

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126780.2 - phase: 0 /pseudo
(297 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
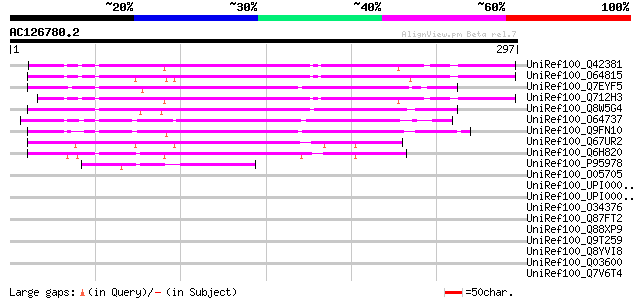
UniRef100_Q42381 Putative N-acetyltransferase hookless1 [Arabido... 167 2e-40
UniRef100_O64815 Similar to hookless1 [Arabidopsis thaliana] 162 1e-38
UniRef100_Q7EYF5 Acetyltransferase-like protein [Oryza sativa] 157 2e-37
UniRef100_Q712H3 Putative N-acetyltransferase hookless1 [Brassic... 157 2e-37
UniRef100_Q8W5G4 Putative acetyl transferase [Oryza sativa] 155 1e-36
UniRef100_O64737 Hookless1-like protein [Arabidopsis thaliana] 148 1e-34
UniRef100_Q9FN10 N-acetyltransferase hookless1-like protein [Ara... 142 8e-33
UniRef100_Q67UR2 GCN5-related N-acetyltransferase-like [Oryza sa... 123 7e-27
UniRef100_Q6H820 GCN5-related N-acetyltransferase-like [Oryza sa... 111 2e-23
UniRef100_P95978 Orf c04040 protein [Sulfolobus solfataricus] 47 5e-04
UniRef100_O05705 N-terminal acetyl transferase [Sulfolobus shiba... 45 0.003
UniRef100_UPI000033E574 UPI000033E574 UniRef100 entry 38 0.28
UniRef100_UPI00002DAC07 UPI00002DAC07 UniRef100 entry 38 0.28
UniRef100_O34376 Putative acetyl transferase [Bacillus subtilis] 38 0.28
UniRef100_Q87FT2 Putative acetyltransferase [Vibrio parahaemolyt... 38 0.28
UniRef100_Q88XP9 Lipoprotein [Lactobacillus plantarum] 38 0.28
UniRef100_Q9T259 Ribosomal protein L5 [Phytophthora infestans] 38 0.37
UniRef100_Q8YVI8 N-terminal acetyltransferase [Anabaena sp.] 37 0.63
UniRef100_Q03600 Integrin alpha ina-1 precursor [Caenorhabditis ... 37 0.63
UniRef100_Q7V6T4 Putative ribosomal-protein-alanine acetyltransf... 36 1.1
>UniRef100_Q42381 Putative N-acetyltransferase hookless1 [Arabidopsis thaliana]
Length = 403
Score = 167 bits (424), Expect = 2e-40
Identities = 110/306 (35%), Positives = 164/306 (52%), Gaps = 35/306 (11%)
Query: 12 VIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVAE 71
V+RE+D RD+ V +ER C E+ + K S+FT+++ GDP+ RIR P ++MLVAE
Sbjct: 3 VVREYDPTRDLVGVEDVERRC-EVGPSGK--LSLFTDLL--GDPICRIRHSPSYLMLVAE 57
Query: 72 M-VESKELVGVVKGCIKSV--------------QTPSGSLFKMGCILGLRVSPIHRRKGV 116
M E KE+VG+++GCIK+V K+ +LGLRVSP HRR+G+
Sbjct: 58 MGTEKKEIVGMIRGCIKTVTCGQKLDLNHKSQNDVVKPLYTKLAYVLGLRVSPFHRRQGI 117
Query: 117 GLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTN 176
G KLV +EEW NGA+Y+++ATE +N AS NLFT KC Y F + I ++P + N
Sbjct: 118 GFKLVKMMEEWFRQNGAEYSYIATENDNQASVNLFTGKCGYSEFRTPSILVNPVYAHRVN 177
Query: 177 HISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVS------YYKD 230
+S++ V + K+ A + Y T E +P D+D +L KLSLGT+V+ Y
