

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.1 + phase: 0 /pseudo
(596 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
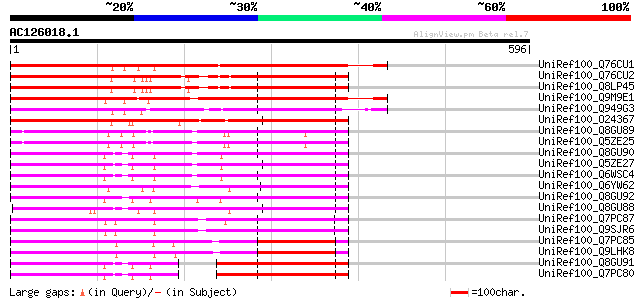
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q76CU1 PDR-type ABC transporter 2 [Nicotiana tabacum] 436 e-120
UniRef100_Q76CU2 PDR-type ABC transporter 1 [Nicotiana tabacum] 429 e-119
UniRef100_Q8LP45 Pleiotropic drug resistance like protein [Nicot... 427 e-118
UniRef100_Q9M9E1 Putative ABC transporter [Arabidopsis thaliana] 419 e-115
UniRef100_Q949G3 ABC1 protein [Nicotiana plumbaginifolia] 402 e-110
UniRef100_O24367 PDR5-like ABC transporter [Spirodela polyrrhiza] 372 e-101
UniRef100_Q8GU89 PDR-like ABC transporter [Oryza sativa] 368 e-100
UniRef100_Q5ZE25 Putative PDR-type ABC transporter 2 [Oryza sativa] 368 e-100
UniRef100_Q8GU90 PDR-like ABC transporter [Oryza sativa] 365 2e-99
UniRef100_Q5ZE27 Putative ABC1 protein [Oryza sativa] 365 2e-99
UniRef100_Q6WSC4 PDR-type ABC transporter 9 [Oryza sativa] 365 3e-99
UniRef100_Q6YW62 Putative PDR-like ABC transporter [Oryza sativa] 363 8e-99
UniRef100_Q8GU92 PDR-like ABC transporter [Oryza sativa] 360 5e-98
UniRef100_Q8GU88 PDR-like ABC transporter [Oryza sativa] 347 7e-94
UniRef100_Q7PC87 PDR6 ABC transporter [Arabidopsis thaliana] 345 2e-93
UniRef100_Q9SJR6 Putative ABC transporter [Arabidopsis thaliana] 343 1e-92
UniRef100_Q7PC85 PDR10 ABC transporter [Arabidopsis thaliana] 292 2e-77
UniRef100_Q9LHK8 ABC transporter-like protein [Arabidopsis thali... 289 1e-76
UniRef100_Q8GU91 PDR-like ABC transporter [Oryza sativa] 233 9e-60
UniRef100_Q7PC80 PDR1 ABC transporter [Oryza sativa] 233 9e-60
>UniRef100_Q76CU1 PDR-type ABC transporter 2 [Nicotiana tabacum]
Length = 1078
Score = 436 bits (1120), Expect = e-120
Identities = 248/469 (52%), Positives = 291/469 (61%), Gaps = 64/469 (13%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI MTLFFRTEM R+ DD G+YAGALFF ++ +MFNGMSE++MTI KLPV+YKQRDL
Sbjct: 185 MALITMTLFFRTEMPRDTTDDGGIYAGALFFVVIMIMFNGMSELAMTIFKLPVFYKQRDL 244
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHES---- 116
LF+PSWAYA+PSWILKIPV+L+EV LWV LTYYVIGFDPN+ R K F ++
Sbjct: 245 LFFPSWAYALPSWILKIPVTLVEVGLWVILTYYVIGFDPNISRFLKHFLLLIVVNQMASG 304
Query: 117 ------------------DGFRIVPSHRIPG*---KHDCCQY-IWVLCTSHI-----SVI 149
F ++ + G + D + IW TS + S++
Sbjct: 305 LFRFIGAMGRTMGVASTFGSFALLLQFALGGFVLSRDDVKSWWIWGYWTSPMMYSVNSIL 364
Query: 150 GWLHSLKNWHNATAD----LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGSR 205
K W++ + LG + +RGFFP AYWYWIGVG + + F
Sbjct: 365 VNEFDGKKWNHIVSGGNETLGTTVVKSRGFFPEAYWYWIGVGALVGFTIVFNFCYSLALA 424
Query: 206 CTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMVLPFEPHSIT 265
P S+ T+ G DS+TES + KKGMVLPFEPHSIT
Sbjct: 425 FLNPFDKPQAVLPEDGENAENVEVSSQITSTDGG--DSITESQNNNKKGMVLPFEPHSIT 482
Query: 266 FDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR 325
FDD+VYSVDMP EMKEQG EDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR
Sbjct: 483 FDDVVYSVDMPQEMKEQGAGEDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR 542
Query: 326 KTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSN 385
KTGGYIDGDIK+SGYPKKQETFARISGYCE NDIHSP+VTV+ESL+YSAWLRLP VD
Sbjct: 543 KTGGYIDGDIKISGYPKKQETFARISGYCEQNDIHSPYVTVYESLVYSAWLRLPQNVDET 602
Query: 386 TTKVSEHVLYSEWIWLHLLTLLNEHDRGGGRFIDHAMLSAELTPLMTSL 434
T K+ F+D M EL PL ++L
Sbjct: 603 TRKM---------------------------FVDEVMELVELRPLRSAL 624
Score = 37.4 bits (85), Expect = 1.3
Identities = 24/106 (22%), Positives = 51/106 (47%), Gaps = 1/106 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRD 59
+ALI T+F+ + D G+++ ++ + S + +A + V+Y++R
Sbjct: 836 IALIFGTMFWDLGTKVSKSQDLLNAMGSMYAAVLFLGVQNASSVQPVVAVERTVFYRERA 895
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y + YA ++IP ++ + + Y +IGF+ +VG+ F
Sbjct: 896 AGMYSAIPYAFGQVSIEIPYIFVQSVFYGIIVYAMIGFEWDVGKFF 941
>UniRef100_Q76CU2 PDR-type ABC transporter 1 [Nicotiana tabacum]
Length = 1434
Score = 429 bits (1104), Expect = e-119
Identities = 245/436 (56%), Positives = 286/436 (65%), Gaps = 62/436 (14%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI MTLFFRTEM R+ DD G+YAGALFF ++ +MFNGMSE++MTI KLPV+YKQRDL
Sbjct: 542 MALITMTLFFRTEMPRDTTDDGGIYAGALFFVVIMIMFNGMSELAMTIFKLPVFYKQRDL 601
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHE---SD 117
LF+PSWAYAIPSWILKIPV+L+EV LWV LTYYVIGFDPN+ R KQF ++ S
Sbjct: 602 LFFPSWAYAIPSWILKIPVTLVEVGLWVILTYYVIGFDPNITRFLKQFLLLIVVNQMASG 661
Query: 118 GFRIVPS-HRIPG*KHDCCQYIWVL-------CTSHISVIGW----------LHSL---- 155
FR + + R G + +L S V W ++S+
Sbjct: 662 MFRFIGAVGRTMGVASTFGSFALLLQFALGGFVLSRDDVKSWWIWGYWISPMMYSVNSIL 721
Query: 156 ------KNWHN----ATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGS- 204
K W++ LG + +RGFFP AYWYWIGVG + + F +C S
Sbjct: 722 VNEFDGKKWNHIVPGGNETLGSTVVKSRGFFPEAYWYWIGVGALVGFTVVFN---FCYSL 778
Query: 205 -----------RCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKK 253
+ P N S+ T+ G DS++ES + KK
Sbjct: 779 ALAYLNPFDKPQAVLPEDGENAENG---------EVSSQITSTDGG--DSISESQNNKK- 826
Query: 254 GMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGA 313
GMVLPFEPHSITFDD+VYSVDMP EMKEQG EDRLVLLKGVSGAFRPGVLTALMGVSGA
Sbjct: 827 GMVLPFEPHSITFDDVVYSVDMPQEMKEQGAGEDRLVLLKGVSGAFRPGVLTALMGVSGA 886
Query: 314 GKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYS 373
GKTTLMDVLAGRKTGGYIDG+IK+SGYPKKQETFARISGYCE NDIHSP+VTV+ESL+YS
Sbjct: 887 GKTTLMDVLAGRKTGGYIDGEIKISGYPKKQETFARISGYCEQNDIHSPYVTVYESLVYS 946
Query: 374 AWLRLPSGVDSNTTKV 389
AWLRLP VD T K+
Sbjct: 947 AWLRLPQDVDEKTRKM 962
Score = 55.1 bits (131), Expect = 6e-06
Identities = 30/91 (32%), Positives = 51/91 (55%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
R+ +L +LK +SG +P +T L+G +GKTTL+ LAG+ + G + +G+
Sbjct: 170 RKRQLTILKDISGIIKPCRMTLLLGPPSSGKTTLLLALAGKLDPALKVTGKVSYNGHELH 229
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y +D+H +TV E+L +SA
Sbjct: 230 EFVPQRTAAYISQHDLHIGEMTVRETLEFSA 260
Score = 37.4 bits (85), Expect = 1.3
Identities = 24/106 (22%), Positives = 51/106 (47%), Gaps = 1/106 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRD 59
+ALI T+F+ + D G+++ ++ + S + +A + V+Y++R
Sbjct: 1192 IALIFGTMFWDLGTKVSKSQDLLNAMGSMYAAVLFLGVQNASSVQPVVAIERTVFYRERA 1251
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y + YA ++IP ++ + + Y +IGF+ +VG+ F
Sbjct: 1252 AGMYSAIPYAFGQVSIEIPYIFVQSVFYGIIVYAMIGFEWDVGKFF 1297
>UniRef100_Q8LP45 Pleiotropic drug resistance like protein [Nicotiana tabacum]
Length = 1434
Score = 427 bits (1098), Expect = e-118
Identities = 244/436 (55%), Positives = 285/436 (64%), Gaps = 62/436 (14%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI MTLFFRTEM R+ DD G+YAGALFF ++ +MFNGMSE++MTI KLPV+YKQRDL
