

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126010.6 + phase: 0
(518 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
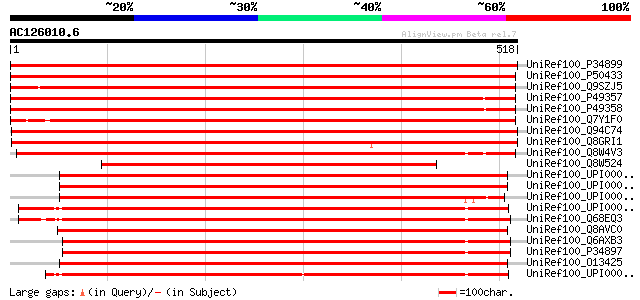
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_P34899 Serine hydroxymethyltransferase, mitochondrial ... 990 0.0
UniRef100_P50433 Serine hydroxymethyltransferase, mitochondrial ... 928 0.0
UniRef100_Q9SZJ5 Serine hydroxymethyltransferase, mitochondrial ... 919 0.0
UniRef100_P49357 Serine hydroxymethyltransferase 1, mitochondria... 918 0.0
UniRef100_P49358 Serine hydroxymethyltransferase 2, mitochondria... 907 0.0
UniRef100_Q7Y1F0 Putative glycine hydroxymethyltransferase [Oryz... 888 0.0
UniRef100_Q94C74 Putative glycine hydroxymethyltransferase [Arab... 881 0.0
UniRef100_Q8GRI1 Putative glycine hydroxymethyltransferase [Arab... 874 0.0
UniRef100_Q8W4V3 Serine hydroxymethyltransferase [Chlamydomonas ... 695 0.0
UniRef100_Q8W524 Serine hydroxymethyltransferase [Zea mays] 650 0.0
UniRef100_UPI0000430024 UPI0000430024 UniRef100 entry 588 e-166
UniRef100_UPI000042D39C UPI000042D39C UniRef100 entry 587 e-166
UniRef100_UPI000023F3A5 UPI000023F3A5 UniRef100 entry 582 e-165
UniRef100_UPI0000337BD6 UPI0000337BD6 UniRef100 entry 580 e-164
UniRef100_Q68EQ3 Shmt2-prov protein [Xenopus tropicalis] 580 e-164
UniRef100_Q8AVC0 Shmt1-prov protein [Xenopus laevis] 580 e-164
UniRef100_Q6AXB3 MGC79128 protein [Xenopus laevis] 579 e-164
UniRef100_P34897 Serine hydroxymethyltransferase, mitochondrial ... 577 e-163
UniRef100_O13425 Serine hydroxymethyltransferase, mitochondrial ... 575 e-162
UniRef100_UPI0000365291 UPI0000365291 UniRef100 entry 575 e-162
>UniRef100_P34899 Serine hydroxymethyltransferase, mitochondrial precursor [Pisum
sativum]
Length = 518
Score = 990 bits (2559), Expect = 0.0
Identities = 494/518 (95%), Positives = 511/518 (98%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMAMALR+LSSS+NKSSRPLFSASS+YYKSSLPDEAVYDKEN RV+WPKQLNS LE ID
Sbjct: 1 MAMAMALRKLSSSVNKSSRPLFSASSLYYKSSLPDEAVYDKENPRVTWPKQLNSPLEVID 60
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 120
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNF VYTALLKPHDRIMALDLPHGGHLS
Sbjct: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFQVYTALLKPHDRIMALDLPHGGHLS 180
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD
Sbjct: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
YARIRKVCDKQKAV+LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 241 YARIRKVCDKQKAVLLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEY+AYQEQV
Sbjct: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYRAYQEQV 360
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
LSN +KFA+ALSEKGY+LVSGGTENHLVLVNLKNKGIDGSRVEKVLE VHIAANKNTVPG
Sbjct: 361 LSNSSKFAKALSEKGYDLVSGGTENHLVLVNLKNKGIDGSRVEKVLELVHIAANKNTVPG 420
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDA+V+LALK+KAESKGTKLKDFVE L
Sbjct: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDAAVSLALKVKAESKGTKLKDFVEAL 480
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
Q+SSYVQSEISKL+HDVEEFAKQFPTIGFEK++MKYNK
Sbjct: 481 QTSSYVQSEISKLKHDVEEFAKQFPTIGFEKATMKYNK 518
>UniRef100_P50433 Serine hydroxymethyltransferase, mitochondrial precursor [Solanum
tuberosum]
Length = 518