Sbjct: 178 -VSRR-VTVIKLEPVDAETLYRIRFSTTEFFPRDIDSVLNNKLSLGTFVAVPRGSCYGSG 235
Query: 231 EGFKLNIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLNHAKDK 290
G + + SW + S+WN C ++ ++ + LR + A DK
Sbjct: 236 SGSWPGSAKFLEY---PPESWAVLSVWN----CKDSFLLEVRGASRLRRVVAKTTRVVDK 288
Query: 291 ICPLFE 296
P +
Sbjct: 289 TLPFLK 294
>UniRef100_O64815 Similar to hookless1 [Arabidopsis thaliana]
Length = 413
Score = 162 bits (409), Expect = 1e-38
Identities = 108/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 11 VVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVA 70
V +RE+D +D+ V +ER C E+ K S+FT+++ GDP+ R+R P ++MLVA
Sbjct: 5 VEVREYDPSKDLATVEDVERRC-EVGPAGK--LSLFTDLL--GDPICRVRHSPSYLMLVA 59
Query: 71 EM--VESKELVGVVKGCIKSVQ----------TPSGS----------LFKMGCILGLRVS 108
E+ E KELVG+++GCIK+V T + S K+ ILGLRVS
Sbjct: 60 EIGPKEKKELVGMIRGCIKTVTCGITTKRLDLTHNKSQNDVVITKPLYTKLAYILGLRVS 119
Query: 109 PIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLH 168
P HRR+G+G KLV ++E+W NGA+Y++ ATE +N+AS NLFT KC Y F + I ++
Sbjct: 120 PTHRRQGIGFKLVKAMEDWFSQNGAEYSYFATENDNHASVNLFTGKCGYAEFRTPSILVN 179
Query: 169 PPTSFPTNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYY 228
P + N IS++ V + K+ A Y T E +P D+D +L KLSLGT+V+
Sbjct: 180 PVYAHRVN-ISRR-VTVIKLEPSDAELLYRLRFSTTEFFPRDIDSVLNNKLSLGTFVAVP 237
Query: 229 KDEGF---KLNIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLN 285
+ + + SW + S+WN C ++ +++ + LR + +
Sbjct: 238 RGSCYGSGSRSWPGSAKFLEYPPDSWAVLSVWN----CKDSFRLEVRGASRLRRVVSKAT 293
Query: 286 HAKDKICPLFE 296
DK P +
Sbjct: 294 RMVDKTLPFLK 304
>UniRef100_Q7EYF5 Acetyltransferase-like protein [Oryza sativa]
Length = 397
Score = 157 bits (398), Expect = 2e-37
Identities = 95/256 (37%), Positives = 148/256 (57%), Gaps = 14/256 (5%)
Query: 11 VVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVA 70
V++REFD RD V +ER C G + +FT+++ GDPL R+R P ++MLVA
Sbjct: 14 VLVREFDGGRDRPGVELVERACEV--GPSGGKLCLFTDLL--GDPLCRVRHSPAYLMLVA 69
Query: 71 EMVESK---ELVGVVKGCIKSVQTPSGSLF-KMGCILGLRVSPIHRRKGVGLKLVTSIEE 126
E V E+VGVV+GC+K+V LF K+ +LGLRVSP HRR+G+G +LV +EE
Sbjct: 70 EAVGGPLGTEIVGVVRGCVKTVACGRSQLFSKVAYLLGLRVSPRHRRRGIGRRLVERMEE 129
Query: 127 WMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHISKKDVKID 186
W GA+YA++AT+++N S LFT C Y F + + +HP F + + +
Sbjct: 130 WFREMGAEYAYVATDRDNEPSVRLFTGACGYAKFRTPSVLVHP--VFGHDLAPSRRAAVV 187
Query: 187 KISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIEDIITHKST 246
++ +A Y R L + E +P D+D +L LSLGT+++ + ++ +E + ++
Sbjct: 188 RLDAREAELLYRRRLGSVEFFPRDIDAVLSNALSLGTFLAVPRGTRWR-GVEGFL---AS 243
Query: 247 TSSSWIIFSLWNTCEA 262
+SW + SLWN +A
Sbjct: 244 PPASWAVASLWNCKDA 259