Sbjct: 542 MALITMTLFFRTEMPRDTTDDGGIYAGALFFVVIMIMFNGMSELAMTIFKLPVFYKQRDL 601
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHE---SD 117
LF+PSWAYAIPSWILKIPV+L+EV LWV LTYYVIGFDPN+ R KQF ++ S
Sbjct: 602 LFFPSWAYAIPSWILKIPVTLVEVGLWVILTYYVIGFDPNITRFLKQFLLLIVVNQMASG 661
Query: 118 GFRIVPS-HRIPG*KHDCCQYIWVL-------CTSHISVIGW----------LHSL---- 155
FR + + R G + +L S V W ++S+
Sbjct: 662 MFRFIGAVGRTMGVASTFGSFALLLQFALGGFVLSRDDVKSWWIWGYWISPMMYSVNSIL 721
Query: 156 ------KNWHN----ATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGS- 204
K W++ LG + +RGFFP AYWYWIGVG + + F +C S
Sbjct: 722 VNEFDGKKWNHIVPGGNETLGSTVVKSRGFFPEAYWYWIGVGALVGFTVVFN---FCYSL 778
Query: 205 -----------RCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKK 253
+ P N S+ + G DS++ES + KK
Sbjct: 779 ALAYLNPFDKPQAVLPEDGENAENG---------EVSSQIPSTDGG--DSISESQNNKK- 826
Query: 254 GMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGA 313
GMVLPFEPHSITFDD+VYSVDMP EMKEQG EDRLVLLKGVSGAFRPGVLTALMGVSGA
Sbjct: 827 GMVLPFEPHSITFDDVVYSVDMPQEMKEQGAGEDRLVLLKGVSGAFRPGVLTALMGVSGA 886
Query: 314 GKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYS 373
GKTTLMDVLAGRKTGGYIDG+IK+SGYPKKQETFARISGYCE NDIHSP+VTV+ESL+YS
Sbjct: 887 GKTTLMDVLAGRKTGGYIDGEIKISGYPKKQETFARISGYCEQNDIHSPYVTVYESLVYS 946
Query: 374 AWLRLPSGVDSNTTKV 389
AWLRLP VD T K+
Sbjct: 947 AWLRLPQDVDEKTRKM 962
Score = 55.1 bits (131), Expect = 6e-06
Identities = 30/91 (32%), Positives = 51/91 (55%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
R+ +L +LK +SG +P +T L+G +GKTTL+ LAG+ + G + +G+
Sbjct: 170 RKRQLTILKDISGIIKPCRMTLLLGPPSSGKTTLLLALAGKLDPALKVTGKVSYNGHELH 229
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y +D+H +TV E+L +SA
Sbjct: 230 EFVPQRTAAYISQHDLHIGEMTVRETLEFSA 260
Score = 37.4 bits (85), Expect = 1.3
Identities = 24/106 (22%), Positives = 51/106 (47%), Gaps = 1/106 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRD 59
+ALI T+F+ + D G+++ ++ + S + +A + V+Y++R
Sbjct: 1192 IALIFGTMFWDLGTKVSKSQDLLNAMGSMYAAVLFLGVQNASSVQPVVAIERTVFYRERA 1251
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y + YA ++IP ++ + + Y +IGF+ +VG+ F
Sbjct: 1252 AGMYSAIPYAFGQVSIEIPYIFVQSVFYGIIVYAMIGFEWDVGKFF 1297
>UniRef100_Q9M9E1 Putative ABC transporter [Arabidopsis thaliana]
Length = 1423
Score = 419 bits (1077), Expect = e-115
Identities = 237/470 (50%), Positives = 289/470 (61%), Gaps = 72/470 (15%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MA + MTLFFRTEM + + D +Y GALFF L+ +MFNGMSE+SMTIAKLPV+YKQRDL
Sbjct: 535 MAFLTMTLFFRTEMQKKTEVDGSLYTGALFFILMMLMFNGMSELSMTIAKLPVFYKQRDL 594
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF------------ 108
LFYP+W Y++P W+LKIP+S ME +L F+TYYVIGFDPNVGR+FKQ+
Sbjct: 595 LFYPAWVYSLPPWLLKIPISFMEAALTTFITYYVIGFDPNVGRLFKQYILLVLMNQMASA 654
Query: 109 ----------RCVVFHESDGFRIVPSHRIPG*---KHDCCQY-IWVLCTSHISVIGWLHS 154
+V + F ++ + G + D ++ IW S I + G
Sbjct: 655 LFKMVAALGRNMIVANTFGAFAMLVFFALGGVVLSRDDIKKWWIWGYWISPI-MYGQNAI 713
Query: 155 LKN------W----HNATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGS 204
L N W N++ LG +L +RGF PHAYWYWIG G + + F G
Sbjct: 714 LANEFFGHSWSRAVENSSETLGVTFLKSRGFLPHAYWYWIGTGALLGFVVLFNFGFTLAL 773
Query: 205 RCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMVLPFEPHSI 264
N+ + + S+ T S R++ V E+ KK+GMVLPFEPHSI
Sbjct: 774 TFL--------NSLGKPQAVIAEEPASDETELQSARSEGVVEAGANKKRGMVLPFEPHSI 825
Query: 265 TFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAG 324
TFD++VYSVDMP EM EQG +EDRLVLLKGV+GAFRPGVLTALMGVSGAGKTTLMDVLAG
Sbjct: 826 TFDNVVYSVDMPQEMIEQGTQEDRLVLLKGVNGAFRPGVLTALMGVSGAGKTTLMDVLAG 885
Query: 325 RKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDS 384
RKTGGYIDG+I +SGYPK Q+TFARISGYCE DIHSPHVTV+ESL+YSAWLRLP VD
Sbjct: 886 RKTGGYIDGNITISGYPKNQQTFARISGYCEQTDIHSPHVTVYESLVYSAWLRLPKEVDK 945
Query: 385 NTTKVSEHVLYSEWIWLHLLTLLNEHDRGGGRFIDHAMLSAELTPLMTSL 434
N K+ FI+ M ELTPL +L
Sbjct: 946 NKRKI---------------------------FIEEVMELVELTPLRQAL 968
Score = 52.0 bits (123), Expect = 5e-05
Identities = 29/91 (31%), Positives = 47/91 (50%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYID-GDIKVSGYPKK 343
R+ + +L VSG +PG + L+G +GKTTL+ LAG+ G + +G+
Sbjct: 163 RKKKFTILNDVSGIVKPGRMALLLGPPSSGKTTLLLALAGKLDQELKQTGRVTYNGHGMN 222
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y ND+H +TV E+ Y+A
Sbjct: 223 EFVPQRTAAYIGQNDVHIGEMTVRETFAYAA 253
Score = 35.8 bits (81), Expect = 3.7
Identities = 27/149 (18%), Positives = 61/149 (40%), Gaps = 8/149 (5%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTI-AKLPVYYKQRD 59
+AL+ T+F+ + D G+++ ++ + + + + + V+Y+++
Sbjct: 1180 IALMFGTMFWDLGGKTKTRQDLSNAMGSMYTAVLFLGLQNAASVQPVVNVERTVFYREQA 1239
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR-------MFKQFRCVV 112
Y + YA ++IP L++ ++ + Y +IGF+ + M+ F
Sbjct: 1240 AGMYSAMPYAFAQVFIEIPYVLVQAIVYGLIVYAMIGFEWTAVKFFWYLFFMYGSFLTFT 1299
Query: 113 FHESDGFRIVPSHRIPG*KHDCCQYIWVL 141
F+ + P+H I IW L
Sbjct: 1300 FYGMMAVAMTPNHHIASVVSSAFYGIWNL 1328
>UniRef100_Q949G3 ABC1 protein [Nicotiana plumbaginifolia]
Length = 1436
Score = 402 bits (1032), Expect = e-110
Identities = 230/472 (48%), Positives = 280/472 (58%), Gaps = 74/472 (15%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
+AL+ MT+FFRT+M R+ +D G+Y+GALFF ++ +MFNG+SE+ MT+ KLPV+YKQRD
Sbjct: 546 IALMTMTIFFRTKMPRDSAEDGGIYSGALFFVVIMIMFNGLSELPMTLYKLPVFYKQRDF 605
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHES---- 116
LFYPSWAYAIPSWILKIPV+ EV +WVFLTYYV+GFDPNVGR FKQF ++
Sbjct: 606 LFYPSWAYAIPSWILKIPVTFAEVGMWVFLTYYVMGFDPNVGRFFKQFLLLLLVNQMASA 665
Query: 117 ------------------DGFRIVPSHRIPG*---KHDCCQY-IWVLCTSHISVI----- 149
F ++ + G ++D + IW TS +
Sbjct: 666 LFRFIAAVGRTMGVASTFGAFALLLQFALGGFILARNDVKDWWIWGYWTSPLMYSVNAIL 725
Query: 150 -------GWLHSLKNWHNATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWC 202
W H + T LG + RGFFP AYWYWIGVG + + F
Sbjct: 726 VNEFDGQKWKHIVAG---GTEPLGAAVVRARGFFPDAYWYWIGVGALAGFIVMFNIAYSV 782
Query: 203 GSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMVLPFEPH 262
P + SE + + + +S KKKGMVLPF+PH
Sbjct: 783 ALAYLNPFDKPQATISDESEN-----NESESSPQITSTQEG-DSASENKKKGMVLPFDPH 836
Query: 263 SITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVL 322
SITFD++VYSVDMP EM+E G ++RLVLLK VSGAFRPGVLTALMGVSGAGKTTLMDVL
Sbjct: 837 SITFDEVVYSVDMPPEMRESGTSDNRLVLLKSVSGAFRPGVLTALMGVSGAGKTTLMDVL 896
Query: 323 AGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
AGRKTGGYIDG IK+SGYPKKQ+TFARISGYCE NDIHSP+VTV ESL+YSAWLRLP V
Sbjct: 897 AGRKTGGYIDGSIKISGYPKKQDTFARISGYCEQNDIHSPYVTVFESLVYSAWLRLPQDV 956
