct: 171 HSDYRATVPARGISVDFVWAPFESNLTRLLHEDLRLAPRTPDVLVLGSGLWHMLHVTDAA 230
Query: 236 DYGGLLGLL---RNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQ 292
YG L + S+ S LPV P PH+FWLG+P L+N MLNT
Sbjct: 231 RYGDALASVVDAAKSLRSPLPVPP-----------------PHMFWLGLPLLVNHMLNTD 273
Query: 293 EKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYE 352
K+V MND + Y+ EV + G+L+ GGPF LLD+G LS CG +CT DGMHYDG VY+
Sbjct: 274 AKKVHMNDTILQAYDLEVEQRGLLQRDGGPFLLLDVGKLSRGCGQQCTADGMHYDGDVYD 333
Query: 353 AELHIMFNALLIESHQKL 370
A LHIM NAL+IES Q++
Sbjct: 334 AVLHIMLNALVIESQQRI 351
>UniRef100_Q9FRK3 Hypothetical protein OSJNBb0064P21.16 [Oryza sativa]
Length = 216
Score = 221 bits (563), Expect = 3e-56
Identities = 113/233 (48%), Positives = 148/233 (63%), Gaps = 21/233 (9%)
Query: 142 MIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYHIDIDEMGLKLDFIWAPYPNN 201
M+ GDSQAR+ L+LL L+LD + LF+RHSDY + G+ +DF+WAP+ +N
Sbjct: 1 MVAGDSQARLLVLALLRLLLDPAAAAAAEPELFRRHSDYRATVPARGISVDFVWAPFESN 60
Query: 202 LTDLVMR-FKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLL---RNSVTSMLPVSPK 257
LT L+ + PDVLV+GSGLWHMLH+T+A+ YG L + S+ S LPV P
Sbjct: 61 LTRLLHEDLRLAPRTPDVLVLGSGLWHMLHVTDAARYGDALASVVDAAKSLRSPLPVPP- 119
Query: 258 FGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMNDVMQGEYEREVRKSGILR 317
PH+FWLG+P L+N MLNT K+V MND + Y+ EV + G+L+
Sbjct: 120 ----------------PHMFWLGLPLLVNHMLNTDAKKVHMNDTILQAYDLEVEQRGLLQ 163
Query: 318 EFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIMFNALLIESHQKL 370
GGPF LLD+G LS CG +CT DGMHYDG VY+A LHIM NAL+IES Q++
Sbjct: 164 RDGGPFLLLDVGKLSRGCGQQCTADGMHYDGDVYDAVLHIMLNALVIESQQRI 216
>UniRef100_UPI0000348884 UPI0000348884 UniRef100 entry
Length = 296
Score = 46.6 bits (109), Expect = 0.001
Identities = 54/220 (24%), Positives = 89/220 (39%), Gaps = 35/220 (15%)
Query: 176 RHSDYHIDIDEM--GLKLDFIWAPYPNNLT---DLVMRFKKKHLYPDVLVMGSGLW---- 226
RH+D + + G L F+WAPY NLT D ++ KK PDV+VMG+GLW
Sbjct: 15 RHADISMVLPPQLGGGNLRFLWAPYAANLTTRFDKLVSSSKK--MPDVVVMGAGLWDALW 72
Query: 227 ---------HMLHITNASDYGGLL-GLLR-----NSVTSMLPVSPKFGNDEPAMGSVSSV 271
H L + ++ + G+ R + + + N A VS +
Sbjct: 73 IRDPVAFKRHALQLQSSKQKQRFVDGIARRQGSKSDILKRVESGKGDLNRRRANLDVSGI 132
Query: 272 RSPHLFWLGMPTLINSMLNTQEKRVKMND-VMQGEYE-REVRKSGILREFGGPFQLLDIG 329
P WL +++ L KR M D +++ +Y V + + P D G
Sbjct: 133 ALPLFMWLDTTAVVDERLAAAPKREHMRDAIVERQYSYHGVNGNAVKFRTQLPLYKPDNG 192
Query: 330 ------SLSLNCGIKCTD-DGMHYDGAVYEAELHIMFNAL 362
+ ++ G + DG+HY VY+ ++ N +
Sbjct: 193 LDGVIEARAVTAGREFDSVDGVHYVDVVYDVLAQVLLNQI 232
>UniRef100_UPI0000365663 UPI0000365663 UniRef100 entry
Length = 777
Score = 45.1 bits (105), Expect = 0.003
Identities = 53/244 (21%), Positives = 95/244 (38%), Gaps = 25/244 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + L G V +GDS+ R S + +LD ER E +H D