>UniRef100_Q712H3 Putative N-acetyltransferase hookless1 [Brassica napus]
Length = 390
Score = 157 bits (398), Expect = 2e-37
Identities = 106/301 (35%), Positives = 156/301 (51%), Gaps = 35/301 (11%)
Query: 17 DEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVAEM-VES 75
D RD+ V +ER C E+ + K S+FT+++ GDP+SRIR P +MLVAE E
Sbjct: 1 DPSRDLAGVEDVERRC-EVGPSGK--LSLFTDLL--GDPISRIRHSPSFLMLVAETGTEE 55
Query: 76 KELVGVVKGCIKSV--------------QTPSGSLFKMGCILGLRVSPIHRRKGVGLKLV 121
KE+VG+++GCIK+V T K+ +LGLRVSP HRR+G+G KLV
Sbjct: 56 KEIVGMIRGCIKTVTCGKKLDLNHKSQNDTVKPLYTKLAYVLGLRVSPSHRRQGIGFKLV 115
Query: 122 TSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHISKK 181
+E W + NGA+Y+++ATE N AS NLFT KC Y F I ++P + N +S++
Sbjct: 116 KMMEAWFMQNGAEYSYIATENENQASVNLFTGKCGYSEFRKPSILVNPVYAHKVN-VSRR 174
Query: 182 DVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVS------YYKDEGFKL 235
V + K+ A S Y T E +P D+D +L LSLGT+V+ Y G
Sbjct: 175 -VTVIKLDPVDAESLYRLRFSTTEFFPRDIDSVLNNNLSLGTFVAVPRGSCYGSGSGSWP 233
Query: 236 NIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLNHAKDKICPLF 295
+ + SW + S+WN C ++ +++ + R + A DK P
Sbjct: 234 GSSKFLEY---PPESWAVLSVWN----CKDSFRLEVRGASLWRRVVAKTTRVVDKTLPFL 286
Query: 296 E 296
+
Sbjct: 287 K 287
>UniRef100_Q8W5G4 Putative acetyl transferase [Oryza sativa]
Length = 399
Score = 155 bits (392), Expect = 1e-36
Identities = 94/266 (35%), Positives = 147/266 (54%), Gaps = 22/266 (8%)
Query: 11 VVIREFDEDRDVKVVGKLERNC-TEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLV 69
V +RE+ EDRD V ++ER C +G + +FT+++ GDPL RIR P ++MLV
Sbjct: 4 VEVREYREDRDRAAVEEVERECEVGSSGGGEAKMCLFTDLL--GDPLCRIRNSPAYLMLV 61
Query: 70 AEMVES------KELVGVVKGCIK------SVQTPSGSLF-KMGCILGLRVSPIHRRKGV 116
AE +E++G+++GC+K SVQ ++ K+ ILGLRVSP +RRKGV
Sbjct: 62 AETANGGGGGNGREIIGLIRGCVKTVVSGGSVQAGKDPIYSKVAYILGLRVSPRYRRKGV 121
Query: 117 GLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTN 176
G KLV +EEW +GA+Y+++ATE++N AS LFT +C Y F + + +HP
Sbjct: 122 GKKLVGRMEEWFRQSGAEYSYMATEQDNEASVRLFTGRCGYSKFRTPSVLVHPVFGHALQ 181
Query: 177 HISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLN 236
++ I K+ +A Y E +P D+D +L ++LSLGT+++ +
Sbjct: 182 --PSRNAAIRKLEPREAELLYRWHFAAVEFFPADIDAVLSKELSLGTFLAVPAGTRW--- 236
Query: 237 IEDIITHKSTTSSSWIIFSLWNTCEA 262
E + +SW + S+WN +A
Sbjct: 237 -ESVEAFMDAPPASWAVMSVWNCMDA 261
>UniRef100_O64737 Hookless1-like protein [Arabidopsis thaliana]
Length = 386
Score = 148 bits (374), Expect = 1e-34
Identities = 93/255 (36%), Positives = 152/255 (59%), Gaps = 26/255 (10%)
Query: 7 IESKVVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHV 66
