

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.1 + phase: 0
(405 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
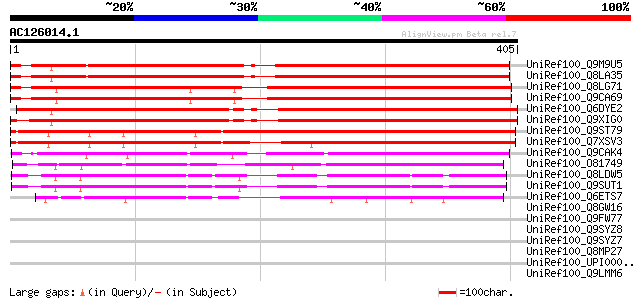
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M9U5 F6A14.15 protein [Arabidopsis thaliana] 479 e-134
UniRef100_Q8LA35 Hypothetical protein [Arabidopsis thaliana] 478 e-133
UniRef100_Q8LG71 Hypothetical protein [Arabidopsis thaliana] 475 e-132
UniRef100_Q9CA69 Hypothetical protein F1M20.13 [Arabidopsis thal... 474 e-132
UniRef100_Q6DYE2 Hypothetical protein [Arabidopsis thaliana] 462 e-128
UniRef100_Q9XIG0 T10P12.8 protein [Arabidopsis thaliana] 461 e-128
UniRef100_Q9ST79 CAA303718.1 protein [Oryza sativa] 407 e-112
UniRef100_Q7XSV3 OSJNBa0039K24.3 protein [Oryza sativa] 342 8e-93
UniRef100_Q9CAK4 Hypothetical protein T12P18.5 [Arabidopsis thal... 268 2e-70
UniRef100_O81749 Hypothetical protein F16G20.230 [Arabidopsis th... 227 5e-58
UniRef100_Q8LDW5 Hypothetical protein [Arabidopsis thaliana] 221 4e-56
UniRef100_Q9SUT1 Hypothetical protein AT4g11300 [Arabidopsis tha... 220 5e-56
UniRef100_Q6ETS7 Hypothetical protein P0017C12.31 [Oryza sativa] 172 2e-41
UniRef100_Q8GW16 Hypothetical protein [Arabidopsis thaliana] 43 0.018
UniRef100_Q9FW77 Hypothetical protein OSJNBa0026L12.27 [Oryza sa... 42 0.024
UniRef100_Q9SYZ8 Hypothetical protein AT4g34330 [Arabidopsis tha... 40 0.090
UniRef100_Q9SYZ7 Hypothetical protein AT4g34320 [Arabidopsis tha... 38 0.45
UniRef100_Q8MP27 Hypothetical protein [Dictyostelium discoideum] 38 0.58
UniRef100_UPI00004367B7 UPI00004367B7 UniRef100 entry 37 0.99
UniRef100_Q9LMM6 Hypothetical protein F22L4.9 [Arabidopsis thali... 37 0.99
>UniRef100_Q9M9U5 F6A14.15 protein [Arabidopsis thaliana]
Length = 382
Score = 479 bits (1234), Expect = e-134
Identities = 243/410 (59%), Positives = 315/410 (76%), Gaps = 39/410 (9%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSM----------EGSTLEIELDSFQQHVTD 50
MPATD+QGS FGRS+LS RR+Q+ S E ST+E+ELDSFQ+ V +
Sbjct: 1 MPATDFQGS-------FGRSLLSLRRDQVDSSTVVSGSSSHHEPSTMEVELDSFQRQVAE 53
Query: 51 RFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERS 110
+F+DL++ +D LLSL+W+GK+LD FL CQEEF+AI+ H+ Q+ + P+DR++S+YFERS
Sbjct: 54 KFIDLNASSND-LLSLEWIGKLLDSFLCCQEEFRAIVFNHRSQISKSPMDRLISDYFERS 112
Query: 111 VKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDG 170
+KALDVCNAIRDG+EQIR W+KL +IV+ ALD R IGEGQ RRAKKALIDL+I MLD+
Sbjct: 113 IKALDVCNAIRDGIEQIRQWEKLADIVISALDSHRPIGEGQLRRAKKALIDLAIGMLDEK 172
Query: 171 KE-SNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTW 229
S ++AHRNRSFGR +D H HRS+G FRSLSWSVSR+W
Sbjct: 173 DHPSGTNLAHRNRSFGRV-----KDSH---------------HRSIGHFRSLSWSVSRSW 212
Query: 230 SAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHF 289
SA++QLQA+ +N+ PR ND++A+NGLA+ VYTM SVLLFVMW LVAAIPCQDRGL V+F
Sbjct: 213 SASKQLQALASNLATPRPNDVVASNGLAVPVYTMTSVLLFVMWVLVAAIPCQDRGLQVNF 272
Query: 290 SIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLT 349
+PR + WA P++ LH++I+EESK+RDRKN CGLL+EI +IEK R+M++L+DS HFPL
Sbjct: 273 FVPRHFQWAAPVMSLHDKIVEESKRRDRKNCCGLLKEIDRIEKSSRLMNELIDSIHFPLN 332
Query: 350 EEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
++KE EV+Q+V E+ +V +AL++GLDP ER+VREVFHRIVRS+TE LDSL
Sbjct: 333 DDKEVEVKQRVDELVQVREALRNGLDPFERKVREVFHRIVRSRTESLDSL 382
>UniRef100_Q8LA35 Hypothetical protein [Arabidopsis thaliana]
Length = 382
Score = 478 bits (1231), Expect = e-133
Identities = 242/410 (59%), Positives = 315/410 (76%), Gaps = 39/410 (9%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSM----------EGSTLEIELDSFQQHVTD 50
