

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
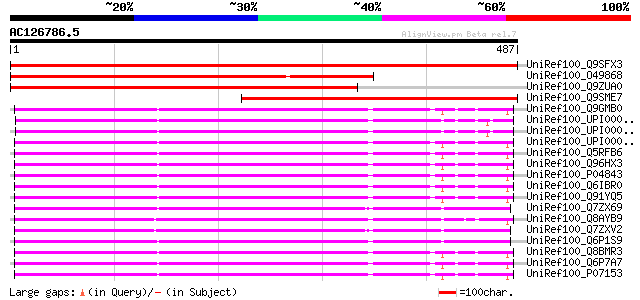
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SFX3 Putative ribophorin I (Dolichyl-diphosphooligos... 719 0.0
UniRef100_O49868 Putative ribophorin I homologue [Hordeum vulgare] 550 e-155
UniRef100_Q9ZUA0 Putative ribophorin I [Arabidopsis thaliana] 406 e-112
UniRef100_Q9SME7 Ribophorin I [Hordeum vulgare] 351 3e-95
UniRef100_Q9GMB0 Ribophorin I precursor [Sus scrofa] 319 1e-85
UniRef100_UPI00003AA8A8 UPI00003AA8A8 UniRef100 entry 319 1e-85
UniRef100_UPI00003AA8A7 UPI00003AA8A7 UniRef100 entry 319 1e-85
UniRef100_UPI000036B734 UPI000036B734 UniRef100 entry 318 2e-85
UniRef100_Q5RFB6 Hypothetical protein DKFZp469O2032 [Pongo pygma... 318 2e-85
UniRef100_Q96HX3 Similar to ribophorin I [Homo sapiens] 316 8e-85
UniRef100_P04843 Dolichyl-diphosphooligosaccharide--protein glyc... 316 8e-85
UniRef100_Q6IBR0 RPN1 protein [Homo sapiens] 315 2e-84
UniRef100_Q91YQ5 Dolichyl-diphosphooligosaccharide--protein glyc... 315 2e-84
UniRef100_Q7ZX69 MGC52894 protein [Xenopus laevis] 313 5e-84
UniRef100_Q8AYB9 Ribophorin I [Brachydanio rerio] 313 9e-84
UniRef100_Q7ZXV2 MGC52808 protein [Xenopus laevis] 312 2e-83
UniRef100_Q6P1S9 Hypothetical protein MGC76266 [Xenopus tropicalis] 311 2e-83
UniRef100_Q8BMR3 Mus musculus adult male testis cDNA, RIKEN full... 311 2e-83
UniRef100_Q6P7A7 Ribophorin I [Rattus norvegicus] 311 2e-83
UniRef100_P07153 Dolichyl-diphosphooligosaccharide--protein glyc... 308 2e-82
>UniRef100_Q9SFX3 Putative ribophorin I (Dolichyl-diphosphooligosaccharide-protein
glycosyltransferase); 43789-46748 [Arabidopsis thaliana]
Length = 614
Score = 719 bits (1855), Expect = 0.0
Identities = 350/487 (71%), Positives = 416/487 (84%)
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
++++ FT++LQPFPEKITQ +I L++ QESAQYLSPY V+ QSL++KLP ARIESYTK
Sbjct: 128 LEVVAAFTNVLQPFPEKITQGEIHLVMLQESAQYLSPYAVESQSLSIKLPNARIESYTKF 187
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
ENTKLQGSELKYGPY+NL +SY PIV+HFE+ FAVA++LVREIE+SHWGNVQ+TE+Y
Sbjct: 188 ENTKLQGSELKYGPYKNLQSYSYSPIVVHFESKAAFAVAEKLVREIEVSHWGNVQVTENY 247
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
N+VH GAQ KGEFSRLD+QARP RGASAFR L A+LPPRAHS+YYRD+IGNISTS +
Sbjct: 248 NVVHRGAQLKGEFSRLDFQARPNPRGASAFRHLLARLPPRAHSIYYRDDIGNISTSEMKS 307
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVIDTL 240
DSKKTEL IEPR+PLFGGWKT FTIGYGLPL DFLF +GKRFLNISFGSPI +LV + L
Sbjct: 308 DSKKTELLIEPRFPLFGGWKTFFTIGYGLPLTDFLFASEGKRFLNISFGSPILDLVTEKL 367
Query: 241 VVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYK 300
+V+VVLPEGSKDIS + PF VK+ E K+SHLDIAGRPVVVLEKNN VP+HN+H QVYYK
Sbjct: 368 IVQVVLPEGSKDISVTTPFAVKQSQEIKYSHLDIAGRPVVVLEKNNVVPDHNQHIQVYYK 427
Query: 301 FNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSI 360
F+ +++L EPLMLISGFF LF+ I+Y AD+SISK+S SYLAK+QW+EV AT+ +V SI
Sbjct: 428 FSNINLLSEPLMLISGFFILFITCIIYTRADISISKSSPSYLAKLQWDEVLATLQEVQSI 487
Query: 361 VSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQ 420
V +CL THDKLEASLRDLSRTGDIQ CKA RKS DS LK+L+KELK + FLQS P A+
Sbjct: 488 VQKCLATHDKLEASLRDLSRTGDIQTCKAARKSTDSLLKDLSKELKPLLGFLQSFPSASH 547
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDL 480
I PKVEE++ KE++LQEKLMAKH+TVV+ YEKK GR+IENRIAS QQKI AL+QEI+DL
Sbjct: 548 ISPKVEELVVKEKELQEKLMAKHTTVVEGYEKKSSGRDIENRIASQQQKIIALRQEIEDL 607
Query: 481 MDLIDEI 487
++ IDEI
Sbjct: 608 LEFIDEI 614
>UniRef100_O49868 Putative ribophorin I homologue [Hordeum vulgare]
Length = 473
Score = 550 bits (1417), Expect = e-155
Identities = 255/349 (73%), Positives = 304/349 (87%), Gaps = 2/349 (0%)
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+D+LTVFTH LQPFPE+ITQAD QL++FQ+S+ YLSPYPVKVQ+L+++LP R+ESYTK
Sbjct: 127 LDVLTVFTHFLQPFPEEITQADSQLVVFQDSSHYLSPYPVKVQTLSIRLPGGRVESYTKY 186
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
