

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.4 + phase: 0
(91 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
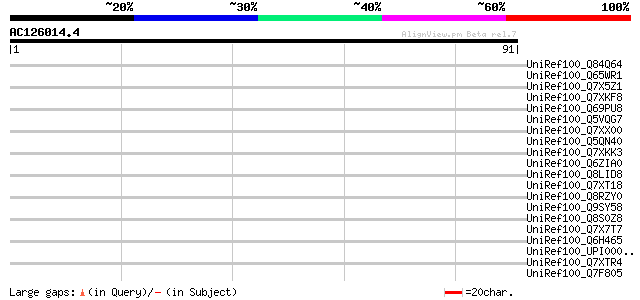
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84Q64 Hypothetical protein OSJNBa0071M09.16 [Oryza sa... 44 0.001
UniRef100_Q65WR1 Putative polyprotein [Oryza sativa] 44 0.001
UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa] 44 0.001
UniRef100_Q7XKF8 OSJNBb0065J09.11 protein [Oryza sativa] 44 0.001
UniRef100_Q69PU8 Hypothetical protein OSJNBa0091L05.16 [Oryza sa... 42 0.004
UniRef100_Q5VQG7 Hypothetical protein OJ1276_B06.35 [Oryza sativa] 40 0.014
UniRef100_Q7XX00 OSJNBa0079F16.5 protein [Oryza sativa] 39 0.031
UniRef100_Q5QN40 Hypothetical protein P0489A05.4 [Oryza sativa] 37 0.15
UniRef100_Q7XKK3 OSJNBa0038O10.4 protein [Oryza sativa] 36 0.20
UniRef100_Q6ZIA0 Cyst nematode resistance protein-like [Oryza sa... 36 0.26
UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [... 35 0.59
UniRef100_Q7XT18 OSJNBb0050O03.4 protein [Oryza sativa] 35 0.59
UniRef100_Q8RZY0 Similar to phosphatidic acid phosphatase beta [... 34 0.77
UniRef100_Q9SY58 F14N23.4 [Arabidopsis thaliana] 33 1.7
UniRef100_Q8S0Z8 P0445E10.23 protein [Oryza sativa] 33 2.2
UniRef100_Q7X7T7 OSJNBa0084K20.4 protein [Oryza sativa] 33 2.2
UniRef100_Q6H465 Hypothetical protein B1250G12.23 [Oryza sativa] 33 2.2
UniRef100_UPI0000281C5A UPI0000281C5A UniRef100 entry 32 2.9
UniRef100_Q7XTR4 OSJNBb0085C12.5 protein [Oryza sativa] 32 2.9
UniRef100_Q7F805 Oryza sativa (japonica cultivar-group) genomic ... 32 2.9
>UniRef100_Q84Q64 Hypothetical protein OSJNBa0071M09.16 [Oryza sativa]
Length = 1143
Score = 43.9 bits (102), Expect = 0.001
Identities = 23/57 (40%), Positives = 30/57 (52%)
Query: 11 NLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
+LA P L+ + VW IWKE N RIF+HK G+LL +K + LW A
Sbjct: 1074 SLANTPGIPKKGLRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIKEEASLWALA 1130
>UniRef100_Q65WR1 Putative polyprotein [Oryza sativa]
Length = 1023
Score = 43.9 bits (102), Expect = 0.001
Identities = 23/57 (40%), Positives = 30/57 (52%)
Query: 11 NLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
+LA P L+ + VW IWKE N RIF+HK G+LL +K + LW A
Sbjct: 954 SLANTPGIPKKGLRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIKEEASLWALA 1010
>UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa]
Length = 1189
Score = 43.9 bits (102), Expect = 0.001
Identities = 23/57 (40%), Positives = 30/57 (52%)
Query: 11 NLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
+LA P L+ + VW IWKE N RIF+HK G+LL +K + LW A
Sbjct: 1120 SLANTPGIPKKGLRSLILLVVWEIWKERNRRIFDHKEMATGVLLAKIKEEASLWALA 1176
>UniRef100_Q7XKF8 OSJNBb0065J09.11 protein [Oryza sativa]
Length = 436
Score = 43.5 bits (101), Expect = 0.001
Identities = 23/57 (40%), Positives = 31/57 (54%)
Query: 11 NLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
+LA P L+ + VW IWKE N RIF++K VGLLL +K + +W A
Sbjct: 367 SLANTPNIPKKGLRSLILLVVWEIWKERNRRIFDNKEMAVGLLLAKIKEEASVWALA 423