++ +VVIR +D+ RD +G++E++C EI + +FT+ + GDP+ RIR P +
Sbjct: 9 VDEEVVIRCYDDRRDRIQMGRMEKSC-EIGHDHQT--LLFTDTL--GDPICRIRNSPFFI 63
Query: 67 MLVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIHRRKGVGLKLVTSIEE 126
MLVA + +LVG ++G +K V+ S+ ++G +LGLRV P +RR+G+G LV +EE
Sbjct: 64 MLVAGV--GNKLVGSIQGSVKPVEFHDKSV-RVGYVLGLRVVPSYRRRGIGSILVRKLEE 120
Query: 127 WMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHIS-KKDVKI 185
W ++ ADYA++ATEK+N AS LF + Y F + I ++P P + D+ I
Sbjct: 121 WFESHNADYAYMATEKDNEASHGLFIGRLGYVVFRNPAILVNPVN--PGRGLKLPSDIGI 178
Query: 186 DKISIDQAISFYTR-ILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIEDIITHK 244
K+ + +A S Y R + T E +P D++ IL+ KLS+GTWV+YY N+++
Sbjct: 179 RKLKVKEAESLYRRNVAATTEFFPDDINKILRNKLSIGTWVAYYN------NVDN----- 227
Query: 245 STTSSSWIIFSLWNT 259
+ SW + S+W++
Sbjct: 228 ---TRSWAMLSVWDS 239
>UniRef100_Q9FN10 N-acetyltransferase hookless1-like protein [Arabidopsis thaliana]
Length = 386
Score = 142 bits (359), Expect = 8e-33
Identities = 98/272 (36%), Positives = 145/272 (53%), Gaps = 34/272 (12%)
Query: 11 VVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVA 70
VV+RE+D RD+ V +LE +C E+ S+ ++M GDPL+RIR P MLVA
Sbjct: 8 VVVREYDPKRDLTSVEELEESC-EVG-------SLLVDLM--GDPLARIRQSPSFHMLVA 57
Query: 71 EMVESKELVGVVKGCIKSVQ------------TPSGSLFKMGCILGLRVSPIHRRKGVGL 118
E+ E+VG+++G IK V +P + K+ + GLRVSP +RR G+GL
Sbjct: 58 EI--GNEIVGMIRGTIKMVTRGVNALRQADDVSPEINTTKLAFVSGLRVSPFYRRMGIGL 115
Query: 119 KLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHI 178
KLV +EEW L N A Y+++ TE +N AS LFT K Y F + ++P F
Sbjct: 116 KLVQRLEEWFLRNDAVYSYVQTENDNIASVKLFTEKSGYSKFRTPTFLVNP--VFNHRVT 173
Query: 179 SKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIE 238
+ VKI K++ A S Y T E +P D++ IL KLSLGT+++ + +
Sbjct: 174 VSRRVKIIKLAPSDAESLYRNRFSTTEFFPSDINSILTNKLSLGTYLAVPRGG------D 227
Query: 239 DIITHKSTTSSSWIIFSLWNTCEACDNNLQVK 270
++ + SW + S+WN+ + LQVK
Sbjct: 228 NVSGSLPDQTGSWAVISIWNSKDV--YRLQVK 257
>UniRef100_Q67UR2 GCN5-related N-acetyltransferase-like [Oryza sativa]
Length = 419
Score = 123 bits (308), Expect = 7e-27
Identities = 90/259 (34%), Positives = 129/259 (49%), Gaps = 45/259 (17%)
Query: 11 VVIREFDEDRDVKVVGKLERNC-TEING--------------------------TTKKGF 43
V +REFD ++D+ V +LER C ++G T KK
Sbjct: 12 VRVREFDVEKDLPAVEELERRCQVGLSGDMAAVHDHADDGDGAAAKEKKKTKTKTKKKKA 71
Query: 44 SIFTNMMSNGDPLSRIRFYPLHVMLVAEM---VESKELVGVVKGCIKSVQTPSGS---LF 97
S+ + GDPL+R+R P HVMLVAE E K++VGV+K C+K+V
Sbjct: 72 SMSLCVEQIGDPLARVRHAPEHVMLVAEYGEEEEKKKVVGVIKACVKTVSRGGKQEKPFV 131
Query: 98 KMGCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNY 157