Query: 383 DSNTTKVSEHVLYSEWIWLHLLTLLNEHDRGGGRFIDHAMLSAELTPLMTSL 434
NE R F++ M ELTPL ++L
Sbjct: 957 -------------------------NEEKR--MMFVEEVMDLVELTPLRSAL 981
Score = 55.5 bits (132), Expect = 4e-06
Identities = 30/91 (32%), Positives = 52/91 (56%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
++ ++ +LK VSG +P +T L+G G+GKTTL+ LAG+ + G + +G+
Sbjct: 174 KKRQVTILKDVSGIVKPCRMTLLLGPPGSGKTTLLLALAGKLDSALKVTGKVTYNGHELH 233
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y +D+H +TV E+L +SA
Sbjct: 234 EFVPQRTAAYISQHDLHIGEMTVRETLEFSA 264
Score = 38.1 bits (87), Expect = 0.74
Identities = 23/114 (20%), Positives = 55/114 (48%), Gaps = 1/114 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRD 59
+ALI T+F+ + D G+++ ++ + S + ++ + V+Y+++
Sbjct: 1193 IALIFGTMFWDIGTKVSRNQDLVNAMGSMYAAVLFLGVQNSSSVQPVVSVERTVFYREKA 1252
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVF 113
Y + YA +++IP ++ +++ + Y +IGF+ V + F F + F
Sbjct: 1253 AGMYSAIPYAFAQVLIEIPYIFVQATVYGLIVYSMIGFEWTVAKFFWDFFFMFF 1306
>UniRef100_O24367 PDR5-like ABC transporter [Spirodela polyrrhiza]
Length = 1441
Score = 372 bits (954), Expect = e-101
Identities = 219/434 (50%), Positives = 274/434 (62%), Gaps = 48/434 (11%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
+ALIAMT+FFRT++ RN +DA ++ GA+F LVT +FNG +E++M+IAKLPV+YKQRDL
Sbjct: 538 LALIAMTVFFRTKLPRNGLEDATIFFGAMFLGLVTHLFNGFAELAMSIAKLPVFYKQRDL 597
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHE---SD 117
LFYP WAYA+P+WILKIP+S +E +W+ +TYYVIGFDPNV RMF+ + +V S
Sbjct: 598 LFYPPWAYALPTWILKIPISFVECGVWIAMTYYVIGFDPNVVRMFRHYLLLVLISQVASG 657
Query: 118 GFRIVPSHRIPG*KHDCCQ-----------------------YIW-------VLCTSHIS 147
FR++ + D +IW + + I+
Sbjct: 658 LFRLLAAVGRDMVVADTFGAFAQLVLLVLGGFIIAREKIKKFWIWGYWSSPLMYAQNAIA 717
Query: 148 VIGWL-HSLKNWHNATAD-LGKDYLDTRGFFPHAYWYWIGVGG-IGWIC---------LS 195
V +L HS +AT LG+ +L RG F WYWIGVG IG++ L
Sbjct: 718 VNEFLGHSWNKLVDATGQTLGERFLRNRGIFVDKNWYWIGVGALIGYMVLFNFLFILFLE 777
Query: 196 FQRGVWCGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGM 255
+ + G N R TR T G + + + +KKGM
Sbjct: 778 WLDPLGKGQTTVSEEALQEKEAN-RTGANVELATRGSAATSDGGSVEIRKDGN--RKKGM 834
Query: 256 VLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGK 315
VLPF P SITFD++ YSVDMP EMK++GV ED+L+LLKGVSGAFRPGVLTALMGVSG GK
Sbjct: 835 VLPFTPLSITFDNVKYSVDMPQEMKDRGVTEDKLLLLKGVSGAFRPGVLTALMGVSGRGK 894
Query: 316 TTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAW 375
TTLMDVLAGRKTGGYI+GDI++SGYPK QETFARISGYCE NDIHSPHVTV+ESLLYSAW
Sbjct: 895 TTLMDVLAGRKTGGYIEGDIRISGYPKNQETFARISGYCEQNDIHSPHVTVYESLLYSAW 954
Query: 376 LRLPSGVDSNTTKV 389
LRLP+ VD K+
Sbjct: 955 LRLPAEVDEKQRKM 968
Score = 54.7 bits (130), Expect = 8e-06
Identities = 30/85 (35%), Positives = 48/85 (56%), Gaps = 1/85 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKKQETFAR 349
+L VSG +P +T L+G GAGKTTL+ LAG+ + G++ +G+ + R
Sbjct: 173 ILHDVSGIIKPCRMTLLLGPPGAGKTTLLLALAGKLDNTLKVTGNVTYNGHGMHEFVPQR 232
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSA 374
S Y +D+H +TV E+L +S+
Sbjct: 233 TSAYISQHDVHIGEMTVRETLAFSS 257
Score = 35.8 bits (81), Expect = 3.7
Identities = 26/103 (25%), Positives = 48/103 (46%), Gaps = 9/103 (8%)
Query: 1 MALIAMTLFFRTEMHRNDQDD-----AGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYY 55
+ALI T+F+ R+ D +YA LF + N + + + V+Y
Sbjct: 1198 IALIFGTIFWDLGKKRSTSLDLINAMGSMYAAVLFIGIQ----NAQTVQPIVDVERTVFY 1253
Query: 56 KQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFD 98
+++ Y + YA ++++P L++ L+ L Y +IGFD
Sbjct: 1254 REKAAGMYSALPYAYAQVLIEVPHILVQTLLYGLLVYSMIGFD 1296
>UniRef100_Q8GU89 PDR-like ABC transporter [Oryza sativa]
Length = 1450
Score = 368 bits (945), Expect = e-100
Identities = 216/447 (48%), Positives = 269/447 (59%), Gaps = 66/447 (14%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI MT FFRT M R+D+D +Y GAL+F L T+MFNG +E++MT+ KLPV++KQRDL
Sbjct: 539 MALIVMTTFFRTSM-RHDRDYGMIYLGALYFALDTVMFNGFAELAMTVMKLPVFFKQRDL 597
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVV-------- 112
LF+P+WAY IPSWIL+IP++ +EV ++VF+TYYVIGFDP+V R FKQ+ ++
Sbjct: 598 LFFPAWAYTIPSWILQIPITFLEVGVYVFITYYVIGFDPSVSRFFKQYLLLLALNQMSSA 657
Query: 113 ---FHESDGFRIVPSHRI------------------PG*KHDCCQYIWV---------LC 142
F G +V SH P K W+ +
Sbjct: 658 LFRFIAGIGRDMVVSHTFGPLSLLAFAALGGFILARPDVKKWWIWGYWISPLSYAQNAIS 717
Query: 143 TSHISVIGWLHSLKNWHNATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWC 202
T+ W L N T LG L +RG F A WYWIG+G + L F
Sbjct: 718 TNEFLGHSWSQILPG-ENVT--LGVSVLKSRGIFTEAKWYWIGLGALLGYTLLFNLLYTV 774
Query: 203 GSRCTRPI**A*CNNN*RFRR*FIYRTRSEFT-TYCSGRADSVT-----ESSH------- 249
P +++ + + T G+ D+ + E SH
Sbjct: 775 ALSVLSPF----TDSHASMSEDALKEKHANLTGEVVEGQKDTKSRKQELELSHIADQNSG 830
Query: 250 -------GKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPG 302
+KGMVLPF P SI+F+D+ YSVDMP MK QG+ EDRL+LLKGVSG+FRPG
Sbjct: 831 INSADSSASRKGMVLPFAPLSISFNDVRYSVDMPEAMKAQGITEDRLLLLKGVSGSFRPG 890
Query: 303 VLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSP 362
VLTALMGVSGAGKTTLMDVLAGRKTGGYI+GDI++SGYPKKQETFARISGYCE NDIHSP
Sbjct: 891 VLTALMGVSGAGKTTLMDVLAGRKTGGYIEGDIRISGYPKKQETFARISGYCEQNDIHSP 950
Query: 363 HVTVHESLLYSAWLRLPSGVDSNTTKV 389
HVTV+ESL++SAWLRLPS VDS K+
Sbjct: 951 HVTVYESLVFSAWLRLPSEVDSEARKM 977
Score = 54.7 bits (130), Expect = 8e-06
Identities = 33/96 (34%), Positives = 53/96 (54%), Gaps = 11/96 (11%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVS------ 338
++ + +L VSG +P +T L+G G+GKTTL+ LAG+ +D D+KVS
Sbjct: 167 KKQPMTVLHDVSGIIKPRRMTLLLGPPGSGKTTLLLALAGK-----LDKDLKVSGKVTYN 221
Query: 339 GYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSA 374
G+ + R + Y +D+H +TV E+L +SA
Sbjct: 222 GHGMHEFVPERTAAYISQHDLHIGEMTVRETLAFSA 257
Score = 37.0 bits (84), Expect = 1.6
Identities = 22/104 (21%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTM-MFNGMSEISMTIAKLPVYYKQRD 59
+AL+ T+F+ Q D G+++ ++ + + N S + + + V+Y++R
Sbjct: 1207 IALMFGTMFWNLGTRTKKQQDLFNAMGSMYAAVLYIGVQNSGSVQPVVVVERTVFYRERA 1266
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR 103
Y ++ YA +++P +++ ++ L Y +IGF+ V +
Sbjct: 1267 AGMYSAFPYAFGQVAIELPYIMVQTLIYGVLVYSMIGFEWTVAK 1310
>UniRef100_Q5ZE25 Putative PDR-type ABC transporter 2 [Oryza sativa]
Length = 1338
Score = 368 bits (945), Expect = e-100
Identities = 216/447 (48%), Positives = 269/447 (59%), Gaps = 66/447 (14%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI MT FFRT M R+D+D +Y GAL+F L T+MFNG +E++MT+ KLPV++KQRDL
Sbjct: 427 MALIVMTTFFRTSM-RHDRDYGMIYLGALYFALDTVMFNGFAELAMTVMKLPVFFKQRDL 485
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVV-------- 112
LF+P+WAY IPSWIL+IP++ +EV ++VF+TYYVIGFDP+V R FKQ+ ++
