

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.14 + phase: 0 /pseudo
(241 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
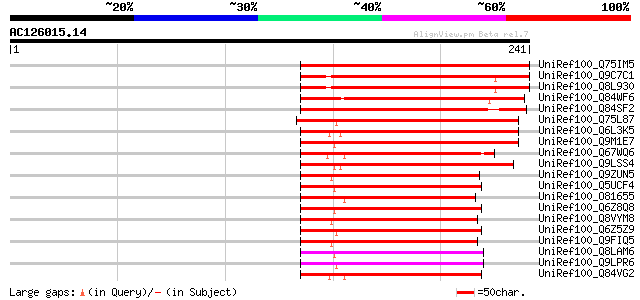
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q75IM5 Putative senescence-associated protein [Oryza s... 152 7e-36
UniRef100_Q9C7C1 Senescence-assocated protein, putative; 28418-2... 143 4e-33
UniRef100_Q8L930 Senescence-assocated protein, putative [Arabido... 143 4e-33
UniRef100_Q84WF6 Hypothetical protein At4g23410 [Arabidopsis tha... 142 1e-32
UniRef100_Q84SF2 Putative senescence-assocated protein [Oryza sa... 128 1e-28
UniRef100_Q75L87 Hypothetical protein OJ1729_E02.7 [Oryza sativa] 125 1e-27
UniRef100_Q6L3K5 Putative senescence-associated protein [Solanum... 115 7e-25
UniRef100_Q9M1E7 Hypothetical protein F9K21.180 [Arabidopsis tha... 108 1e-22
UniRef100_Q67WQ6 Putative senescence-associated protein 5 [Oryza... 108 1e-22
UniRef100_Q9LSS4 Similarity to senescence-associated protein [Ar... 105 1e-21
UniRef100_Q9ZUN5 Putative senescence-associated protein 5 [Arabi... 102 6e-21
UniRef100_Q5UCF4 Senescence-associated protein DH [Zea mays] 102 8e-21
UniRef100_O81655 Senescence-associated protein 5 [Hemerocallis sp.] 100 4e-20
UniRef100_Q6Z8Q8 Putative senescence-associated protein [Oryza s... 99 1e-19
UniRef100_Q8VYM8 Putative senescence-associated protein 5 [Arabi... 97 4e-19
UniRef100_Q6Z5Z9 Putative senescence-associated protein [Oryza s... 97 4e-19
UniRef100_Q9FIQ5 Senescence-associated protein 5-like protein [A... 97 4e-19
UniRef100_Q8LAM6 Hypothetical protein [Arabidopsis thaliana] 97 5e-19
UniRef100_Q9LPR6 F15H18.1 [Arabidopsis thaliana] 95 2e-18
UniRef100_Q84VG2 Putative senescence-associated protein 5 [Oryza... 95 2e-18
>UniRef100_Q75IM5 Putative senescence-associated protein [Oryza sativa]
Length = 286
Score = 152 bits (384), Expect = 7e-36
Identities = 61/106 (57%), Positives = 83/106 (77%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC Y+ + QD DC++W+N +LCY C+SC+AGV+E +R +WHK+SVL
Sbjct: 168 SGCCKPPTACAYSGGVAVGAQDEDCFRWNNAAGILCYGCESCRAGVMEKVREDWHKISVL 227
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDYH 241
V +L++LI I + GCCAFRNARR+ ++YPYG NRM K+ PRWDY+
Sbjct: 228 NVMVLVVLICICACGCCAFRNARRSVSEYPYGVNRMHKIHPRWDYY 273
>UniRef100_Q9C7C1 Senescence-assocated protein, putative; 28418-29806 [Arabidopsis
thaliana]
Length = 282
Score = 143 bits (360), Expect = 4e-33
Identities = 62/107 (57%), Positives = 81/107 (74%), Gaps = 3/107 (2%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC Y EA ++ DC++W+N +LCYECD+CKAGVLE+IR +W KLSV+
Sbjct: 165 SGCCKPPTACTY--EAGVVDGGGDCFRWNNGVEMLCYECDACKAGVLEEIRLDWRKLSVV 222
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDY-PYGENRMTKVRPRWDYH 241
+ +L+LLI +Y+ GCCAF N R A Y P +NRMT+VRPRWDY+
Sbjct: 223 NILVLVLLIAVYAAGCCAFHNTRHAAHPYHPSDDNRMTRVRPRWDYY 269
>UniRef100_Q8L930 Senescence-assocated protein, putative [Arabidopsis thaliana]
Length = 120
Score = 143 bits (360), Expect = 4e-33
Identities = 62/107 (57%), Positives = 81/107 (74%), Gaps = 3/107 (2%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC Y EA ++ DC++W+N +LCYECD+CKAGVLE+IR +W KLSV+
Sbjct: 3 SGCCKPPTACTY--EAGVVDGGGDCFRWNNGVEMLCYECDACKAGVLEEIRLDWRKLSVV 60
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDY-PYGENRMTKVRPRWDYH 241
+ +L+LLI +Y+ GCCAF N R A Y P +NRMT+VRPRWDY+