K+ +LGLRVSP HRR G+G LV EEW + GA++A +AT ++N AS LFT + Y
Sbjct: 132 KVANLLGLRVSPSHRRLGIGTALVRRAEEWCVARGAEHATMATTESNAASLALFTGRFGY 191
Query: 158 FNFTSLIIFLHPPTSFPTNHISKKDV----KIDKISIDQAISFYTRIL--KTKELYPLDM 211
F HP H + V ++ ++ + A + Y R+L + E P DM
Sbjct: 192 APFRRPEFIGHPV------HAHRLPVARGHRVFQLPPEVAAAAYARLLPPQDAEFLPADM 245
Query: 212 DIILKEKLSLGTWVSYYKD 230
+L KL+LGT+V+ D
Sbjct: 246 PALLAHKLTLGTFVAVAAD 264
>UniRef100_Q6H820 GCN5-related N-acetyltransferase-like [Oryza sativa]
Length = 421
Score = 111 bits (278), Expect = 2e-23
Identities = 82/252 (32%), Positives = 121/252 (47%), Gaps = 40/252 (15%)
Query: 11 VVIREFDEDRDVKVVGKLERNC----TEINGT-------TKKGFSIFTNMMSNGDPLSRI 59
V +REF ++D+ V +LER C + NG K+G S++ + GDP +R+
Sbjct: 15 VRVREFIMEKDLPAVEELERLCQAGLSGDNGAGGGGGKKKKRGMSLYAEQI--GDPFARV 72
Query: 60 RFYPLHVMLVAEMVESKELVGVVKGCIKSV-----------QTPSGSLFKMGCILGLRVS 108
R P HV+LVAE + E+VGV+K C++ V +T + K C+LGLRVS
Sbjct: 73 RHAPDHVILVAECGD--EVVGVIKACVRMVTRGSSSSLRKTKTKTNKFVKAACLLGLRVS 130
Query: 109 PIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLH 168
P HRR G+ +LV EEW GA YA +AT +N AS LF + Y F H
Sbjct: 131 PSHRRLGIATELVRRAEEWCAARGAAYATMATTASNAASLALFQGRFKYALFRKPRFLGH 190
Query: 169 P----PTSFPTNHISKKDVKIDKISIDQAISFYTRIL----KTKELYPLDMDIILKEKLS 220
P P H ++ ++ A + Y +L E P D+ +L KL+
Sbjct: 191 PVHRHRARVPRAH------RVLQLPPPLAAAAYAALLPAAAAAPEFVPADLPALLAHKLT 244
Query: 221 LGTWVSYYKDEG 232
GT+++ + G
Sbjct: 245 RGTYLAVERSPG 256
>UniRef100_P95978 Orf c04040 protein [Sulfolobus solfataricus]
Length = 151
Score = 47.4 bits (111), Expect = 5e-04
Identities = 35/105 (33%), Positives = 52/105 (49%), Gaps = 13/105 (12%)
Query: 43 FSIFTNMMSNGDPLSRIRFYPL---HVMLVAEMVESKELVGVVKGCIKSVQTPSGSLFKM 99
F I T + P S +R Y + LVA+ E ++VG + G I+
Sbjct: 16 FQIETESFEDPYPYSLLRAYLFLANKLYLVAKQRE--KVVGYIIGIIQYGYR-------- 65
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNN 144
G I+ + V PI+R++G+G KL+ IEE NGA Y++L NN
Sbjct: 66 GHIVSIAVEPIYRKQGIGAKLLNEIEERFKLNGARYSYLEVNTNN 110
>UniRef100_O05705 N-terminal acetyl transferase [Sulfolobus shibatae]
Length = 87
Score = 44.7 bits (104), Expect = 0.003
Identities = 20/45 (44%), Positives = 29/45 (64%)
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNN 144
G I+ + V P +R+KG+G KL++ IEE NGA Y++L NN
Sbjct: 2 GHIVSIAVEPAYRKKGIGTKLLSEIEERFKLNGAKYSYLEVNINN 46
>UniRef100_UPI000033E574 UPI000033E574 UniRef100 entry
Length = 169
Score = 38.1 bits (87), Expect = 0.28
Identities = 27/97 (27%), Positives = 47/97 (47%), Gaps = 9/97 (9%)
Query: 52 NGDPLSRIRFYPLHVMLVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIH 111