NTKL SELKYGPYE++PPFSY PI++HFENN PFAVAKEL+REIEISHWGNVQITEHY
Sbjct: 187 GNTKLVESELKYGPYEDVPPFSYNPIIVHFENNNPFAVAKELIREIEISHWGNVQITEHY 246
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
N+VHGGA+ KGEFSRLDYQ+RP RG S+FR L A+LPPRAHS+YYRDEIGNISTS LW
Sbjct: 247 NIVHGGARLKGEFSRLDYQSRPNARGVSSFRHLIARLPPRAHSIYYRDEIGNISTSHLWS 306
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVIDTL 240
DSKKT+LEIEPR+PLFGGW+T FTIGYGLPL+DF+F DG+RFLNI+FGSP+ E++I+ L
Sbjct: 307 DSKKTQLEIEPRFPLFGGWQTTFTIGYGLPLEDFVFSSDGRRFLNITFGSPMEEILIEKL 366
Query: 241 VVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYK 300
+VKVVLPEGSKD S PFP + E +SHLDIAGRPV++LEK + +PEHN HFQVYYK
Sbjct: 367 IVKVVLPEGSKDFDVSAPFPANQWQE--YSHLDIAGRPVLILEKADVIPEHNLHFQVYYK 424
Query: 301 FNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEE 349
FN +S+L EP+MLI+GFF LF+A I YMH D+SISK S SYLAK+QW+E
Sbjct: 425 FNNISLLIEPMMLITGFFLLFVACIAYMHTDMSISKNSPSYLAKLQWDE 473
>UniRef100_Q9ZUA0 Putative ribophorin I [Arabidopsis thaliana]
Length = 464
Score = 406 bits (1043), Expect = e-112
Identities = 193/335 (57%), Positives = 250/335 (74%), Gaps = 1/335 (0%)
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+++L + TH L+PFP +ITQ++ QL+ + +SA LSPY VK Q+ +K P R+ES+T +
Sbjct: 127 LEVLYILTHSLEPFPVEITQSESQLVYYHDSAVILSPYHVKQQTTFIKTPSTRVESFTSI 186
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
E G E+KYGPYEN +SY P++IHFENN PFAV +ELVREIEISHWG++QITE+Y
Sbjct: 187 EPANRAGKEIKYGPYENRASYSYTPVIIHFENNSPFAVVEELVREIEISHWGSLQITENY 246
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
L HGGA+ KG FSR+DYQ++ V GAS+F L A LPPR +SVYYRDEIGNISTS L
Sbjct: 247 RLTHGGARHKGVFSRVDYQSKRSVSGASSFNALLAVLPPRVNSVYYRDEIGNISTSHLRT 306
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLF-GVDGKRFLNISFGSPINELVIDT 239
+K+ELE EPRYPLFGGW F IGY +PL+D+LF DG+R+LN +FG P+ E +++
Sbjct: 307 GFRKSELEFEPRYPLFGGWSATFIIGYRVPLEDYLFEASDGRRYLNFTFGCPLVETIVNK 366
Query: 240 LVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYY 299
L +KVVLPEGSKD S +PF V + + K+S+LDI GR VVVL+K+N VP HN FQVYY
Sbjct: 367 LTIKVVLPEGSKDPSAVLPFTVNQELQVKYSYLDIVGRTVVVLQKDNVVPTHNVPFQVYY 426
Query: 300 KFNRLSMLREPLMLISGFFFLFLASIVYMHADLSI 334
F + ML EP ML+S FF +F+AS+ Y+H DL+I
Sbjct: 427 TFKPIYMLAEPFMLVSAFFLVFVASLAYVHIDLNI 461
>UniRef100_Q9SME7 Ribophorin I [Hordeum vulgare]
Length = 265
Score = 351 bits (900), Expect = 3e-95
Identities = 166/265 (62%), Positives = 213/265 (79%)
Query: 223 FLNISFGSPINELVIDTLVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVL 282
FLNI+FGSP+ E++I+ L+VKVVLPEGSKD S PFP + E K+SHLDIAGRPV++L
Sbjct: 1 FLNITFGSPMEEILIEKLIVKVVLPEGSKDFDVSAPFPANQWQEVKYSHLDIAGRPVLIL 60
Query: 283 EKNNAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYL 342
EK + +PEHN HFQVYYKFN +S+L EP+MLI+GFF LF+A I YMH D+SISK S SYL
Sbjct: 61 EKADVIPEHNLHFQVYYKFNNISLLIEPMMLITGFFLLFVACIAYMHTDMSISKNSPSYL 120
Query: 343 AKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELT 402
AK+QW+E+QAT+ Q+ I +CL HDKLE SL DLSRTGD + CKATRK+ D+ KEL
Sbjct: 121 AKLQWDEMQATVQQIQGIFEQCLAVHDKLEISLHDLSRTGDTKSCKATRKAADAQFKELA 180
Query: 403 KELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENR 462
KELK + +QSSPQ+ I PK+E+++AKER++QEKLMA+H+TVVD +EKK G++IENR
Sbjct: 181 KELKPLLLSVQSSPQSYLIWPKLEDLVAKEREMQEKLMARHATVVDSFEKKQRGQDIENR 240
Query: 463 IASHQQKITALKQEIDDLMDLIDEI 487
IAS QQKI AL+QE++ L++ + EI
Sbjct: 241 IASQQQKIAALRQEVESLLEYLSEI 265
>UniRef100_Q9GMB0 Ribophorin I precursor [Sus scrofa]
Length = 608
Score = 319 bits (818), Expect = 1e-85
Identities = 180/488 (36%), Positives = 282/488 (56%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+LQP+P +ITQ++ Q ++F+ + + SPYP K QS+ VKL +ESYTKL N
Sbjct: 134 TVYTHVLQPYPTQITQSEKQFVVFEGNHYFYSPYPTKSQSMRVKLASRNVESYTKLGNPT 193
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++PP+S +H ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 194 RSEDLLDYGPFRDVPPYSQDTFKVHSENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 253
