

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.9 + phase: 0
(233 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
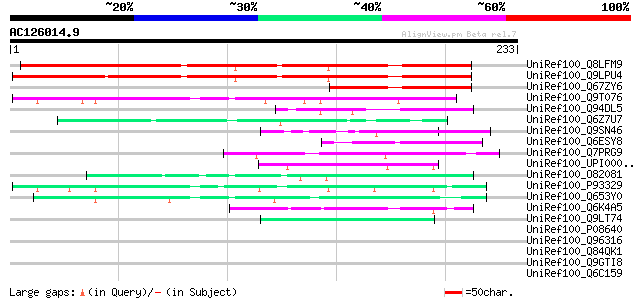
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LFM9 Hypothetical protein [Arabidopsis thaliana] 217 2e-55
UniRef100_Q9LPU4 T22I11.8 [Arabidopsis thaliana] 215 8e-55
UniRef100_Q67ZY6 Hypothetical protein At1g21090 [Arabidopsis tha... 77 3e-13
UniRef100_Q9T076 Early nodulin-like protein 2 precursor [Arabido... 61 2e-08
UniRef100_Q94DL5 P0483G10.28 protein [Oryza sativa] 53 7e-06
UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa] 52 1e-05
UniRef100_Q9SN46 Extensin-like protein [Arabidopsis thaliana] 52 2e-05
UniRef100_Q6ESY8 Hypothetical protein P0472F10.18 [Oryza sativa] 51 2e-05
UniRef100_Q7PRG9 ENSANGP00000024130 [Anopheles gambiae str. PEST] 50 4e-05
UniRef100_UPI00004574FB UPI00004574FB UniRef100 entry 50 6e-05
UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana] 48 2e-04
UniRef100_P93329 Early nodulin 20 precursor [Medicago truncatula] 48 2e-04
UniRef100_Q653Y0 Putative phytocyanin protein, PUP2 [Oryza sativa] 47 5e-04
UniRef100_Q6K4A5 Hypothetical protein OJ1595_D08.12-1 [Oryza sat... 46 9e-04
UniRef100_Q9LT74 Similarity to late embryogenesis abundant prote... 46 9e-04
UniRef100_P08640 Glucoamylase S1/S2 precursor [Saccharomyces cer... 45 0.001
UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis th... 45 0.002
UniRef100_Q84QK1 Putative arabinagalactan protein [Lotus japonicus] 45 0.002
UniRef100_Q9GTI8 Hypothetical esophageal gland cell secretory pr... 45 0.002
UniRef100_Q6C159 Similarity [Yarrowia lipolytica] 45 0.002
>UniRef100_Q8LFM9 Hypothetical protein [Arabidopsis thaliana]
Length = 232
Score = 217 bits (552), Expect = 2e-55
Identities = 113/210 (53%), Positives = 145/210 (68%), Gaps = 13/210 (6%)
Query: 6 FFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQKAFNLC 65
F++F LSLF S S T LVDG S WK+PTV GDS+ F+HK Y+LYIF+N+ AFN+C
Sbjct: 4 FYFFCFLSLFSCPSLSATFLVDGVSVWKSPTVHTGDSVIFRHKYGYDLYIFRNKDAFNVC 63
Query: 66 NFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSL--KACQDSQKLAIKVTPTKASAP 123
NFTQA LLT P++T S+TW+PSR G +YF+F+N++ K CQ +QKL ++V AS P
Sbjct: 64 NFTQATLLTKPNST--SFTWYPSRTGSYYFSFTNNTSLPKTCQLNQKLTVQVILAAASPP 121
Query: 124 EASSPMPTTPGPSSGGDIQSSP-SFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSG 182
S P T P P S G + SSP S+PWP P +GS+ P P+P + +TVP
Sbjct: 122 --SQPPATAPVPVSEGGVISSPSSYPWPLGPREGSAFSPGPSPSEITSVTVP------GK 173
Query: 183 DGMPFINSNPAVPLPTGEVDSATIHPLATS 212
DG+PFINSNPAVPLPTG+VDS +I+PL TS
Sbjct: 174 DGVPFINSNPAVPLPTGDVDSTSINPLPTS 203
>UniRef100_Q9LPU4 T22I11.8 [Arabidopsis thaliana]
Length = 233
Score = 215 bits (547), Expect = 8e-55
Identities = 114/214 (53%), Positives = 148/214 (68%), Gaps = 14/214 (6%)
Query: 2 SLPIFFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQKA 61
S+ F++F LSLF + S S T LVDG S WK+PTV GDS++ KHK Y+LYIF+N+ A
Sbjct: 10 SMLFFYFFCFLSLFSRPSLSATFLVDGVSVWKSPTVHTGDSVS-KHKYGYDLYIFRNKDA 68
Query: 62 FNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSL--KACQDSQKLAIKVTPTK 119
FN+CNFTQA LLT P++T S+TW+PSR G +YF+F+N++ K CQ +QKL ++V
Sbjct: 69 FNVCNFTQATLLTKPNST--SFTWYPSRTGSYYFSFTNNTSLPKTCQLNQKLTVQVILAA 126
Query: 120 ASAPEASSPMPTTPGPSSGGDIQSSP-SFPWPFHPHQGSSPGPAPTPEASSPITVPLVPY 178
AS P S P T P P S G + SSP S+PWP P +GS+ P P+P + +TVP
Sbjct: 127 ASPP--SQPPATAPVPVSEGGVISSPSSYPWPLGPREGSAFSPGPSPSEITSVTVP---- 180
Query: 179 KGSGDGMPFINSNPAVPLPTGEVDSATIHPLATS 212
DG+PFINSNPAVPLPTG+VDS +I+PL TS
Sbjct: 181 --GKDGVPFINSNPAVPLPTGDVDSTSINPLPTS 212
>UniRef100_Q67ZY6 Hypothetical protein At1g21090 [Arabidopsis thaliana]
Length = 88
Score = 77.4 bits (189), Expect = 3e-13
Identities = 36/65 (55%), Positives = 45/65 (68%), Gaps = 6/65 (9%)
