

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
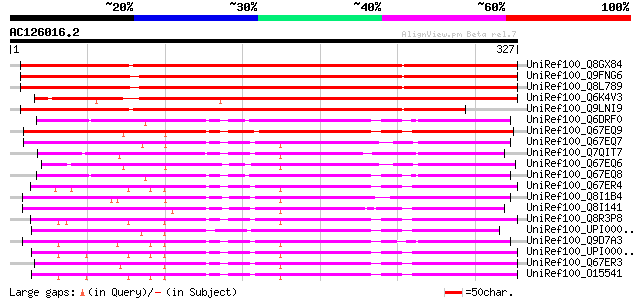
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GX84 Hypothetical protein At1g01350/F6F3_27 [Arabido... 476 e-133
UniRef100_Q9FNG6 Similarity to zinc finger protein [Arabidopsis ... 473 e-132
UniRef100_Q8L789 Hypothetical protein At5g06420 [Arabidopsis tha... 470 e-131
UniRef100_Q6K4V3 Putative zinc finger protein [Oryza sativa] 422 e-117
UniRef100_Q9LNI9 Putative zinc finger protein [Arabidopsis thali... 416 e-115
UniRef100_Q6DRF0 Zinc finger protein 183-like 1 [Brachydanio rerio] 248 2e-64
UniRef100_Q67EQ9 Zinc finger protein 183 [Oncorhynchus mykiss] 243 4e-63
UniRef100_Q67EQ7 Zinc finger protein 183 [Oryzias latipes] 234 2e-60
UniRef100_Q7QIT7 ENSANGP00000021479 [Anopheles gambiae str. PEST] 234 2e-60
UniRef100_Q67EQ6 Zinc finger protein 183 [Ciona intestinalis] 233 4e-60
UniRef100_Q67EQ8 Zinc finger protein 183 [Brachydanio rerio] 231 2e-59
UniRef100_Q67ER4 Zinc finger protein 183 [Bos taurus] 231 2e-59
UniRef100_Q8I1B4 CG4973-PA [Drosophila pseudoobscura] 231 3e-59
UniRef100_Q8I141 CG4973-PA [Drosophila willistoni] 231 3e-59
UniRef100_Q8R3P8 RIKEN cDNA 2810428C21 [Mus musculus] 230 5e-59
UniRef100_UPI000036445E UPI000036445E UniRef100 entry 229 6e-59
UniRef100_Q9D7A3 Mus musculus adult male tongue cDNA, RIKEN full... 229 6e-59
UniRef100_UPI000036F7CE UPI000036F7CE UniRef100 entry 228 2e-58
UniRef100_Q67ER3 Zinc finger protein 183 [Gallus gallus] 228 2e-58
UniRef100_O15541 Zinc finger protein 183 [Homo sapiens] 228 2e-58
>UniRef100_Q8GX84 Hypothetical protein At1g01350/F6F3_27 [Arabidopsis thaliana]
Length = 343
Score = 476 bits (1225), Expect = e-133
Identities = 223/320 (69%), Positives = 266/320 (82%), Gaps = 3/320 (0%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
+ + Q++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+KL+FS
Sbjct: 20 AAQETQSQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSKLYFS 79
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G SKSS + + E+ FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 80 SGPSKSSTTT--SGAPERSVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 137
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A E+LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKAGGSHGPLRASAHIRVSA
Sbjct: 138 KADEALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAGGSHGPLRASAHIRVSA 197
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYKSGWQ+EKEW+EAEK RK A G +
Sbjct: 198 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKSGWQIEKEWEEAEKVRKRNKAMGVED 257
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
E++ A D D+DE+ALPFACFICR PFVDPV TKCKHYFCEHCALK H KNKKCFVCNQ
Sbjct: 258 EDDEAD-KDSDEDENALPFACFICREPFVDPVVTKCKHYFCEHCALKRHTKNKKCFVCNQ 316
Query: 308 PTLGIFNVAHEIRKKMAEDK 327
PT+GIFN AHEI+K+MAE++
Sbjct: 317 PTMGIFNAAHEIKKRMAEER 336
>UniRef100_Q9FNG6 Similarity to zinc finger protein [Arabidopsis thaliana]
Length = 378
Score = 473 bits (1217), Expect = e-132
Identities = 219/320 (68%), Positives = 265/320 (82%), Gaps = 6/320 (1%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
++ NQ+++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+ L+FS
Sbjct: 58 NSNNQESQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSNLYFS 117
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G S ++ A E+P FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 118 SGPSTRTSGAP-----ERPVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 172
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A +LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKAGGSHGPLRASAHIRVSA