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 254 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 312
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + + +E VID+L VK
Sbjct: 313 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRLVDHVFDEQVIDSLTVK 372
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 373 IILPEGAKNIQVDSPYEISRAPDELHYTYLDTFGRPVIVAHKKNLVEQHIQDIVVHYTFN 432
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF IVY+ D SI+K A+ +V QV ++V+
Sbjct: 433 KVLMLQEPLLVVAAFYILFFTVIVYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 488
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ D+ + +KS++ K LT E V LQS + +
Sbjct: 489 KRIGLYRHFDETINRYKQSRDVSTLNSGKKSLELEHKALTSE----VALLQSRLKTEGSD 544
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ KV E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 545 LCDKVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 600
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 601 IDHILDAL 608
>UniRef100_UPI00003AA8A8 UPI00003AA8A8 UniRef100 entry
Length = 541
Score = 319 bits (817), Expect = 1e-85
Identities = 181/486 (37%), Positives = 283/486 (57%), Gaps = 19/486 (3%)
Query: 6 VFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTKL 65
VFTH+LQP+P ITQ++ Q ++F+ + + SPY K Q+ VKL +ESYTKL N
Sbjct: 68 VFTHVLQPYPTHITQSEKQFVVFEGNHYFYSPYFTKTQTTRVKLASRNVESYTKLGNPSR 127
Query: 66 QGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVHG 125
++YGP++++PP+S + +H+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 128 TEDMIEYGPFKDIPPYSQDTLKVHYENNSPFLTITSMTRVIEVSHWGNIAVEETVDLKHT 187
Query: 126 GAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKKT 185
GA KG FSR DYQ +P G S+ + LP A VYYRDEIGNISTS L
Sbjct: 188 GAVLKGPFSRYDYQRQP-DSGISSVKSFKTILPAAAQDVYYRDEIGNISTSHLLVLDDSV 246
Query: 186 ELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVKV 244
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + F + +E V D+L VK+
Sbjct: 247 EMEIRPRFPLFGGWKTHYIIGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVTDSLTVKI 306
Query: 245 VLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFNR 303
VLPEG+K+I P+ + + E +++LD GRPV+V KNN V +H + V+Y FN+
Sbjct: 307 VLPEGAKNIHVDSPYEINRASDELHYTYLDTFGRPVIVAHKNNLVEQHIQDIVVHYTFNK 366
Query: 304 LSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSR 363
+ ML+EPL+++ F+ LF I+Y+ D SI+K A+ +V QV ++V++
Sbjct: 367 ILMLQEPLLVVGAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVNK 422
Query: 364 CLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILP 423
L + + ++ ++ DI + +KS+++ K LT E+ + + L++ + + +
Sbjct: 423 RLGLYRHFDEAVNKYKQSRDISTLNSGKKSLETEHKALTNEIAALQSKLKT--EGSDLCD 480
Query: 424 KVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGR-----EIENRIASHQQKITALKQEID 478
KV E+ ++ D Q K + S+V E+ + G+ IEN H K L +ID
Sbjct: 481 KVSEI--QKLDGQVKELVLKSSVE--AERLVAGKLKKDTYIENE-KMHSNKRQDLVTKID 535
Query: 479 DLMDLI 484
+++D +
Sbjct: 536 NILDAL 541
>UniRef100_UPI00003AA8A7 UPI00003AA8A7 UniRef100 entry
Length = 574
Score = 319 bits (817), Expect = 1e-85
Identities = 181/486 (37%), Positives = 283/486 (57%), Gaps = 19/486 (3%)
Query: 6 VFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTKL 65
VFTH+LQP+P ITQ++ Q ++F+ + + SPY K Q+ VKL +ESYTKL N
Sbjct: 101 VFTHVLQPYPTHITQSEKQFVVFEGNHYFYSPYFTKTQTTRVKLASRNVESYTKLGNPSR 160
Query: 66 QGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVHG 125
++YGP++++PP+S + +H+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 161 TEDMIEYGPFKDIPPYSQDTLKVHYENNSPFLTITSMTRVIEVSHWGNIAVEETVDLKHT 220
Query: 126 GAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKKT 185
GA KG FSR DYQ +P G S+ + LP A VYYRDEIGNISTS L
Sbjct: 221 GAVLKGPFSRYDYQRQP-DSGISSVKSFKTILPAAAQDVYYRDEIGNISTSHLLVLDDSV 279
Query: 186 ELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVKV 244
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + F + +E V D+L VK+