MPATD+QGS FGRS+LS RR+Q+ S E ST+E+ELDSFQ+ V +
Sbjct: 1 MPATDFQGS-------FGRSLLSLRRDQVDSSTVVSGSSSHHEPSTMEVELDSFQRQVAE 53
Query: 51 RFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERS 110
+F+DL++ +D LLSL+W+GK+LD FL CQEEF+AI+ H+ Q+ + P+DR++S+YFERS
Sbjct: 54 KFIDLNASSND-LLSLEWIGKLLDSFLCCQEEFRAIVFNHRSQISKSPMDRLISDYFERS 112
Query: 111 VKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDG 170
+KALDVCNAIRDG+EQIR W+KL +IV+ ALD R IGEGQ RRAKKALIDL+I MLD+
Sbjct: 113 IKALDVCNAIRDGIEQIRQWEKLADIVISALDSHRPIGEGQLRRAKKALIDLAIGMLDEK 172
Query: 171 KE-SNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTW 229
S ++AHRNRSFGR +D H HRS+G FRSL+WSVSR+W
Sbjct: 173 DHPSGTNLAHRNRSFGRV-----KDSH---------------HRSIGHFRSLTWSVSRSW 212
Query: 230 SAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHF 289
SA++QLQA+ +N+ PR ND++A+NGLA+ VYTM SVLLFVMW LVAAIPCQDRGL V+F
Sbjct: 213 SASKQLQALASNLATPRPNDVVASNGLAVPVYTMTSVLLFVMWVLVAAIPCQDRGLQVNF 272
Query: 290 SIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLT 349
+PR + WA P++ LH++I+EESK+RDRKN CGLL+EI +IEK R+M++L+DS HFPL
Sbjct: 273 FVPRHFQWAAPVMSLHDKIVEESKRRDRKNCCGLLKEIDRIEKSSRLMNELIDSIHFPLN 332
Query: 350 EEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
++KE EV+Q+V E+ +V +AL++GLDP ER+VREVFHRIVRS+TE LDSL
Sbjct: 333 DDKEVEVKQRVDELVQVREALRNGLDPFERKVREVFHRIVRSRTESLDSL 382
>UniRef100_Q8LG71 Hypothetical protein [Arabidopsis thaliana]
Length = 397
Score = 475 bits (1222), Expect = e-132
Identities = 246/418 (58%), Positives = 316/418 (74%), Gaps = 43/418 (10%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQ-IHSMEGST-------LEIELDSFQQHVTDRF 52
MPAT+YQ S FGRS L+ RR+ ++S+E +T +E EL SFQ+ V +RF
Sbjct: 1 MPATEYQRS-------FGRSFLNLRRDTAVNSVESTTVTPELTQMEAELVSFQRKVAERF 53
Query: 53 VDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVK 112
+DL++ ++LLSL+WVGK+LD FL CQEEF++I+ H+ + +PP+DR+VS+YFERSVK
Sbjct: 54 IDLNASSCEDLLSLEWVGKLLDSFLSCQEEFRSIVINHRSMITKPPMDRLVSDYFERSVK 113
Query: 113 ALDVCNAIRDGVEQIRVWQKLLEIVLYALDH-------QRSIGEGQFRRAKKALIDLSIS 165
ALDVCNAIRDGVEQIR WQKL+EIV+ A ++ +R +GEGQFRRA+K LI+L+I
Sbjct: 114 ALDVCNAIRDGVEQIRQWQKLIEIVICAFNNNAGGSSGKRPLGEGQFRRARKTLIELAIG 173
Query: 166 MLDDGKESNASVA--HRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSW 223
MLD+ S++SV+ HRNRSFGR N HR++G FRSLSW
Sbjct: 174 MLDEKDSSSSSVSSQHRNRSFGR-------------------NKEQLHHRTIGHFRSLSW 214
Query: 224 SVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDR 283
SVSR+WSA +QLQAIGNN+ PRA+D+ ATNGL + VYTM SVLLFVMWALVAAIPCQDR
Sbjct: 215 SVSRSWSAXKQLQAIGNNLATPRASDITATNGLIVPVYTMTSVLLFVMWALVAAIPCQDR 274
Query: 284 GLNVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDS 343
GL VHF++PR+Y W L+ LH+RI+EESKKR+RKN CGLL+EI Q EK R+M++LVDS
Sbjct: 275 GLQVHFNVPRNYQWGGSLMSLHDRIIEESKKRERKNTCGLLKEIHQFEKTSRLMNELVDS 334
Query: 344 AHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGR 401
FPL+EEKE EVR++V E+ K+ +ALK+GLDP ER+VREVFHRIVRS+TEGLD++G+
Sbjct: 335 VQFPLSEEKEMEVRERVEELGKLQEALKNGLDPFERKVREVFHRIVRSRTEGLDTVGK 392
>UniRef100_Q9CA69 Hypothetical protein F1M20.13 [Arabidopsis thaliana]
Length = 397
Score = 474 bits (1220), Expect = e-132
Identities = 245/418 (58%), Positives = 317/418 (75%), Gaps = 43/418 (10%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQ-IHSMEGST-------LEIELDSFQQHVTDRF 52
MPAT+YQ S FGRS L+ RR+ ++S+E +T +E EL SFQ+ V +RF
Sbjct: 1 MPATEYQRS-------FGRSFLNLRRDTAVNSVESTTVTPELTQMEAELVSFQRKVAERF 53