Sbjct: 173 KADHALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAGGSHGPLRASAHIRVSA 232
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYK GWQ+EKEW+EAEK RK A G +
Sbjct: 233 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKPGWQIEKEWEEAEKVRKRNKAMGVED 292
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
+++ A D D+DE+ALPFACFICR PF+DPV TKCKHYFCEHCALKHH KNKKCFVCNQ
Sbjct: 293 DDDEAD-KDSDEDENALPFACFICREPFLDPVVTKCKHYFCEHCALKHHTKNKKCFVCNQ 351
Query: 308 PTLGIFNVAHEIRKKMAEDK 327
PT+GIFN AHEI+K+MAE++
Sbjct: 352 PTMGIFNAAHEIKKRMAEER 371
>UniRef100_Q8L789 Hypothetical protein At5g06420 [Arabidopsis thaliana]
Length = 378
Score = 470 bits (1210), Expect = e-131
Identities = 218/320 (68%), Positives = 265/320 (82%), Gaps = 6/320 (1%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
++ NQ+++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+ L+FS
Sbjct: 58 NSNNQESQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSNLYFS 117
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G S ++ A E+P FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 118 SGPSTRTSGAP-----ERPVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 172
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A +LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKA GSHGPLRASAHIRVSA
Sbjct: 173 KADHALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAEGSHGPLRASAHIRVSA 232
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYK GWQ+EKEW+EAEK RK A G +
Sbjct: 233 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKPGWQIEKEWEEAEKVRKRNKAMGVED 292
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
+++ A D D+DE+ALPFACFICR+PF+DPV TKCKHYFCEHCALKHH KNKKCFVCNQ
Sbjct: 293 DDDEAD-KDSDEDENALPFACFICRDPFLDPVVTKCKHYFCEHCALKHHTKNKKCFVCNQ 351
Query: 308 PTLGIFNVAHEIRKKMAEDK 327
PT+GIFN AHEI+K+MAE++
Sbjct: 352 PTMGIFNAAHEIKKRMAEER 371
>UniRef100_Q6K4V3 Putative zinc finger protein [Oryza sativa]
Length = 326
Score = 422 bits (1086), Expect = e-117
Identities = 210/319 (65%), Positives = 241/319 (74%), Gaps = 20/319 (6%)
Query: 17 VCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQ--KKNTKADNKLFFSTGSSKSS 74
VCSF RKP KN+RKR +++D + + KK + +KLFFS+ S
Sbjct: 18 VCSFVRKPP--KNIRKRPTAPAGSDDDDEDGSGAIAAARAKKAPSSTSKLFFSSADGSS- 74
Query: 75 ASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQATESLK 134
E F +ESS+ IQ DS+ATATLETET+F RDARAIRER LKQA ESLK
Sbjct: 75 ---------EPRRFQYESSRTIQASTDSRATATLETETEFDRDARAIRERQLKQAEESLK 125
Query: 135 ------GKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSAR 188
S+ S ++YKGI+ YTD+KAGFRRE T++SEKAGGSHGPLRASAHIR+SAR
Sbjct: 126 KNPSAPASSSGSGSGEVYKGIHGYTDYKAGFRREHTVSSEKAGGSHGPLRASAHIRLSAR 185
Query: 189 FDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAE 248
FDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQ+EKEW+EAEKARK R+A G D
Sbjct: 186 FDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQIEKEWEEAEKARKRRIAMGGDGS 245
Query: 249 EEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQP 308
+ A D+DDDE+ALPFAC+ICR PFVDPV TKCKHYFCEHCALKHH+KNKKCFVCN+P
Sbjct: 246 DYEAGEEDDDDDEEALPFACYICREPFVDPVVTKCKHYFCEHCALKHHSKNKKCFVCNKP 305
Query: 309 TLGIFNVAHEIRKKMAEDK 327
TLGIFN A EIRKKMA+DK
Sbjct: 306 TLGIFNAAQEIRKKMAQDK 324
>UniRef100_Q9LNI9 Putative zinc finger protein [Arabidopsis thaliana]
Length = 304
Score = 416 bits (1068), Expect = e-115
Identities = 197/287 (68%), Positives = 235/287 (81%), Gaps = 3/287 (1%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
+ + Q++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+KL+FS