Sbjct: 61 NILVLVLLIAVYAAGCCAFHNTRHAAHPYHPSDDNRMTRVRPRWDYY 107
>UniRef100_Q84WF6 Hypothetical protein At4g23410 [Arabidopsis thaliana]
Length = 281
Score = 142 bits (357), Expect = 1e-32
Identities = 58/105 (55%), Positives = 82/105 (77%), Gaps = 2/105 (1%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPT+C YN + V+ QD DCY+W+N T+LCY+CD+C+AGVLE +RR+WHKLS++
Sbjct: 165 SGCCKPPTSCVYNTDTVIQ-QDPDCYRWNNAATVLCYDCDTCRAGVLETVRRDWHKLSLV 223
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAE-TDYPYGENRMTKVRPRWD 239
V ++I LI +Y +GCCAF+NA+R + +PYG M+K RP W+
Sbjct: 224 NVIVVIFLIAVYCVGCCAFKNAKRPQHYGFPYGRYGMSKSRPGWE 268
>UniRef100_Q84SF2 Putative senescence-assocated protein [Oryza sativa]
Length = 298
Score = 128 bits (321), Expect = 1e-28
Identities = 55/105 (52%), Positives = 70/105 (66%), Gaps = 5/105 (4%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC Y +A Q DCY+WSN P +LCY CDSCKAGVLE +RR+WH +++L
Sbjct: 178 SGCCKPPTACTYYDDAQQQQQQPDCYRWSNAPGVLCYGCDSCKAGVLEQLRRHWHNVTIL 237
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDY 240
V +L+LLI YS CCAFRN A + + + PRW+Y
Sbjct: 238 NVVLLLLLILFYSCACCAFRNTATATS-----SKTIFHLHPRWEY 277
>UniRef100_Q75L87 Hypothetical protein OJ1729_E02.7 [Oryza sativa]
Length = 339
Score = 125 bits (313), Expect = 1e-27
Identities = 55/113 (48%), Positives = 74/113 (64%), Gaps = 10/113 (8%)
Query: 134 SNTGCCKPPTACNYNME----------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLE 183
S +GCCKPPT+C YN + D DC KWSN+ LC++CDSCKAGVL
Sbjct: 222 SKSGCCKPPTSCAYNYVNETFWTANPGVPTVVNDVDCSKWSNDQQTLCFQCDSCKAGVLA 281
Query: 184 DIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
I+++W K+++L + +LI+L+ +Y GC AFRNARR E D P+G RMTK +P
Sbjct: 282 GIKKSWRKVAILNIVVLIILVIVYVAGCAAFRNARRIENDEPFGMARMTKTQP 334
>UniRef100_Q6L3K5 Putative senescence-associated protein [Solanum demissum]
Length = 232
Score = 115 bits (289), Expect = 7e-25
Identities = 52/109 (47%), Positives = 71/109 (64%), Gaps = 8/109 (7%)
Query: 136 TGCCKPPTACNY--NMEAV------MMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C Y E V ++ D DC KWSN+ LCY CDSCKAGVL +++
Sbjct: 119 SGCCKPPTECGYVYQNETVWIPGGGLVGADPDCVKWSNDQEQLCYNCDSCKAGVLASLKK 178
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
+W K+SV+ + +LILL+ +Y + AFR+ +R + D PYGE RM K +P
Sbjct: 179 SWRKVSVINIVILILLVIMYMVAIAAFRHNKRIDNDEPYGETRMEKTQP 227
>UniRef100_Q9M1E7 Hypothetical protein F9K21.180 [Arabidopsis thaliana]
Length = 285
Score = 108 bits (270), Expect = 1e-22
Identities = 47/109 (43%), Positives = 69/109 (63%), Gaps = 8/109 (7%)
Query: 136 TGCCKPPTACNYNM--------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C ++ M+ + DC WSN+ ++LCY+C SCKAGVL +++
Sbjct: 172 SGCCKPPTDCGFSYVNETGWDTRGGMIGPNQDCMVWSNDQSMLCYQCSSCKAGVLGSLKK 231
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
+W K+SV+ + +LI+L+ Y I A+RN +R + D P GE RMTK P
Sbjct: 232 SWRKVSVINIVVLIILVIFYVIAYAAYRNVKRIDNDEPAGEARMTKSHP 280
>UniRef100_Q67WQ6 Putative senescence-associated protein 5 [Oryza sativa]
Length = 269
Score = 108 bits (270), Expect = 1e-22
Identities = 50/99 (50%), Positives = 64/99 (64%), Gaps = 10/99 (10%)
Query: 136 TGCCKPPTACN--YNMEAVMM-------TQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
+GCCKPPT CN Y E V T D DC WSN+ T LCY+C SCKAGVL +++
Sbjct: 169 SGCCKPPTGCNFAYVSETVWTKPSGFNSTDDPDCTTWSNDQTALCYDCQSCKAGVLANLK 228
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYP 225