Query: 148 PWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIH 207
PWP P +GS+ P P+P + +TVP DG+PFINSNPAVPLPTG+VDS +I+
Sbjct: 1 PWPLGPREGSAFSPGPSPSEITSVTVP------GKDGVPFINSNPAVPLPTGDVDSTSIN 54
Query: 208 PLATS 212
PL TS
Sbjct: 55 PLPTS 59
>UniRef100_Q9T076 Early nodulin-like protein 2 precursor [Arabidopsis thaliana]
Length = 349
Score = 61.2 bits (147), Expect = 2e-08
Identities = 63/230 (27%), Positives = 96/230 (41%), Gaps = 32/230 (13%)
Query: 2 SLPIFFYFLI-LSLFFKLSHSTTILVDGSSEW-KNPTVS-----------IGDSITFKHK 48
SL FF L+ LS F +S++ V GS W NP + + D++ F +
Sbjct: 8 SLSFFFTILLSLSTLFTISNARKFNVGGSGAWVTNPPENYESWSGKNRFLVHDTLYFSYA 67
Query: 49 QNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDS 108
+ + + N+ ++ CN D + S R G FYF N+ C+
Sbjct: 68 KGADSVLEVNKADYDACNTKNPIKRVDDGDSEISL----DRYGPFYFISGNED--NCKKG 121
Query: 109 QKLAIKVT----PTKASAPEASSPMPTTPG---PSSGGDI--QSSPSFPWPFHPHQGSSP 159
QKL + V P+ A +P A++P +TPG P G SSP P P + P
Sbjct: 122 QKLNVVVISARIPSTAQSPHAAAPGSSTPGSMTPPGGAHSPKSSSPVSPTTSPPGSTTPP 181
Query: 160 GPAPTPEASSPITVPLVP----YKGSGDGMPFINSNPAVPLPTGEVDSAT 205
G A +P++SS ++ P SG + S PA P T V ++
Sbjct: 182 GGAHSPKSSSAVSPATSPPGSMAPKSGSPVSPTTSPPAPPKSTSPVSPSS 231
Score = 35.4 bits (80), Expect = 1.2
Identities = 26/103 (25%), Positives = 45/103 (43%), Gaps = 2/103 (1%)
Query: 117 PTKASAP--EASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
P K+++P +S+PM + P P + + P P GS + +P ++SP P
Sbjct: 220 PPKSTSPVSPSSAPMTSPPAPMAPKSSSTIPPSSAPMTSPPGSMAPKSSSPVSNSPTVSP 279
Query: 175 LVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQ 217
+ GS P + + + P+G+ SA +G GQ
Sbjct: 280 SLAPGGSTSSSPSDSPSGSAMGPSGDGPSAAGDISTPAGAPGQ 322
>UniRef100_Q94DL5 P0483G10.28 protein [Oryza sativa]
Length = 365
Score = 52.8 bits (125), Expect = 7e-06
Identities = 36/101 (35%), Positives = 47/101 (45%), Gaps = 28/101 (27%)
Query: 123 PEASSPMPTTPGPSSGGDI---------QSSPSFPWPFHPHQG-SSPGPAPTPEASSPIT 172
P S+P P P +G D Q+ P F +P P G ++P APT
Sbjct: 71 PPLSAP---APAPIAGADDLPAFGRAPKQAPPHFGFPLQPTFGVAAPPVAPT-------- 119
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSG 213
+G+G PFI SNP VPLPTG D++T+ PL G
Sbjct: 120 -------AAGEGYPFIGSNPTVPLPTGMTDTSTVLPLPDRG 153
>UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa]
Length = 246
Score = 52.0 bits (123), Expect = 1e-05
Identities = 50/193 (25%), Positives = 76/193 (38%), Gaps = 30/193 (15%)
Query: 23 TILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCS 82
TI VD +S + + +GDS+ FK+ + + + + C AN L S+ S +
Sbjct: 37 TIGVDYTSWAGSKSFKVGDSLVFKYASGAHTVVEVSAAGYLAC--AAANALGSDSSGSTT 94
Query: 83 YTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKASAP--------------EASSP 128
+F T + C K+ + V+ + +S+ SSP
Sbjct: 95 VALKTPGKHYFICTIAGH----CAGGMKMEVDVSGSSSSSSGGGGGGGGGGSTPSSPSSP 150
Query: 129 MPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFI 188
PTTP PS+ PS P P P+ S+P PT +P T P P G G
Sbjct: 151 TPTTPNPSTPTPTTPYPSTPMPTTPYP-STPMTTPT----TPYTTPTSPACSGGAG---- 201
Query: 189 NSNPAVPLPTGEV 201
+ P P+ G V
Sbjct: 202 -ATPVTPVTPGTV 213
>UniRef100_Q9SN46 Extensin-like protein [Arabidopsis thaliana]
Length = 839
Score = 51.6 bits (122), Expect = 2e-05
Identities = 39/106 (36%), Positives = 49/106 (45%), Gaps = 11/106 (10%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TPT +P +S PTTP P GG SSP+ P P G SP P+P+ S PITVP
Sbjct: 487 TPTPGGSPPSS---PTTPTP--GGSPPSSPTTPSP-----GGSP-PSPSISPSPPITVPS 535
Query: 176 VPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVPIL 221
P + G P S+P P + + P S Q PI+
Sbjct: 536 PPSTPTSPGSPPSPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPII 581
Score = 45.8 bits (107), Expect = 9e-04
Identities = 29/85 (34%), Positives = 36/85 (42%), Gaps = 3/85 (3%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEA---SSPIT 172
TPT +P +S P+ G I SP P P +SPG P+P + SSPI
