

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.6 - phase: 0
(135 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
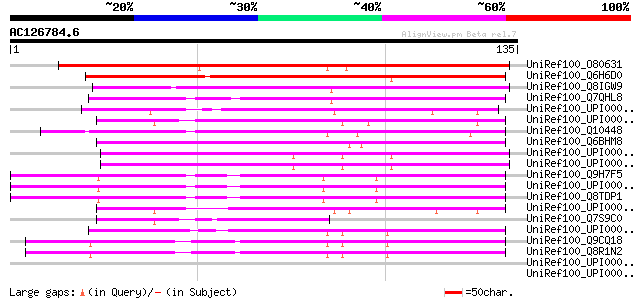
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O80631 Hypothetical protein At2g39440 [Arabidopsis tha... 133 1e-30
UniRef100_Q6H6D0 Hypothetical protein P0669G09.16 [Oryza sativa] 108 3e-23
UniRef100_Q8IGW9 RE11282p [Drosophila melanogaster] 64 7e-10
UniRef100_Q7QHL8 ENSANGP00000015645 [Anopheles gambiae str. PEST] 63 1e-09
UniRef100_UPI000021BE1A UPI000021BE1A UniRef100 entry 58 5e-08
UniRef100_UPI0000235380 UPI0000235380 UniRef100 entry 57 1e-07
UniRef100_Q10448 Hypothetical protein C12B10.15c in chromosome I... 56 2e-07
UniRef100_Q6BHM8 Similar to CA1323|IPF6675 Candida albicans IPF6... 55 4e-07
UniRef100_UPI000042E59D UPI000042E59D UniRef100 entry 54 6e-07
UniRef100_UPI000042E4BB UPI000042E4BB UniRef100 entry 54 6e-07
UniRef100_Q9H7F5 Hypothetical protein FLJ20974 [Homo sapiens] 51 6e-06
UniRef100_UPI000036EEAC UPI000036EEAC UniRef100 entry 50 8e-06
UniRef100_Q8TDP1 RNase H1 small subunit [Homo sapiens] 50 8e-06
UniRef100_UPI000023E713 UPI000023E713 UniRef100 entry 49 2e-05
UniRef100_Q7S9C0 Predicted protein [Neurospora crassa] 47 7e-05
UniRef100_UPI00001C757F UPI00001C757F UniRef100 entry 47 9e-05
UniRef100_Q9CQ18 Mus musculus adult male cerebellum cDNA, RIKEN ... 47 9e-05
UniRef100_Q8R1N2 AYP1 [Mus musculus] 47 9e-05
UniRef100_UPI000042C016 UPI000042C016 UniRef100 entry 39 0.019
UniRef100_UPI0000452A64 UPI0000452A64 UniRef100 entry 36 0.16
>UniRef100_O80631 Hypothetical protein At2g39440 [Arabidopsis thaliana]
Length = 773
Score = 133 bits (334), Expect = 1e-30
Identities = 67/142 (47%), Positives = 87/142 (61%), Gaps = 22/142 (15%)
Query: 14 DGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGV----------VGEDGLPLQQSHFR 63
+ I DLS +VH LPCCI+ DGP EVS YFKPK + V DG+ +++HFR
Sbjct: 83 NSIVDLSSQVHQLPCCIRFDGPVEVSHYFKPKSSVRLMWKAVEFVDVEIDGVKTEEAHFR 142
Query: 64 GRLLEGTTLPLPHNYSGFVL---------GKKNS---VESSNSWETSVTFNDITYWNHDS 111
GR L+G T+ LP YSGFVL GK+ + E + WE FN++TYWNHDS
Sbjct: 143 GRKLQGATISLPSGYSGFVLRQASNLNANGKRKASMPTEDNQCWEVKAKFNNLTYWNHDS 202
Query: 112 VPSNNDDFSRAFHWIPVAEAVS 133
+PS +D F R+FHW +AEAV+
Sbjct: 203 LPSKDDTFFRSFHWFSIAEAVT 224
>UniRef100_Q6H6D0 Hypothetical protein P0669G09.16 [Oryza sativa]
Length = 161
Score = 108 bits (270), Expect = 3e-23
Identities = 56/119 (47%), Positives = 73/119 (61%), Gaps = 8/119 (6%)
Query: 21 GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSG 80
GRVHLLPC IK +G VS YFKPK TGV E G+ ++++ FRGR L+G T+ LP Y G
Sbjct: 24 GRVHLLPCGIKQNGAAAVSDYFKPKDTGVEVE-GIRVEEAFFRGRKLQGATISLPDGYRG 82
Query: 81 FVLGKKNSVESSNSWETSVT-------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+VL K++ + E V+ F +ITYWNHD+ PS D R FH + VA A+
Sbjct: 83 YVLEKRSGGKDMKKLEGEVSNFKSRAEFQNITYWNHDTTPSAEDPLPRCFHLLTVANAM 141