Score = 928 bits (2399), Expect = 0.0
Identities = 455/516 (88%), Positives = 497/516 (96%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMA+ALRRLS++++K + L++ S+YY SSLP+EAVYDKE S V+WPKQLN+ LE +D
Sbjct: 1 MAMAIALRRLSATVDKPVKSLYNGGSLYYMSSLPNEAVYDKEKSGVAWPKQLNAPLEVVD 60
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIE EKARQWKGLELIPSENFTS+SVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 61 PEIADIIEHEKARQWKGLELIPSENFTSVSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 120
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDPAKWGVNVQPLSGSP+NF VYTALLKPH+RIMALDLPHGGHLS
Sbjct: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPANFQVYTALLKPHERIMALDLPHGGHLS 180
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD
Sbjct: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Y RIRKVC+KQKA++LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIF+R
Sbjct: 241 YDRIRKVCNKQKAILLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFYR 300
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KG+KEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEY+AYQEQV
Sbjct: 301 KGVKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYRAYQEQV 360
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
LSN +KFAQAL EKGYELVSGGT+NHLVLVN+KNKGIDGSRVEKVLEAVHIAANKNTVPG
Sbjct: 361 LSNSSKFAQALGEKGYELVSGGTDNHLVLVNMKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGF+EEDFVKVA++FDA+V +A+K+KAE++GTKLKDFV TL
Sbjct: 421 DVSAMVPGGIRMGTPALTSRGFLEEDFVKVADFFDAAVKIAVKVKAETQGTKLKDFVATL 480
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
+SS+ ++SEI+KLRHDVEE+AKQFPTIGFEK +MKY
Sbjct: 481 ESSAPIKSEIAKLRHDVEEYAKQFPTIGFEKETMKY 516
>UniRef100_Q9SZJ5 Serine hydroxymethyltransferase, mitochondrial precursor
[Arabidopsis thaliana]
Length = 517
Score = 919 bits (2376), Expect = 0.0
Identities = 458/516 (88%), Positives = 489/516 (94%), Gaps = 1/516 (0%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMAMALRRLSSSI+K RPL ++S Y SSLP EAV +KE SRV+WPKQLN+ LEE+D
Sbjct: 1 MAMAMALRRLSSSIDKPIRPLIRSTSCYM-SSLPSEAVDEKERSRVTWPKQLNAPLEEVD 59
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIE EKARQWKGLELIPSENFTS+SVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 60 PEIADIIEHEKARQWKGLELIPSENFTSVSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 119
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDP KWGVNVQPLSGSP+NFHVYTALLKPH+RIMALDLPHGGHLS
Sbjct: 120 MAETLCQKRALEAFRLDPEKWGVNVQPLSGSPANFHVYTALLKPHERIMALDLPHGGHLS 179
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQ+EKSATLFRPKLIVAGASAYARLYD
Sbjct: 180 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQMEKSATLFRPKLIVAGASAYARLYD 239
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
YARIRKVC+KQKAVMLADMAHISGLVAA VIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 240 YARIRKVCNKQKAVMLADMAHISGLVAANVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 299
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KG+KE+NKQGKEV YD+EDKINQAVFPGLQGGPHNHTITGLAVALKQATT EYKAYQEQV
Sbjct: 300 KGVKEINKQGKEVLYDFEDKINQAVFPGLQGGPHNHTITGLAVALKQATTSEYKAYQEQV 359
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
LSN AKFAQ L E+GYELVSGGT+NHLVLVNLK KGIDGSRVEKVLEAVHIA+NKNTVPG
Sbjct: 360 LSNSAKFAQTLMERGYELVSGGTDNHLVLVNLKPKGIDGSRVEKVLEAVHIASNKNTVPG 419