Sbjct: 500 TPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPSTPTSPGSPPSPSSPTPSSPIP 559
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLP 197
P P P NS P +P P
Sbjct: 560 SPPTPSTPPTPISPGQNSPPIIPSP 584
Score = 39.3 bits (90), Expect = 0.083
Identities = 30/100 (30%), Positives = 39/100 (39%), Gaps = 16/100 (16%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPG-------PAPTPEAS 168
TPT +P P P++P PSS PS P P P SPG P+P
Sbjct: 539 TPTSPGSP----PSPSSPTPSS-----PIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGP 589
Query: 169 SPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHP 208
SP + P P P + P+ P P+ +HP
Sbjct: 590 SPPSSPSPPLPPVIPSPPIVGPTPSSPPPSTPTPGTLLHP 629
Score = 37.0 bits (84), Expect = 0.41
Identities = 31/96 (32%), Positives = 39/96 (40%), Gaps = 9/96 (9%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAP---TPEASSPIT 172
+P +P S P P+ PS + S P+ P P S P+P TP SP T
Sbjct: 412 SPPTTPSPGGSPPSPSI-SPSPPITVPSPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPT 470
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHP 208
P P G G P S+P P P G S+ P
Sbjct: 471 SPTTPTPG---GSP--PSSPTTPTPGGSPPSSPTTP 501
>UniRef100_Q6ESY8 Hypothetical protein P0472F10.18 [Oryza sativa]
Length = 213
Score = 51.2 bits (121), Expect = 2e-05
Identities = 30/74 (40%), Positives = 39/74 (52%), Gaps = 10/74 (13%)
Query: 144 SPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDS 203
SPS W +P AP+P+AS+ P G G FI S+PAVP+P G D+
Sbjct: 119 SPSSSW--------APARAPSPDASASAPDSAAP--GQSGGTAFIRSSPAVPVPRGVTDT 168
Query: 204 ATIHPLATSGHQGQ 217
ATI P+ T G + Q
Sbjct: 169 ATILPMPTPGDKHQ 182
>UniRef100_Q7PRG9 ENSANGP00000024130 [Anopheles gambiae str. PEST]
Length = 262
Score = 50.4 bits (119), Expect = 4e-05
Identities = 42/134 (31%), Positives = 57/134 (42%), Gaps = 17/134 (12%)
Query: 99 NDSLKACQDSQKLA-----IKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHP 153
ND++K C + + I T TKAS EA +P PTT P+ +P+ P
Sbjct: 81 NDTIKRCDAPENVQCEDEQIPETTTKASTTEAPTPAPTTEAPT------PAPTTEVPTPA 134
Query: 154 HQGSSPGPAPTPEASSPI--TVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLAT 211
+P PAPT EA +P T P + P ++ P PT S T P
Sbjct: 135 PTTEAPTPAPTTEAPTPAPSTEAPTPAPTTEAPTPAPSTEAPTPAPTTSTPSNTFLPEV- 193
Query: 212 SGHQGQVPILFSHS 225
+G +PILF S
Sbjct: 194 ---EGALPILFPGS 204
>UniRef100_UPI00004574FB UPI00004574FB UniRef100 entry
Length = 293
Score = 49.7 bits (117), Expect = 6e-05
Identities = 32/93 (34%), Positives = 40/93 (42%), Gaps = 13/93 (13%)
Query: 115 VTPTKASAPEAS----SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
+TP SAP + SP P TP P S + P P P P GS+P P P + +P
Sbjct: 91 LTPVPGSAPPLTPAPGSPAPLTPAPGSPAPLTPVPGSPAPLTPVPGSAPPLTPAPGSPAP 150
Query: 171 IT------VPLVPYKGSGDGMPFINSNPAVPLP 197
+T PL P GS P + P P P
Sbjct: 151 LTPAPGSPAPLTPVPGSA---PPLTPAPGSPAP 180
Score = 45.8 bits (107), Expect = 9e-04
Identities = 31/95 (32%), Positives = 42/95 (43%), Gaps = 12/95 (12%)
Query: 115 VTPTKASAPEAS----SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
+TP SAP + S P TP P S + P P P P GS+P P P + +P
Sbjct: 191 LTPVPGSAPPLTPVPGSAPPLTPAPGSPAPLTPVPGSPAPLTPVPGSAPPLTPAPGSPAP 250
Query: 171 IT------VPLVPYKGSGDGMPFINSNPA--VPLP 197
+T PL P GS + + +PA P+P
Sbjct: 251 LTPAPGSPAPLTPAPGSPAPLTPVPGSPAPLTPVP 285
Score = 45.4 bits (106), Expect = 0.001
Identities = 29/91 (31%), Positives = 40/91 (43%), Gaps = 14/91 (15%)
Query: 115 VTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPIT-- 172
+TP SAP P TP P S + +P P P P GS+P P P + +P+T
Sbjct: 131 LTPVPGSAP------PLTPAPGSPAPLTPAPGSPAPLTPVPGSAPPLTPAPGSPAPLTPV 184
Query: 173 ----VPLVPYKGSGDGMPFI--NSNPAVPLP 197
PL P GS + + ++ P P P
Sbjct: 185 PGSPAPLTPVPGSAPPLTPVPGSAPPLTPAP 215
Score = 45.1 bits (105), Expect = 0.002
Identities = 31/93 (33%), Positives = 39/93 (41%), Gaps = 13/93 (13%)