>UniRef100_Q69PU8 Hypothetical protein OSJNBa0091L05.16 [Oryza sativa]
Length = 76
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/55 (38%), Positives = 33/55 (59%), Gaps = 1/55 (1%)
Query: 17 RSTHTFLQVIWHACVWL-IWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKANKL 70
RS L+ + CV +WKETN+R+F+++ K VGLL ++K +W +A L
Sbjct: 17 RSRRGSLRYLGSLCVLDDVWKETNARVFHNREKSVGLLFGAIKEEVIIWKEAGVL 71
>UniRef100_Q5VQG7 Hypothetical protein OJ1276_B06.35 [Oryza sativa]
Length = 76
Score = 40.0 bits (92), Expect = 0.014
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Query: 11 NLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
N+ G P+ L ++ VW IWKE N RI++HK LL +K LW+ A
Sbjct: 10 NMCGVPKKGLRSLILL---VVWEIWKERNRRIYDHKEMATSFLLTKIKEEVGLWVLA 63
>UniRef100_Q7XX00 OSJNBa0079F16.5 protein [Oryza sativa]
Length = 153
Score = 38.9 bits (89), Expect = 0.031
Identities = 13/36 (36%), Positives = 23/36 (63%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
W++WKE N+R+F ++ + G L S+K +W +A
Sbjct: 106 WMVWKERNARVFQNQRRSAGFLFGSIKEEVAIWKEA 141
>UniRef100_Q5QN40 Hypothetical protein P0489A05.4 [Oryza sativa]
Length = 247
Score = 36.6 bits (83), Expect = 0.15
Identities = 15/40 (37%), Positives = 22/40 (54%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKANKLT 71
W +WKE NSR+F+ V ++L+S+ LW A T
Sbjct: 40 WQLWKERNSRVFDSALSSVSVVLESIHSEGHLWSLAGIAT 79
>UniRef100_Q7XKK3 OSJNBa0038O10.4 protein [Oryza sativa]
Length = 1045
Score = 36.2 bits (82), Expect = 0.20
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 3/66 (4%)
Query: 2 VRDNLHRFGNLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTS 61
V+D +++G P+ L+ + V +WKE N RIF+HK LL +K +
Sbjct: 970 VKDWWEMLASVSGVPKKG---LRTLILLIVLEVWKERNRRIFDHKEAATSYLLSKIKEEA 1026
Query: 62 FLWLKA 67
+W A
Sbjct: 1027 GMWALA 1032
>UniRef100_Q6ZIA0 Cyst nematode resistance protein-like [Oryza sativa]
Length = 158
Score = 35.8 bits (81), Expect = 0.26
Identities = 14/36 (38%), Positives = 21/36 (57%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
W IWKE N+RIF H +K L +++ +W +A
Sbjct: 116 WFIWKERNARIFEHVSKSATQLFWAIREEIIIWREA 151
>UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [Oryza sativa]
Length = 210
Score = 34.7 bits (78), Expect = 0.59
Identities = 14/36 (38%), Positives = 21/36 (57%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
WLIWKE N+RIF + + L++ +K +W A
Sbjct: 166 WLIWKERNARIFEQRLRSPEQLVEDIKEEINVWRSA 201
>UniRef100_Q7XT18 OSJNBb0050O03.4 protein [Oryza sativa]
Length = 1199
Score = 34.7 bits (78), Expect = 0.59
Identities = 15/53 (28%), Positives = 27/53 (50%)
Query: 16 PRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKAN 68
P H+ + W +WKE NS++F+ V ++L+S++ LW A+
Sbjct: 1117 PEHLHSGFDSLVLLVSWQLWKERNSKVFDSALASVVVVLESIRSEGQLWSLAS 1169
>UniRef100_Q8RZY0 Similar to phosphatidic acid phosphatase beta [Oryza sativa]
Length = 328
Score = 34.3 bits (77), Expect = 0.77
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 3/66 (4%)
Query: 2 VRDNLHRFGNLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTS 61
V+D +++G P+ L ++ VW +WK N RIF++K LL +K +