Query: 53 VDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVK 112
+DL++ ++LLSL+WVGK+LD FL CQEEF++I+ H+ + +PP+DR+VS+YFERSVK
Sbjct: 54 IDLNASSCEDLLSLEWVGKLLDSFLSCQEEFRSIVINHRSMITKPPMDRLVSDYFERSVK 113
Query: 113 ALDVCNAIRDGVEQIRVWQKLLEIVLYALDH-------QRSIGEGQFRRAKKALIDLSIS 165
ALDVCNAIRDGVEQIR WQKL+EIV+ A ++ +R +GEGQFRRA+K LI+L+I
Sbjct: 114 ALDVCNAIRDGVEQIRQWQKLIEIVICAFNNNGGGSSGKRPLGEGQFRRARKTLIELAIG 173
Query: 166 MLDDGKESNASVA--HRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSW 223
MLD+ S++SV+ HRNRSFGR N HR++G FRSLSW
Sbjct: 174 MLDEKDSSSSSVSSQHRNRSFGR-------------------NKEQLHHRTIGHFRSLSW 214
Query: 224 SVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDR 283
SVSR+WSA++QLQAIGNN+ PRA+D+ ATNGL + VYTM +VLLFVMWALVAAIPCQDR
Sbjct: 215 SVSRSWSASKQLQAIGNNLATPRASDITATNGLIVPVYTMTTVLLFVMWALVAAIPCQDR 274
Query: 284 GLNVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDS 343
GL VHF++PR+Y W L+ LH+RI+EESKKR+RKN CGLL+EI Q EK R+M++LVDS
Sbjct: 275 GLQVHFNVPRNYQWGGSLMSLHDRIIEESKKRERKNTCGLLKEIHQFEKTSRLMNELVDS 334
Query: 344 AHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGR 401
FPL+EEKE EVR++V E+ K+ +ALK+GLDP ER+VREVFHRIVRS+TEGLD++G+
Sbjct: 335 VQFPLSEEKEMEVRERVEELGKLQEALKNGLDPFERKVREVFHRIVRSRTEGLDTVGK 392
>UniRef100_Q6DYE2 Hypothetical protein [Arabidopsis thaliana]
Length = 413
Score = 462 bits (1188), Expect = e-128
Identities = 236/410 (57%), Positives = 309/410 (74%), Gaps = 34/410 (8%)
Query: 6 YQGSSPSSLTNFGRSILSFRREQIHSM------EGSTLEIELDSFQQHVTDRFVDLS-SV 58
Y P + +FGRS LS RR+Q H M E T+E+ELDSFQ+ V ++F+DL+ S
Sbjct: 27 YISKMPETEYSFGRSFLSLRRDQAHLMDPTSFSEPMTMEVELDSFQRQVAEKFIDLNASA 86
Query: 59 PHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCN 118
E+LSL+W+GK+LD FL CQE+F+ I+ HK Q+++ P+DR++ EYFERSVKALDVCN
Sbjct: 87 DEAEILSLEWIGKLLDSFLCCQEDFRVIIFNHKPQLLKQPMDRLIEEYFERSVKALDVCN 146
Query: 119 AIRDGVEQIRVWQKLLEIVLYALD-HQRSIGEGQFRRAKKALIDLSISMLDDGKESNASV 177
AIRDG+EQIR WQKL+EIV+ ALD +QR +GEG+ RAKKALIDL+I MLD+ SN
Sbjct: 147 AIRDGIEQIRQWQKLIEIVISALDTNQRQLGEGEIHRAKKALIDLAIGMLDEKDSSNT-- 204
Query: 178 AHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQA 237
HRNRSF R+ +DH+QH +G RSLSWSVSR+WSA+RQLQ
Sbjct: 205 -HRNRSFTRN-----KDHNQH----------------IGYIRSLSWSVSRSWSASRQLQG 242
Query: 238 IGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIPRSYTW 297
IGNN+ PRA+D+MATNGLA++VYTM S+LLFV W LVAAIPCQDRGL+VHF PR + W
Sbjct: 243 IGNNLATPRASDVMATNGLALTVYTMTSILLFVTWVLVAAIPCQDRGLHVHFYFPRHFQW 302
Query: 298 AIPLLLLHERIMEESKKRD-RKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEK-EGE 355
A+P++ LH++IM+ESKKRD +K CGLLREI QIE+ R++SDL+DS +F LT+EK E
Sbjct: 303 AVPVMSLHDKIMDESKKRDKKKKGCGLLREINQIERNSRMLSDLIDSDNFSLTDEKCTLE 362
Query: 356 VRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE 405
V+++V E+ VC+A+K+GLDP +R+VR+VFH+IVR++TE LDSLG+ N+
Sbjct: 363 VKERVQELMNVCEAIKEGLDPFDRKVRDVFHQIVRTRTEALDSLGKLQNQ 412
>UniRef100_Q9XIG0 T10P12.8 protein [Arabidopsis thaliana]
Length = 383
Score = 461 bits (1187), Expect = e-128
Identities = 238/415 (57%), Positives = 311/415 (74%), Gaps = 43/415 (10%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSM------EGSTLEIELDSFQQHVTDRFVD 54
MP T+Y +FGRS LS RR+Q H M E T+E+ELDSFQ+ V ++F+D
Sbjct: 1 MPETEY---------SFGRSFLSLRRDQAHLMDPTSFSEPMTMEVELDSFQRQVAEKFID 51
Query: 55 LS-SVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKA 113
L+ S E+LSL+W+GK+LD FL CQE+F+ I+ HK Q+++ P+DR++ EYFERSVKA
Sbjct: 52 LNASADEAEILSLEWIGKLLDSFLCCQEDFRVIIFNHKPQLLKQPMDRLIEEYFERSVKA 111