Sbjct: 486 LFFPAWAYTIPSWILQIPITFLEVGVYVFITYYVIGFDPSVSRFFKQYLLLLALNQMSSA 545
Query: 113 ---FHESDGFRIVPSHRI------------------PG*KHDCCQYIWV---------LC 142
F G +V SH P K W+ +
Sbjct: 546 LFRFIAGIGRDMVVSHTFGPLSLLAFAALGGFILARPDVKKWWIWGYWISPLSYAQNAIS 605
Query: 143 TSHISVIGWLHSLKNWHNATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWC 202
T+ W L N T LG L +RG F A WYWIG+G + L F
Sbjct: 606 TNEFLGHSWSQILPG-ENVT--LGVSVLKSRGIFTEAKWYWIGLGALLGYTLLFNLLYTV 662
Query: 203 GSRCTRPI**A*CNNN*RFRR*FIYRTRSEFT-TYCSGRADSVT-----ESSH------- 249
P +++ + + T G+ D+ + E SH
Sbjct: 663 ALSVLSPF----TDSHASMSEDALKEKHANLTGEVVEGQKDTKSRKQELELSHIADQNSG 718
Query: 250 -------GKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPG 302
+KGMVLPF P SI+F+D+ YSVDMP MK QG+ EDRL+LLKGVSG+FRPG
Sbjct: 719 INSADSSASRKGMVLPFAPLSISFNDVRYSVDMPEAMKAQGITEDRLLLLKGVSGSFRPG 778
Query: 303 VLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSP 362
VLTALMGVSGAGKTTLMDVLAGRKTGGYI+GDI++SGYPKKQETFARISGYCE NDIHSP
Sbjct: 779 VLTALMGVSGAGKTTLMDVLAGRKTGGYIEGDIRISGYPKKQETFARISGYCEQNDIHSP 838
Query: 363 HVTVHESLLYSAWLRLPSGVDSNTTKV 389
HVTV+ESL++SAWLRLPS VDS K+
Sbjct: 839 HVTVYESLVFSAWLRLPSEVDSEARKM 865
Score = 54.7 bits (130), Expect = 8e-06
Identities = 33/96 (34%), Positives = 53/96 (54%), Gaps = 11/96 (11%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVS------ 338
++ + +L VSG +P +T L+G G+GKTTL+ LAG+ +D D+KVS
Sbjct: 55 KKQPMTVLHDVSGIIKPRRMTLLLGPPGSGKTTLLLALAGK-----LDKDLKVSGKVTYN 109
Query: 339 GYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSA 374
G+ + R + Y +D+H +TV E+L +SA
Sbjct: 110 GHGMHEFVPERTAAYISQHDLHIGEMTVRETLAFSA 145
Score = 37.0 bits (84), Expect = 1.6
Identities = 22/104 (21%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTM-MFNGMSEISMTIAKLPVYYKQRD 59
+AL+ T+F+ Q D G+++ ++ + + N S + + + V+Y++R
Sbjct: 1095 IALMFGTMFWNLGTRTKKQQDLFNAMGSMYAAVLYIGVQNSGSVQPVVVVERTVFYRERA 1154
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR 103
Y ++ YA +++P +++ ++ L Y +IGF+ V +
Sbjct: 1155 AGMYSAFPYAFGQVAIELPYIMVQTLIYGVLVYSMIGFEWTVAK 1198
>UniRef100_Q8GU90 PDR-like ABC transporter [Oryza sativa]
Length = 1457
Score = 365 bits (937), Expect = 2e-99
Identities = 216/452 (47%), Positives = 269/452 (58%), Gaps = 74/452 (16%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++LIAMTLFFRT+M R+ G+Y GALFF ++ +MFNG SE+++T+ KLPV++KQRDL
Sbjct: 545 VSLIAMTLFFRTKMKRDSVTSGGIYMGALFFGVLMIMFNGFSELALTVFKLPVFFKQRDL 604
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LFYP+W+Y IPSWILKIP++ +EV +VFLTYYVIGFD NVG FKQ
Sbjct: 605 LFYPAWSYTIPSWILKIPITFIEVGGYVFLTYYVIGFDSNVGSFFKQYLLMLAINQMAGS 664
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF I+ ++ +IW +
Sbjct: 665 LFRFIGGAARNMIVANVFASFMLLIFMVLGGF-ILAREQVKK------WWIWGYWISPMM 717
Query: 141 LCTSHISVIGWL-HSLKNWHNATAD---LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSF 196
+ ISV + HS N++A LG L +RG FP A WYWIG G + + F
Sbjct: 718 YAQNAISVNELMGHSWNKIVNSSASNETLGVQVLKSRGVFPEARWYWIGFGAMIGFTILF 777
Query: 197 QRGVWCGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRA--------------- 241
RP N+ + R+ G
Sbjct: 778 NALFTLALTYLRPY----GNSRQSVSEEELKEKRANLNGEIVGDVHLSSGSTRRPMGNGT 833
Query: 242 --DS--VTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
DS V + + ++GMVLPF P S++FD++ YSVDMP EMK QGV +DRL LLKGVSG
Sbjct: 834 ENDSTIVDDDTEVTQRGMVLPFTPLSLSFDNVRYSVDMPQEMKAQGVADDRLELLKGVSG 893
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHN 357
+FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI+G I +SGYPKKQETFAR+SGYCE N
Sbjct: 894 SFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGSINISGYPKKQETFARVSGYCEQN 953
Query: 358 DIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
DIHSP VTV+ESLL+SAWLRLP VDSNT K+
Sbjct: 954 DIHSPQVTVYESLLFSAWLRLPEDVDSNTRKM 985
Score = 54.3 bits (129), Expect = 1e-05
Identities = 31/91 (34%), Positives = 50/91 (54%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
R+ + +L VSG +P +T L+G G+GKTTL+ LAGR G + +G+ +
Sbjct: 173 RKQTMPVLHDVSGIIKPRRMTLLLGPPGSGKTTLLLALAGRLGKDLKASGKVTYNGHGME 232
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y +D+H +TV E+L +SA
Sbjct: 233 EFVPERTAAYISQHDLHIGEMTVRETLAFSA 263
Score = 39.7 bits (91), Expect = 0.25
Identities = 22/90 (24%), Positives = 46/90 (50%), Gaps = 4/90 (4%)
Query: 24 VYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLME 83
+YA LF ++ N S + + V+Y++R Y ++ YA +++IP +L++
Sbjct: 1243 MYAAVLFIGVM----NCTSVQPVVAVERTVFYRERAAGMYSAFPYAFGQVVIEIPYTLVQ 1298
Query: 84 VSLWVFLTYYVIGFDPNVGRMFKQFRCVVF 113
+++ + Y +IGF+ + F +VF
Sbjct: 1299 ATVYGIIVYAMIGFEWTAAKFFWYLFFMVF 1328
>UniRef100_Q5ZE27 Putative ABC1 protein [Oryza sativa]
Length = 1281
Score = 365 bits (937), Expect = 2e-99
Identities = 216/452 (47%), Positives = 269/452 (58%), Gaps = 74/452 (16%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++LIAMTLFFRT+M R+ G+Y GALFF ++ +MFNG SE+++T+ KLPV++KQRDL
Sbjct: 369 VSLIAMTLFFRTKMKRDSVTSGGIYMGALFFGVLMIMFNGFSELALTVFKLPVFFKQRDL 428
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LFYP+W+Y IPSWILKIP++ +EV +VFLTYYVIGFD NVG FKQ
Sbjct: 429 LFYPAWSYTIPSWILKIPITFIEVGGYVFLTYYVIGFDSNVGSFFKQYLLMLAINQMAGS 488
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF I+ ++ +IW +
Sbjct: 489 LFRFIGGAARNMIVANVFASFMLLIFMVLGGF-ILAREQVKK------WWIWGYWISPMM 541
Query: 141 LCTSHISVIGWL-HSLKNWHNATAD---LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSF 196
+ ISV + HS N++A LG L +RG FP A WYWIG G + + F
Sbjct: 542 YAQNAISVNELMGHSWNKIVNSSASNETLGVQVLKSRGVFPEARWYWIGFGAMIGFTILF 601
Query: 197 QRGVWCGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRA--------------- 241
RP N+ + R+ G
Sbjct: 602 NALFTLALTYLRPY----GNSRQSVSEEELKEKRANLNGEIVGDVHLSSGSTRRPMGNGT 657
Query: 242 --DS--VTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
DS V + + ++GMVLPF P S++FD++ YSVDMP EMK QGV +DRL LLKGVSG
Sbjct: 658 ENDSTIVDDDTEVTQRGMVLPFTPLSLSFDNVRYSVDMPQEMKAQGVADDRLELLKGVSG 717
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHN 357
+FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI+G I +SGYPKKQETFAR+SGYCE N
Sbjct: 718 SFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGSINISGYPKKQETFARVSGYCEQN 777
Query: 358 DIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
DIHSP VTV+ESLL+SAWLRLP VDSNT K+
Sbjct: 778 DIHSPQVTVYESLLFSAWLRLPEDVDSNTRKM 809
Score = 52.4 bits (124), Expect = 4e-05
Identities = 30/85 (35%), Positives = 47/85 (55%), Gaps = 1/85 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKKQETFAR 349
+L VSG +P +T L+G G+GKTTL+ LAGR G + +G+ ++ R
Sbjct: 3 VLHDVSGIIKPRRMTLLLGPPGSGKTTLLLALAGRLGKDLKASGKVTYNGHGMEEFVPER 62
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSA 374