Sbjct: 20 AAQETQSQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSKLYFS 79
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G SKSS + + E+ FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 80 SGPSKSSTTT--SGAPERSVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 137
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A E+LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKAGGSHGPLRASAHIRVSA
Sbjct: 138 KADEALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAGGSHGPLRASAHIRVSA 197
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYK GWQ+EKEW+EAEK RK A G +
Sbjct: 198 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKPGWQIEKEWEEAEKVRKRNKAMGVED 257
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK 294
E++ A D D+DE+ALPFACFICR PFVDPV TKCKHYFCEHCALK
Sbjct: 258 EDDEAD-KDSDEDENALPFACFICREPFVDPVVTKCKHYFCEHCALK 303
>UniRef100_Q6DRF0 Zinc finger protein 183-like 1 [Brachydanio rerio]
Length = 321
Score = 248 bits (633), Expect = 2e-64
Identities = 140/314 (44%), Positives = 189/314 (59%), Gaps = 28/314 (8%)
Query: 18 CSFFRKPVNRK-NMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSAS 76
C+F K N+K + RKR + D E S +E S + V+KK + A N + T + A
Sbjct: 12 CTFIFKKSNKKFSARKRNASDSDKEKSSGDEGSAV-VRKKKSAAVNPMIQKTKKVEREAV 70
Query: 77 AKPNEESEKP------SFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQAT 130
+ E EK ++ S + + D ATA E +T+ +DA+AI ER+ K
Sbjct: 71 SSSESEEEKEEKSVTVAYKSTRSAKPEGPDDMGATAVYELDTERDKDAQAIFERSQK-IQ 129
Query: 131 ESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSH-GPLRASAHIRVSARF 189
E L GK ED K+Y+GINNY HK ++ T+ + +G GP+RA H+R + R+
Sbjct: 130 EELTGK----EDDKIYRGINNY--HKFIKPKDSTMGNASSGMVRKGPIRAPEHLRATVRW 183
Query: 190 DYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEE 249
DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E +E R +D
Sbjct: 184 DYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELEEG------RYGANDDENY 237
Query: 250 EGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPT 309
E +S DDED LPF CFICR F +P+ TKC+HYFCE CAL+H+ K+K+C+VCNQ T
Sbjct: 238 EVSS-----DDED-LPFKCFICRESFKNPIITKCRHYFCEACALQHYRKSKRCYVCNQQT 291
Query: 310 LGIFNVAHEIRKKM 323
G+FN A E+ KM
Sbjct: 292 NGVFNPAKELMAKM 305
>UniRef100_Q67EQ9 Zinc finger protein 183 [Oncorhynchus mykiss]
Length = 320
Score = 243 bits (621), Expect = 4e-63
Identities = 133/323 (41%), Positives = 195/323 (60%), Gaps = 26/323 (8%)
Query: 10 ENQQTEQVCSF-FRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFST 68
E+ + C+F F+K R N RKR + D + +S+ +++ + ++K T + N + T
Sbjct: 3 ESGEPNATCTFLFKKSTKRCNARKRKASDSDTDGNSDEDKNSVVRKEKKTGSANPMIQRT 62
Query: 69 GSSK---SSASAKPNEESEKPSFHFESSKEIQVQ--HDSKATATLETETDFSRDARAIRE 123
+ SSA + +E K + ++S++ + + D ATA E +T DA+AI E
Sbjct: 63 KKVEREVSSAESDEEKEKNKVTVAYKSTRSAKPEGPDDMGATAIYELDTAKDNDAQAIFE 122
Query: 124 RALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSH-GPLRASAH 182
R+ K E L GK ED K+Y+G+NNY H ++ T+ + +G GP+RA H
Sbjct: 123 RSQK-IQEELTGK----EDDKIYRGMNNYKKHIKP--KDSTMGNASSGMVRKGPIRAPEH 175
Query: 183 IRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLA 242
+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE R
Sbjct: 176 LRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG------RYG 229
Query: 243 TGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKC 302
+D E +S DE+ +PF CFICR F +PV TKC+HYFCE CAL+H+ K+++C
Sbjct: 230 ANDDENYEVSS------DEEDMPFKCFICRESFKNPVITKCRHYFCETCALQHYRKSQRC 283
Query: 303 FVCNQPTLGIFNVAHEIRKKMAE 325
+VCN T G+FN A E+ K+A+
Sbjct: 284 YVCNVQTNGVFNPAKELAAKIAK 306