Query: 115 VTPTKASAPEAS----SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
+TP SAP + SP P TP P S + P P P GS+P P P + +P
Sbjct: 161 LTPVPGSAPPLTPAPGSPAPLTPVPGSPAPLTPVPGSAPPLTPVPGSAPPLTPAPGSPAP 220
Query: 171 IT------VPLVPYKGSGDGMPFINSNPAVPLP 197
+T PL P GS P + P P P
Sbjct: 221 LTPVPGSPAPLTPVPGSA---PPLTPAPGSPAP 250
Score = 42.7 bits (99), Expect = 0.008
Identities = 33/91 (36%), Positives = 39/91 (42%), Gaps = 7/91 (7%)
Query: 123 PEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAP-TPEASSPITVPLVPYKGS 181
P SP P TP P S + +P P P P GS PAP TP SP PL P GS
Sbjct: 83 PAPGSPAPLTPVPGSAPPLTPAPGSPAPLTPAPGS---PAPLTPVPGSP--APLTPVPGS 137
Query: 182 GDGMPFINSNPAVPLPTGEVDSATIHPLATS 212
+ +PA PL A + P+ S
Sbjct: 138 APPLTPAPGSPA-PLTPAPGSPAPLTPVPGS 167
Score = 41.6 bits (96), Expect = 0.017
Identities = 26/77 (33%), Positives = 32/77 (40%), Gaps = 10/77 (12%)
Query: 115 VTPTKAS----APEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
+TP S P SP P TP P S + +P P P P GS P P + +P
Sbjct: 211 LTPAPGSPAPLTPVPGSPAPLTPVPGSAPPLTPAPGSPAPLTPAPGSPAPLTPAPGSPAP 270
Query: 171 IT------VPLVPYKGS 181
+T PL P GS
Sbjct: 271 LTPVPGSPAPLTPVPGS 287
Score = 41.2 bits (95), Expect = 0.022
Identities = 22/62 (35%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Query: 115 VTPTKASAPEAS----SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
+TP SAP + SP P TP P S + +P P P P GS P P ++ P
Sbjct: 231 LTPVPGSAPPLTPAPGSPAPLTPAPGSPAPLTPAPGSPAPLTPVPGSPAPLTPVPGSAPP 290
Query: 171 IT 172
+T
Sbjct: 291 LT 292
Score = 34.7 bits (78), Expect = 2.1
Identities = 29/102 (28%), Positives = 38/102 (36%), Gaps = 21/102 (20%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPS-------------FPWPFHPHQGSSPGPAP 163
P+ P+A P TPGP SPS P P P GS P
Sbjct: 34 PSPQRGPQAPFPAARTPGPLPRSAEPRSPSPQRGAQVPFPAARSPAPLTPAPGSPAPLTP 93
Query: 164 TPEASSPIT------VPLVPYKGSGDGMPFINSNPA--VPLP 197
P ++ P+T PL P GS + + +PA P+P
Sbjct: 94 VPGSAPPLTPAPGSPAPLTPAPGSPAPLTPVPGSPAPLTPVP 135
Score = 34.3 bits (77), Expect = 2.7
Identities = 19/53 (35%), Positives = 22/53 (40%), Gaps = 4/53 (7%)
Query: 115 VTPTKAS----APEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAP 163
+TP S P SP P TP P S + P P P P GS+P P
Sbjct: 241 LTPAPGSPAPLTPAPGSPAPLTPAPGSPAPLTPVPGSPAPLTPVPGSAPPLTP 293
>UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana]
Length = 261
Score = 47.8 bits (112), Expect = 2e-04
Identities = 47/185 (25%), Positives = 73/185 (39%), Gaps = 15/185 (8%)
Query: 36 TVSIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYF 95
T ++GD++ F + ++ + + F+ C + L+T + S P G YF
Sbjct: 48 TFAVGDNLVFSYPAAFHDVVEVTKPEFDSCQAVKP-LITFANGNSLVPLTTP---GKRYF 103
Query: 96 TFSNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTT------PGPSSGGDIQSS-PSFP 148
C KL + V PT AP A P+P T P PSS IQ P P
Sbjct: 104 ICGMPG--HCSQGMKLEVNVVPTATVAPTA--PLPNTVPSLNAPSPSSVLPIQPLLPLNP 159
Query: 149 WPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHP 208
P S+P P+ + P++ L P +G +P +P T + P
Sbjct: 160 VPVLSPSSSTPLPSSSLPLIPPLSPALSPATAAGTSLPLFPGSPGSSSSTTSTKTVGTFP 219
Query: 209 LATSG 213
+T+G
Sbjct: 220 SSTTG 224
>UniRef100_P93329 Early nodulin 20 precursor [Medicago truncatula]
Length = 268
Score = 47.8 bits (112), Expect = 2e-04
Identities = 59/246 (23%), Positives = 95/246 (37%), Gaps = 40/246 (16%)
Query: 2 SLPIFFYFLI-LSLFFKLSHSTTILV-DGSSEWKNPTVS--------------IGDSITF 45
S PI F+ + + S ST LV D + WK P + +GD+ITF
Sbjct: 4 SSPILLMFIFSIWMLISYSESTDYLVGDSENSWKFPLPTRHALTRWASNYQFIVGDTITF 63
Query: 46 KHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKAC 105
++ ++ ++ C ++ T + G +F + + C
Sbjct: 64 QYNNKTESVHEVEEEDYDRCGIRGEHVDHYDGNTMVVL----KKTGIHHFI--SGKKRHC 117
Query: 106 QDSQKLAI--KVTPTKASAPEASSPMPTTPGPSSGGDIQSSP--SFPWPFHPHQGSSPGP 161
+ KLA+ V P +S P P P+ P P S I P S P P P SP P