+ Y +D+H +TV E+L +SA
Sbjct: 63 TAAYISQHDLHIGEMTVRETLAFSA 87
Score = 39.7 bits (91), Expect = 0.25
Identities = 22/90 (24%), Positives = 46/90 (50%), Gaps = 4/90 (4%)
Query: 24 VYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLME 83
+YA LF ++ N S + + V+Y++R Y ++ YA +++IP +L++
Sbjct: 1067 MYAAVLFIGVM----NCTSVQPVVAVERTVFYRERAAGMYSAFPYAFGQVVIEIPYTLVQ 1122
Query: 84 VSLWVFLTYYVIGFDPNVGRMFKQFRCVVF 113
+++ + Y +IGF+ + F +VF
Sbjct: 1123 ATVYGIIVYAMIGFEWTAAKFFWYLFFMVF 1152
>UniRef100_Q6WSC4 PDR-type ABC transporter 9 [Oryza sativa]
Length = 1457
Score = 365 bits (936), Expect = 3e-99
Identities = 216/452 (47%), Positives = 269/452 (58%), Gaps = 74/452 (16%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++LIAMTLFFRT+M R+ G+Y GALFF ++ +MFNG SE+++T+ KLPV++KQRDL
Sbjct: 545 VSLIAMTLFFRTKMKRDSVTSGGIYMGALFFGVLMIMFNGFSELALTVFKLPVFFKQRDL 604
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LFYP+W+Y IPSWILKIP++ +EV +VFLTYYVIGFD NVG FKQ
Sbjct: 605 LFYPAWSYTIPSWILKIPITFIEVGGYVFLTYYVIGFDSNVGSFFKQYLLMLAINQMAGS 664
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF I+ ++ +IW +
Sbjct: 665 LFRFIGGAARNMIVANVFASFMLLIFMVLGGF-ILAREQVKK------WWIWGYWISPMM 717
Query: 141 LCTSHISVIGWL-HSLKNWHNATAD---LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSF 196
+ ISV + HS N++A LG L +RG FP A WYWIG G + + F
Sbjct: 718 YAQNAISVNELMGHSWNKIVNSSASNETLGVQVLKSRGVFPEARWYWIGFGAMIGFTILF 777
Query: 197 QRGVWCGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRA--------------- 241
RP N+ + R+ G
Sbjct: 778 NALFTLALTYLRPY----GNSRQSVSEEEMKEKRANLNGEIVGDVHLSSGSTRRPMGNGT 833
Query: 242 --DS--VTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
DS V + + ++GMVLPF P S++FD++ YSVDMP EMK QGV +DRL LLKGVSG
Sbjct: 834 ENDSTIVDDDTEVTQRGMVLPFTPLSLSFDNVRYSVDMPQEMKAQGVADDRLELLKGVSG 893
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHN 357
+FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI+G I +SGYPKKQETFAR+SGYCE N
Sbjct: 894 SFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGSINISGYPKKQETFARVSGYCEQN 953
Query: 358 DIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
DIHSP VTV+ESLL+SAWLRLP VDSNT K+
Sbjct: 954 DIHSPQVTVYESLLFSAWLRLPEDVDSNTRKM 985
Score = 54.3 bits (129), Expect = 1e-05
Identities = 31/91 (34%), Positives = 50/91 (54%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
R+ + +L VSG +P +T L+G G+GKTTL+ LAGR G + +G+ +
Sbjct: 173 RKQTMPVLHDVSGIIKPRRMTLLLGPPGSGKTTLLLALAGRLGKDLKASGKVTYNGHGME 232
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R + Y +D+H +TV E+L +SA
Sbjct: 233 EFVPERTAAYISQHDLHIGEMTVRETLAFSA 263
Score = 39.7 bits (91), Expect = 0.25
Identities = 22/90 (24%), Positives = 46/90 (50%), Gaps = 4/90 (4%)
Query: 24 VYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLME 83
+YA LF ++ N S + + V+Y++R Y ++ YA +++IP +L++
Sbjct: 1243 MYAAVLFIGVM----NCTSVQPVVAVERTVFYRERAAGMYSAFPYAFGQVVIEIPYTLVQ 1298
Query: 84 VSLWVFLTYYVIGFDPNVGRMFKQFRCVVF 113
+++ + Y +IGF+ + F +VF
Sbjct: 1299 ATVYGIIVYAMIGFEWTAAKFFWYLFFMVF 1328
>UniRef100_Q6YW62 Putative PDR-like ABC transporter [Oryza sativa]
Length = 1489
Score = 363 bits (932), Expect = 8e-99
Identities = 211/442 (47%), Positives = 264/442 (58%), Gaps = 61/442 (13%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
+ +I MTLF RT MH + D VY GALFF +V MFNG SE++M KLPV++KQRD
Sbjct: 583 ITIIVMTLFLRTNMHHETRTDGIVYLGALFFAMVAHMFNGFSELAMATIKLPVFFKQRDY 642
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVV-------- 112
LF+PSWAY IP+WILKIP+S EV++ VFL+YYVIGFDPNVGR+FKQ+ ++
Sbjct: 643 LFFPSWAYTIPTWILKIPISCFEVAITVFLSYYVIGFDPNVGRLFKQYLLLLLVNQMAAA 702
Query: 113 ---FHESDGFRIVPSHRIPG*KHDCCQYIWVLCTSHISVIGW------LHSLKNWHNATA 163
F + G +V ++ + + SH V W + L+ NA A
Sbjct: 703 LFRFIAALGRTMVVANTLASFALLVLLVLSGFILSHHDVKKWWIWGYWISPLQYAMNAIA 762
Query: 164 ------------------DLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGSR 205
LG + L +RG F A WYWIGVG + + F
Sbjct: 763 VNEFLGHKWNRLVQGTNTTLGIEVLKSRGMFTEAKWYWIGVGALFGYVIVFNILFTIALG 822
Query: 206 CTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRA--DSVTESSHGK------------ 251
+P + + ++ E +G D +S G+
Sbjct: 823 YLKP--------SGKAQQILSEEALKEKHANITGETINDPRNSASSGQTTNTRRNAAPGE 874
Query: 252 ----KKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTAL 307
++GMVLPF P ++ F++I YSVDMP EMK QGV +DRL+LLKGVSG+FRPGVLTAL
Sbjct: 875 ASENRRGMVLPFAPLAVAFNNIRYSVDMPPEMKAQGVDQDRLLLLKGVSGSFRPGVLTAL 934
Query: 308 MGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVH 367
MGVSGAGKTTLMDVLAGRKTGGYI+GDI +SGYPKKQETFAR+SGYCE NDIHSP+VTV+
Sbjct: 935 MGVSGAGKTTLMDVLAGRKTGGYIEGDISISGYPKKQETFARVSGYCEQNDIHSPNVTVY 994
Query: 368 ESLLYSAWLRLPSGVDSNTTKV 389
ESL YSAWLRLPS VDS T K+
Sbjct: 995 ESLAYSAWLRLPSDVDSETRKM 1016
Score = 57.8 bits (138), Expect = 9e-07
Identities = 31/87 (35%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 289 LVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKKQETF 347
L +L V G +P +T L+G G+GKTTL+ LAG+ + G + +GY +
Sbjct: 215 LNILHDVHGVIKPRRMTLLLGPPGSGKTTLLLALAGKLGSDLKVSGKVTYNGYGMDEFVA 274
Query: 348 ARISGYCEHNDIHSPHVTVHESLLYSA 374
R + Y +D+H P +TV E+L +SA
Sbjct: 275 QRSAAYISQHDLHIPEMTVRETLAFSA 301
Score = 43.5 bits (101), Expect = 0.018
Identities = 25/106 (23%), Positives = 56/106 (52%), Gaps = 1/106 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRD 59
+AL+ T+F+R R+ Q D G+++ ++ M + S + +A + V+Y++R
Sbjct: 1246 VALMFGTIFWRLGSKRSRQQDLFNAMGSMYAAVLFMGISYSSSVQPVVAVERTVFYRERA 1305
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y + YA ++++P L++ +++ + Y +IGF+ + F
Sbjct: 1306 AGMYSALPYAFGQVVVELPYVLVQSAVYGVIVYAMIGFEWEAKKFF 1351
>UniRef100_Q8GU92 PDR-like ABC transporter [Oryza sativa]
Length = 1464
Score = 360 bits (925), Expect = 5e-98
Identities = 211/446 (47%), Positives = 268/446 (59%), Gaps = 64/446 (14%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++ IAMT+FFRT+MHR+ D ++ GALFF+++ +MFNG+SE+ +TI KLPV++KQRDL
Sbjct: 554 VSAIAMTVFFRTKMHRDSVTDGVIFMGALFFSVMMIMFNGLSELPLTIFKLPVFFKQRDL 613
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LF+P+W Y IPSWILKIP+S +EV +VF++YYVIGFDP+ GR FKQ
Sbjct: 614 LFFPAWTYTIPSWILKIPMSFIEVGGFVFMSYYVIGFDPSAGRFFKQYLLMLAINQMAAA 673
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF +V +IW +
Sbjct: 674 LFRFVGGAARNMIVANVFGSFMLLIFMVLGGFILVREKVKKW-------WIWGYWISPMM 726
Query: 141 LCTSHISVIGWL-HSLKNWHN---ATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSF 196
+ ISV +L HS N + LG L +RG FP A WYWIG G + + F
Sbjct: 727 YAQNAISVNEFLGHSWDKVLNNSLSNETLGVQALRSRGVFPEAKWYWIGFGALLGFIMLF 786
Query: 197 QRGVWCGSRCTRPI**A*---CNNN*RFRR*FIYRTRSEFTTYCSGR----------ADS 243
+P + + ++ I + T S +