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDF KVAEYFD +V +ALK+K+E++GTKLKDFV +
Sbjct: 420 DVSAMVPGGIRMGTPALTSRGFVEEDFAKVAEYFDKAVTIALKVKSEAQGTKLKDFVSAM 479
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
+SSS +QSEI+KLRH+VEEFAKQFPTIGFEK +MKY
Sbjct: 480 ESSSTIQSEIAKLRHEVEEFAKQFPTIGFEKETMKY 515
>UniRef100_P49357 Serine hydroxymethyltransferase 1, mitochondrial precursor
[Flaveria pringlei]
Length = 517
Score = 918 bits (2372), Expect = 0.0
Identities = 459/516 (88%), Positives = 486/516 (93%), Gaps = 1/516 (0%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMA+ALRRLSSS +K + LF+ +Y SSLP EAVY+KE V+WPKQLN+ LE +D
Sbjct: 1 MAMALALRRLSSSADKPLQRLFNGGHLYSMSSLPSEAVYEKERPGVTWPKQLNAPLEVVD 60
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 120
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDPAKWGVNVQPLSGSP+NFHVYTALLK HDRIMALDLPHGGHLS
Sbjct: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPANFHVYTALLKAHDRIMALDLPHGGHLS 180
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRL+ESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD
Sbjct: 181 HGYQTDTKKISAVSIFFETMPYRLNESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
YARIRKVCDKQKA+MLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 241 YARIRKVCDKQKAIMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATT EYKAYQEQV
Sbjct: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTAEYKAYQEQV 360
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
+SN AKFA+ L + GYELVSGGTENHLVLVNLKNKGIDGS+VEKVLEAVHIAANKNTVPG
Sbjct: 361 MSNSAKFAETLVKSGYELVSGGTENHLVLVNLKNKGIDGSKVEKVLEAVHIAANKNTVPG 420
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDF KVA +FD +V LA+KIK E+KGTKLKDFV +
Sbjct: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFAKVAYFFDLAVKLAVKIKGEAKGTKLKDFVTAM 480
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
+SS+ +QSEISKLRHDVEE+AKQFPTIGFEK +MKY
Sbjct: 481 ESSA-IQSEISKLRHDVEEYAKQFPTIGFEKETMKY 515
>UniRef100_P49358 Serine hydroxymethyltransferase 2, mitochondrial precursor
[Flaveria pringlei]
Length = 517
Score = 907 bits (2344), Expect = 0.0
Identities = 457/516 (88%), Positives = 481/516 (92%), Gaps = 1/516 (0%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMA ALRRLSSS NK + LF+ +Y SSLP EAVY+KE V+WPKQLN+ LE D
Sbjct: 1 MAMASALRRLSSSSNKPLQRLFNGGHLYSMSSLPSEAVYEKERPGVTWPKQLNAPLEVGD 60
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIELEKARQWKGLELI SENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 61 PEIADIIELEKARQWKGLELILSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 120
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLD AKWGVNVQPLSGSP+NFHVYTALLK HDRIMALDLPHGGHLS
Sbjct: 121 MAETLCQKRALEAFRLDAAKWGVNVQPLSGSPANFHVYTALLKAHDRIMALDLPHGGHLS 180
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRL+ESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD
Sbjct: 181 HGYQTDTKKISAVSIFFETMPYRLNESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
YARIRKVCDKQKA++LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 241 YARIRKVCDKQKAILLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KG+KEVNKQGKEV YDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATT EYKAYQEQV
Sbjct: 301 KGVKEVNKQGKEVLYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTAEYKAYQEQV 360