>UniRef100_Q8IGW9 RE11282p [Drosophila melanogaster]
Length = 165
Score = 63.9 bits (154), Expect = 7e-10
Identities = 43/114 (37%), Positives = 52/114 (44%), Gaps = 4/114 (3%)
Query: 23 VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
VH LP I DG V YF T E G + + RG L G L +P Y G V
Sbjct: 30 VHYLPAKIDGDGEANVKNYFN-NYTREATEFGTGILTNALRGFPLMGEKLKVPEGYCGLV 88
Query: 83 LG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
L K S S + F D TYWN+D VPSN D + +A VA+A+S
Sbjct: 89 LQETEKPISDTSDRKLRLTGVFQDFTYWNYDKVPSNGDPYRQALPLPDVAQALS 142
>UniRef100_Q7QHL8 ENSANGP00000015645 [Anopheles gambiae str. PEST]
Length = 129
Score = 63.2 bits (152), Expect = 1e-09
Identities = 38/114 (33%), Positives = 53/114 (46%), Gaps = 7/114 (6%)
Query: 22 RVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGF 81
++ +P IK DGP + Q+F P DG RG L G LP Y+G
Sbjct: 3 KLQYIPATIKGDGPANLEQFFTPYTE--TQRDGTLCNA--LRGYPLRGKATSLPAGYTGV 58
Query: 82 VLG---KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+L K S E + + F D TYWN+D PS ND ++A W+ +AE +
Sbjct: 59 MLQETKKPLSDEDDRTLTFAGAFRDFTYWNYDRQPSRNDPMAKALGWLQLAEVL 112
>UniRef100_UPI000021BE1A UPI000021BE1A UniRef100 entry
Length = 331
Score = 57.8 bits (138), Expect = 5e-08
Identities = 39/134 (29%), Positives = 61/134 (45%), Gaps = 29/134 (21%)
Query: 20 SGRVHLLPCCIKHDGPT-EVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNY 78
S VHLLPC I HDGP +V Y+KP + +DG + + FRGR L G + +P Y
Sbjct: 17 SAAVHLLPCTIHHDGPVDQVQSYWKPTES----QDG--KKTAFFRGRKLHGKAIRVPEGY 70
Query: 79 SGFVLGK--------------------KNSVESSNSWETSVTFNDITYWNHDS-VPSNND 117
G V+ K ++ + +T F ++ W H+S +++D
Sbjct: 71 KGIVVEKGPEDGPAETRADEPVTVDVDAEEKVATTALDTKAEFEEMVIWGHESTADASSD 130
Query: 118 DFSRAF-HWIPVAE 130
+ R W+ +AE
Sbjct: 131 PYVRGVEEWVSLAE 144
>UniRef100_UPI0000235380 UPI0000235380 UniRef100 entry
Length = 159
Score = 56.6 bits (135), Expect = 1e-07
Identities = 38/128 (29%), Positives = 57/128 (43%), Gaps = 23/128 (17%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
++LPC I +DG E +++Y+ P E LQ ++FRGR L G + LP Y G V
Sbjct: 25 NILPCRIHYDGSVESLNRYWTPS----TDEKDKDLQTAYFRGRRLRGRRVQLPEGYEGVV 80
Query: 83 LGKKN------------SVESSNS-----WETSVTFNDITYWNHDSVPSNNDDFSRAF-H 124
+ S E N+ E TF D W H+ +P+ +D F +
Sbjct: 81 VTHTEREIPNATDNVTVSKEGENNKAVKLLEKQATFGDYVVWGHEVIPAADDSFVKGVEE 140
Query: 125 WIPVAEAV 132
W+ AE +
Sbjct: 141 WVKFAEVM 148
>UniRef100_Q10448 Hypothetical protein C12B10.15c in chromosome I
[Schizosaccharomyces pombe]
Length = 147
Score = 55.8 bits (133), Expect = 2e-07
Identities = 42/135 (31%), Positives = 62/135 (45%), Gaps = 14/135 (10%)
Query: 9 IDLKRDGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLE 68
+ L++ G +L + LLPC I +DGP V +YF K + + RGR L
Sbjct: 8 VKLEQAGSSNLE-KASLLPCHISYDGPAPVFEYFHDKIQ--ISNNTHSKTTVCLRGRELN 64
Query: 69 GTTLPLPHNYSGFVL----GKKNSVES-----SNSWETSVTFNDITYWNHDSVPSNNDDF 119
G L LP NY+G V+ G +N S ++W + TF + WN D++ S+ D