Sbjct: 280 EMEIRPRFPLFGGWKTHYIIGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVTDSLTVKI 339
Query: 245 VLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFNR 303
VLPEG+K+I P+ + + E +++LD GRPV+V KNN V +H + V+Y FN+
Sbjct: 340 VLPEGAKNIHVDSPYEINRASDELHYTYLDTFGRPVIVAHKNNLVEQHIQDIVVHYTFNK 399
Query: 304 LSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSR 363
+ ML+EPL+++ F+ LF I+Y+ D SI+K A+ +V QV ++V++
Sbjct: 400 ILMLQEPLLVVGAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVNK 455
Query: 364 CLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILP 423
L + + ++ ++ DI + +KS+++ K LT E+ + + L++ + + +
Sbjct: 456 RLGLYRHFDEAVNKYKQSRDISTLNSGKKSLETEHKALTNEIAALQSKLKT--EGSDLCD 513
Query: 424 KVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGR-----EIENRIASHQQKITALKQEID 478
KV E+ ++ D Q K + S+V E+ + G+ IEN H K L +ID
Sbjct: 514 KVSEI--QKLDGQVKELVLKSSVE--AERLVAGKLKKDTYIENE-KMHSNKRQDLVTKID 568
Query: 479 DLMDLI 484
+++D +
Sbjct: 569 NILDAL 574
>UniRef100_UPI000036B734 UPI000036B734 UniRef100 entry
Length = 607
Score = 318 bits (816), Expect = 2e-85
Identities = 177/488 (36%), Positives = 285/488 (58%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+LQP+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 133 TVYTHVLQPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 192
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 193 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 252
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 253 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 311
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 312 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 371
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 372 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 431
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 432 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 487
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +KS+++ K LT E ++ LQS + +
Sbjct: 488 KRIGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKALTSE----ISLLQSRLKTEGSD 543
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 544 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 599
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 600 IDHILDAL 607
>UniRef100_Q5RFB6 Hypothetical protein DKFZp469O2032 [Pongo pygmaeus]
Length = 607
Score = 318 bits (815), Expect = 2e-85
Identities = 177/488 (36%), Positives = 284/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+LQP+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 133 TVYTHVLQPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 192
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 193 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 252
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 253 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 311
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 312 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 371
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 372 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 431
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 432 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 487
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +KS+++ K LT E + LQS + +
Sbjct: 488 KRIGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKALTSE----IALLQSRLKTEGSD 543
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 544 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 599
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 600 IDHILDAL 607
>UniRef100_Q96HX3 Similar to ribophorin I [Homo sapiens]
Length = 568
Score = 316 bits (810), Expect = 8e-85
Identities = 176/488 (36%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 94 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 153
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 154 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 213