Sbjct: 69 CMMHKYKSTEAKSCLAGKRVAFLGDSRIRQLFYSFIK-ILDPERREDGN-----KHEDIS 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTD--LVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ + +DF+W P NN LV ++ PDV+V+ + W + + +S+
Sbjct: 123 FEDPGSSVNVDFLWHPEANNSMKERLVSWTQETSPKPDVVVLAAAAWSIKLHSGSSE--- 179
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L R ++T+M A + ++W+ + L+ K +
Sbjct: 180 ALQQYRANLTAM------------AAHLEQLAQHGEVYWVLQDPVNEEALSDSRKMITNQ 227
Query: 300 DV-MQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E EV K G + G +LL + I +DDG+H + ++
Sbjct: 228 QLELYNEAAVEVLKPG-PADGGSRVKLLAATRQAAMATIAQSDDGLHLPESTRSVGAAVL 286
Query: 359 FNAL 362
N++
Sbjct: 287 MNSV 290
>UniRef100_UPI00003AB9C2 UPI00003AB9C2 UniRef100 entry
Length = 756
Score = 42.7 bits (99), Expect = 0.016
Identities = 47/244 (19%), Positives = 92/244 (37%), Gaps = 29/244 (11%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + L ++ +GDS+ R S + ++ + E +H +
Sbjct: 27 CMMHKYKNSEAKSCLIDKHIVFLGDSRIRQLFYSFIKIINPQIKEEG------NKHGNIP 80
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRFKK----KHLYPDVLVMGSGLWHMLHITNASDY 237
+ +K+DF+W P N + R K P ++V G+ W + I N S+
Sbjct: 81 FEDKSASIKVDFLWYPEVNG--SMRQRIKSWTEGSVAKPHIIVAGAATW-SIKIHNGSNE 137
Query: 238 GGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVK 297
L + ++TS+ P+ K +S ++W+ + ML+ K +
Sbjct: 138 A--LTQYKINITSIAPLLEKL------------AKSSDVYWILQDPVYEDMLSESRKMI- 182
Query: 298 MNDVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELH 356
N+ + E VR + R ++ + L I + DG+H + +
Sbjct: 183 TNEKIDAYNEAAVRILNSSSRNSKAKVKVFSVSKLIAQETIMKSSDGLHLPESSRDMNAM 242
Query: 357 IMFN 360
I+ N
Sbjct: 243 ILMN 246
>UniRef100_UPI000023EF23 UPI000023EF23 UniRef100 entry
Length = 851
Score = 42.0 bits (97), Expect = 0.027
Identities = 32/119 (26%), Positives = 56/119 (46%), Gaps = 16/119 (13%)
Query: 120 SECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSD 179
S C + K++++ + G ++ VGDS R + L LD+E+ ++VR+ K S
Sbjct: 69 SGCMLHKYKKEEIEQCMEGRHMLFVGDSTTRQVFYGMARL-LDAEKAQTVRAKSQKHESH 127
Query: 180 YHIDIDEMGLKLDFIWAPYPNN---LTDLVMRFKKKHL---------YPDVLVMGSGLW 226
D + G++L IW PY N + D + + ++ L P V +G G+W
Sbjct: 128 ---DEEFGGIRLKSIWDPYGNESDIVRDELSLYHQEKLDRVPVEEQKGPAVAFVGMGVW 183
>UniRef100_Q7TN73 O-acetyltransferase [Mus musculus]
Length = 797
Score = 38.9 bits (89), Expect = 0.23
Identities = 45/242 (18%), Positives = 91/242 (37%), Gaps = 25/242 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + L + +GDS+ R S + ++ + E +H +
Sbjct: 69 CMMHKYKISEAKTCLVDKHIAFIGDSRIRQLFYSFVKIINPQFKEEG------NKHENIP 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRF--KKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ +K+DF+W P N ++ + L P V+V G+ W + I N S+