Query: 114 LDVCNAIRDGVEQIRVWQKLLEIVLYALD-HQRSIGEGQFRRAKKALIDLSISMLDDGKE 172
LDVCNAIRDG+EQIR WQKL+EIV+ ALD +QR +GEG+ RAKKALIDL+I MLD+
Sbjct: 112 LDVCNAIRDGIEQIRQWQKLIEIVISALDTNQRQLGEGEIHRAKKALIDLAIGMLDEKDS 171
Query: 173 SNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAA 232
SN HRNRSF R+ +DH+QH +G RSLSWSVSR+WSA+
Sbjct: 172 SNT---HRNRSFTRN-----KDHNQH----------------IGYIRSLSWSVSRSWSAS 207
Query: 233 RQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIP 292
RQLQ IGNN+ PRA+D+MATNGLA++VYTM S+LLFV W LVAAIPCQDRGL+VHF P
Sbjct: 208 RQLQGIGNNLATPRASDVMATNGLALTVYTMTSILLFVTWVLVAAIPCQDRGLHVHFYFP 267
Query: 293 RSYTWAIPLLLLHERIMEESKKRD-RKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEE 351
R + WA+P++ LH++IM+ESKKRD +K CGLLREI QIE+ R++SDL+DS +F LT+E
Sbjct: 268 RHFQWAVPVMSLHDKIMDESKKRDKKKKGCGLLREINQIERNSRMLSDLIDSDNFSLTDE 327
Query: 352 K-EGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE 405
K EV+++V E+ VC+A+K+GLDP +R+VR+VFH+IVR++TE LDSLG+ N+
Sbjct: 328 KCTLEVKERVQELMNVCEAIKEGLDPFDRKVRDVFHQIVRTRTEALDSLGKLQNQ 382
>UniRef100_Q9ST79 CAA303718.1 protein [Oryza sativa]
Length = 425
Score = 407 bits (1047), Expect = e-112
Identities = 219/425 (51%), Positives = 292/425 (68%), Gaps = 23/425 (5%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQI-----------HSMEGSTLEIELDSFQQHVT 49
MPATD S+ + LT+FGRS LS RR+QI H+ S+ ++E+D+F +H
Sbjct: 1 MPATD-SSSAAAPLTSFGRSFLSHRRDQIPPPPPDHHSHSHTQHPSSSDLEIDAFHRHAA 59
Query: 50 DRFVDLSSVPHDE-----LLSLKWVGKMLDCFLICQEEFKAILHT--HKGQVVRPPLDRM 102
D DL S + + LLSL W ++LD FLIC EEF+AIL + RPPLDR+
Sbjct: 60 DLLHDLLSDSNSDPSAPDLLSLAWTRRLLDSFLICLEEFRAILFALADSQPLSRPPLDRL 119
Query: 103 VSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSI--GEGQFRRAKKALI 160
+ ++ +R+VKALD+CNA+RDG++ IR W+K L I AL + GE Q RRA+KAL
Sbjct: 120 LLDFLDRAVKALDLCNALRDGLDLIRQWRKHLAIAAAALAPAPAAQRGEAQIRRARKALT 179
Query: 161 DLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRS 220
DL+I MLDD K++ V RNRSFGR+ RD H HG+ +++ + S RS
Sbjct: 180 DLTILMLDD-KDAGGVVGQRNRSFGRAGTTRDSLPHGHGHHRRSSSGGSSGSGSGSHLRS 238
Query: 221 LSWSVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPC 280
LSWSVSRTWSAARQLQAIG + PRAND+ AT GLA +VY M +VL V WALVAAIPC
Sbjct: 239 LSWSVSRTWSAARQLQAIGGGLTVPRANDIAATGGLASAVYAMGAVLFVVTWALVAAIPC 298
Query: 281 QDRGLNVHFS-IPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSD 339
QDRGL H + +PR++ WA PL+ L +RI++ESKK+DRK++CGLL+EI QIE+C R + +
Sbjct: 299 QDRGLQAHLTAVPRTFPWAGPLITLFDRILDESKKKDRKHSCGLLKEIHQIERCSRQLME 358
Query: 340 LVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
+ D+A FPL ++K+ EV++ E+ +VC +LKDGLDPLERQVRE+FHR+VR++TE LD L
Sbjct: 359 VTDAAEFPLADDKDSEVQEATQELVQVCGSLKDGLDPLERQVREMFHRVVRTRTEILDYL 418
Query: 400 GRPNN 404
RP+N
Sbjct: 419 SRPHN 423
>UniRef100_Q7XSV3 OSJNBa0039K24.3 protein [Oryza sativa]
Length = 403
Score = 342 bits (878), Expect = 8e-93
Identities = 195/427 (45%), Positives = 271/427 (62%), Gaps = 49/427 (11%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQI-----------HSMEGSTLEIELDSFQQHVT 49
MPATD S+ + LT+FGRS LS RR+QI H+ S+ ++E+D+F +H
Sbjct: 1 MPATD-SSSAAAPLTSFGRSFLSHRRDQIPPPPPDHHSHSHTQHPSSSDLEIDAFHRHAA 59
Query: 50 DRFVDLSSVPHDE-----LLSLKWVGKMLDCFLICQEEFKAILHT--HKGQVVRPPLDRM 102
D DL S + + LLSL W ++LD FLIC EEF+AIL + RPPLDR+
Sbjct: 60 DLLHDLLSDSNSDPSAPDLLSLAWTRRLLDSFLICLEEFRAILFALADSQPLSRPPLDRL 119
Query: 103 VSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSI--GEGQFRRAKKALI 160
+ ++ +R+VKALD+CNA+RDG++ IR W+K L I AL + GE Q RRA+KAL
Sbjct: 120 LLDFLDRAVKALDLCNALRDGLDLIRQWRKHLAIAAAALATAPAAQRGEAQIRRARKALT 179