Sbjct: 65 GEELDLPENYTGQVVLCDDGLENDETSEKEIPESTWCINSTFEKVMLWNRDNMRSSEKDQ 124
Query: 120 SR--AFHWIPVAEAV 132
+ WI A V
Sbjct: 125 WKRGVLEWIQFASKV 139
>UniRef100_Q6BHM8 Similar to CA1323|IPF6675 Candida albicans IPF6675 [Debaryomyces
hansenii]
Length = 138
Score = 54.7 bits (130), Expect = 4e-07
Identities = 29/120 (24%), Positives = 58/120 (48%), Gaps = 11/120 (9%)
Query: 24 HLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFVL 83
+++PC I+++G YF P G ++ +++RG L G + + ++YSG+++
Sbjct: 17 NIVPCHIQYNGAANTKDYFTPSKKQDTLPTGEQVEVAYYRGCKLVGKNMNISNDYSGYLI 76
Query: 84 GKKNSV-----ESS------NSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
K S+ E+S N++ F +IT + HD+VP + W ++E +
Sbjct: 77 NKSESLARVEDETSEDYLTVNTYTPVAKFEEITVYGHDTVPELTSQWGLIQEWKDISEVI 136
>UniRef100_UPI000042E59D UPI000042E59D UniRef100 entry
Length = 137
Score = 54.3 bits (129), Expect = 6e-07
Identities = 34/119 (28%), Positives = 59/119 (49%), Gaps = 10/119 (8%)
Query: 25 LLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPL-PHNYSGFVL 83
++P I++ GP YF P T DG L ++FRG L G TL L ++ G+V+
Sbjct: 18 VVPMNIQYSGPANTEDYFAPSKTQETQPDGTLLDVAYFRGCKLVGKTLELDKYDVKGYVI 77
Query: 84 GKKN---SVESSNSWETSVT------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
K S + + +T+VT F +T + HD++PS + +S ++ ++ V+
Sbjct: 78 NKSEHLVSDKETGEVKTAVTYIPVGNFQKLTVYGHDTLPSRTNQWSLIPEYLSISNIVN 136
>UniRef100_UPI000042E4BB UPI000042E4BB UniRef100 entry
Length = 137
Score = 54.3 bits (129), Expect = 6e-07
Identities = 34/119 (28%), Positives = 59/119 (49%), Gaps = 10/119 (8%)
Query: 25 LLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPL-PHNYSGFVL 83
++P I++ GP YF P T DG L ++FRG L G TL L ++ G+V+
Sbjct: 18 VVPMNIQYSGPANTEDYFAPSKTQETQPDGTLLDVAYFRGCKLVGKTLELDKYDVKGYVI 77
Query: 84 GKKN---SVESSNSWETSVT------FNDITYWNHDSVPSNNDDFSRAFHWIPVAEAVS 133
K S + + +T+VT F +T + HD++PS + +S ++ ++ V+
Sbjct: 78 NKSEHLVSDKETGEVKTAVTYIPVGNFQKLTVYGHDTLPSRTNQWSLIPEYLSISNIVN 136
>UniRef100_Q9H7F5 Hypothetical protein FLJ20974 [Homo sapiens]
Length = 163
Score = 50.8 bits (120), Expect = 6e-06
Identities = 42/161 (26%), Positives = 67/161 (41%), Gaps = 34/161 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 1 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV--------LGKKNSVESSNSWE---------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 59 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKVSMGKPDPLRDSGTDDQEEEPLERDF 115
Query: 97 -----TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+ F+ T W +++P + A W +A A+
Sbjct: 116 DRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAAI 156
>UniRef100_UPI000036EEAC UPI000036EEAC UniRef100 entry
Length = 164
Score = 50.4 bits (119), Expect = 8e-06
Identities = 42/162 (25%), Positives = 67/162 (40%), Gaps = 35/162 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 1 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV---------LGKKNSVESSNSWE--------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 59 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKEVSMGKPDPLRDSGTDDQEEEPLERD 115