Sbjct: 123 FEDKAASVKVDFLWHPEVNGSMKQCIKVWTEDSVLKPHVIVAGAATW-SIKIHNGSEEA- 180
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L + ++TS+ P+ K ++ ++W+ + +L+ K + N
Sbjct: 181 -LAQYKMNITSIAPLLEKL------------AKTSDVYWVLQDPVYEDLLSENRKMI-TN 226
Query: 300 DVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E V + R ++ + L I + DG+H + E I+
Sbjct: 227 EKIDAYNEAAVSILNSSTRTSKSNVKMFSVSKLIAQETIMESLDGLHLPESSRETSAMIL 286
Query: 359 FN 360
N
Sbjct: 287 MN 288
>UniRef100_Q84JH9 Hypothetical protein At1g01430 [Arabidopsis thaliana]
Length = 456
Score = 38.9 bits (89), Expect = 0.23
Identities = 22/88 (25%), Positives = 36/88 (40%), Gaps = 6/88 (6%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C+ R + +D + W+ +GDS +R SLL ++ E VE +
Sbjct: 142 WRWQP----RDCDLPRFNPEQFLDNMRNKWLAFIGDSISRNHVQSLLCILSQVEEVEDI- 196
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
F K + L IW+P+
Sbjct: 197 -FHDKEYKSRIWRFPSYNFTLSVIWSPF 223
>UniRef100_Q9LNI0 F6F3.23 protein [Arabidopsis thaliana]
Length = 442
Score = 38.9 bits (89), Expect = 0.23
Identities = 22/88 (25%), Positives = 36/88 (40%), Gaps = 6/88 (6%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C+ R + +D + W+ +GDS +R SLL ++ E VE +
Sbjct: 128 WRWQP----RDCDLPRFNPEQFLDNMRNKWLAFIGDSISRNHVQSLLCILSQVEEVEDI- 182
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
F K + L IW+P+
Sbjct: 183 -FHDKEYKSRIWRFPSYNFTLSVIWSPF 209
>UniRef100_Q8LJS7 Homeodomain protein GhHOX2 [Gossypium hirsutum]
Length = 775
Score = 38.5 bits (88), Expect = 0.30
Identities = 19/56 (33%), Positives = 32/56 (56%)
Query: 195 WAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLRNSVTS 250
W PYP++ ++R +++ +VL G+ L + HI N S G + LLR +V+S
Sbjct: 585 WLPYPHHHVFDLLRDERRRAQLEVLSNGNALHEVAHIANGSHPGNCISLLRINVSS 640
>UniRef100_Q7XAU0 Homeodomain protein BNLGHi6863 [Gossypium hirsutum]
Length = 762
Score = 38.5 bits (88), Expect = 0.30
Identities = 19/56 (33%), Positives = 32/56 (56%)
Query: 195 WAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLRNSVTS 250
W PYP++ ++R +++ +VL G+ L + HI N S G + LLR +V+S
Sbjct: 572 WLPYPHHHVFDLLRDERRRAQLEVLSNGNALHEVAHIANGSHPGNCISLLRINVSS 627
>UniRef100_Q9EVG2 3-isopropylmalate dehydratase large subunit [Buchnera aphidicola]
Length = 442
Score = 37.7 bits (86), Expect = 0.51
Identities = 30/115 (26%), Positives = 56/115 (48%), Gaps = 9/115 (7%)
Query: 58 DFMSSVKSSND---DVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNNSLNYDKFWSWS 114
DF ++KS D D +F D +L +I W P +V + NY+ F S +
Sbjct: 260 DFWQTLKSDKDAFFDKIFTIDIS---NLAPQITWGTNPDQVIS--IDEKIPNYETFSSLT 314
Query: 115 RLN-SVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVES 168