Query: 161 DLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRS 220
DL+I MLDD K++ V RNRSFG ++ R F +
Sbjct: 180 DLTILMLDD-KDAGGVVGQRNRSFGSASTTRT------------------------PFPT 214
Query: 221 LSWSVSRTWSAARQLQAIG--NNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAI 278
+ + + +A A+ + + PRAND+ AT GLA +VY M +VL V WALVAAI
Sbjct: 215 ATATTAGAAAAGAPGPALAPISGLTVPRANDIAATGGLASAVYAMGAVLFVVTWALVAAI 274
Query: 279 PCQDRGLNVHFS-IPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVM 337
PCQDRGL H + +PR++ WA PL+ L +RI++ESKK+DRK++CGLL+EI QIE+C R +
Sbjct: 275 PCQDRGLQAHLTAVPRTFPWAGPLITLFDRILDESKKKDRKHSCGLLKEIHQIERCSRQL 334
Query: 338 SDLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLD 397
++ D+A FPL ++K+ EV++ E+ +VC +LKDGLDPLERQVRE+FHR+VR++TE LD
Sbjct: 335 MEVTDAAEFPLADDKDSEVQEATQELVQVCGSLKDGLDPLERQVREMFHRVVRTRTEILD 394
Query: 398 SLGRPNN 404
L RP+N
Sbjct: 395 YLSRPHN 401
>UniRef100_Q9CAK4 Hypothetical protein T12P18.5 [Arabidopsis thaliana]
Length = 415
Score = 268 bits (685), Expect = 2e-70
Identities = 153/428 (35%), Positives = 250/428 (57%), Gaps = 56/428 (13%)
Query: 2 PATDYQGSSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPH- 60
PA D QGS GR +S RR Q + + +L+ FQ+H+ DRF +L S P
Sbjct: 3 PAQDNQGSF------LGR--ISIRRNQFVDVNNEQEQEDLELFQKHIADRFTELLSPPQP 54
Query: 61 ---------------DELLSLKWVGKMLDCFLICQEEFKAILHTHKG--QVVRPPLDRMV 103
++++S+ W+ K++D FL C+ EFKAIL + Q+ +PP DR+V
Sbjct: 55 PPSDEINTVASVAATEQIMSVTWLRKLMDVFLCCEAEFKAILLMGRDPTQISKPPFDRLV 114
Query: 104 SEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLS 163
E +RS+KALD+C A+ +G++ +R +Q+L EI + AL+ QR +G+G RRAK+AL +L
Sbjct: 115 PEMLDRSIKALDICTAVVNGIDSVRHYQRLAEIAVTALE-QRPLGDGNVRRAKRALANLV 173
Query: 164 ISMLDDGKESNAS----------VAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHR 213
+++ + KE+ + R+ SFGR +GG ++ +
Sbjct: 174 VALSLEDKENVSGGGGGGGGGNKTTERSWSFGRRSGG--------------SSAASKGGA 219
Query: 214 SMGQFRSLSWSVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWA 273
++GQ +S SW+V R WSAA+Q+ A+ N+ PPR N+ GL ++ M++V++FVMW
Sbjct: 220 TIGQLKSSSWAVGRNWSAAKQIHAMTANLTPPRGNEAA---GLPQPMFIMSTVMVFVMWV 276
Query: 274 LVAAIPCQDR-GLNVHFSIPRSY-TWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIE 331
L AA+PCQ+R GL H +P + WA L+ +HE+I +E KK+++K + GL+ E+ ++E
Sbjct: 277 LTAAVPCQERSGLANHLPVPPKHLNWAQSLIGIHEKIGDEWKKKEKKGSAGLMEEMTRME 336
Query: 332 KCVRVMSDLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRS 391
K + + D H+P ++ +V E++++C +++ L PL++Q+REVFHRIVRS
Sbjct: 337 KLGHSLMEFADGFHYPAEKDAAESAAVQVAEMAEICRRMEEELVPLQQQIREVFHRIVRS 396
Query: 392 KTEGLDSL 399
+ E L+ L
Sbjct: 397 RAEILEVL 404
>UniRef100_O81749 Hypothetical protein F16G20.230 [Arabidopsis thaliana]
Length = 396
Score = 227 bits (578), Expect = 5e-58
Identities = 151/418 (36%), Positives = 233/418 (55%), Gaps = 62/418 (14%)
Query: 3 ATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS---TLEIELDSFQQHVTDRFVDLS--- 56
AT++QGS S + S RR QI SM+ + LE EL+ FQ+HV +RF DL
Sbjct: 4 ATEFQGSFLSRI--------SIRRNQIVSMDVNHEQELE-ELEYFQKHVAERFSDLITSP 54
Query: 57 -----------SVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPP-LDRMVS 104
S P D +LS+ W+ +LD F+ C+ EFKA+L T Q+ + P L+R++
Sbjct: 55 SPPPSSSSSAVSQPSDPILSIPWLQNLLDVFMSCEAEFKAVLSTT--QISKSPSLERVLP 112
Query: 105 EYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSI 164
E +R +KALD+CNA+ +G++ +R ++ EI + AL QR + +G RRAK+AL L I
Sbjct: 113 EMLDRILKALDLCNAVVNGIDSVRQSRRFAEIAVTALK-QRPLCDGSVRRAKRALTSLLI 171