>UniRef100_Q67EQ7 Zinc finger protein 183 [Oryzias latipes]
Length = 319
Score = 234 bits (598), Expect = 2e-60
Identities = 127/322 (39%), Positives = 191/322 (58%), Gaps = 28/322 (8%)
Query: 10 ENQQTEQVCSF-FRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFST 68
E+++ ++ C+F F+K + RKR + D + +S E+S++ + + N + T
Sbjct: 3 ESEEPKKSCTFLFKKSAKKFCGRKRKASDSDKDANSEEEQSVVVRRDRKEGRVNPMIQRT 62
Query: 69 GSSKSSASAKPNEESE---KPSFHFESSKEIQVQ--HDSKATATLETETDFSRDARAIRE 123
+ A + + E + K + ++SS+ + + D ATAT E +T+ +DA+AI E
Sbjct: 63 KKVERDAVSSSDSEDDNEGKITVAYKSSRSAKPEGPEDMGATATYELDTEKDKDAQAIFE 122
Query: 124 RALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPLRASA 181
R+ K E L GK ED K+Y+GINNY + + T + G GP+RA
Sbjct: 123 RSQK-IQEELTGK----EDDKIYRGINNYQKF---IKPKDTSMGNASSGMVRKGPIRAPE 174
Query: 182 HIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRL 241
H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E +E
Sbjct: 175 HLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELEEGRYGAN--- 231
Query: 242 ATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKK 301
+EE ++ +D+D +PF CFICR + +P+ TKCKHYFCE CAL+H+ K+K+
Sbjct: 232 ------DEENYEVSSDDED---VPFKCFICRESYKNPIVTKCKHYFCEACALQHYRKSKR 282
Query: 302 CFVCNQPTLGIFNVAHEIRKKM 323
C+VCN T G+FN A E+ K+
Sbjct: 283 CYVCNTQTNGVFNPAKELMAKL 304
>UniRef100_Q7QIT7 ENSANGP00000021479 [Anopheles gambiae str. PEST]
Length = 315
Score = 234 bits (597), Expect = 2e-60
Identities = 127/311 (40%), Positives = 187/311 (59%), Gaps = 25/311 (8%)
Query: 19 SFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTG--------S 70
+F ++ + K RKR +E +E + + S++ VQ + KA+ + ++ S
Sbjct: 3 AFVKRNIKNKGARKRQKSSESDEAEEESS-SVVVVQDRRKKANPNVQSTSALRKKQARSS 61
Query: 71 SKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQAT 130
+ S+ + +EES S+ + S + + D ATA LE ET+ RDA+AI ++++
Sbjct: 62 NADSSHSSDDEESAGLSYKSKRSAQPEGPRDQGATAELEIETEKDRDAQAIYQKSI-DIN 120
Query: 131 ESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPLRASAHIRVSAR 188
+ L+GK ED K+Y+G+NNYT + F+++ + A G GP+RA A+IR + R
Sbjct: 121 KELEGK----EDDKVYRGLNNYTQY---FKKKDSAQGNAASGMVRKGPIRAPANIRSTVR 173
Query: 189 FDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAE 248
+DYQPDICKDYKETGYCG+GDSCKF+HDR DYK GWQME+ E A G+D++
Sbjct: 174 WDYQPDICKDYKETGYCGFGDSCKFLHDRSDYKHGWQMEQ-----EGAGSGHNHGGDDSD 228
Query: 249 EEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQP 308
+ DDE+ LPF C++CR FVDP+ TKCKHYFCE CAL + K+ +C +C
Sbjct: 229 GDDTKYEIHSDDEE-LPFKCYVCRESFVDPIVTKCKHYFCERCALAQYKKSSRCAICGVQ 287
Query: 309 TLGIFNVAHEI 319
T G+FN A E+
Sbjct: 288 TNGMFNPAKEL 298
>UniRef100_Q67EQ6 Zinc finger protein 183 [Ciona intestinalis]
Length = 325
Score = 233 bits (595), Expect = 4e-60
Identities = 127/316 (40%), Positives = 186/316 (58%), Gaps = 31/316 (9%)
Query: 21 FRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSK------SS 74
F++ RK ++R +E N + S++EE V+K+ + N + ST K SS
Sbjct: 20 FKRGGGRKRPQRRVVEK--NNSSSSSEEESAVVRKEKRQILNHMKQSTNKMKKKVKYSSS 77
Query: 75 ASAKPNEESEKPSFHFESSKEIQVQ--HDSKATATLETETDFSRDARAIRERALKQATES 132
+S++ +++++ + ++S++ + D AT E +T RDA+A+ ERA K E
Sbjct: 78 SSSESDQKAKAIAVLYKSTQSAKASGPDDMGATKIFELDTSKDRDAQAVFERAQKLQEEL 137
Query: 133 LKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPLRASAHIRVSARFD 190
G++ D K+Y+G Y + + + T + G GP+RA H+R + R+D
Sbjct: 138 KSGEA----DDKVYRGAAGY---RKFIKPKDTALGNASSGMVRKGPIRAPEHLRATVRWD 190
Query: 191 YQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEE 250
YQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E E R +EE
Sbjct: 191 YQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELTEGTYGRN---------QEE 241