Sbjct: 118 RLGLKLAVVVMVAPVLSSPP----PPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSP 173
Query: 162 APTPE-ASSPITVPLVPYKGSGDGMPFINSNPA-------VPLPTGEVDSATIHPLATSG 213
+P+P S+PI P S P ++ +P+ P P+ V A++ P ++
Sbjct: 174 SPSPSPRSTPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSPSDSV--ASLAPSSSPS 231
Query: 214 HQGQVP 219
+ P
Sbjct: 232 DESPSP 237
>UniRef100_Q653Y0 Putative phytocyanin protein, PUP2 [Oryza sativa]
Length = 279
Score = 46.6 bits (109), Expect = 5e-04
Identities = 53/234 (22%), Positives = 80/234 (33%), Gaps = 45/234 (19%)
Query: 12 LSLFFKLSHSTTILVDGSSEWKNPTVS--------------IGDSITFKHKQNYNLYIFK 57
L L + +T V G W P + IGD + F + + + +
Sbjct: 15 LGLMAAAASATQFRVGGGRGWSVPDANAEPYNSWAGRMRFQIGDQLLFVYPKEMDAVVVV 74
Query: 58 NQKAFNLCNFTQANL-----LTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLA 112
+Q A++ CN + + D T ++ R G F+F N++ C+ +KL
Sbjct: 75 DQGAYDACNTSSSVAGGGGGRYDDGNTVFTF----DRSGPFFFISGNEA--NCRAGEKLV 128
Query: 113 IKVTPTKA-------SAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP 165
+ V + S P P+ P PS SSP P P +P P T
Sbjct: 129 VVVMADRGGRHAPPPSPPAVPPPVAPVPMPSPA----SSPPSPAPAAATPSLAPSPVATT 184
Query: 166 EASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
+ SP P+ P P PA P+ P T G Q P
Sbjct: 185 PSPSPSVSPMAPAPAPTTSTPSSPPAPAAMAPS---------PSTTPGGVAQPP 229
>UniRef100_Q6K4A5 Hypothetical protein OJ1595_D08.12-1 [Oryza sativa]
Length = 131
Score = 45.8 bits (107), Expect = 9e-04
Identities = 40/118 (33%), Positives = 53/118 (44%), Gaps = 16/118 (13%)
Query: 102 LKACQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGP 161
L A +Q A TP ++P ++P P TP P S Q+ P+ P P + P
Sbjct: 4 LFAAASAQAPAATPTPAPKASPPPATP-PPTPAPVSAPPAQA-PATPPPAPAPAPKASAP 61
Query: 162 APTPEASSPITVPLVPYKGSGDGMPFINSNPA------VPLPTGEVDSATIHPLATSG 213
AP P+AS+P VP + P I+S PA P PT EV T P A +G
Sbjct: 62 APAPKASAPAPVP-----AAAAPTPEISSPPAPSPAGLAPSPTAEV---TPPPSAAAG 111
>UniRef100_Q9LT74 Similarity to late embryogenesis abundant protein [Arabidopsis
thaliana]
Length = 631
Score = 45.8 bits (107), Expect = 9e-04
Identities = 28/80 (35%), Positives = 31/80 (38%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TP+ S SP P TP PS P+ P P P S P P PTP SP V
Sbjct: 213 TPSVPSPTPPVSPPPPTPTPSVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVPSPPDVTP 272
Query: 176 VPYKGSGDGMPFINSNPAVP 195
P S P + P P
Sbjct: 273 TPPTPSVPSPPDVTPTPPTP 292
Score = 42.0 bits (97), Expect = 0.013
Identities = 29/88 (32%), Positives = 36/88 (39%), Gaps = 5/88 (5%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TPT S P + P+PT P PS + P P P P PTP SP V
Sbjct: 229 TPTP-SVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVPSPPDVTPTPPTPSVPSPPDVTP 287
Query: 176 VPYKGSGDGMPFIN----SNPAVPLPTG 199
P S P + + P+VP P+G
Sbjct: 288 TPPTPSVPSPPDVTPTPPTPPSVPTPSG 315
Score = 39.7 bits (91), Expect = 0.064
Identities = 31/89 (34%), Positives = 39/89 (42%), Gaps = 14/89 (15%)
Query: 116 TPTKASAPEASSPMPTTPGPS---SGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPIT 172
+PT +P +P P+ P P+ S +PS P P P S P P PTP SP T
Sbjct: 164 SPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPP--VSPPPPTPTPSVPSP-T 220
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEV 201
P+ P P P+VP PT V
Sbjct: 221 PPVSP--------PPPTPTPSVPSPTPPV 241
Score = 38.5 bits (88), Expect = 0.14
Identities = 28/86 (32%), Positives = 37/86 (42%), Gaps = 7/86 (8%)
Query: 116 TPTKASAPEASSPMPTTPGPS---SGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPIT 172
+PT +P +P P+ P P+ S +PS P P P SP P PTP S P
Sbjct: 182 SPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPP---VSP-PPPTPTPSVPSP 237
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPT 198
P VP P ++ P P P+
Sbjct: 238 TPPVPTDPMPSPPPPVSPPPPTPTPS 263
Score = 37.4 bits (85), Expect = 0.32
Identities = 30/91 (32%), Positives = 32/91 (34%), Gaps = 5/91 (5%)