Query: 165 SMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWS 224
+ N RD + G+ SN T + S G +++
Sbjct: 172 GL---------------------NADERRDRNSGGSGCSNQRRTTSRSWSFGTRSNVTGG 210
Query: 225 ------VSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAI 278
VS+ WSA++Q+QA+ N+ PR + +G AM VY M+SV++ VMW LVAA+
Sbjct: 211 GLYGQVVSKNWSASKQIQAMVANLVLPRGAE---ASGPAMPVYIMSSVMVLVMWVLVAAV 267
Query: 279 PCQDRGLNVH-FSIPRSYTWAIPLLLLHERIMEESKKRDRK-NACGLLREIQQIEKCVRV 336
PCQ + V +P+ WA + + ERI EE K+++++ GL+ E+Q++EK
Sbjct: 268 PCQTSSVLVAPLPLPKHQNWASAAMSIQERIGEEIKRKEKRCGGGGLMEEMQRMEKIGLS 327
Query: 337 MSDLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTE 394
+ + + FP EE+E EV +KV E+ ++C ++ GL+ L+RQVR+VFHR+VRS+ E
Sbjct: 328 LMEFAERFRFPADEEEEVEVAEKVDEMEEICRRMEVGLEDLQRQVRQVFHRLVRSRIE 385
>UniRef100_Q8LDW5 Hypothetical protein [Arabidopsis thaliana]
Length = 371
Score = 221 bits (562), Expect = 4e-56
Identities = 145/411 (35%), Positives = 230/411 (55%), Gaps = 61/411 (14%)
Query: 2 PATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS--TLEIELDSFQQHVTDRFVDL---S 56
PAT++Q S S L+ RR Q+ SME + + EL+ FQ+HV +RF +L S
Sbjct: 3 PATEHQSSFLSRLS---------RRNQVVSMEVNHEQEQEELEDFQKHVAERFAELLPPS 53
Query: 57 SVPHD-ELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALD 115
P +LS++W+ K+LD F+ + EF ++L ++ Q+ +PPLD++V E +R VKALD
Sbjct: 54 DSPESYPILSIQWLRKLLDVFMSIESEFHSVLTSNPSQISKPPLDKLVPEMLDRIVKALD 113
Query: 116 VCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNA 175
+C A+ +GV+ +R Q+ EI + AL Q + +G RRAK+AL L ++ L+ K S +
Sbjct: 114 ICTAVVNGVDSVRQIQRCAEIAVTAL-KQTPLSDGSVRRAKRALTSL-LAALNADKNSGS 171
Query: 176 SVAHRNR--------SFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSR 227
S R SFGR +GG S G + VS+
Sbjct: 172 SGGGSGRRSSTDQWSSFGRRSGG-----------------------SSGGGGGGAGCVSK 208
Query: 228 TWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQ-DRGLN 286
WSAA+Q+QA+ N+ PR G A +Y M+SV++ VMW LV A+PCQ GL
Sbjct: 209 NWSAAKQIQAMTANLVAPR-------GGEASPMYIMSSVMVMVMWTLVVAVPCQTSNGLM 261
Query: 287 VHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHF 346
VH +P++ WA + + ER+ EE K+++ + GL+ E+Q++E+ + + + F
Sbjct: 262 VHVPLPKNQVWASAAVSISERVGEEMKRKETRGG-GLMEEMQRMERIWLKLMEFSEGFRF 320
Query: 347 PLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLD 397
E +V +V E+ ++C ++DGL+ L+R+VREVFHR+V+S++E L+
Sbjct: 321 ----NGEEDVVAEVAEMEEICRKMEDGLEGLQRRVREVFHRLVKSRSEILE 367
>UniRef100_Q9SUT1 Hypothetical protein AT4g11300 [Arabidopsis thaliana]
Length = 371
Score = 220 bits (561), Expect = 5e-56
Identities = 145/411 (35%), Positives = 230/411 (55%), Gaps = 61/411 (14%)
Query: 2 PATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS--TLEIELDSFQQHVTDRFVDL---S 56
PAT++Q S S L+ RR Q+ SME + + EL+ FQ+HV +RF +L S
Sbjct: 3 PATEHQSSFLSRLS---------RRNQVVSMEVNHEQEQEELEDFQKHVAERFAELLPPS 53
Query: 57 SVPHD-ELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALD 115
P +LS++W+ K+LD F+ + EF ++L ++ Q+ +PPLD++V E +R VKALD
Sbjct: 54 DSPESYPILSIQWLRKLLDVFMSIESEFHSVLTSNPSQISKPPLDKLVPEMLDRIVKALD 113
Query: 116 VCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNA 175
+C A+ +GV+ +R Q+ EI + AL Q + +G RRAK+AL L ++ L+ K S +
Sbjct: 114 ICTAVVNGVDSVRQIQRCAEIAVTAL-KQTPLSDGSVRRAKRALTSL-LAALNADKNSGS 171
Query: 176 SVAHRNR--------SFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSR 227
S R SFGR +GG S G + VS+
Sbjct: 172 SGGGSGRRSSTDQWSSFGRRSGG-----------------------SSGGGGGGAGCVSK 208
Query: 228 TWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQ-DRGLN 286