N PL F +++A+ +E G + GC +++ SG + ++G ++ VSP++
Sbjct: 30 NAHPLDHQSFTQSRAVILAKTLE-----GEMVGCA-AIKAGSGPVGELGYLV---VSPLY 80
Query: 112 RRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASK 148
RR+G+ L E T G F + NNAS+
Sbjct: 81 RRRGIAQGLTLKRIEVAKTQGIALLFATIREENNASR 117
>UniRef100_UPI00002DAC07 UPI00002DAC07 UniRef100 entry
Length = 481
Score = 38.1 bits (87), Expect = 0.28
Identities = 19/56 (33%), Positives = 30/56 (52%)
Query: 98 KMGCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTN 153
K G + L +SP+ RRKG+G +L ++ E+ G L T + A+ NL+ N
Sbjct: 346 KQGELRRLSISPLFRRKGLGSRLTQTVTEFCEERGFSELVLQTSASRTAAVNLYKN 401
>UniRef100_O34376 Putative acetyl transferase [Bacillus subtilis]
Length = 247
Score = 38.1 bits (87), Expect = 0.28
Identities = 18/52 (34%), Positives = 29/52 (55%)
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLF 151
G + + V+ HR KG G +++ + EW NGA+ FL K N A+ +L+
Sbjct: 179 GGLSNIVVAEEHRGKGAGTQVIRVLTEWAKNNGAERMFLQVMKENLAAVSLY 230
>UniRef100_Q87FT2 Putative acetyltransferase [Vibrio parahaemolyticus]
Length = 162
Score = 38.1 bits (87), Expect = 0.28
Identities = 22/89 (24%), Positives = 43/89 (47%), Gaps = 1/89 (1%)
Query: 76 KELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADY 135
+++VG + G ++ + + ++G + L VS HR G+GL L+ IE + G +
Sbjct: 69 RQVVGFISGTVRDIDSMVSVAKRVGFVNELVVSESHRNLGIGLSLMDKIESDLCNQGIEE 128
Query: 136 AFLATEKNNNASKNLFTNKCNYFNFTSLI 164
L N+ ++ F +K Y T ++
Sbjct: 129 IGLTVASFNHEGED-FYHKMGYRVMTKMM 156
>UniRef100_Q88XP9 Lipoprotein [Lactobacillus plantarum]
Length = 391
Score = 38.1 bits (87), Expect = 0.28
Identities = 22/77 (28%), Positives = 37/77 (47%), Gaps = 1/77 (1%)
Query: 122 TSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHISKK 181
TSI W N +Y F AT ++ N+ N ++ ++ F S + P S T + K
Sbjct: 256 TSISNWTSLNWKNYVFPATSESKNSGAN-SNDESDFSQFKSHVQNFFPNLSGVTANAHYK 314
Query: 182 DVKIDKISIDQAISFYT 198
D + K+S++ FY+
Sbjct: 315 DKSLSKLSVNITTKFYS 331
>UniRef100_Q9T259 Ribosomal protein L5 [Phytophthora infestans]
Length = 177
Score = 37.7 bits (86), Expect = 0.37
Identities = 30/117 (25%), Positives = 55/117 (46%), Gaps = 6/117 (5%)
Query: 103 LGLRVSPIHRRKGVG----LKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYF 158
+G + + I ++K + LKL+T+ + + + + FL +KN+ + K N F
Sbjct: 39 IGFKNANIEKKKLINIILLLKLITNQKPIITKSKKNNIFLKIKKNSIIGCKITLRKKNIF 98
Query: 159 NF-TSLIIFLHPP-TSFPTNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDI 213
NF ++IF+ P T N +K ++ Q T LK K++ P+D+ I
Sbjct: 99 NFLAKILIFILPNLTKINFNFTNKNIFNFQIQNVLQFFELKTEFLKFKDIPPIDVSI 155
>UniRef100_Q8YVI8 N-terminal acetyltransferase [Anabaena sp.]