+ N + S C++ LK++ + ++ V I + ARI L L + +L ++++ +
Sbjct: 315 QRNLAKSACKYMGLKKESYLTNISIDKVFIGSXTNARIEDLRLAAKILKNKKISN 369
>UniRef100_UPI0000365664 UPI0000365664 UniRef100 entry
Length = 149
Score = 37.0 bits (84), Expect = 0.88
Identities = 23/80 (28%), Positives = 34/80 (41%), Gaps = 6/80 (7%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + L G V +GDS+ R S + +LD ER E +H D
Sbjct: 24 CMMHKYKSTEAKSCLAGKRVAFLGDSRIRQLFYSFIK-ILDPERREDGN-----KHEDIS 77
Query: 182 IDIDEMGLKLDFIWAPYPNN 201
+ + +DF+W P NN
Sbjct: 78 FEDPGSSVNVDFLWHPEANN 97
>UniRef100_UPI000036DE1E UPI000036DE1E UniRef100 entry
Length = 797
Score = 37.0 bits (84), Expect = 0.88
Identities = 44/242 (18%), Positives = 90/242 (37%), Gaps = 25/242 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + + L + +GDS+ R S + ++ + E +H +
Sbjct: 69 CMMHKYKISEAKNCLVDKHIAFIGDSRIRQLFYSFVKIINPQFKEEG------NKHENIP 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRF--KKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ +K+DF+W P N ++ + P V+V G+ W + I N S
Sbjct: 123 FEDKTASVKVDFLWHPEVNGSMKQCIKVWTEDSIAKPHVIVAGAATW-SIKIHNGSSEA- 180
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L + ++TS+ P+ K ++ ++W+ + +L+ K + N
Sbjct: 181 -LSQYKMNITSIAPLLEKL------------AKTSDVYWVLQDPVYEDLLSENRKMI-TN 226
Query: 300 DVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E V + R ++ + L I + DG+H + E I+
Sbjct: 227 EKIDAYNEAAVSILNSSTRNSKSNVKMFSVSKLIAQETIMESLDGLHLPESSRETTAMIL 286
Query: 359 FN 360
N
Sbjct: 287 MN 288
>UniRef100_Q6P4R4 O-acetyltransferase [Homo sapiens]
Length = 797
Score = 37.0 bits (84), Expect = 0.88
Identities = 44/242 (18%), Positives = 90/242 (37%), Gaps = 25/242 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + + L + +GDS+ R S + ++ + E +H +
Sbjct: 69 CMMHKYKISEAKNCLVDKHIAFIGDSRIRQLFYSFVKIINPQFKEEG------NKHENIP 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRF--KKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ +K+DF+W P N ++ + P V+V G+ W + I N S
Sbjct: 123 FEDKTASVKVDFLWHPEVNGSMKQCIKVWTEDSIAKPHVIVAGAATW-SIKIHNGSSEA- 180
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L + ++TS+ P+ K ++ ++W+ + +L+ K + N
Sbjct: 181 -LSQYKMNITSIAPLLEKL------------AKTSDVYWVLQDPVYEDLLSENRKMI-TN 226
Query: 300 DVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E V + R ++ + L I + DG+H + E I+
Sbjct: 227 EKIDAYNEAAVSILNSSTRNSKSNVKMFSVSKLIAQETIMESLDGLHLPESSRETTAMIL 286
Query: 359 FN 360
N
Sbjct: 287 MN 288
>UniRef100_Q96PB1 C7ORF12 [Homo sapiens]
Length = 797
Score = 37.0 bits (84), Expect = 0.88
Identities = 44/242 (18%), Positives = 90/242 (37%), Gaps = 25/242 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + + L + +GDS+ R S + ++ + E +H +