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
+SNCAKFA+ L + GYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG
Sbjct: 361 MSNCAKFAETLVKSGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDF KVA FD +V LA+KIK E++GTKLKDFV +
Sbjct: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFAKVAYLFDLAVKLAVKIKGEAQGTKLKDFVAAM 480
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
QSS++ QSEISKLRHDVEE+AKQFPTIGFEK +MKY
Sbjct: 481 QSSAF-QSEISKLRHDVEEYAKQFPTIGFEKETMKY 515
>UniRef100_Q7Y1F0 Putative glycine hydroxymethyltransferase [Oryza sativa]
Length = 557
Score = 888 bits (2294), Expect = 0.0
Identities = 442/516 (85%), Positives = 480/516 (92%), Gaps = 5/516 (0%)
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMA ALR+LSS + +PL + +YY +SLP +E S V+WPKQLN+ LEE+D
Sbjct: 45 MAMATALRKLSSDALRR-QPLSRITPLYYMASLPAT----EERSGVTWPKQLNAPLEEVD 99
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIE EKARQWKGLELIPSENFTS+SVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 100 PEIADIIEHEKARQWKGLELIPSENFTSVSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 159
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAE+LCQKRALEAFRLDPAKWGVNVQPLSGSP+NFHVYTALLKPH+RIMALDLPHGGHLS
Sbjct: 160 MAESLCQKRALEAFRLDPAKWGVNVQPLSGSPANFHVYTALLKPHERIMALDLPHGGHLS 219
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTG IDYDQ+EKSA LFRPKLIVAGASAYARLYD
Sbjct: 220 HGYQTDTKKISAVSIFFETMPYRLDESTGLIDYDQMEKSAVLFRPKLIVAGASAYARLYD 279
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
Y R+RKVCDKQKA++LADMAHISGLVAAGV+PSPFDYADVVTTTTHKSLRGPRGAMIF+R
Sbjct: 280 YDRMRKVCDKQKAILLADMAHISGLVAAGVVPSPFDYADVVTTTTHKSLRGPRGAMIFYR 339
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KG+K VNKQGKEV YD+EDKIN AVFPGLQGGPHNHTITGLAVALKQATTPEY+AYQEQV
Sbjct: 340 KGVKGVNKQGKEVMYDFEDKINAAVFPGLQGGPHNHTITGLAVALKQATTPEYRAYQEQV 399
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
+SNCAKFAQ+L+ KGYELVSGGT+NHLVLVNLK+KGIDGSRVEKVLE VHIAANKNTVPG
Sbjct: 400 MSNCAKFAQSLTAKGYELVSGGTDNHLVLVNLKSKGIDGSRVEKVLENVHIAANKNTVPG 459
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDF KVA++FDA+VNLALK+KA + GTKLKDFV TL
Sbjct: 460 DVSAMVPGGIRMGTPALTSRGFVEEDFAKVADFFDAAVNLALKVKAAAGGTKLKDFVATL 519
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
QS S +QSEI+KLRHDVEE+AKQFPTIGFEK +MKY
Sbjct: 520 QSDSNIQSEIAKLRHDVEEYAKQFPTIGFEKETMKY 555
>UniRef100_Q94C74 Putative glycine hydroxymethyltransferase [Arabidopsis thaliana]
Length = 517
Score = 881 bits (2276), Expect = 0.0
Identities = 440/517 (85%), Positives = 476/517 (91%), Gaps = 1/517 (0%)
Query: 3 MAMALRRLSSSINKSSRPLFS-ASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDP 61
MA+ALRRLSSS+ K L S S+ + SSL A+ + E SR SW KQLN+SL+EIDP
Sbjct: 1 MALALRRLSSSVKKPISLLSSNGGSLRFMSSLSTAAMAESEKSRSSWIKQLNASLDEIDP 60
Query: 62 EIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDM 121
E+ADIIELEKARQWKG ELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYIDM
Sbjct: 61 EVADIIELEKARQWKGFELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYIDM 120
Query: 122 AETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSH 181
AETLCQKRALEAF+LDP+KWGVNVQ LSGSP+NF VYTALLKPH+RIMALDLPHGGHLSH
Sbjct: 121 AETLCQKRALEAFQLDPSKWGVNVQSLSGSPANFQVYTALLKPHERIMALDLPHGGHLSH 180