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 214 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 272
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 273 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 332
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 333 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 392
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 393 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 448
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +KS+++ K LT E + LQS + +
Sbjct: 449 KRIGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKALTSE----IALLQSRLKTEGSD 504
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 505 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 560
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 561 IDHILDAL 568
>UniRef100_P04843 Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 67
kDa subunit precursor [Homo sapiens]
Length = 607
Score = 316 bits (810), Expect = 8e-85
Identities = 176/488 (36%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 133 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 192
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 193 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 252
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 253 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 311
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 312 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 371
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 372 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 431
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 432 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 487
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +KS+++ K LT E + LQS + +
Sbjct: 488 KRIGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKALTSE----IALLQSRLKTEGSD 543
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 544 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 599
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 600 IDHILDAL 607
>UniRef100_Q6IBR0 RPN1 protein [Homo sapiens]
Length = 607
Score = 315 bits (807), Expect = 2e-84
Identities = 175/488 (35%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 133 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 192
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 193 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 252
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 253 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 311
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 312 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 371
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 372 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 431
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 432 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 487
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +K++++ K LT E + LQS + +
Sbjct: 488 KRIGLYRHFDETVNRYKQSRDISTLNSGKKNLETEHKALTSE----IALLQSRLKTEGSD 543
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 544 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 599
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 600 IDHILDAL 607
>UniRef100_Q91YQ5 Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 67
kDa subunit precursor [Mus musculus]
Length = 608
Score = 315 bits (807), Expect = 2e-84
Identities = 175/488 (35%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 134 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPS 193
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 194 RSEDVLDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 253
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 254 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 312