Query: 251 GASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPTL 310
++D DD+ LPF CFICR FVDP+ST+CKHYFCE CAL H+ K+++CFVC + T
Sbjct: 242 NWEVSDSDDE---LPFKCFICRKSFVDPISTRCKHYFCEQCALNHYRKSRRCFVCGEQTN 298
Query: 311 GIFNVAHEIRKKMAED 326
G+FN A EI KM +
Sbjct: 299 GVFNPAKEIVSKMENE 314
>UniRef100_Q67EQ8 Zinc finger protein 183 [Brachydanio rerio]
Length = 321
Score = 231 bits (590), Expect = 2e-59
Identities = 135/314 (42%), Positives = 184/314 (57%), Gaps = 28/314 (8%)
Query: 18 CSFFRKPVNRK-NMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSAS 76
C+F K N+K + RKR + D E S +E S + V+KK + A N + T + A
Sbjct: 12 CTFIFKKSNKKFSARKRNASDSDKEKSSGDEGSAV-VRKKKSAAVNPMIQKTKKVEREAV 70
Query: 77 AKPNEESEKP------SFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQAT 130
+ E EK ++ S + + D ATA E +T+ +DA+AI ER+ K
Sbjct: 71 SSSESEEEKEEKSVTVAYKSTRSAKPEGPDDMGATAVYELDTERDKDAQAIFERSQK-IQ 129
Query: 131 ESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSH-GPLRASAHIRVSARF 189
E L GK ED K+Y+GINNY HK ++ T+ + G GP+RA H+R + R+
Sbjct: 130 EELTGK----EDDKIYRGINNY--HKFIKPKDSTMGNASPGMVRIGPIRAPQHLRATVRW 183
Query: 190 DYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEE 249
DYQPDIC+D+ ETG+ G DSCKF+HDR DYK GWQ+E+E +E R +D
Sbjct: 184 DYQPDICQDHIETGFSGIADSCKFLHDRSDYKHGWQIERELEEG------RYGAYDDDNY 237
Query: 250 EGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPT 309
E +S DDED LPF CFICR F +P+ TKC+HYFCE CAL+H+ K+K+C+VCNQ T
Sbjct: 238 EVSS-----DDED-LPFKCFICRESFKNPIITKCRHYFCEACALQHYRKSKRCYVCNQQT 291
Query: 310 LGIFNVAHEIRKKM 323
G+FN A E+ KM
Sbjct: 292 NGVFNPAKELMAKM 305
>UniRef100_Q67ER4 Zinc finger protein 183 [Bos taurus]
Length = 343
Score = 231 bits (589), Expect = 2e-59
Identities = 135/331 (40%), Positives = 190/331 (56%), Gaps = 37/331 (11%)
Query: 14 TEQVCSFFRKPVNRK---NMRKRTIENE---DNENDSNNEESLMHVQKKNTKADNKLFFS 67
T+QVC+F K RK RKR I N+ D+ + S+ +++ +KK + + +
Sbjct: 11 TDQVCTFLFKKPGRKVAAGRRKRPICNQESGDSSSSSDEGNTVVRPEKKRAVHNPMIQKT 70
Query: 68 TGSSKSSA-----SAKPNEESEKPSFH--FESSKEIQV--QHDSKATATLETETDFSRDA 118
GS K S++ EE++ S ++S++ + D ATA E +T+ RDA
Sbjct: 71 RGSGKQKVAYGDLSSEEEEENKSESLGVVYKSTRSAKPVGPEDMGATAVYELDTEKERDA 130
Query: 119 RAIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGP 176
+AI ER+ K E L+G+ ED K+Y+GINNY + + T + G GP
Sbjct: 131 QAIFERSQK-IQEELRGQ----EDDKIYRGINNYQKF---MKPKDTSMGNASSGMVRKGP 182
Query: 177 LRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKA 236
+RA H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE
Sbjct: 183 IRAPEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG--- 239
Query: 237 RKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHH 296
R ED E S DE+ +PF CFICR F +PV TKC+HYFCE CAL+H
Sbjct: 240 ---RYGVYEDENYEVGS------DEEEIPFKCFICRQTFQNPVVTKCRHYFCESCALQHF 290
Query: 297 AKNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C+VC+Q T G+FN A E+ K+ + +
Sbjct: 291 RTTPRCYVCDQQTNGVFNPAKELIAKLEKHR 321
>UniRef100_Q8I1B4 CG4973-PA [Drosophila pseudoobscura]
Length = 358
Score = 231 bits (588), Expect = 3e-59
Identities = 129/337 (38%), Positives = 191/337 (56%), Gaps = 39/337 (11%)
Query: 9 TENQQTEQVCSFFRKPVNR-KNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKL--- 64
+E T F ++ +NR MR++ + N++ +++E + +A+N+
Sbjct: 2 SEEPPTTSAFGFKKRNINRGAAMRRKKESSSSNDSAKSSDEETQNKASALVRAENRRKRT 61
Query: 65 ---FFST----------GSSKSSASAKPNEESEKPSFHFESSKEIQVQ--HDSKATATLE 109
F ST G + + S+ E + ++S +E D AT+ E
Sbjct: 62 NPNFQSTKTVGKAKRLAGVATEATSSDGKSEDDGLGVAYKSKREALPSGPQDQGATSVNE 121
Query: 110 TETDFSRDARAIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEK 169