Length = 213
Score = 37.0 bits (84), Expect = 0.63
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 7/87 (8%)
Query: 68 LVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIHRRKGVGLKLVTSIEEW 127
LVAE EL G + G I + + + G IL L V+P +R+GV KLV +
Sbjct: 60 LVAET--DGELAGFILGTIITKAS-----WTYGYILWLAVNPKFQRQGVADKLVDKVVAR 112
Query: 128 MLTNGADYAFLATEKNNNASKNLFTNK 154
M+ +GA + + T+ N + F K
Sbjct: 113 MIEDGARFMLVDTDPTNTPAVKFFNRK 139
>UniRef100_Q03600 Integrin alpha ina-1 precursor [Caenorhabditis elegans]
Length = 1139
Score = 37.0 bits (84), Expect = 0.63
Identities = 37/137 (27%), Positives = 62/137 (45%), Gaps = 11/137 (8%)
Query: 159 NFTSLIIFLHPPTSFPTNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEK 218
NF +L+ P S T H +K + + ID+ ++ K +PLDM I + E+
Sbjct: 488 NFVALLRS-RPVISIETKHKMEKRM----VDIDKGVNCPRG---AKTCFPLDMVIYVDEE 539
Query: 219 LSLGTWVSYYKDEGFKLNIEDIITHKSTTSSSWIIFS-LWNTCEACDNNLQVKTKLFQPL 277
G + + + F N+E I TT+ +I S N C +N V+ + ++ L
Sbjct: 540 TKRGAELVDFSSDVFMCNLEAIPFRADTTARGFIEGSHSHNYSWPCGSNSHVQKRTYRQL 599
Query: 278 RFLHATLNHAKDKICPL 294
+L + +KD I PL
Sbjct: 600 IYL--PVQESKDWITPL 614
>UniRef100_Q7V6T4 Putative ribosomal-protein-alanine acetyltransferase
[Prochlorococcus marinus]
Length = 161
Score = 36.2 bits (82), Expect = 1.1
Identities = 25/93 (26%), Positives = 47/93 (49%), Gaps = 19/93 (20%)
Query: 69 VAEMVESKEL-VGVVKGCIKSVQTPSGSLFKMGC---------ILGLRVSPIHRRKGVGL 118
+ E+++S+ L +GV+K S +L + C + + V P HRR+G+
Sbjct: 40 IQELIDSRSLCMGVLK---------SSTLLALACGWLVVDELHLTAIGVHPQHRRQGLAR 90
Query: 119 KLVTSIEEWMLTNGADYAFLATEKNNNASKNLF 151
L++ + E GA +A L +NN+A++ L+
Sbjct: 91 LLLSKLLEQGQRTGAIHATLEVARNNSAARGLY 123
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 505,838,719
Number of Sequences: 2790947
Number of extensions: 20995566
Number of successful extensions: 59802
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 31
Number of HSP's that attempted gapping in prelim test: 59717
Number of HSP's gapped (non-prelim): 85
length of query: 297
length of database: 848,049,833
effective HSP length: 126
effective length of query: 171
effective length of database: 496,390,511
effective search space: 84882777381
effective search space used: 84882777381
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC126780.2