Sbjct: 253 VKDWWKILASVSGVPKKGLCTLILL---IVWEVWKVRNRRIFDYKEATTSYLLSKIKEEA 309
Query: 62 FLWLKA 67
+W A
Sbjct: 310 GMWALA 315
>UniRef100_Q9SY58 F14N23.4 [Arabidopsis thaliana]
Length = 1161
Score = 33.1 bits (74), Expect = 1.7
Identities = 13/47 (27%), Positives = 27/47 (56%)
Query: 13 AGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKL 59
A R+ ++++HA V+ IW+E N RI ++ + L++ ++L
Sbjct: 1077 ASRDRNITLITKLLFHASVYFIWRERNLRIHSNSVRPAHLIIKEIQL 1123
>UniRef100_Q8S0Z8 P0445E10.23 protein [Oryza sativa]
Length = 484
Score = 32.7 bits (73), Expect = 2.2
Identities = 12/27 (44%), Positives = 17/27 (62%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVK 58
WL+WKE N+R+F+ K LL +K
Sbjct: 398 WLVWKERNARVFDQKFNTADLLSADIK 424
>UniRef100_Q7X7T7 OSJNBa0084K20.4 protein [Oryza sativa]
Length = 181
Score = 32.7 bits (73), Expect = 2.2
Identities = 10/33 (30%), Positives = 19/33 (57%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLW 64
W +WKE NSR+F+ + ++L+ + +W
Sbjct: 135 WRLWKERNSRVFDSALSSISVVLEFIHSEGHMW 167
>UniRef100_Q6H465 Hypothetical protein B1250G12.23 [Oryza sativa]
Length = 222
Score = 32.7 bits (73), Expect = 2.2
Identities = 13/37 (35%), Positives = 18/37 (48%)
Query: 31 VWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKA 67
VW IWKE RIF H + + +K ++ W A
Sbjct: 175 VWEIWKERYQRIFEHNEATLSFIFAKIKEEAWTWATA 211
>UniRef100_UPI0000281C5A UPI0000281C5A UniRef100 entry
Length = 217
Score = 32.3 bits (72), Expect = 2.9
Identities = 16/41 (39%), Positives = 20/41 (48%)
Query: 34 IWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKANKLTSAF 74
IWK S I N K + LDS+ S +WL +TS F
Sbjct: 26 IWKSNRSIIINSKNNQQDSHLDSINFESLIWLVFLGITSYF 66
>UniRef100_Q7XTR4 OSJNBb0085C12.5 protein [Oryza sativa]
Length = 1715
Score = 32.3 bits (72), Expect = 2.9
Identities = 16/53 (30%), Positives = 23/53 (43%)
Query: 16 PRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLKAN 68
P S I W IWKE N+R+FN K + + ++ + LW N
Sbjct: 1644 PSSQRRGFDTIATLAAWTIWKEWNNRVFNQKQRSWTEVARAMAEEAALWRLTN 1696
>UniRef100_Q7F805 Oryza sativa (japonica cultivar-group) genomic DNA, chromosome 1,
PAC clone:P0433F09 [Oryza sativa]
Length = 985
Score = 32.3 bits (72), Expect = 2.9
Identities = 16/61 (26%), Positives = 26/61 (42%)
Query: 7 HRFGNLAGAPRSTHTFLQVIWHACVWLIWKETNSRIFNHKTKEVGLLLDSVKLTSFLWLK 66
H + + +P + L+ + W IW E N+RIF +L +K W+K
Sbjct: 910 HWWNLVVKSPNTPRAPLRTLLMLVTWEIWGERNARIFRSTASTPASILAKIKEEGRAWVK 969
Query: 67 A 67
A
Sbjct: 970 A 970
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.137 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 157,851,286
Number of Sequences: 2790947
Number of extensions: 4963128
Number of successful extensions: 13528
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 13500
Number of HSP's gapped (non-prelim): 35
length of query: 91
length of database: 848,049,833
effective HSP length: 67
effective length of query: 24
effective length of database: 661,056,384
effective search space: 15865353216
effective search space used: 15865353216
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC126014.4