+T+ RDA+AI RALK E L+GK+ D K+Y+GINNY + ++++ T A
Sbjct: 122 MDTELDRDAQAIHARALK-INEELEGKA----DDKIYRGINNYAQY---YKKQDTAAGNA 173
Query: 170 AGGS--HGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQME 227
+ G GP+RA AH+R + R+DYQPDICKDYKETGYCG+GDSCKF+HDR DYK+GWQ+E
Sbjct: 174 SSGMVRSGPIRAPAHLRATVRWDYQPDICKDYKETGYCGFGDSCKFLHDRSDYKAGWQLE 233
Query: 228 KEWDEAEKARKMRLATGEDAEEEGASLNDE-DDDEDALPFACFICRNPFVDPVSTKCKHY 286
+ + + D E +G E DE++LPF C ICR FV+PV TKCKHY
Sbjct: 234 ADHEAQRRG---------DCESDGDDGKYEIHSDEESLPFKCHICRQSFVNPVVTKCKHY 284
Query: 287 FCEHCALKHHAKNKKCFVCNQPTLGIFNVAHEIRKKM 323
FCE CAL + K+++C +C+Q T GIFN A E+ +++
Sbjct: 285 FCEKCALAQYKKSQRCIICSQQTNGIFNPAKELIERL 321
>UniRef100_Q8I141 CG4973-PA [Drosophila willistoni]
Length = 357
Score = 231 bits (588), Expect = 3e-59
Identities = 132/333 (39%), Positives = 192/333 (57%), Gaps = 38/333 (11%)
Query: 9 TENQQTEQVCSFFRKPVNRKNMRKRT-IENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
+E T F ++ +N+ R+R + NE+ +++E +A+N+ +
Sbjct: 2 SEEAPTTSAFGFKKRNINKGAARRRKKSSSSSNESVKSSDEEQQRKTSAVVRAENRRKRT 61
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSK-------------------ATATL 108
+ +S+ S ++ + SS+E + +HD+ AT+
Sbjct: 62 NPNFQSTKSGVAKRQAGAGIANANSSEEAEEEHDNLGVAYKSKREALPTGPQDQGATSIN 121
Query: 109 ETETDFSRDARAIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASE 168
E +T+ RDA+AI RALK E L+GK+ D K+Y+GINNY + ++++ T A
Sbjct: 122 EMDTELDRDAQAIHARALK-INEELEGKA----DDKIYRGINNYAQY---YKKQDTAAGN 173
Query: 169 KAGGS--HGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQM 226
+ G GP+RA AH+R + R+DYQPDICKDYKETGYCG+GDSCKF+HDR DYK+GWQ+
Sbjct: 174 ASSGMVRSGPIRAPAHLRATVRWDYQPDICKDYKETGYCGFGDSCKFLHDRSDYKAGWQL 233
Query: 227 EKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHY 286
E + +AEK R + G+D + E S DE+ LPF C ICR FV+PV TKCKHY
Sbjct: 234 EMD-HQAEK-RGDCDSDGDDGKYEIHS------DEETLPFKCHICRQSFVNPVVTKCKHY 285
Query: 287 FCEHCALKHHAKNKKCFVCNQPTLGIFNVAHEI 319
FCE CAL + K+++C VC+Q T GIFN A E+
Sbjct: 286 FCEKCALAQYKKSQRCIVCSQQTNGIFNPAKEL 318
>UniRef100_Q8R3P8 RIKEN cDNA 2810428C21 [Mus musculus]
Length = 341
Score = 230 bits (586), Expect = 5e-59
Identities = 133/329 (40%), Positives = 188/329 (56%), Gaps = 35/329 (10%)
Query: 14 TEQVCSFFRKPVNRKNM---RKRTI---ENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
T QVC+F K RK RKR + + D+ + S+ S++ +KK + + +
Sbjct: 11 TRQVCTFLFKKPGRKGAAGRRKRRLCDAGSGDSCSSSDEGSSVVRPEKKQATHNPMIQKT 70
Query: 68 TGSSKSSA-----SAKPNEESEKPSFHFESSKEIQV--QHDSKATATLETETDFSRDARA 120
GS+K A S++ E E ++S++ + D ATA E +T+ RDA++
Sbjct: 71 RGSAKQKATYGGLSSEDENEPEGLGVVYKSTRSAKPVGPEDMGATAVYELDTEKDRDAQS 130
Query: 121 IRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPLR 178
I ER+ K E L+GK ED K+Y+GINNY + + + T + G GP+R
Sbjct: 131 IFERSQK-IQEELRGK----EDDKIYRGINNYQKY---VKPKDTSMGNASSGMVRKGPIR 182
Query: 179 ASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARK 238
A H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE
Sbjct: 183 APEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG----- 237
Query: 239 MRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAK 298
R ED E S D++ +PF CFICR F +PV TKC+HYFCE CAL+H
Sbjct: 238 -RYGVYEDENYEVGS------DDEEIPFKCFICRQTFQNPVVTKCRHYFCERCALQHFRT 290
Query: 299 NKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C+VC+Q T G+FN A E+ K+ + +
Sbjct: 291 TSRCYVCDQQTNGVFNPAKELIAKLGKHR 319
>UniRef100_UPI000036445E UPI000036445E UniRef100 entry
Length = 293
Score = 229 bits (585), Expect = 6e-59
Identities = 126/309 (40%), Positives = 185/309 (59%), Gaps = 26/309 (8%)