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 313 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 372
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 373 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 432
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 433 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 488
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 489 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 544
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 545 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYLENEKLSSGKRQELVTK 600
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 601 IDHILDAL 608
>UniRef100_Q7ZX69 MGC52894 protein [Xenopus laevis]
Length = 595
Score = 313 bits (803), Expect = 5e-84
Identities = 171/479 (35%), Positives = 275/479 (56%), Gaps = 9/479 (1%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TVFTH+LQP+P +ITQA+ Q + F+ + + SPYP K Q+ VKL +E+YTKL N
Sbjct: 121 TVFTHVLQPYPTQITQAEKQFVTFEGNHYFFSPYPTKSQTTRVKLASKNVETYTKLGNPT 180
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
++YGP++++ P+S P+ +H+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 181 RSEELIEYGPFKDIAPWSQDPLKVHYENNSPFLTITSMTRLIEVSHWGNIAVEETVDLKH 240
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ + LP A VYYRDEIGNISTS L
Sbjct: 241 TGAYLKGPFSRYDYQRQP-DSGISSVKSFKTILPAAAQDVYYRDEIGNISTSHLLILDDS 299
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + + ++ V+D+L VK
Sbjct: 300 VEMEIRPRFPLFGGWKTHYLIGYNLPSYEYLYNLGDQYALKMRLIDHVFDDQVVDSLTVK 359
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GR V+V KNN V +H + V+Y FN
Sbjct: 360 IILPEGAKNIHLDNPYDINRAPDELHYTYLDTFGRAVIVAHKNNLVEQHIQDIVVHYSFN 419
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
+ ML+EPL++++ F+ LF I+Y+ D SI+K A+ AK+ + Q+ ++V+
Sbjct: 420 NILMLQEPLLVVAAFYILFFTVILYVRLDFSITKDPAAE-AKM---KAACITEQILTLVN 475
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQIL 422
+ L + + ++ + DI + +K++++ K LT E+ S + L++ + + +
Sbjct: 476 KRLGLYRHFDEAVNKYKLSRDISSLNSGKKTLETEHKNLTNEIASLQSKLKA--EGSDLC 533
Query: 423 PKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLM 481
KV E+ + ++E ++ S KL H + L +ID+++
Sbjct: 534 DKVAEIQKLDTQVKEIVLKCSSEAERLVSGKLKKESYIENEKQHSNRRQELVNKIDNIL 592
>UniRef100_Q8AYB9 Ribophorin I [Brachydanio rerio]
Length = 598
Score = 313 bits (801), Expect = 9e-84
Identities = 174/486 (35%), Positives = 283/486 (57%), Gaps = 17/486 (3%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TVF+H+L+PFP ITQA+ QL++FQ + SPYP + Q+ V+L +ESYTKL N
Sbjct: 124 TVFSHVLKPFPTHITQAERQLVVFQGNHYLYSPYPTRSQTTRVRLASKTVESYTKLGNPT 183
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
++YGP++++PPFS + IH+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 184 KSDETIEYGPFKDIPPFSQDVMKIHYENNSPFLTISSITRTIEVSHWGNIAVEETIDLRH 243
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ R G S+ + LP A VYYRDEIGNISTS L
Sbjct: 244 TGAHLKGPFSRYDYQ-RQSDSGISSVKSFKTILPASAQDVYYRDEIGNISTSHLQVLDDS 302
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + + ++ VI L V+
Sbjct: 303 VEVEIRPRFPLFGGWKTHYIIGYNLPSYEYLYNLGDQYALKMRLVDHVYDDQVIXQLTVR 362
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+++I P+P+ + + E +++LD GRPV+V KNN V +H + V+Y FN
Sbjct: 363 LILPEGARNIHMDTPYPISRSQDELHYTYLDTFGRPVLVATKNNLVEQHIQDVVVHYTFN 422
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL+++ F+ LF I+Y+ D SI+K A+ + +V + QV ++V+
Sbjct: 423 KILMLQEPLLVVGAFYILFFTVIIYVRLDFSITKDPAAEVRM----KVASITEQVLTLVN 478
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQIL 422
+ L + ++ + ++ D + RK++++ + LT ++ + T L++ + + +
Sbjct: 479 KRLGLYRHMDEVVNRYKQSRDTGALNSGRKTLEAEHRTLTNDITALQTRLKA--EGSDLA 536
Query: 423 PKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE----ID 478
KV E+ + L++ L+ + S E+ + G+ ++ + +T + E ID
Sbjct: 537 EKVGEIQKLDGQLKD-LVCRSSLDA---ERLVAGKVKKDAYIDSDKNLTGKRHELVNRID 592
Query: 479 DLMDLI 484
L+D +
Sbjct: 593 SLLDAL 598
>UniRef100_Q7ZXV2 MGC52808 protein [Xenopus laevis]