Sbjct: 787 NGLFTLALTYLKPYGKSQPSVSEEELKEKQANINGNVLDVDTMASSTNLAIVDNTETSSE 846
Query: 244 VTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGV 303
+ ++S ++GMVLPF P S+TFD+I YSVDMP EMK G+ EDRL LLKGVSG+FRPGV
Sbjct: 847 IADNSQPTQRGMVLPFAPLSLTFDNIKYSVDMPQEMKAHGIVEDRLELLKGVSGSFRPGV 906
Query: 304 LTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPH 363
LTALMGVSGAGKTTLMDVLAGRKTGGYI+G+I +SGYPKKQETFAR+SGYCE NDIHSP
Sbjct: 907 LTALMGVSGAGKTTLMDVLAGRKTGGYIEGNITISGYPKKQETFARVSGYCEQNDIHSPQ 966
Query: 364 VTVHESLLYSAWLRLPSGVDSNTTKV 389
VTV ESLL+SAWLRLP VDSNT K+
Sbjct: 967 VTVSESLLFSAWLRLPKDVDSNTRKM 992
Score = 53.1 bits (126), Expect = 2e-05
Identities = 30/91 (32%), Positives = 49/91 (52%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKK 343
++ + +L VSG +P +T L+G G+GKTTL+ LAGR G + +G+ +
Sbjct: 182 KKQTMPILHDVSGIVKPRRMTLLLGPPGSGKTTLLLALAGRLGKDIKFSGQVTYNGHQME 241
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
R + Y +D+H +TV E+L +SA
Sbjct: 242 DFVPQRTAAYISQHDLHIGEMTVRETLSFSA 272
Score = 35.4 bits (80), Expect = 4.8
Identities = 30/154 (19%), Positives = 63/154 (40%), Gaps = 8/154 (5%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTM-MFNGMSEISMTIAKLPVYYKQRD 59
+AL+ T+F+ D G+++ ++ + + N S + + V+Y++R
Sbjct: 1222 IALLFGTIFWDLGGKTGKSQDLFNAMGSMYSAVLFIGVLNSQSVQPVVSVERTVFYRERA 1281
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR-------MFKQFRCVV 112
Y ++ YA ++ P +L++ ++ + Y +IGF + MF F
Sbjct: 1282 AGMYSAFPYAFGQVAIEFPYTLVQSIIYGIIVYSMIGFKWTAAKFFWYLFFMFFTFLYFT 1341
Query: 113 FHESDGFRIVPSHRIPG*KHDCCQYIWVLCTSHI 146
F+ + PS+ + IW L + I
Sbjct: 1342 FYGMMAVGLTPSYHVASIVSSAFYGIWNLFSGFI 1375
>UniRef100_Q8GU88 PDR-like ABC transporter [Oryza sativa]
Length = 1444
Score = 347 bits (889), Expect = 7e-94
Identities = 207/442 (46%), Positives = 263/442 (58%), Gaps = 63/442 (14%)
Query: 4 IAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFY 63
I MT+F RT+MHR +D ++ GA+F LVT +FNG +E++M+IAKLP++YKQRDLLFY
Sbjct: 537 IGMTVFLRTKMHRRSVEDGAIFLGAMFLGLVTHLFNGFAELAMSIAKLPIFYKQRDLLFY 596
Query: 64 PSWAYAIPSWILKIPVSLMEVSLWVFLT---------------YYVI------------- 95
PSWAYA+P+W+LKIP+S +E ++W+ +T +YV+
Sbjct: 597 PSWAYALPTWVLKIPISFLECAVWICMTYYVMGFDPNIERFFRHYVLLVLISQMASGLFR 656
Query: 96 -----GFDPNVGRMFKQFRCVVFHESDGFRIVPSHRIPG*KHDCCQYIWVLCTSHI---- 146
G + V F F ++ GF ++ I +IW +S +
Sbjct: 657 LLAALGREMVVADTFGSFAQLILLVLGGF-LISRENIKK------WWIWGYWSSPLMYAQ 709
Query: 147 SVIGWLHSLKNWHNATAD-------LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRG 199
+ I L + N D LG L RG F A WYWIGVG + + F
Sbjct: 710 NAIAVNEFLGHSWNKVVDPTQSNDTLGVQVLKVRGIFVDANWYWIGVGALLGYIMLFNIL 769
Query: 200 VWCGSRCTRPI**A*CN-NN*RFRR*FIYRTRS--EFTTYCSGRADSVTESSHGK----- 251
P+ + R + RT E T + +S ++++ G+
Sbjct: 770 FILFLEWLDPLGKGQAVVSEEELREKHVNRTGENVELLTLGTDSQNSPSDANAGRGEITG 829
Query: 252 ----KKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTAL 307
K+GMVLPF P SITFD+I YSVDMP EMK++GV EDRL+LLKGVSGAFRPGVLTAL
Sbjct: 830 ADTRKRGMVLPFTPLSITFDNIRYSVDMPQEMKDKGVTEDRLLLLKGVSGAFRPGVLTAL 889
Query: 308 MGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVH 367
MGVSGAGKTTLMDVLAGRKTGGYI+GDI +SGYPKKQETFARI+GYCE NDIHSPHVTV+
Sbjct: 890 MGVSGAGKTTLMDVLAGRKTGGYIEGDISISGYPKKQETFARIAGYCEQNDIHSPHVTVY 949
Query: 368 ESLLYSAWLRLPSGVDSNTTKV 389
ESLLYSAWLRLPS VDS K+
Sbjct: 950 ESLLYSAWLRLPSEVDSEARKM 971
Score = 54.3 bits (129), Expect = 1e-05
Identities = 29/85 (34%), Positives = 48/85 (56%), Gaps = 1/85 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKKQETFAR 349
+L +SG RPG ++ L+G G+GKT+L+ LAG+ + G + +G+ + R
Sbjct: 169 ILHDISGIIRPGRMSLLLGPPGSGKTSLLLALAGKLDSTLKVSGRVTYNGHDMDEFVPQR 228
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSA 374
S Y +D+H +TV E+L +SA
Sbjct: 229 TSAYIGQHDLHIGEMTVRETLAFSA 253
Score = 37.7 bits (86), Expect = 0.96
Identities = 27/110 (24%), Positives = 50/110 (44%), Gaps = 9/110 (8%)
Query: 1 MALIAMTLFFRTEMHRNDQDD-----AGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYY 55
+ALI T+F N + D +YA LF + NG + + + V+Y
Sbjct: 1201 IALIFGTIFLNLGKKINKRLDLFNSLGSMYAAVLFIGIQ----NGQTVQPIVDVERTVFY 1256
Query: 56 KQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
+++ Y + YA +++IP ++ ++ + Y +IGFD V + F
Sbjct: 1257 REKAAGMYSALPYAFAQVLIEIPHIFLQTVVYGLIVYSLIGFDWTVEKFF 1306
>UniRef100_Q7PC87 PDR6 ABC transporter [Arabidopsis thaliana]
Length = 1453
Score = 345 bits (885), Expect = 2e-93
Identities = 193/432 (44%), Positives = 254/432 (58%), Gaps = 51/432 (11%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
M+LIAMT++FRTEMH D + GALFF+L+ +MFNGM+E++ T+ +LPV++KQRD
Sbjct: 554 MSLIAMTVYFRTEMHVGTVQDGQKFYGALFFSLINLMFNGMAELAFTVMRLPVFFKQRDF 613
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF------------ 108
LFYP WA+A+P ++LKIP+SL+E +W+ LTYY IGF P+ R F+Q
Sbjct: 614 LFYPPWAFALPGFLLKIPLSLIESVIWIALTYYTIGFAPSAARFFRQLLAYFCVNQMALS 673
Query: 109 ---------RCVVFHESDGFR-----------IVPSHRIPG*KHDCCQYIWVLCTSHISV 148
R V S G I+ IP C Y + ++
Sbjct: 674 LFRFLGALGRTEVIANSGGTLALLVVFVLGGFIISKDDIPSWL-TWCYYTSPMMYGQTAL 732
Query: 149 IGWLHSLKNWHNATAD-------LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVW 201
+ + W + D +G+ L +RGFF YW+WI +G + + F
Sbjct: 733 VINEFLDERWGSPNNDTRINAKTVGEVLLKSRGFFTEPYWFWICIGALLGFTVLFNFCYI 792
Query: 202 CGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTE----SSHGKKKGMVL 257
P+ + + + + SG SV E SSHG KKGMVL
Sbjct: 793 IALMYLNPLGNSKATT-------VVEEGKDKHKGSHSGTGGSVVELTSTSSHGPKKGMVL 845
Query: 258 PFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTT 317
PF+P S+ F+++ Y VDMP EMK QGV DRL LL+ V GAFRPGVLTAL+GVSGAGKTT
Sbjct: 846 PFQPLSLAFNNVNYYVDMPAEMKAQGVEGDRLQLLRDVGGAFRPGVLTALVGVSGAGKTT 905
Query: 318 LMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLR 377
LMDVLAGRKTGGY++G I +SGYPK Q TFAR+SGYCE NDIHSPHVTV+ESL+YSAWLR
Sbjct: 906 LMDVLAGRKTGGYVEGSINISGYPKNQATFARVSGYCEQNDIHSPHVTVYESLIYSAWLR 965
Query: 378 LPSGVDSNTTKV 389
L + +D+ T ++
Sbjct: 966 LSADIDTKTREM 977
Score = 51.6 bits (122), Expect = 6e-05
Identities = 28/90 (31%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGG-YIDGDIKVSGYPKK 343
++ ++ +LK +SG +P +T L+G +GKTTL+ LAG+ + G I G+ +
Sbjct: 182 KKRKIEILKDISGIIKPSRMTLLLGPPSSGKTTLLQALAGKLDDTLQMSGRITYCGHEFR 241
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYS 373
+ + Y +D+H +TV ESL +S
Sbjct: 242 EFVPQKTCAYISQHDLHFGEMTVRESLDFS 271
Score = 39.3 bits (90), Expect = 0.33
Identities = 27/130 (20%), Positives = 61/130 (46%), Gaps = 8/130 (6%)
Query: 8 LFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRDLLFYPSW 66
LF++T + D + GA++ ++ + + + +A + V+Y+++ Y +
Sbjct: 1214 LFWQTGTKIEKEQDLNNFFGAMYAAVLFLGATNAATVQPAVAIERTVFYREKAAGMYSAI 1273