Query: 182 GYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDY 241
GYQTDTKKISAVSIFFETMPYRLDE+TGYIDYDQLEKSA LFRPKLIVAGASAYARLYDY
Sbjct: 181 GYQTDTKKISAVSIFFETMPYRLDENTGYIDYDQLEKSAVLFRPKLIVAGASAYARLYDY 240
Query: 242 ARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRK 301
ARIRKVC+KQKAVMLADMAHISGLVAAGVIPSPF+YADVVTTTTHKSLRGPRGAMIFFRK
Sbjct: 241 ARIRKVCNKQKAVMLADMAHISGLVAAGVIPSPFEYADVVTTTTHKSLRGPRGAMIFFRK 300
Query: 302 GLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVL 361
GLKE+NKQGKEV YDYED+INQAVFPGLQGGPHNHTITGLAVALKQA TPEYKAYQ+QVL
Sbjct: 301 GLKEINKQGKEVMYDYEDRINQAVFPGLQGGPHNHTITGLAVALKQARTPEYKAYQDQVL 360
Query: 362 SNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGD 421
NC+KFA+ L KGY+LVSGGT+NHLVLVNLKNKGIDGSRVEKVLE VHIAANKNTVPGD
Sbjct: 361 RNCSKFAETLLAKGYDLVSGGTDNHLVLVNLKNKGIDGSRVEKVLELVHIAANKNTVPGD 420
Query: 422 VSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQ 481
VSAMVPGGI MGTPALTSRGF+EEDF KVAEYFD +V +ALKIKAES+GTKLKDFV T+Q
Sbjct: 421 VSAMVPGGIHMGTPALTSRGFIEEDFAKVAEYFDLAVKIALKIKAESQGTKLKDFVATMQ 480
Query: 482 SSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
S+ +QSE+SKLR VEE+AKQFPTIGFEK +M+Y +
Sbjct: 481 SNEKLQSEMSKLREMVEEYAKQFPTIGFEKETMRYKE 517
>UniRef100_Q8GRI1 Putative glycine hydroxymethyltransferase [Arabidopsis thaliana]
Length = 533
Score = 874 bits (2257), Expect = 0.0
Identities = 442/533 (82%), Positives = 477/533 (88%), Gaps = 17/533 (3%)
Query: 3 MAMALRRLSSSINKSSRPLFS-ASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDP 61
MA+ALRRLSSS+ K L S S+ + SSL A+ + E SR SW KQLN+SL+EIDP
Sbjct: 1 MALALRRLSSSVKKPISLLSSNGGSLRFMSSLSTAAMAESEKSRSSWIKQLNASLDEIDP 60
Query: 62 EIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDM 121
E+ADIIELEKARQWKG ELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYIDM
Sbjct: 61 EVADIIELEKARQWKGFELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYIDM 120
Query: 122 AETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSH 181
AETLCQKRALEAF+LDP+KWGVNVQ LSGSP+NF VYTALLKPH+RIMALDLPHGGHLSH
Sbjct: 121 AETLCQKRALEAFQLDPSKWGVNVQSLSGSPANFQVYTALLKPHERIMALDLPHGGHLSH 180
Query: 182 GYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDY 241
GYQTDTKKISAVSIFFETMPYRLDE+TGYIDYDQLEKSA LFRPKLIVAGASAYARLYDY
Sbjct: 181 GYQTDTKKISAVSIFFETMPYRLDENTGYIDYDQLEKSAVLFRPKLIVAGASAYARLYDY 240
Query: 242 ARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRK 301
ARIRKVC+KQKAVMLADMAHISGLVAAGVIPSPF+YADVVTTTTHKSLRGPRGAMIFFRK
Sbjct: 241 ARIRKVCNKQKAVMLADMAHISGLVAAGVIPSPFEYADVVTTTTHKSLRGPRGAMIFFRK 300
Query: 302 GLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVL 361
GLKE+NKQGKEV YDYED+INQAVFPGLQGGPHNHTITGLAVALKQA TPEYKAYQ+QVL
Sbjct: 301 GLKEINKQGKEVMYDYEDRINQAVFPGLQGGPHNHTITGLAVALKQARTPEYKAYQDQVL 360
Query: 362 SNCAKFA----------------QALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKV 405
NC+KFA Q L KGY+LVSGGT+NHLVLVNLKNKGIDGSRVEKV
Sbjct: 361 RNCSKFAELDIRPTVIISYGLSMQTLLAKGYDLVSGGTDNHLVLVNLKNKGIDGSRVEKV 420
Query: 406 LEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIK 465
LE VHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGF+EEDF KVAEYFD +V +ALKIK
Sbjct: 421 LELVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFIEEDFAKVAEYFDLAVKIALKIK 480