Query: 120 ASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQG---SSPGPAPTPEASSPI--TVP 174
A P S P PT PS + P P P P S P P PTP SP P
Sbjct: 148 APVPPVSPPPPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSP 207
Query: 175 LVPYKGSGDGMPFINSNPAVPLPTGEVDSAT 205
P P +P P PT V S T
Sbjct: 208 PPPTPTPSVPSPTPPVSPPPPTPTPSVPSPT 238
Score = 32.7 bits (73), Expect = 7.8
Identities = 32/111 (28%), Positives = 41/111 (36%), Gaps = 12/111 (10%)
Query: 103 KACQDSQKLAIKVTPTKASAPEAS-SPM--PTTPGPSSGGDIQSS-------PSFPWPFH 152
K C + + + K P AS P+ P +PG GGD P+ P
Sbjct: 95 KHCYNLEHVCPKFCPDSCHVECASCKPICGPPSPGDDGGGDDSGGDDGGYTPPAPVPPVS 154
Query: 153 PHQGSSPGPAPTPEASSP--ITVPLVPYKGSGDGMPFINSNPAVPLPTGEV 201
P + P+PTP S P P VP P P+VP PT V
Sbjct: 155 PPPPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPV 205
Score = 32.7 bits (73), Expect = 7.8
Identities = 31/89 (34%), Positives = 40/89 (44%), Gaps = 19/89 (21%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TP+ S P+ + P P TP S D+ +P P P P S P PTP T P
Sbjct: 261 TPSVPSPPDVT-PTPPTPSVPSPPDVTPTP--PTPSVP---SPPDVTPTPP-----TPPS 309
Query: 176 VPYKGSGDGMPFINSNPAVPLPTGEVDSA 204
VP + G P P VP P+ E ++A
Sbjct: 310 VP---TPSGSP-----PYVPPPSDEEEAA 330
>UniRef100_P08640 Glucoamylase S1/S2 precursor [Saccharomyces cerevisiae]
Length = 1367
Score = 45.4 bits (106), Expect = 0.001
Identities = 39/120 (32%), Positives = 52/120 (42%), Gaps = 16/120 (13%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TP+ ++ +S+P+PT PSS SS P P SS P PTP +SS IT
Sbjct: 766 TPSSSTTESSSAPVPT---PSSSTTESSSAPVPTPSSSTTESSVAPVPTPSSSSNIT--- 819
Query: 176 VPYKGSGDGMPFINS--NPAVPLP-----TGEVDSATIHPLATSGHQGQVPILFSHSCFT 228
+ PF +S + +VP+P T E SA + T VP S S T
Sbjct: 820 ---SSAPSSTPFSSSTESSSVPVPTPSSSTTESSSAPVSSSTTESSVAPVPTPSSSSNIT 876
Score = 37.7 bits (86), Expect = 0.24
Identities = 30/97 (30%), Positives = 44/97 (44%), Gaps = 9/97 (9%)
Query: 116 TPTKASAPEASS-PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
TP +S E+SS P+PT PSS SS P P SS PAPTP +S+ +
Sbjct: 567 TPVTSSTTESSSAPVPT---PSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSS 623
Query: 175 LVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLAT 211
+ + +S+ VP P+ ++ P+ T
Sbjct: 624 APVTSSTTE-----SSSAPVPTPSSSTTESSSAPVPT 655
Score = 35.4 bits (80), Expect = 1.2
Identities = 27/101 (26%), Positives = 42/101 (40%), Gaps = 10/101 (9%)
Query: 118 TKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASS--PITVPL 175
T ++ +S+P+PT PSS SS P P SS P PTP +S+ + P+
Sbjct: 627 TSSTTESSSAPVPT---PSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPV 683
Query: 176 VPYKGSGDGMPFI-----NSNPAVPLPTGEVDSATIHPLAT 211
P +S+ VP P+ ++ P+ T
Sbjct: 684 TSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPT 724
Score = 34.3 bits (77), Expect = 2.7
Identities = 28/96 (29%), Positives = 42/96 (43%), Gaps = 9/96 (9%)
Query: 117 PTKASAPEASS-PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
P +S E+SS P+PT PSS SS P P SS P PTP +S+ +
Sbjct: 694 PVTSSTTESSSAPVPT---PSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSA 750
Query: 176 VPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLAT 211
+ + +S+ VP P+ ++ P+ T
Sbjct: 751 PVTSSTTE-----SSSAPVPTPSSSTTESSSAPVPT 781
Score = 33.9 bits (76), Expect = 3.5
Identities = 36/153 (23%), Positives = 51/153 (32%), Gaps = 11/153 (7%)
Query: 76 PSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTPGP 135
P T S T S T + S + + + PT +S+ SS P P P
Sbjct: 667 PVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAP-VPTP 725
Query: 136 SSGGDIQSSPSFPWPFHPHQGSSPGP----------APTPEASSPITVPLVPYKGSGDGM 185
SS SS P P SS P AP P SS T +
Sbjct: 726 SSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSS 785
Query: 186 PFINSNPAVPLPTGEVDSATIHPLATSGHQGQV 218
+S+ VP P+ +++ P+ T +
Sbjct: 786 TTESSSAPVPTPSSSTTESSVAPVPTPSSSSNI 818
>UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis thaliana]
Length = 222
Score = 45.1 bits (105), Expect = 0.002
Identities = 49/189 (25%), Positives = 77/189 (39%), Gaps = 21/189 (11%)
Query: 36 TVSIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYF 95
T +GD++ F + ++++ + ++ ++ C+ + A T T VG +F
Sbjct: 46 TFRVGDTLEFVYGLSHSVSVV-DKAGYDNCDSSGATQNFADGDTKIDLT----TVGTMHF 100
Query: 96 ---TFSNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFH 152
TF + C++ KLA+ V S SSP T PSS S+PS P
Sbjct: 101 LCPTFGH-----CKNGMKLAVPVLAAAPSPSTPSSPPSTPSTPSSPPSTPSTPSSP---- 151
Query: 153 PHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATS 212
P S P P+ P + P P P G+ D + P PLP +A + +
Sbjct: 152 PSPPSPPSPSLPPSSLPPSASP--PTNGTPDSETL--TPPPAPLPPSLSPNAASKGVMSY 207
Query: 213 GHQGQVPIL 221
G G IL
Sbjct: 208 GIIGVTMIL 216
>UniRef100_Q84QK1 Putative arabinagalactan protein [Lotus japonicus]
Length = 149
Score = 45.1 bits (105), Expect = 0.002
Identities = 32/84 (38%), Positives = 41/84 (48%), Gaps = 7/84 (8%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASS-PITVPL 175
P A+ P A++P PTT P++ +SP P P +SP APTP ASS P +P
Sbjct: 46 PPPATPPPAATPAPTTTPPAATPAPSASPPAPTP-----TASPTGAPTPSASSPPAPIPS 100
Query: 176 VPYKGSGDGM-PFINSNPAVPLPT 198
P G G P NS P P+
Sbjct: 101 GPASGPGPAAGPGPNSADTPPPPS 124
Score = 33.1 bits (74), Expect = 6.0
Identities = 27/91 (29%), Positives = 37/91 (39%), Gaps = 3/91 (3%)
Query: 111 LAIKVTPTKASAPEASSPMP--TTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEAS 168
L + T A AP A+ TTP P P+ P P ++ PA TP S
Sbjct: 12 LGLLATSCVAQAPGAAPTQAPTTTPPPPPAAAPAPPPATPPPAATPAPTTTPPAATPAPS 71
Query: 169 SPITVPLVPYKGSGDGMPFINSNPAVPLPTG 199
+ P +G P +S PA P+P+G
Sbjct: 72 ASPPAPTPTASPTGAPTPSASSPPA-PIPSG 101
>UniRef100_Q9GTI8 Hypothetical esophageal gland cell secretory protein 10 [Heterodera
glycines]
Length = 196
Score = 45.1 bits (105), Expect = 0.002
Identities = 34/106 (32%), Positives = 41/106 (38%), Gaps = 5/106 (4%)
Query: 116 TPTKASAPEASSPMPT-TPGPSSGGDIQS-SPSFPWPFHPHQGSSPGPAPTPEASSPITV 173
T P S+P PT TPGPS+ + PS P P S+P P TP S+P
Sbjct: 85 TTAPTGTPGPSTPAPTGTPGPSTPAPTGTPGPSTPTPTGTPGPSTPAPTGTPGPSTPAPT 144
Query: 174 PLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
G P P+ P PTG +T P T G P
Sbjct: 145 GT---PGPXTPAPTGTPGPSTPAPTGTPGPSTPAPTGTPGPSTPAP 187
Score = 40.4 bits (93), Expect = 0.037
Identities = 29/86 (33%), Positives = 35/86 (39%), Gaps = 16/86 (18%)
Query: 116 TPTKASAPEASSPMPT-TPGPSSGGDIQS-SPSFPWPFHPHQGSSPGPAPTPEASSPITV 173
TPT P S+P PT TPGPS+ + P P P S+P P TP S+P
Sbjct: 118 TPTPTGTPGPSTPAPTGTPGPSTPAPTGTPGPXTPAPTGTPGPSTPAPTGTPGPSTP--- 174
Query: 174 PLVPYKGSGDGMPFINSNPAVPLPTG 199
P P+ P PTG
Sbjct: 175 -----------APTGTPGPSTPAPTG 189
Score = 39.7 bits (91), Expect = 0.064
Identities = 32/105 (30%), Positives = 37/105 (34%), Gaps = 14/105 (13%)
Query: 116 TPTKASAPEASSPMPT-TPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
TP P S+ PT TPGPS+ P P S+P P TP S+P
Sbjct: 74 TPAPTGTPGQSTTAPTGTPGPST----------PAPTGTPGPSTPAPTGTPGPSTPTPTG 123
Query: 175 LVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
G P P+ P PTG T P T G P
Sbjct: 124 T---PGPSTPAPTGTPGPSTPAPTGTPGPXTPAPTGTPGPSTPAP 165
Score = 35.4 bits (80), Expect = 1.2
Identities = 32/112 (28%), Positives = 37/112 (32%), Gaps = 13/112 (11%)
Query: 118 TKASAPEASSPMPTTP----------GPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEA 167
T S P SS PTT P + G S P P P S+ P TP
Sbjct: 35 TCESTPGVSSTAPTTTVGPTGTTTTGTPLTTGQGTSGPXTPAPTGTPGQSTTAPTGTPGP 94
Query: 168 SSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
S+P G P P+ P PTG +T P T G P
Sbjct: 95 STPAPTGT---PGPSTPAPTGTPGPSTPTPTGTPGPSTPAPTGTPGPSTPAP 143
Score = 34.7 bits (78), Expect = 2.1
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 2/57 (3%)