Query: 67 AYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF----KQFRCVVFHESDGFRIV 122
YAI ++I + ++ ++ + Y +IG+D V + F C V+ G +V
Sbjct: 1274 PYAISQVAVEIMYNTIQTGVYTLILYSMIGYDWTVVKFFWFYYYMLTCFVYFTLYGMMLV 1333
Query: 123 ---PSHRIPG 129
P+++I G
Sbjct: 1334 ALTPNYQIAG 1343
>UniRef100_Q9SJR6 Putative ABC transporter [Arabidopsis thaliana]
Length = 1450
Score = 343 bits (879), Expect = 1e-92
Identities = 191/429 (44%), Positives = 253/429 (58%), Gaps = 48/429 (11%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
M+LIAMT++FRTEMH D + GALFF+L+ +MFNGM+E++ T+ +LPV++KQRD
Sbjct: 554 MSLIAMTVYFRTEMHVGTVQDGQKFYGALFFSLINLMFNGMAELAFTVMRLPVFFKQRDF 613
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF------------ 108
LFYP WA+A+P ++LKIP+SL+E +W+ LTYY IGF P+ R F+Q
Sbjct: 614 LFYPPWAFALPGFLLKIPLSLIESVIWIALTYYTIGFAPSAARFFRQLLAYFCVNQMALS 673
Query: 109 ---------RCVVFHESDGFR-----------IVPSHRIPG*KHDCCQYIWVLCTSHISV 148
R V S G I+ IP C Y + ++
Sbjct: 674 LFRFLGALGRTEVIANSGGTLALLVVFVLGGFIISKDDIPSWL-TWCYYTSPMMYGQTAL 732
Query: 149 IGWLHSLKNWHNATAD-------LGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVW 201
+ + W + D +G+ L +RGFF YW+WI +G + + F
Sbjct: 733 VINEFLDERWGSPNNDTRINAKTVGEVLLKSRGFFTEPYWFWICIGALLGFTVLFNFCYI 792
Query: 202 CGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTE-SSHGKKKGMVLPFE 260
P+ + + + + SG +T SSHG KKGMVLPF+
Sbjct: 793 IALMYLNPLGNSKATT-------VVEEGKDKHKGSHSGTGVELTSTSSHGPKKGMVLPFQ 845
Query: 261 PHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMD 320
P S+ F+++ Y VDMP EMK QGV DRL LL+ V GAFRPGVLTAL+GVSGAGKTTLMD
Sbjct: 846 PLSLAFNNVNYYVDMPAEMKAQGVEGDRLQLLRDVGGAFRPGVLTALVGVSGAGKTTLMD 905
Query: 321 VLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPS 380
VLAGRKTGGY++G I +SGYPK Q TFAR+SGYCE NDIHSPHVTV+ESL+YSAWLRL +
Sbjct: 906 VLAGRKTGGYVEGSINISGYPKNQATFARVSGYCEQNDIHSPHVTVYESLIYSAWLRLSA 965
Query: 381 GVDSNTTKV 389
+D+ T ++
Sbjct: 966 DIDTKTREM 974
Score = 51.6 bits (122), Expect = 6e-05
Identities = 28/90 (31%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGG-YIDGDIKVSGYPKK 343
++ ++ +LK +SG +P +T L+G +GKTTL+ LAG+ + G I G+ +
Sbjct: 182 KKRKIEILKDISGIIKPSRMTLLLGPPSSGKTTLLQALAGKLDDTLQMSGRITYCGHEFR 241
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYS 373
+ + Y +D+H +TV ESL +S
Sbjct: 242 EFVPQKTCAYISQHDLHFGEMTVRESLDFS 271
Score = 39.3 bits (90), Expect = 0.33
Identities = 27/130 (20%), Positives = 61/130 (46%), Gaps = 8/130 (6%)
Query: 8 LFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIA-KLPVYYKQRDLLFYPSW 66
LF++T + D + GA++ ++ + + + +A + V+Y+++ Y +
Sbjct: 1211 LFWQTGTKIEKEQDLNNFFGAMYAAVLFLGATNAATVQPAVAIERTVFYREKAAGMYSAI 1270
Query: 67 AYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF----KQFRCVVFHESDGFRIV 122
YAI ++I + ++ ++ + Y +IG+D V + F C V+ G +V
Sbjct: 1271 PYAISQVAVEIMYNTIQTGVYTLILYSMIGYDWTVVKFFWFYYYMLTCFVYFTLYGMMLV 1330
Query: 123 ---PSHRIPG 129
P+++I G
Sbjct: 1331 ALTPNYQIAG 1340
>UniRef100_Q7PC85 PDR10 ABC transporter [Arabidopsis thaliana]
Length = 1418
Score = 292 bits (747), Expect = 2e-77
Identities = 167/426 (39%), Positives = 242/426 (56%), Gaps = 45/426 (10%)
Query: 2 ALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLL 61
A++ +F++ + + + +D +Y GA++ + ++F+G E+ MTI KLPV+YKQR
Sbjct: 528 AILIGVVFWQQKNYPSTVEDGIIYMGAIYLEVQMIVFSGFFELPMTIDKLPVFYKQRHFS 587
Query: 62 FYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHESDGFRI 121
FYPSWA+++P+ I+ P+S +EV + V +TY+ IG+D V K + + + +
Sbjct: 588 FYPSWAFSLPTSIITFPLSFVEVFIVVLITYFTIGYDLTVPSFLKHYLVLALCGQMSYGL 647
Query: 122 ------VPSHRIPG*KHDCCQYIWVLCTS-HISVIGWLHSLKNWHNATA----------- 163
V + + C +W++ S ++ +H W T+
Sbjct: 648 FRCIAAVTRNHVVSNTMGCLAVMWLMTFSGYVLSRNQVHKWLTWAYWTSPMMYIQTAVSV 707
Query: 164 ----------DLGKDYLDTRGFFPHAYWYWIGV----------GGIGWICLSFQRGVWCG 203
LG L +RGFF YWYWIG+ I +CL+F +
Sbjct: 708 NEFRSESWKDGLGVAVLKSRGFFVETYWYWIGLLALILSTILSNIITSLCLAFLKQYGIS 767
Query: 204 SRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMVLPFEPHS 263
P ++N R + T F D V + K + +PF+P
Sbjct: 768 KTAVLPDEREEADSNNTTGRDYTGTTMERF-------FDRVVTTRTCNDKKLRIPFKPLY 820
Query: 264 ITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLA 323
+TF++I YSVD P EMKE+G+RE++LVLL G+SGAFRPGVLTALMGVSGAGKTTLMDVLA
Sbjct: 821 MTFENITYSVDTPKEMKEKGIRENKLVLLNGLSGAFRPGVLTALMGVSGAGKTTLMDVLA 880
Query: 324 GRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVD 383
GRK GYI G+I VSG+PKKQ++FAR+SGYCE +DIHSP +TV+ESLLYSAWLRLP +D
Sbjct: 881 GRKNTGYIQGEIYVSGFPKKQDSFARVSGYCEQSDIHSPLLTVYESLLYSAWLRLPPDID 940
Query: 384 SNTTKV 389
++T ++
Sbjct: 941 THTREL 946
Score = 67.0 bits (162), Expect = 1e-09
Identities = 35/91 (38%), Positives = 56/91 (61%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR-KTGGYIDGDIKVSGYPKK 343
R+ R+ +L VSG +PG LT L+G G+GK+TL+ L+G+ +TG G + +G+
Sbjct: 155 RKKRISILNDVSGIIKPGRLTLLLGPPGSGKSTLLKALSGKTETGLRSTGKVTYNGHELH 214
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R +GY + D+H P +TV E+L +SA
Sbjct: 215 EFVPERTAGYIDQYDVHLPDLTVRETLKFSA 245
>UniRef100_Q9LHK8 ABC transporter-like protein [Arabidopsis thaliana]
Length = 1405
Score = 289 bits (740), Expect = 1e-76
Identities = 167/433 (38%), Positives = 242/433 (55%), Gaps = 52/433 (12%)
Query: 2 ALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLL 61
A++ +F++ + + + +D +Y GA++ + ++F+G E+ MTI KLPV+YKQR
Sbjct: 528 AILIGVVFWQQKNYPSTVEDGIIYMGAIYLEVQMIVFSGFFELPMTIDKLPVFYKQRHFS 587
Query: 62 FYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHESDGFRI 121
FYPSWA+++P+ I+ P+S +EV + V +TY+ IG+D V K + + + +
Sbjct: 588 FYPSWAFSLPTSIITFPLSFVEVFIVVLITYFTIGYDLTVPSFLKHYLVLALCGQMSYGL 647
Query: 122 ------VPSHRIPG*KHDCCQYIWVLCTS-HISVIGWLHSLKNWHNATAD---------- 164
V + + C +W++ S ++ +H W T+
Sbjct: 648 FRCIAAVTRNHVVSNTMGCLAVMWLMTFSGYVLSRNQVHKWLTWAYWTSPMMYIQTAVSV 707
Query: 165 ------------------LGKDYLDTRGFFPHAYWYWIGVGG----------IGWICLSF 196
LG L +RGFF YWYWIG+ I +CL+F
Sbjct: 708 NEFRSESWKDVISKKPQGLGVAVLKSRGFFVETYWYWIGLLALILSTILSNIITSLCLAF 767
Query: 197 QRGVWCGSRCTRPI**A*CNNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMV 256
+ P ++N R + T F D V + K +
Sbjct: 768 LKQYGISKTAVLPDEREEADSNNTTGRDYTGTTMERFF-------DRVVTTRTCNDKKLR 820
Query: 257 LPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKT 316
+PF+P +TF++I YSVD P EMKE+G+RE++LVLL G+SGAFRPGVLTALMGVSGAGKT
Sbjct: 821 IPFKPLYMTFENITYSVDTPKEMKEKGIRENKLVLLNGLSGAFRPGVLTALMGVSGAGKT 880
Query: 317 TLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWL 376
TLMDVLAGRK GYI G+I VSG+PKKQ++FAR+SGYCE +DIHSP +TV+ESLLYSAWL
Sbjct: 881 TLMDVLAGRKNTGYIQGEIYVSGFPKKQDSFARVSGYCEQSDIHSPLLTVYESLLYSAWL 940
Query: 377 RLPSGVDSNTTKV 389
RLP +D++T ++