Query: 15 EQVCSF-FRKPVNRKNMRKRTIENEDNENDSN-NEESLMHVQKKNTKADNKLFFSTGSSK 72
E C+F F+K + RKR + D + S+ N S++ + K K + + + K
Sbjct: 3 ENSCAFLFKKSTKKFAGRKRKASDSDKDGSSDENPTSVVRRENKGAKVNPMIQKTKKVEK 62
Query: 73 SSASAKPNEES--EKPSFHFESSKEIQV--QHDSKATATLETETDFSRDARAIRERALKQ 128
+ S+ +EE +K + ++S++ + D ATAT + +T+ DA+AI ER+ K
Sbjct: 63 KAVSSSDSEEEKEDKITVAYKSTRSAKPVGPDDMGATATYQLDTERDNDAQAIFERSQK- 121
Query: 129 ATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSH-GPLRASAHIRVSA 187
++ + T +D K+Y+GINNY K ++ T+ + +G GP+RA H+R +
Sbjct: 122 ----IQEERTGKDDDKIYRGINNYV--KFIKPKDTTMGNASSGMVRKGPIRAPEHLRATV 175
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E +E R G D
Sbjct: 176 RWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELEEG------RYGAGNDE 229
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
E +S DE+ LPF CFIC+ F +P+ TKCKHYFCE CAL+H+ K+K+C+VCN
Sbjct: 230 NYEVSS------DEEDLPFKCFICKESFKNPIVTKCKHYFCEVCALQHYRKSKRCYVCNT 283
Query: 308 PTLGIFNVA 316
T G+FN A
Sbjct: 284 QTNGVFNPA 292
>UniRef100_Q9D7A3 Mus musculus adult male tongue cDNA, RIKEN full-length enriched
library, clone:2310020H19 product:hypothetical RING
finger domain, C3HC4 structure containing protein, full
insert sequence [Mus musculus]
Length = 345
Score = 229 bits (585), Expect = 6e-59
Identities = 139/332 (41%), Positives = 188/332 (55%), Gaps = 37/332 (11%)
Query: 9 TENQQTEQVCSFFRKPVNRKNM---RKR-TIENEDNENDSNNEESLMHVQ-KKNTKADNK 63
++ + +QVC+F K RK RKR + E E+ S+++E V+ +K A N
Sbjct: 6 SQGKSEDQVCTFLFKKPGRKGSAGRRKRPACDPESGESGSSSDEGCSVVRPEKKRAAHNP 65
Query: 64 LFFST---GSSKSSASAKPNEESEKPSFH-----FESSKEIQV--QHDSKATATLETETD 113
+ T G K + +EE EK ++S++ + D ATA E +T+
Sbjct: 66 MIQKTSGSGKQKGAYCDLSSEEEEKAGNESLGVVYKSTRSAKPVGPEDMGATAVYELDTE 125
Query: 114 FSRDARAIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS 173
RDA+AI ER+ K E L+GK ED K+Y+GINNY + + + T + G
Sbjct: 126 KERDAQAIFERSQK-IQEELRGK----EDDKIYRGINNYQKY---MKPKDTSMGNASSGM 177
Query: 174 --HGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWD 231
GP+RA H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E D
Sbjct: 178 VRKGPIRAPEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELD 237
Query: 232 EAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHC 291
E R ED E E DDE+ +PF CFICR F +PV TKCKHYFCE C
Sbjct: 238 EG------RYGVYEDENYE-----VESDDEE-IPFKCFICRQTFQNPVVTKCKHYFCETC 285
Query: 292 ALKHHAKNKKCFVCNQPTLGIFNVAHEIRKKM 323
AL+H +C+VC Q T G+FN A E+ K+
Sbjct: 286 ALQHFRTTPRCYVCEQQTHGVFNPAKELIAKL 317
>UniRef100_UPI000036F7CE UPI000036F7CE UniRef100 entry
Length = 343
Score = 228 bits (581), Expect = 2e-58
Identities = 135/330 (40%), Positives = 191/330 (56%), Gaps = 37/330 (11%)
Query: 15 EQVCSFFRKPVNRKNM---RKR-TIENEDNENDSNNEE--SLMHVQKKNTKADNKLFFST 68
+QVC+F K RK RKR + E E+ S+++E +++ +KK + + +
Sbjct: 12 DQVCTFLFKKPGRKGAAGRRKRPACDPEPGESGSSSDEGCTVVRPEKKRVTHNPMIQKTR 71
Query: 69 GSSKSSA-----SAKPNEESEKPSFH--FESSKEIQV--QHDSKATATLETETDFSRDAR 119
S K A S++ EE+E S ++S++ + D ATA E +T+ RDA+
Sbjct: 72 DSGKQKAAYGDLSSEEEEENEPESLGVVYKSTRSAKPVGPEDMGATAVYELDTEKERDAQ 131
Query: 120 AIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPL 177
AI ER+ K E L+GK ED K+Y+GINNY + + + T + G GP+
Sbjct: 132 AIFERSQK-IQEELRGK----EDDKIYRGINNYQKY---MKPKDTSMGNASSGMVRKGPI 183
Query: 178 RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKAR 237
RA H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE
Sbjct: 184 RAPEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG---- 239
Query: 238 KMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHA 297
R ED E S D++ +PF CFICR F +PV TKC+HYFCE CAL+H