Length = 595
Score = 312 bits (799), Expect = 2e-83
Identities = 170/479 (35%), Positives = 274/479 (56%), Gaps = 9/479 (1%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TVFTH+LQP+P +ITQA+ Q + F+ S + SPYP K Q+ VKL +E+YTKL N
Sbjct: 121 TVFTHVLQPYPTQITQAEKQFVTFEGSHYFYSPYPTKSQTTRVKLASRNVETYTKLGNPT 180
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
++YGP++++ +S P+ +H+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 181 RSEDLIEYGPFKDIAAWSQDPLKVHYENNSPFLTITSMTRLIEVSHWGNIAVEETVDLKH 240
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ R G S+ + LP A VYYRDEIGNISTS L
Sbjct: 241 TGAYLKGPFSRYDYQ-RQTDSGISSVKSFKTILPASAQDVYYRDEIGNISTSHLLILDDS 299
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + F + ++ V+D+L VK
Sbjct: 300 VEMEIRPRFPLFGGWKTHYMIGYNLPSYEYLYNLGDQYALKMRFIDHVFDDQVVDSLTVK 359
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K++ P+ + + E +++LD GR V++ KNN V +H + V+Y FN
Sbjct: 360 IILPEGAKNLQVDNPYEINRAPDELHYTYLDTFGRTVIIAHKNNLVEQHIQDIVVHYSFN 419
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL+++ F+ LF I+Y+ D SI+K A+ AK+ + Q+ ++V+
Sbjct: 420 KILMLQEPLLVVGAFYILFFTVILYVRLDFSITKDPAAE-AKM---KAACITEQILTLVN 475
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQIL 422
+ L + + ++ + DI + +K++++ K LT E+ S + L++ + + +
Sbjct: 476 KRLGLYRHFDEAVNKYKLSRDISTLNSGKKTLETEHKNLTNEIASLQSKLKA--EGSDLC 533
Query: 423 PKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLM 481
KV E+ + ++E ++ S KL H + L +ID+++
Sbjct: 534 DKVAEIQKLDAQVKEIVLKSSSDAERLVSGKLKKESYIENEKQHSSRRQELVNKIDNIL 592
>UniRef100_Q6P1S9 Hypothetical protein MGC76266 [Xenopus tropicalis]
Length = 595
Score = 311 bits (798), Expect = 2e-83
Identities = 169/479 (35%), Positives = 272/479 (56%), Gaps = 9/479 (1%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TVFTH+LQP+P +ITQA+ Q + F+ + + SPYP K Q+ VKL +E+YTKL N
Sbjct: 121 TVFTHVLQPYPTQITQAEKQFVTFEGNHYFFSPYPTKSQTTRVKLASRNVETYTKLGNPT 180
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
++YGP++++ P+S + +H+ENN PF + R IE+SHWGN+ + E +L H
Sbjct: 181 RSEDLIEYGPFKDIAPWSQDLLKVHYENNSPFLTITSMTRLIEVSHWGNIAVEETVDLKH 240
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ + LP A VYYRDEIGNISTS L
Sbjct: 241 TGAYLKGPFSRYDYQRQP-DSGISSVKSFKTILPAAAQDVYYRDEIGNISTSHLLILDDS 299
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + IGY LP ++L+ + + L + F + ++ V+D+L VK
Sbjct: 300 VEMEIRPRFPLFGGWKTHYIIGYNLPSYEYLYNLGDQYALKMRFIDHVFDDQVVDSLTVK 359
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
+VLPEG+K++ P+ + + E +++LD GR V+V KNN V +H + V+Y FN
Sbjct: 360 IVLPEGAKNVHVDNPYDINRAPDELHYTYLDTFGRTVIVAHKNNLVEQHIQDIVVHYSFN 419
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL+++ F+ LF I+Y+ D SI+K A+ + Q+ ++V+
Sbjct: 420 KILMLQEPLLVVGAFYILFFTVILYVRLDFSITKDPAAEARM----KAACITEQILTLVN 475
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQIL 422
+ L + + ++ + DI + +K++++ K LT E+ S + L++ + + +
Sbjct: 476 KRLGLYRHFDEAVNKYKLSRDIATLNSGKKTLETEHKNLTNEIASLQSKLKA--EGSDLC 533
Query: 423 PKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLM 481
KV E+ + ++E ++ S KL H + L +ID+++
Sbjct: 534 DKVAEIQKLDSQVKEIVLKSSSEAERLVSGKLKKETYIENEKQHSSRRQELVNKIDNIL 592
>UniRef100_Q8BMR3 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4932403A10 product:DOLICHYL-
DIPHOSPHOOLIGOSACCHARIDE-- PROTEIN GLYCOSYLTRANSFERASE
67 kDa SUBUNIT (EC 2.4.1.119) (RIBOPHORIN I) (RPN-I)
homolog [Mus musculus]
Length = 608
Score = 311 bits (798), Expect = 2e-83
Identities = 174/488 (35%), Positives = 282/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 134 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPS 193
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWG + + E+ +L H
Sbjct: 194 RSEDVLDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGIIAVEENVDLKH 253
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 254 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 312