+W K++ + + LI LI +YS+GCCAFRN RR + YP
Sbjct: 229 NDWKKIATVNIIFLIFLIIVYSVGCCAFRNNRR-DNSYP 266
>UniRef100_Q9LSS4 Similarity to senescence-associated protein [Arabidopsis thaliana]
Length = 327
Score = 105 bits (262), Expect = 1e-21
Identities = 47/107 (43%), Positives = 68/107 (62%), Gaps = 8/107 (7%)
Query: 136 TGCCKPPTACNYNM--EAV------MMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C Y E V M+ + DC W+N+ LLCY+C SCKAGVL +++
Sbjct: 172 SGCCKPPTDCGYTYVNETVWIPGGEMVGPNPDCMLWNNDQRLLCYQCSSCKAGVLGSLKK 231
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKV 234
+W K+SV+ + ++I+L+ Y I C A++N +R D P GE RMT +
Sbjct: 232 SWRKVSVINIVVVIILVIFYVIACAAYQNVKRMYNDEPVGEARMTNL 278
>UniRef100_Q9ZUN5 Putative senescence-associated protein 5 [Arabidopsis thaliana]
Length = 270
Score = 102 bits (255), Expect = 6e-21
Identities = 42/90 (46%), Positives = 61/90 (67%), Gaps = 7/90 (7%)
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPTAC YN + M D+DCY WSN+ + LCY C+SCKAG+L ++R+
Sbjct: 168 SGCCKPPTACGYNFVNPTLWLNPTNMAADADCYLWSNDQSQLCYNCNSCKAGLLGNLRKE 227
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W K +++ + +++LI +Y I C AFRNA+
Sbjct: 228 WRKANLILIITVVVLIWVYVIACSAFRNAQ 257
>UniRef100_Q5UCF4 Senescence-associated protein DH [Zea mays]
Length = 273
Score = 102 bits (254), Expect = 8e-21
Identities = 42/91 (46%), Positives = 58/91 (63%), Gaps = 7/91 (7%)
Query: 136 TGCCKPPTACNYNM-------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT+C Y + D DC WSN+ + LCY C SCKAGV+ +RN
Sbjct: 170 SGCCKPPTSCGYTYVGGTDWTPVTTNSTDPDCKTWSNDASALCYNCQSCKAGVVATFQRN 229
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
W +++V+ + L+ +I +YS+GCCAFRN RR
Sbjct: 230 WKRVAVVCIVFLVFIIIVYSVGCCAFRNNRR 260
>UniRef100_O81655 Senescence-associated protein 5 [Hemerocallis sp.]
Length = 275
Score = 100 bits (248), Expect = 4e-20
Identities = 38/90 (42%), Positives = 60/90 (66%), Gaps = 9/90 (10%)
Query: 136 TGCCKPPTACNYNMEAVMM---------TQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
+GCCKPPT C + ++ + + ++DC W N+PT+LCY+C SCK GV+ +++
Sbjct: 172 SGCCKPPTECGFTYQSPTVWNKPATGFTSNNTDCATWENDPTILCYDCQSCKGGVIANLK 231
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRN 216
W K++V+ + LI +I +YS+GCCAFRN
Sbjct: 232 SKWKKVAVVNIIFLIFIIIVYSVGCCAFRN 261
>UniRef100_Q6Z8Q8 Putative senescence-associated protein [Oryza sativa]
Length = 277
Score = 98.6 bits (244), Expect = 1e-19
Identities = 39/92 (42%), Positives = 62/92 (67%), Gaps = 8/92 (8%)
Query: 136 TGCCKPPTACNYNM--------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT+CN+ A + + D DC +WSN+ +CY C SCKAGV+ ++R
Sbjct: 170 SGCCKPPTSCNFTYGGGTRWGKTARLSSADPDCDEWSNDADEVCYGCRSCKAGVVAALKR 229
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
+W +++++ V L ++ +YS+GCCAF+N+RR
Sbjct: 230 DWKRVAIVNVVFLAFIVVVYSVGCCAFKNSRR 261
>UniRef100_Q8VYM8 Putative senescence-associated protein 5 [Arabidopsis thaliana]
Length = 269
Score = 97.1 bits (240), Expect = 4e-19
Identities = 43/89 (48%), Positives = 56/89 (62%), Gaps = 7/89 (7%)
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT C + + + M+ D DC WSN+ LCY CDSCKAG+L +I+ +
Sbjct: 167 SGCCKPPTKCGFTFVNPTYWVSPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVD 226
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNA 217
W K + + LI LI +Y IGCCAFRNA
Sbjct: 227 WLKADIFLLLALIGLIIVYIIGCCAFRNA 255
>UniRef100_Q6Z5Z9 Putative senescence-associated protein [Oryza sativa]