Sbjct: 240 --RYGVYEDENYEVGS------DDEEIPFKCFICRQSFQNPVVTKCRHYFCESCALQHFR 291
Query: 298 KNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C+VC+Q T G+FN A E+ K+ + +
Sbjct: 292 TTPRCYVCDQQTNGVFNPAKELIAKLEKHR 321
>UniRef100_Q67ER3 Zinc finger protein 183 [Gallus gallus]
Length = 328
Score = 228 bits (581), Expect = 2e-58
Identities = 129/323 (39%), Positives = 191/323 (58%), Gaps = 32/323 (9%)
Query: 17 VCSF-FRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTK-ADNKLFFSTGS---- 70
VCSF F+K +R + D E +S+ EE V+K+ + N + T
Sbjct: 7 VCSFVFKKRGLAAGRGRRKRPSSDQEQESSGEEGSTVVRKERRRETPNPMIQKTRRCTKE 66
Query: 71 --SKSSASAKPNEESEKPSFHFESSKEIQV--QHDSKATATLETETDFSRDARAIRERAL 126
S + +S+ ++ S++ ++S++ + D ATA E +T+ +DA+AI ER+
Sbjct: 67 RPSYALSSSDDDDPSKEIGVTYKSTRSAKPVGPEDMGATAVYELDTEKDKDAQAIFERSQ 126
Query: 127 KQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPLRASAHIR 184
K E L+GK ED K+Y+GINNY + + + T + G GP+RA H+R
Sbjct: 127 K-IQEELRGK----EDDKIYRGINNYQKY---VKPKDTSMGNASSGMVRKGPIRAPEHLR 178
Query: 185 VSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATG 244
+ R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE R
Sbjct: 179 ATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG------RYGVN 232
Query: 245 EDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFV 304
++ E +S DE+ +PF CFICR+ F +PV TKC+HYFCE CAL+H+ K+++C+V
Sbjct: 233 DEENYEVSS------DEEDMPFKCFICRSSFKNPVVTKCRHYFCESCALQHYRKSQRCYV 286
Query: 305 CNQPTLGIFNVAHEIRKKMAEDK 327
C++ T G+FN A E+ K+ + K
Sbjct: 287 CDKQTNGVFNPAKELMAKLEKHK 309
>UniRef100_O15541 Zinc finger protein 183 [Homo sapiens]
Length = 343
Score = 228 bits (581), Expect = 2e-58
Identities = 135/330 (40%), Positives = 191/330 (56%), Gaps = 37/330 (11%)
Query: 15 EQVCSFFRKPVNRKNM---RKR-TIENEDNENDSNNEE--SLMHVQKKNTKADNKLFFST 68
+QVC+F K RK RKR + E E+ S+++E +++ +KK + + +
Sbjct: 12 DQVCTFLFKKPGRKGAAGRRKRPACDPEPGESGSSSDEGCTVVRPEKKRVTHNPMIQKTR 71
Query: 69 GSSKSSA-----SAKPNEESEKPSFH--FESSKEIQV--QHDSKATATLETETDFSRDAR 119
S K A S++ EE+E S ++S++ + D ATA E +T+ RDA+
Sbjct: 72 DSGKQKAAYGDLSSEEEEENEPESLGVVYKSTRSAKPVGPEDMGATAVYELDTEKERDAQ 131
Query: 120 AIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPL 177
AI ER+ K E L+GK ED K+Y+GINNY + + + T + G GP+
Sbjct: 132 AIFERSQK-IQEELRGK----EDDKIYRGINNYQKY---MKPKDTSMGNASSGMVRKGPI 183
Query: 178 RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKAR 237
RA H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE
Sbjct: 184 RAPEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG---- 239
Query: 238 KMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHA 297
R ED E S D++ +PF CFICR F +PV TKC+HYFCE CAL+H
Sbjct: 240 --RYGVYEDENYEVGS------DDEEIPFKCFICRQSFQNPVVTKCRHYFCESCALQHFR 291
Query: 298 KNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C+VC+Q T G+FN A E+ K+ + +
Sbjct: 292 TTPRCYVCDQQTNGVFNPAKELIAKLEKHR 321
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 544,687,246
Number of Sequences: 2790947
Number of extensions: 23076323
Number of successful extensions: 137087
Number of sequences better than 10.0: 2545
Number of HSP's better than 10.0 without gapping: 1228
Number of HSP's successfully gapped in prelim test: 1362
Number of HSP's that attempted gapping in prelim test: 130510
Number of HSP's gapped (non-prelim): 6009
length of query: 327
length of database: 848,049,833
effective HSP length: 127
effective length of query: 200
effective length of database: 493,599,564
effective search space: 98719912800
effective search space used: 98719912800
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC126016.2