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 313 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 372
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 373 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 432
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 433 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 488
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 489 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 544
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 545 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYLENEKLSSGKRQELVTK 600
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 601 IDHILDAL 608
>UniRef100_Q6P7A7 Ribophorin I [Rattus norvegicus]
Length = 606
Score = 311 bits (798), Expect = 2e-83
Identities = 173/488 (35%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ V+L +ES+TKL N
Sbjct: 132 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVRLASRNVESHTKLGNPS 191
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 192 RSEDILDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 251
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 252 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 310
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 311 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 370
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 371 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 430
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 431 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 486
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 487 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 542
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 543 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLSSGKRQELVTK 598
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 599 IDHILDAL 606
>UniRef100_P07153 Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 67
kDa subunit precursor [Rattus norvegicus]
Length = 605
Score = 308 bits (789), Expect = 2e-82
Identities = 172/488 (35%), Positives = 282/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ V+L +ES+TKL N
Sbjct: 131 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVRLASRNVESHTKLGNPS 190
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 191 RSEDILDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 250
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 251 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 309
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+ LFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 310 VEMEIRPRFGLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 369
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 370 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 429
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 430 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 485
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 486 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 541
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 542 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLSSGKRQELVTK 597
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 598 IDHILDAL 605
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 794,120,024
Number of Sequences: 2790947
Number of extensions: 33746830
Number of successful extensions: 95629
Number of sequences better than 10.0: 176
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 113
Number of HSP's that attempted gapping in prelim test: 95302
Number of HSP's gapped (non-prelim): 256
length of query: 487
length of database: 848,049,833
effective HSP length: 131
effective length of query: 356
effective length of database: 482,435,776
effective search space: 171747136256
effective search space used: 171747136256
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC126786.5