Query: 97 ------TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+ F+ T W +++P + A W +A A+
Sbjct: 116 FDRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAAI 157
>UniRef100_Q8TDP1 RNase H1 small subunit [Homo sapiens]
Length = 164
Score = 50.4 bits (119), Expect = 8e-06
Identities = 42/162 (25%), Positives = 67/162 (40%), Gaps = 35/162 (21%)
Query: 1 MEKGGEAKIDLKRDGIEDLSGR------VHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDG 54
ME G EA I+ R + + R +HLLPC + DGP V ++F P G +G
Sbjct: 1 MESGDEAAIERHRVHLRSATLRDAVPATLHLLPCEVAVDGPAPVGRFFTPAIR--QGPEG 58
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFV---------LGKKNSVESSNSWE--------- 96
L + FRGR L G + +P G+V +GK + + S + +
Sbjct: 59 LEVS---FRGRCLRGEEVAVPPGLVGYVMVTEEKKVSMGKPDPLRDSGTDDQEEEPLERD 115
Query: 97 ------TSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+ F+ T W +++P + A W +A A+
Sbjct: 116 FDRFIGATANFSRFTLWGLETIPGPDAKVRGALTWPSLAAAI 157
>UniRef100_UPI000023E713 UPI000023E713 UniRef100 entry
Length = 149
Score = 48.9 bits (115), Expect = 2e-05
Identities = 35/129 (27%), Positives = 61/129 (47%), Gaps = 31/129 (24%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
++LPC I HDGP E S Y+ P T ++FRGR L+G + LP + G V
Sbjct: 20 NILPCRIHHDGPIEPTSTYWTPAAT-----------DAYFRGRKLQGKIVKLPKDCRGIV 68
Query: 83 LGK------KNSV-----------ESSNSWETSVTFNDITYWNHDSV-PSNNDDFSRAF- 123
+ + K +V E + + + F+++ W H++V ++ D + R+
Sbjct: 69 VERVSQQDPKTAVEEIVEDEDAEQEQTGRMQVTAEFDEMVVWGHEAVADASADPYVRSME 128
Query: 124 HWIPVAEAV 132
W+ VA+ +
Sbjct: 129 EWLQVADKI 137
>UniRef100_Q7S9C0 Predicted protein [Neurospora crassa]
Length = 169
Score = 47.4 bits (111), Expect = 7e-05
Identities = 28/63 (44%), Positives = 35/63 (55%), Gaps = 6/63 (9%)
Query: 24 HLLPCCIKHDGPTE-VSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGFV 82
+LLPC I H G E ++ P+ +DG L+ S+FRGR L G TLPLP Y G V
Sbjct: 26 NLLPCRIHHSGSVEPTDSFWNPR----CEQDG-NLKVSYFRGRKLHGKTLPLPEGYKGVV 80
Query: 83 LGK 85
K
Sbjct: 81 ASK 83
>UniRef100_UPI00001C757F UPI00001C757F UniRef100 entry
Length = 166
Score = 47.0 bits (110), Expect = 9e-05
Identities = 39/137 (28%), Positives = 58/137 (41%), Gaps = 31/137 (22%)
Query: 22 RVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPHNYSGF 81
++HLLPC + P V ++F P G DGL Q FRGR L G + +P ++GF
Sbjct: 28 KLHLLPCDVLVSRPAPVERFFTPAVRH--GADGL---QVSFRGRGLRGEDVAVPPGFAGF 82
Query: 82 VL----------GKKN--------SVESSNSWETSV--------TFNDITYWNHDSVPSN 115
V+ GK N + E+ E +F+ T W ++VP
Sbjct: 83 VMVTEEMGEGLIGKLNFSGDAEDKADEAQEPLERDFDRFIGATGSFSHFTLWGLETVPGP 142
Query: 116 NDDFSRAFHWIPVAEAV 132
+ RA W +A A+
Sbjct: 143 DAKVHRALGWPSLAAAI 159
>UniRef100_Q9CQ18 Mus musculus adult male cerebellum cDNA, RIKEN full-length enriched