WSAA+Q+QA+ N+ PR G A +Y M+SV++ VMW LV A+PCQ GL
Sbjct: 209 NWSAAKQIQAMTANLVAPR-------GGEASPMYIMSSVMVMVMWTLVVAVPCQTSNGLM 261
Query: 287 VHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHF 346
VH +P++ WA + + ER+ EE K+++ + GL+ E+Q++E+ + + + F
Sbjct: 262 VHVPLPKNQVWANAAVSISERVGEEMKRKETRGG-GLMEEMQRMERIGLKLMEFSEGFRF 320
Query: 347 PLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLD 397
E +V +V E+ ++C ++DGL+ L+R+VREVFHR+V+S++E L+
Sbjct: 321 ----NGEEDVVAEVAEMEEICRKMEDGLEGLQRRVREVFHRLVKSRSEILE 367
>UniRef100_Q6ETS7 Hypothetical protein P0017C12.31 [Oryza sativa]
Length = 417
Score = 172 bits (436), Expect = 2e-41
Identities = 128/404 (31%), Positives = 197/404 (48%), Gaps = 70/404 (17%)
Query: 21 ILSFRRE-----QIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDC 75
+LSFRR + L++ L + Q HV DR LS H LLSL ++ K+LD
Sbjct: 18 LLSFRRSATAVASFDPAQDDELQV-LHALQAHVADRLAALSH--HPPLLSLAFLSKLLDA 74
Query: 76 FLICQEEFKAILHTHK--GQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKL 133
L + F+ +L + RPP DR+ ++ +R+VK LD+ NA+ + +R +
Sbjct: 75 VLSSDDAFREVLGIGPVAAALSRPPADRLAADLLDRTVKTLDILNAVSLTLASLRGSHRA 134
Query: 134 LEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHRNRSFGRSNGGRDR 193
L + F RA++A+ L D + A+ + R+
Sbjct: 135 ALTAASCL-LAPPLHRAHFGRARRAISRL----FPDAAKLAAAPSPSCRA---------- 179
Query: 194 DHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQAIGNNINPPRANDLMAT 253
G R+LS+SVSR WS+ R + A+ ++ PP + A+
Sbjct: 180 ----------------------GPARALSFSVSRNWSSGRHVHAMAAHLAPPPQSPTSAS 217
Query: 254 NG----LAMSVYTMNSVLLFVMWALVAAIPCQDR---GLNVHFSIPRSYTWAIPLLLLHE 306
G L +++YTM+SVL+F MWALVAA+PCQDR N + P+ WA P+ L E
Sbjct: 218 PGAGCGLGLALYTMSSVLVFSMWALVAAVPCQDRSSAATNPPVAPPKQVQWAAPMCALQE 277
Query: 307 RIMEESKKRDRKN-------ACGLLREIQQIEKCVRVMSDLVDSAH---------FPLTE 350
RI +E +K+D+K A GLL E+Q +E+ R +S L++ T+
Sbjct: 278 RIADEWRKKDKKGSSSGSAAATGLLAEMQAVERAARELSSLLEEVAEEEEEEQLVMGATD 337
Query: 351 EKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTE 394
E+ +V ++ ++ C AL++GL PLERQVR VFHR+V S+ E
Sbjct: 338 ERARDVVERAEALAAACRALEEGLAPLERQVRAVFHRVVASRGE 381
>UniRef100_Q8GW16 Hypothetical protein [Arabidopsis thaliana]
Length = 412
Score = 42.7 bits (99), Expect = 0.018
Identities = 31/107 (28%), Positives = 55/107 (50%), Gaps = 2/107 (1%)
Query: 42 DSFQQHVTDRFVD-LSSVPHDELLSLKWVGKMLDCFL-ICQEEFKAILHTHKGQVVRPPL 99
DS T+R ++ L+S LS + ++ C L + QE + I+ + + L
Sbjct: 59 DSSLHQRTNRVINSLASGAQTRSLSFDALIEVSGCLLEMNQEVVRFIIESKEDVWDNKDL 118
Query: 100 DRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRS 146
+V+ YF+ S+K LD CNA+ + V++ R+ Q LL+ L + + S
Sbjct: 119 TCLVNAYFDSSIKTLDFCNAVDNCVKRARIGQMLLQFALKQFEMESS 165
>UniRef100_Q9FW77 Hypothetical protein OSJNBa0026L12.27 [Oryza sativa]
Length = 485
Score = 42.4 bits (98), Expect = 0.024
Identities = 26/119 (21%), Positives = 55/119 (45%), Gaps = 1/119 (0%)
Query: 226 SRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGL 285
+R S + LQ + N++ + + L ++Y + SV +FV VA + + L
Sbjct: 166 ARLVSCSATLQELAGNLSLMKVKNSAKGKVLMRALYGIESVTVFVCSIFVAVLSGSPKPL 225
Query: 286 NVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSA 344
V +P + W+ LH + EE ++ + ++E++++E C R + L ++
Sbjct: 226 -VELHVPEKFGWSQAFNDLHTAVSEELTRQLSGGSVAAVKELEEVEACARRLHVLASTS 283
Score = 38.9 bits (89), Expect = 0.26
Identities = 28/106 (26%), Positives = 52/106 (48%), Gaps = 16/106 (15%)
Query: 62 ELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRP---PL----DRMVSEYFERSVKAL 114
++L+L W+ +DC + LHT+ ++ P+ D+ V Y SVK L