Query: 466 AESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
AES+GTKLKDFV T+QS+ +QSE+SKLR VEE+AKQFPTIGFEK +M+Y +
Sbjct: 481 AESQGTKLKDFVATMQSNEKLQSEMSKLREMVEEYAKQFPTIGFEKETMRYKE 533
>UniRef100_Q8W4V3 Serine hydroxymethyltransferase [Chlamydomonas reinhardtii]
Length = 520
Score = 695 bits (1794), Expect = 0.0
Identities = 349/510 (68%), Positives = 415/510 (80%), Gaps = 5/510 (0%)
Query: 8 RRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDPEIADII 67
R LSS+ + + F+A D + V+WPK LN+ L E+DP++ DII
Sbjct: 13 RALSSAHAVAGQRCFAAQPALSNEEEYSRFSQDASRAHVTWPKVLNAGLAEVDPDLFDII 72
Query: 68 ELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQ 127
E EK RQ+KGLELIPSENF S SVM+AVGS+MTNKYSEGYPGARYYGGNE+ID AE LCQ
Sbjct: 73 EKEKNRQFKGLELIPSENFVSASVMEAVGSVMTNKYSEGYPGARYYGGNEFIDQAERLCQ 132
Query: 128 KRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDT 187
+RAL+AF LDPA+WGVNVQ LSGSPSNF VYTALL+PHDRIMALDLPHGGHLSHGYQTDT
Sbjct: 133 ERALKAFHLDPAQWGVNVQSLSGSPSNFQVYTALLQPHDRIMALDLPHGGHLSHGYQTDT 192
Query: 188 KKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKV 247
KKISA SI+FE MPYRL+E TG IDYD LEK+A LFRPKLIVAGASAY R YDYAR+R +
Sbjct: 193 KKISATSIYFEQMPYRLNEETGLIDYDMLEKTAVLFRPKLIVAGASAYTRHYDYARMRAI 252
Query: 248 CDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVN 307
DK A +LADMAHISGLVAA ++PSPF +ADVVTTTTHKSLRGPRGAMIF+RKG++ +
Sbjct: 253 ADKVGAWLLADMAHISGLVAADLVPSPFGFADVVTTTTHKSLRGPRGAMIFYRKGVRRTD 312
Query: 308 -KQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAK 366
K GK + YD EDKIN AVFPGLQGGPHNHTI GLA ALKQA TPE+K+YQ+QVLSN
Sbjct: 313 AKTGKPINYDIEDKINFAVFPGLQGGPHNHTIAGLACALKQAATPEFKSYQQQVLSNSQA 372
Query: 367 FAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMV 426
A AL+++G++LVSGGT+NH+VLV+L+ KG+DGSRVE+VLE HIAANKNTVPGDVSA+V
Sbjct: 373 LAGALAKRGFKLVSGGTDNHIVLVDLRPKGVDGSRVERVLELAHIAANKNTVPGDVSALV 432
Query: 427 PGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYV 486
PGG+RMG+PALTSRGFVE+DF +VAE+ D +VN+A+ +K K KLK+F E + S
Sbjct: 433 PGGLRMGSPALTSRGFVEKDFEQVAEFVDRAVNIAVDLK--KKYPKLKEFREAMAKES-- 488
Query: 487 QSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
+I+ L+ DVE FA +FPTIGF+K++M+Y
Sbjct: 489 TPDINALKKDVETFAMRFPTIGFDKAAMRY 518
>UniRef100_Q8W524 Serine hydroxymethyltransferase [Zea mays]
Length = 343
Score = 650 bits (1678), Expect = 0.0
Identities = 313/343 (91%), Positives = 334/343 (97%)
Query: 94 AVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPS 153
AVGS+MTNKYSEGYPGARYYGGNE+IDMAE+LCQKRALEAFRLDPAKWGVNVQPLSGSP+
Sbjct: 1 AVGSVMTNKYSEGYPGARYYGGNEFIDMAESLCQKRALEAFRLDPAKWGVNVQPLSGSPA 60
Query: 154 NFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDY 213
NFHVYTALLKPH+RIMALDLPHGGHLSHGYQTDTKKISA SIFFETMPYRLDESTG IDY
Sbjct: 61 NFHVYTALLKPHERIMALDLPHGGHLSHGYQTDTKKISATSIFFETMPYRLDESTGLIDY 120
Query: 214 DQLEKSATLFRPKLIVAGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPS 273
DQL+KSA LFRPKLI+AGASAYARLYDY R+RK+C KQKA++LADMAHISGLVAAGV+PS
Sbjct: 121 DQLKKSAVLFRPKLIIAGASAYARLYDYDRMRKICTKQKAILLADMAHISGLVAAGVVPS 180
Query: 274 PFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGP 333
PFDYADVVTTTTHKSLRGPRGAMIF+RKG+KE+NKQGKEV YD+EDKIN AVFPGLQGGP
Sbjct: 181 PFDYADVVTTTTHKSLRGPRGAMIFYRKGVKEINKQGKEVMYDFEDKINAAVFPGLQGGP 240
Query: 334 HNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLK 393