Sbjct: 941 RLPPDIDTHTREL 953
Score = 67.0 bits (162), Expect = 1e-09
Identities = 35/91 (38%), Positives = 56/91 (61%), Gaps = 1/91 (1%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR-KTGGYIDGDIKVSGYPKK 343
R+ R+ +L VSG +PG LT L+G G+GK+TL+ L+G+ +TG G + +G+
Sbjct: 155 RKKRISILNDVSGIIKPGRLTLLLGPPGSGKSTLLKALSGKTETGLRSTGKVTYNGHELH 214
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSA 374
+ R +GY + D+H P +TV E+L +SA
Sbjct: 215 EFVPERTAGYIDQYDVHLPDLTVRETLKFSA 245
>UniRef100_Q8GU91 PDR-like ABC transporter [Oryza sativa]
Length = 1479
Score = 233 bits (595), Expect = 9e-60
Identities = 113/152 (74%), Positives = 128/152 (83%)
Query: 238 SGRADSVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
+G + ++S ++GMVLPF P S+TF+DI YSVDMP EMK G+ EDRL LLKGVSG
Sbjct: 845 TGTGSEIADNSQPTQRGMVLPFTPLSLTFEDIKYSVDMPQEMKAHGIVEDRLELLKGVSG 904
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHN 357
FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI+G+I +SGYPKKQETFAR+SGYCE N
Sbjct: 905 CFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGNISISGYPKKQETFARVSGYCEQN 964
Query: 358 DIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
DIHSP VTV ESLL+SAWLRLP VDSNT K+
Sbjct: 965 DIHSPQVTVSESLLFSAWLRLPKDVDSNTRKM 996
Score = 153 bits (386), Expect = 2e-35
Identities = 94/239 (39%), Positives = 131/239 (54%), Gaps = 52/239 (21%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++ +AMT+FFRT+MHR+ D ++ GALFF ++ +M NG+SE+ +TI KLPV++KQRDL
Sbjct: 558 VSAMAMTVFFRTKMHRDSVADGVIFMGALFFAVMMIMLNGLSELPLTIFKLPVFFKQRDL 617
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LF+P+W Y IPSWILK P+S +EV + F++YYVIGFDPNVGR FKQ
Sbjct: 618 LFFPAWTYTIPSWILKSPMSFIEVGGFCFMSYYVIGFDPNVGRFFKQYLLMLAVSQMAAA 677
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF I+ ++ +IW +
Sbjct: 678 LFRFVGGAARNLIVANVFGSFMLLIFMVLGGF-ILARDKVNK------WWIWGYWISPMM 730
Query: 141 LCTSHISVIGWL-HSLKNWHN---ATADLGKDYLDTRGFFPHAYWYWIGVGG-IGWICL 194
+ +SV +L HS N + LG L +RG FP A WYWIG G +G+I L
Sbjct: 731 YAQNAVSVNEFLGHSWDKVLNNSLSNETLGVQALMSRGIFPEAKWYWIGFGALLGFIML 789
Score = 57.0 bits (136), Expect = 2e-06
Identities = 30/90 (33%), Positives = 48/90 (53%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQ 344
R+ L +L +SG +P +T L+G G+GKTT + LAGR G + +G+ +
Sbjct: 187 RKQTLRILHDISGIIKPKRMTLLLGPPGSGKTTFLLALAGRLKDLKFSGQVTYNGHQMED 246
Query: 345 ETFARISGYCEHNDIHSPHVTVHESLLYSA 374
R + Y +D+H +TV E+L +SA
Sbjct: 247 FVPQRTAAYISQHDLHIGEMTVRETLSFSA 276
Score = 39.7 bits (91), Expect = 0.25
Identities = 35/158 (22%), Positives = 62/158 (39%), Gaps = 16/158 (10%)
Query: 1 MALIAMTLFFRTEMHRNDQDD-----AGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYY 55
+ALI T+F+ D +YA LF ++ NG S + + V+Y
Sbjct: 1226 IALIFGTIFWDLGGKMGQSQDLFNAMGSMYAAVLFIGVL----NGQSVQPVVSVERTVFY 1281
Query: 56 KQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR-------MFKQF 108
++R Y + YA ++ P +L++ ++ + Y +IGF V + MF
Sbjct: 1282 RERAAGMYSALPYAFGQVAIEFPYTLVQSVIYSIIVYSMIGFQWTVAKFFWYLFFMFFTL 1341
Query: 109 RCVVFHESDGFRIVPSHRIPG*KHDCCQYIWVLCTSHI 146
F+ + PS+ + IW L T +
Sbjct: 1342 LYFTFYGMMAVGLTPSYHVASIVSSAFYAIWNLFTGFV 1379
>UniRef100_Q7PC80 PDR1 ABC transporter [Oryza sativa]
Length = 1468
Score = 233 bits (595), Expect = 9e-60
Identities = 113/152 (74%), Positives = 128/152 (83%)
Query: 238 SGRADSVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
+G + ++S ++GMVLPF P S+TF+DI YSVDMP EMK G+ EDRL LLKGVSG
Sbjct: 845 TGTGSEIADNSQPTQRGMVLPFTPLSLTFEDIKYSVDMPQEMKAHGIVEDRLELLKGVSG 904
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHN 357
FRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI+G+I +SGYPKKQETFAR+SGYCE N
Sbjct: 905 CFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGNISISGYPKKQETFARVSGYCEQN 964
Query: 358 DIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
DIHSP VTV ESLL+SAWLRLP VDSNT K+
Sbjct: 965 DIHSPQVTVSESLLFSAWLRLPKDVDSNTRKM 996
Score = 153 bits (386), Expect = 2e-35
Identities = 94/239 (39%), Positives = 131/239 (54%), Gaps = 52/239 (21%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
++ +AMT+FFRT+MHR+ D ++ GALFF ++ +M NG+SE+ +TI KLPV++KQRDL
Sbjct: 558 VSAMAMTVFFRTKMHRDSVADGVIFMGALFFAVMMIMLNGLSELPLTIFKLPVFFKQRDL 617
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQ------------- 107
LF+P+W Y IPSWILK P+S +EV + F++YYVIGFDPNVGR FKQ
Sbjct: 618 LFFPAWTYTIPSWILKSPMSFIEVGGFCFMSYYVIGFDPNVGRFFKQYLLMLAVSQMAAA 677
Query: 108 --------------------FRCVVFHESDGFRIVPSHRIPG*KHDCCQYIW-------V 140
F ++F GF I+ ++ +IW +
Sbjct: 678 LFRFVGGAARNLIVANVFGSFMLLIFMVLGGF-ILARDKVNK------WWIWGYWISPMM 730
Query: 141 LCTSHISVIGWL-HSLKNWHN---ATADLGKDYLDTRGFFPHAYWYWIGVGG-IGWICL 194
+ +SV +L HS N + LG L +RG FP A WYWIG G +G+I L
Sbjct: 731 YAQNAVSVNEFLGHSWDKVLNNSLSNETLGVQALMSRGIFPEAKWYWIGFGALLGFIML 789
Score = 57.0 bits (136), Expect = 2e-06
Identities = 30/90 (33%), Positives = 48/90 (53%)
Query: 285 REDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQ 344
R+ L +L +SG +P +T L+G G+GKTT + LAGR G + +G+ +
Sbjct: 187 RKQTLRILHDISGIIKPKRMTLLLGPPGSGKTTFLLALAGRLKDLKFSGQVTYNGHQMED 246
Query: 345 ETFARISGYCEHNDIHSPHVTVHESLLYSA 374
R + Y +D+H +TV E+L +SA
Sbjct: 247 FVPQRTAAYISQHDLHIGEMTVRETLSFSA 276
Score = 39.7 bits (91), Expect = 0.25
Identities = 35/158 (22%), Positives = 62/158 (39%), Gaps = 16/158 (10%)
Query: 1 MALIAMTLFFRTEMHRNDQDD-----AGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYY 55
+ALI T+F+ D +YA LF ++ NG S + + V+Y
Sbjct: 1226 IALIFGTIFWDLGGKMGQSQDLFNAMGSMYAAVLFIGVL----NGQSVQPVVSVERTVFY 1281
Query: 56 KQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGR-------MFKQF 108
++R Y + YA ++ P +L++ ++ + Y +IGF V + MF
Sbjct: 1282 RERAAGMYSALPYAFGQVAIEFPYTLVQSVIYSIIVYSMIGFQWTVAKFFWYLFFMFFTL 1341
Query: 109 RCVVFHESDGFRIVPSHRIPG*KHDCCQYIWVLCTSHI 146
F+ + PS+ + IW L T +
Sbjct: 1342 LYFTFYGMMAVGLTPSYHVASIVSSAFYAIWNLFTGFV 1379
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.331 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,031,656,561
Number of Sequences: 2790947
Number of extensions: 48708493
Number of successful extensions: 241950
Number of sequences better than 10.0: 5464
Number of HSP's better than 10.0 without gapping: 2266
Number of HSP's successfully gapped in prelim test: 3321
Number of HSP's that attempted gapping in prelim test: 222412
Number of HSP's gapped (non-prelim): 12150
length of query: 596
length of database: 848,049,833
effective HSP length: 133
effective length of query: 463
effective length of database: 476,853,882
effective search space: 220783347366
effective search space used: 220783347366
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC126018.1