Sbjct: 57 DVLTLSWMRLAVDCL--------SELHTNIANLITDLELPVSDWDDKWVDIYLNSSVKLL 108
Query: 115 DVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSI-GEGQFRRAKKAL 159
D+C A+ + ++ Q LL+ L+ L + + + Q +RA+ +L
Sbjct: 109 DICIALSSELSRLDQGQLLLQYALHVLGSESGVPSQEQLKRAEPSL 154
>UniRef100_Q9SYZ8 Hypothetical protein AT4g34330 [Arabidopsis thaliana]
Length = 354
Score = 40.4 bits (93), Expect = 0.090
Identities = 32/124 (25%), Positives = 56/124 (44%), Gaps = 8/124 (6%)
Query: 9 SSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRF---VDLSSVPHDELLS 65
S S N+ + S+ ME + + + + HV V++ S+ D L +
Sbjct: 9 SKEKSGRNYTTELRSYEAACKEDMEIQSFDTRMQARTSHVISTLATGVEVRSLSFDSLKA 68
Query: 66 LKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVE 125
+ +G +LD + QE K IL K + V YFE S+K LD NA++ G++
Sbjct: 69 V--IGSLLD---MNQEVAKVILDCKKDIWKNQEMFEFVEAYFETSLKTLDFFNALKRGLQ 123
Query: 126 QIRV 129
+++
Sbjct: 124 GVQI 127
>UniRef100_Q9SYZ7 Hypothetical protein AT4g34320 [Arabidopsis thaliana]
Length = 374
Score = 38.1 bits (87), Expect = 0.45
Identities = 25/88 (28%), Positives = 41/88 (46%), Gaps = 1/88 (1%)
Query: 64 LSLKWVGKMLDCFL-ICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRD 122
LS + ++ C L + QE K IL K + +V +YFE S+K LD C A+
Sbjct: 63 LSFDSLKEVTQCLLEMNQEVVKVILDCKKDIWKNQEMFELVEDYFENSLKTLDFCAALEK 122
Query: 123 GVEQIRVWQKLLEIVLYALDHQRSIGEG 150
G+ + R L+ + L + + + G
Sbjct: 123 GLRRARDSHLLILVALQQFEDESLVQGG 150
>UniRef100_Q8MP27 Hypothetical protein [Dictyostelium discoideum]
Length = 1880
Score = 37.7 bits (86), Expect = 0.58
Identities = 22/98 (22%), Positives = 45/98 (45%)
Query: 162 LSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSL 221
LS S +D GK + + N + +N + +++ + N+++NNN N + + S
Sbjct: 824 LSESDIDHGKGGTGPIKNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNHSS 883
Query: 222 SWSVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMS 259
+ S + + S++ NN N + + N L+ S
Sbjct: 884 NKSRNISTSSSTSTSTSTNNTNNNSNSSTIGGNSLSSS 921
>UniRef100_UPI00004367B7 UPI00004367B7 UniRef100 entry
Length = 610
Score = 37.0 bits (84), Expect = 0.99
Identities = 21/75 (28%), Positives = 37/75 (49%)
Query: 134 LEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHRNRSFGRSNGGRDR 193
+E +YA+ G G ++++K + +D SN +RN S R+N +
Sbjct: 459 MENQIYAIPENIMRGCGSEQQSEKQRNHNNNHSNNDNNHSNKDNNNRNNSNNRNNVNHNN 518
Query: 194 DHHQHGNSHSNNNIN 208
+H ++ N+HSN N N
Sbjct: 519 NHSKNNNNHSNKNNN 533
>UniRef100_Q9LMM6 Hypothetical protein F22L4.9 [Arabidopsis thaliana]
Length = 349
Score = 37.0 bits (84), Expect = 0.99
Identities = 26/113 (23%), Positives = 52/113 (46%), Gaps = 1/113 (0%)
Query: 30 HSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHT 89
+S+ S L L++F+ ++ L ++L++ W+ + ++ K ++ T
Sbjct: 25 NSVLSSKLLPLLNNFETNLASSISKLVPKEKSDILTVSWMKQAMESLCETHNGIKTLI-T 83
Query: 90 HKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALD 142
V D+ V Y + SVK LD+CNA + ++ LL+ L+ L+
Sbjct: 84 DLELPVSDWEDKWVDVYLDISVKLLDLCNAFSSELTRLNQGHLLLQFALHNLE 136
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 656,440,361
Number of Sequences: 2790947
Number of extensions: 26101463
Number of successful extensions: 85824
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 85506
Number of HSP's gapped (non-prelim): 202
length of query: 405
length of database: 848,049,833
effective HSP length: 130
effective length of query: 275
effective length of database: 485,226,723
effective search space: 133437348825
effective search space used: 133437348825
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC126014.1