HNHTITGLAVALKQATTPEY+AYQEQV+SNCAKFAQ+L KGYELVSGGT+NHLVLVNLK
Sbjct: 241 HNHTITGLAVALKQATTPEYRAYQEQVISNCAKFAQSLISKGYELVSGGTDNHLVLVNLK 300
Query: 394 NKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPA 436
NKGIDGSRVEKVLE+VHIAANKNTVPGDVSAMVPGGIRMGTPA
Sbjct: 301 NKGIDGSRVEKVLESVHIAANKNTVPGDVSAMVPGGIRMGTPA 343
>UniRef100_UPI0000430024 UPI0000430024 UniRef100 entry
Length = 493
Score = 588 bits (1516), Expect = e-166
Identities = 282/457 (61%), Positives = 366/457 (79%)
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
++ S++++DPE+ADI+ E+ RQ + LIPSENFTS +VM +GS M NKYSEGYPG R
Sbjct: 35 ISKSVQDVDPEMADILNQERTRQKNSITLIPSENFTSKAVMDLLGSEMQNKYSEGYPGER 94
Query: 112 YYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMAL 171
YYGGNE ID AE LCQKRALEAF LDP++WGVNVQPLSG+P+N + Y+A+L+ DRIM L
Sbjct: 95 YYGGNEIIDKAEALCQKRALEAFGLDPSQWGVNVQPLSGAPANLYAYSAILEVGDRIMGL 154
Query: 172 DLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
DLPHGGHLSHGYQT+T KIS +S +F+TMPYRL+E TG IDYD LEK+A LFRPK+IVAG
Sbjct: 155 DLPHGGHLSHGYQTNTTKISYISKYFQTMPYRLNEETGIIDYDTLEKNAQLFRPKVIVAG 214
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
ASAY+R+ DY R+R++ DK A +L+DMAHISGLV+AGV SPF Y+D+VTTTTHKSLRG
Sbjct: 215 ASAYSRVIDYKRMRQIADKVGAYLLSDMAHISGLVSAGVTDSPFPYSDIVTTTTHKSLRG 274
Query: 292 PRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTP 351
PRGAMIFFRKG+++V K+GKE+ Y+ E KIN +VFPG QGGPHNHTI+ LAVALKQ T P
Sbjct: 275 PRGAMIFFRKGIRKVTKKGKEIPYELERKINFSVFPGHQGGPHNHTISALAVALKQCTEP 334
Query: 352 EYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHI 411
EY YQ++V+SN FA AL KG++LVS GT+ HL+LV+L+++ IDG+RVE VLE +I
Sbjct: 335 EYVKYQQEVVSNAKHFADALVSKGFKLVSDGTDTHLILVDLRSRNIDGARVEAVLERANI 394
Query: 412 AANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGT 471
AANKNTVPGDVSA+ P G+R+GTPA+T+RGF E+F KVAE+ D +VN+A+++KA+ +G
Sbjct: 395 AANKNTVPGDVSALFPSGLRVGTPAMTTRGFGPEEFDKVAEFIDQAVNIAIELKAQEQGK 454
Query: 472 KLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIG 508
K+ + + + + ++ +L +V + ++P G
Sbjct: 455 VPKELLASFKKLADESDKVKQLDKEVVSWVSKYPVPG 491
>UniRef100_UPI000042D39C UPI000042D39C UniRef100 entry
Length = 493
Score = 587 bits (1512), Expect = e-166
Identities = 281/457 (61%), Positives = 365/457 (79%)
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
++ S++++DPE+ADI+ E+ RQ + LIPSENFTS +VM +GS M NKYSEGYPG R
Sbjct: 35 ISKSVQDVDPEMADILNQERTRQKNSITLIPSENFTSKAVMDLLGSEMQNKYSEGYPGER 94
Query: 112 YYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMAL 171
YYGGNE ID AE LCQKRALEAF LDP++WGVNVQPLSG+P+N + Y+A+L+ DRIM L
Sbjct: 95 YYGGNEIIDKAEALCQKRALEAFGLDPSQWGVNVQPLSGAPANLYAYSAILEVGDRIMGL 154
Query: 172 DLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
DLPHGGHLSHGYQT+T KIS +S +F+TMPYRL+E TG IDYD LEK+A LFRPK+IVAG
Sbjct: 155 DLPHGGHLSHGYQTNTTKISYISKYFQTMPYRLNEETGIIDYDTLEKNAQLFRPKVIVAG 214
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
ASAY+R+ DY R+R++ DK A +L+DMAHISGLV+AGV SPF Y+D+VTTTTHKSLRG
Sbjct: 215 ASAYSRVIDYKRMRQIADKVGAYLLSDMAHISGLVSAGVTDSPFPYSDIVTTTTHKSLRG 274
Query: 292 PRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTP 351
PRGAMIFFRKG+++V K+GKE+ Y+ E KIN +VFPG QGGPHNHTI+ LAVALKQ T P
Sbjct: 275 PRGAMIFFRKGIRKVTKKGKEIPYELERKINFSVFPGHQGGPHNHTISALAVALKQCTEP 334