library, clone:1500026D16 product:hypothetical Copper
amine oxidase, domains 1 and 2 structure containing
protein, full insert sequence [Mus musculus]
Length = 166
Score = 47.0 bits (110), Expect = 9e-05
Identities = 42/155 (27%), Positives = 66/155 (42%), Gaps = 32/155 (20%)
Query: 5 GEAKIDLKRDGIEDLS-GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFR 63
G+ +I L+ + + ++HLLPC + P V ++F P V D LQ S FR
Sbjct: 10 GKQRIHLRPGSLRGAAPAKLHLLPCDVLVSRPAPVDRFFTP----AVRHDADGLQAS-FR 64
Query: 64 GRLLEGTTLPLPHNYSGFVL----------GKKN--------SVESSNSWETSV------ 99
GR L G + +P ++GFV+ GK N + E+ E
Sbjct: 65 GRGLRGEEVAVPPGFAGFVMVTEEKGEGLIGKLNFSGDAEDKADEAQEPLERDFDRLIGA 124
Query: 100 --TFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+F+ T W ++VP + RA W +A A+
Sbjct: 125 TGSFSHFTLWGLETVPGPDAKVHRALGWPSLAAAI 159
>UniRef100_Q8R1N2 AYP1 [Mus musculus]
Length = 166
Score = 47.0 bits (110), Expect = 9e-05
Identities = 42/155 (27%), Positives = 66/155 (42%), Gaps = 32/155 (20%)
Query: 5 GEAKIDLKRDGIEDLS-GRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFR 63
G+ +I L+ + + ++HLLPC + P V ++F P V D LQ S FR
Sbjct: 10 GKQRIHLRPGSLRGAAPAKLHLLPCDVLVSRPAPVDRFFTP----AVRHDADGLQAS-FR 64
Query: 64 GRLLEGTTLPLPHNYSGFVL----------GKKN--------SVESSNSWETSV------ 99
GR L G + +P ++GFV+ GK N + E+ E
Sbjct: 65 GRGLRGEEVAVPPGFAGFVMVTEEKGEGLIGKLNFSGDAEDKADEAQEPLERDFDRLIGA 124
Query: 100 --TFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
+F+ T W ++VP + RA W +A A+
Sbjct: 125 TGSFSHFTLWGLETVPGPDAKVHRALGWPSLAAAI 159
>UniRef100_UPI000042C016 UPI000042C016 UniRef100 entry
Length = 122
Score = 39.3 bits (90), Expect = 0.019
Identities = 18/57 (31%), Positives = 28/57 (48%)
Query: 62 FRGRLLEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVPSNNDD 118
FRGR L G + L + ++ KN+ + +FN +TYWN+D +DD
Sbjct: 48 FRGRKLNGRIIHLSKSSHSILILSKNNSQKKMDLIIDKSFNQVTYWNYDDEIKQSDD 104
>UniRef100_UPI0000452A64 UPI0000452A64 UniRef100 entry
Length = 122
Score = 36.2 bits (82), Expect = 0.16
Identities = 17/57 (29%), Positives = 26/57 (44%)
Query: 62 FRGRLLEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVPSNNDD 118
FRGR L G + L + ++ KN+ + N +TYWN+D +DD
Sbjct: 48 FRGRKLNGRIIHLSKSSHSILILSKNNSRKKMDLIIDKSLNQVTYWNYDDEIKQSDD 104
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 264,027,598
Number of Sequences: 2790947
Number of extensions: 11269585
Number of successful extensions: 21547
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 88
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 21421
Number of HSP's gapped (non-prelim): 143
length of query: 135
length of database: 848,049,833
effective HSP length: 111
effective length of query: 24
effective length of database: 538,254,716
effective search space: 12918113184
effective search space used: 12918113184
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 67 (30.4 bits)
Medicago: description of AC126784.6