Sbjct: 69 CMMHKYKISEAKNCLVDKHIAFIGDSRIRQLFYSFVKIINPQFKEEG------NKHENIP 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRF--KKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ +K+DF+W P N ++ + P V+V G+ W + I N S
Sbjct: 123 FEDKTASVKVDFLWHPEVNGSMKQCIKVWTEDSIAKPHVIVAGAATW-SIKIHNGSSEA- 180
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L + ++TS+ P+ K ++ ++W+ + +L+ K + N
Sbjct: 181 -LSQYKMNITSIAPLLEKL------------AKTSDVYWVLQDPVYEDLLSENRKMI-TN 226
Query: 300 DVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E V + R ++ + L I + DG+H + E I+
Sbjct: 227 EKIDAYNEAAVSILNSSTRNSKSNVKMFSVSKLIAQETIMESLDGLHLPESSRETTAMIL 286
Query: 359 FN 360
N
Sbjct: 287 MN 288
>UniRef100_Q8WZ77 O-acetyltransferase [Homo sapiens]
Length = 797
Score = 37.0 bits (84), Expect = 0.88
Identities = 44/242 (18%), Positives = 90/242 (37%), Gaps = 25/242 (10%)
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYH 181
C + K + + L + +GDS+ R S + ++ + E +H +
Sbjct: 69 CMMHKYKISEAKNCLVDKHIAFIGDSRIRQLFYSFVKIINPQFKEEG------NKHENIP 122
Query: 182 IDIDEMGLKLDFIWAPYPNNLTDLVMRF--KKKHLYPDVLVMGSGLWHMLHITNASDYGG 239
+ +K+DF+W P N ++ + P V+V G+ W + I N S
Sbjct: 123 FEDKTASVKVDFLWHPEVNGSMKQCIKVWTEDSIAKPHVIVAGAATW-SIKIHNGSSEA- 180
Query: 240 LLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEKRVKMN 299
L + ++TS+ P+ K ++ ++W+ + +L+ K + N
Sbjct: 181 -LSQYKMNITSIAPLLEKL------------AKTSDVYWVLQDPVYEDLLSENRKMI-TN 226
Query: 300 DVMQGEYEREVR-KSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAELHIM 358
+ + E V + R ++ + L I + DG+H + E I+
Sbjct: 227 EKIDAYNEAAVSILNSSTRNSKSNVKMFSVSKLIAQETIMESLDGLHLPESSRETTAMIL 286
Query: 359 FN 360
N
Sbjct: 287 MN 288
>UniRef100_Q8X227 O-acetyltransferase [Cryptococcus neoformans var. neoformans]
Length = 960
Score = 37.0 bits (84), Expect = 0.88
Identities = 25/91 (27%), Positives = 46/91 (50%), Gaps = 7/91 (7%)
Query: 142 MIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYHIDI-DEMG---LKLDFIWAP 197
+ VGDS R + V + + + ++H+D + + D +G L+L+F W P
Sbjct: 110 LFVGDSTVRQLYFAAARKVGKTSKAWELEG---EKHTDRSLLVSDPLGGPSLELEFWWDP 166
Query: 198 YPNNLTDLVMRFKKKHLYPDVLVMGSGLWHM 228
Y N+ + + + + +LVMGSGLW++
Sbjct: 167 YLNSSKTIGLLSGQSSVPSSLLVMGSGLWYL 197
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 629,746,658
Number of Sequences: 2790947
Number of extensions: 26554636
Number of successful extensions: 56501
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 56467
Number of HSP's gapped (non-prelim): 50
length of query: 370
length of database: 848,049,833
effective HSP length: 129
effective length of query: 241
effective length of database: 488,017,670
effective search space: 117612258470
effective search space used: 117612258470
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC126787.14