Length = 273
Score = 97.1 bits (240), Expect = 4e-19
Identities = 42/93 (45%), Positives = 57/93 (61%), Gaps = 9/93 (9%)
Query: 136 TGCCKPPTACNYNME---------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
+GCCKPPTACN+ + D DC WSN+ + LCY C SCKAGVL ++R
Sbjct: 173 SGCCKPPTACNFTYQNETYWIKPPTPSNYSDPDCNSWSNDQSELCYGCQSCKAGVLGNLR 232
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
+W K++ + + LL+ +YS+GCCA RN RR
Sbjct: 233 SSWKKIAFVNAAFVALLLVVYSLGCCALRNNRR 265
>UniRef100_Q9FIQ5 Senescence-associated protein 5-like protein [Arabidopsis thaliana]
Length = 269
Score = 97.1 bits (240), Expect = 4e-19
Identities = 43/89 (48%), Positives = 56/89 (62%), Gaps = 7/89 (7%)
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT C + + + M+ D DC WSN+ LCY CDSCKAG+L +I+ +
Sbjct: 167 SGCCKPPTKCGFTFVNPTYWISPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVD 226
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNA 217
W K + + LI LI +Y IGCCAFRNA
Sbjct: 227 WLKADIFLLLALIGLIIVYIIGCCAFRNA 255
>UniRef100_Q8LAM6 Hypothetical protein [Arabidopsis thaliana]
Length = 272
Score = 96.7 bits (239), Expect = 5e-19
Identities = 44/97 (45%), Positives = 57/97 (58%), Gaps = 12/97 (12%)
Query: 136 TGCCKPPTACNYNM------------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLE 183
+GCCKPP+ CN+ E + ++ DC WSN T LC+ C++CKAGVL
Sbjct: 170 SGCCKPPSDCNFEFRNATFWIPPSKNETAVAAENGDCGTWSNVQTELCFNCNACKAGVLT 229
Query: 184 DIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRA 220
+IR W L V + +LILLI +YS GCCA RN R A
Sbjct: 230 NIREKWRNLLVFNICLLILLITVYSCGCCARRNNRTA 266
>UniRef100_Q9LPR6 F15H18.1 [Arabidopsis thaliana]
Length = 271
Score = 94.7 bits (234), Expect = 2e-18
Identities = 43/96 (44%), Positives = 56/96 (57%), Gaps = 11/96 (11%)
Query: 136 TGCCKPPTACNYNME-----------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLED 184
+GCCKPP+ CN+ + ++ DC WSN T LC+ C++CKAGVL +
Sbjct: 170 SGCCKPPSDCNFEFRNATFWIPPSKNETAVAENGDCGTWSNVQTELCFNCNACKAGVLAN 229
Query: 185 IRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRA 220
IR W L V + +LILLI +YS GCCA RN R A
Sbjct: 230 IREKWRNLLVFNICLLILLITVYSCGCCARRNNRTA 265
>UniRef100_Q84VG2 Putative senescence-associated protein 5 [Oryza sativa]
Length = 276
Score = 94.7 bits (234), Expect = 2e-18
Identities = 39/91 (42%), Positives = 60/91 (65%), Gaps = 7/91 (7%)
Query: 136 TGCCKPPTACNY------NMEAVMM-TQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPP++CN+ N V + D DC W ++ T LCY C SCKAG + ++R+
Sbjct: 170 SGCCKPPSSCNFLYVSGTNWTKVPTNSSDPDCNTWVDDGTQLCYNCQSCKAGAVATLKRD 229
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
W +++V+ + L+ ++ +YS+GCCAFRN RR
Sbjct: 230 WKRVAVVCIVFLVFIVIVYSLGCCAFRNNRR 260
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.343 0.149 0.575
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 392,389,616
Number of Sequences: 2790947
Number of extensions: 14330539
Number of successful extensions: 55045
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 54965
Number of HSP's gapped (non-prelim): 44
length of query: 241
length of database: 848,049,833
effective HSP length: 124
effective length of query: 117
effective length of database: 501,972,405
effective search space: 58730771385
effective search space used: 58730771385
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 73 (32.7 bits)
Medicago: description of AC126015.14