Query: 116 TPTKASAPEASSPMPT-TPGPSSGGDIQS-SPSFPWPFHPHQGSSPGPAPTPEASSP 170
TP P +P PT TPGPS+ + PS P P S+P P TP +P
Sbjct: 140 TPAPTGTPGPXTPAPTGTPGPSTPAPTGTPGPSTPAPTGTPGPSTPAPTGTPGPXTP 196
>UniRef100_Q6C159 Similarity [Yarrowia lipolytica]
Length = 823
Score = 45.1 bits (105), Expect = 0.002
Identities = 31/102 (30%), Positives = 44/102 (42%), Gaps = 7/102 (6%)
Query: 105 CQDSQKLAIKVTPTKASAPEASSPMPTTPGPSS-GGDIQSSP-----SFPWPFHPHQGSS 158
C + + + T A PE+S+P P P S G + +P S P P + S+
Sbjct: 588 CDTTTEAPVPETSAPAPKPESSAPPAPAPAPESPAGKPEPTPAPAPSSAPAPAPKPESSA 647
Query: 159 PGPAPTPEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGE 200
P PAP PE+S+P P P + P S+ P P E
Sbjct: 648 PAPAPKPESSAPAPAP-KPESSAPAPAPKPESSAPAPAPKPE 688
Score = 43.5 bits (101), Expect = 0.004
Identities = 31/97 (31%), Positives = 39/97 (39%), Gaps = 4/97 (4%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSP--SFPWPFHPHQGSSPGPAPTPEASSPITVP 174
P A PE+S+P P SS P S P P + S+P PAP PE S+P P
Sbjct: 648 PAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAPKPETSAPAPAP 707
Query: 175 LVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLAT 211
+S P P P E S + P +T
Sbjct: 708 KPEESAPAPAPKPESSAPPAPAPAPE--SPAVKPEST 742
Score = 40.8 bits (94), Expect = 0.029
Identities = 23/60 (38%), Positives = 29/60 (48%), Gaps = 2/60 (3%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSP--SFPWPFHPHQGSSPGPAPTPEASSPITVP 174
P A PE+S+P P SS P S P P + S+P PAP PE+S+P P
Sbjct: 637 PAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAP 696
Score = 40.4 bits (93), Expect = 0.037
Identities = 29/90 (32%), Positives = 40/90 (44%), Gaps = 6/90 (6%)
Query: 114 KVTPTKASAPE---ASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
K PT A AP A +P P + P+ +SS P P + S+P PAP PE+S+P
Sbjct: 624 KPEPTPAPAPSSAPAPAPKPESSAPAPAPKPESSAPAPAP--KPESSAPAPAPKPESSAP 681
Query: 171 ITVPLVPYKGSGDGMPFINSNPAVPLPTGE 200
P P + P ++ P P E
Sbjct: 682 APAP-KPESSAPAPAPKPETSAPAPAPKPE 710
Score = 38.5 bits (88), Expect = 0.14
Identities = 22/59 (37%), Positives = 26/59 (43%), Gaps = 5/59 (8%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFH-----PHQGSSPGPAPTPEASSP 170
P A PE+S+P P P S S P P P + P PAP PEAS+P
Sbjct: 714 PAPAPKPESSAPPAPAPAPESPAVKPESTPAPAPAPAPAPAPESSAPPAPAPKPEASAP 772
Score = 34.3 bits (77), Expect = 2.7
Identities = 29/104 (27%), Positives = 35/104 (32%), Gaps = 8/104 (7%)
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
T T+A PE S+P P P S S+P P P P P P P SS
Sbjct: 590 TTTEAPVPETSAPAPK---PES-----SAPPAPAPAPESPAGKPEPTPAPAPSSAPAPAP 641
Query: 176 VPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
P + P S+ P P E + P S P
Sbjct: 642 KPESSAPAPAPKPESSAPAPAPKPESSAPAPAPKPESSAPAPAP 685
Score = 32.7 bits (73), Expect = 7.8
Identities = 20/62 (32%), Positives = 26/62 (41%), Gaps = 2/62 (3%)
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWP-FHPHQGSSPGPAPTPEASSPITVPL 175
P A PE S+P P P P S +P+ P P +P PAP P + + P
Sbjct: 703 PAPAPKPEESAPAPA-PKPESSAPPAPAPAPESPAVKPESTPAPAPAPAPAPAPESSAPP 761
Query: 176 VP 177
P
Sbjct: 762 AP 763
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 456,922,103
Number of Sequences: 2790947
Number of extensions: 22605585
Number of successful extensions: 134849
Number of sequences better than 10.0: 1989
Number of HSP's better than 10.0 without gapping: 240
Number of HSP's successfully gapped in prelim test: 1890
Number of HSP's that attempted gapping in prelim test: 120958
Number of HSP's gapped (non-prelim): 9074
length of query: 233
length of database: 848,049,833
effective HSP length: 123
effective length of query: 110
effective length of database: 504,763,352
effective search space: 55523968720
effective search space used: 55523968720
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 73 (32.7 bits)
Medicago: description of AC126014.9