Query: 352 EYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHI 411
EY YQ++V+SN FA AL KG++LVS GT+ HL+LV+L+++ IDG+RVE VLE +I
Sbjct: 335 EYVKYQQEVVSNAKHFADALVSKGFKLVSDGTDTHLILVDLRSRNIDGARVEAVLERANI 394
Query: 412 AANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGT 471
A NKNTVPGDVSA+ P G+R+GTPA+T+RGF E+F KVAE+ D +VN+A+++KA+ +G
Sbjct: 395 ATNKNTVPGDVSALFPSGLRVGTPAMTTRGFGPEEFDKVAEFIDQAVNIAIELKAQEQGK 454
Query: 472 KLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIG 508
K+ + + + + ++ +L +V + ++P G
Sbjct: 455 VPKELLASFKKLADESDKVKQLDKEVVSWVSKYPVPG 491
>UniRef100_UPI000023F3A5 UPI000023F3A5 UniRef100 entry
Length = 502
Score = 582 bits (1500), Expect = e-165
Identities = 293/464 (63%), Positives = 363/464 (78%), Gaps = 11/464 (2%)
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
L SSLE+ DPEI I++ E+ RQ + LIPSENFTS SV+ A+GS+M NKYSEGYPGAR
Sbjct: 34 LGSSLEQGDPEIHAILKREEKRQNHFINLIPSENFTSRSVLDALGSVMQNKYSEGYPGAR 93
Query: 112 YYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMAL 171
YYGGNE+ID AE LCQ RALE FRLDP KWGVNVQPLSGSP+N + Y+A+L HDRIM L
Sbjct: 94 YYGGNEHIDEAERLCQSRALETFRLDPEKWGVNVQPLSGSPANLYAYSAILNTHDRIMGL 153
Query: 172 DLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
DLPHGGHLSHGYQ KKIS +S ++ET PYRL+E TG IDY++L ++A L+RPK+IVAG
Sbjct: 154 DLPHGGHLSHGYQIPGKKISMISKYYETFPYRLNEETGLIDYEKLRENALLYRPKVIVAG 213
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
SAY+RL DY R+R + ++ A +L+DMAH+SGLVAAGVI +PFD +D+VTTTTHKSLRG
Sbjct: 214 TSAYSRLIDYERMRAIANEAGAYLLSDMAHVSGLVAAGVIGTPFDDSDIVTTTTHKSLRG 273
Query: 292 PRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTP 351
PRGAMIF+RKG++ +K+GK++ YD E IN +VFPG QGGPHNHTIT LAVALKQA TP
Sbjct: 274 PRGAMIFYRKGVRSTDKKGKQIMYDLEGPINASVFPGHQGGPHNHTITALAVALKQAQTP 333
Query: 352 EYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHI 411
E+K YQE+VL+N A L++ GY LVSGGT+NHLVLV+LK KGIDG+RVE+VLE V +
Sbjct: 334 EFKDYQEKVLANSQAMANQLTDLGYSLVSGGTDNHLVLVDLKPKGIDGARVERVLELVGV 393
Query: 412 AANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKI------K 465
A+NKNTVPGD SA+ PGG+R+GTPA+T+RGF EDF +VA+ D V + L +
Sbjct: 394 ASNKNTVPGDRSALKPGGLRLGTPAMTTRGFNGEDFKRVADIVDRGVKITLAVDKDARAA 453
Query: 466 AESKGTK----LKDFVETLQSSSYVQSEISKLRHDVEEFAKQFP 505
AE+KG K +K+F+E L S V+ EI+ LR +V E+ FP
Sbjct: 454 AEAKGAKNPGTVKNFLEFLGDGSSVK-EIAALRDEVAEWVGGFP 496
>UniRef100_UPI0000337BD6 UPI0000337BD6 UniRef100 entry
Length = 501
Score = 580 bits (1496), Expect = e-164
Identities = 294/502 (58%), Positives = 365/502 (72%), Gaps = 7/502 (1%)
Query: 10 LSSSINKSSRPLFSA--SSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDPEIADII 67
LS ++ K +RPL + + + + A + E R W Q SL + DPE+ ++
Sbjct: 2 LSLALRKLARPLCQRVPACLSVRGQQSNAATHAVEEDR-PWTGQ--ESLAQDDPEMWGLL 58
Query: 68 ELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQ 127
+ EK RQ +GLELI SENF S + ++A+GS + NKYSEGYPG RYYGG E +D E LCQ
Sbjct: 59 QKEKDRQCRGLELIASENFCSRAALEALGSCLNNKYSEGYPGRRYYGGEEVVDQIELLCQ 118
Query: 128 KRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDT 187
KRAL+AF LDPA WGVNVQP SGSP+NF VYTA+LKPHDRIM LDLP GGHL+HGY +D
Sbjct: 119 KRALQAFDLDPALWGVNVQPYSGSPANFAVYTAVLKPHDRIMGLDLPDGGHLTHGYMSDV 178
Query: 188 KKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKV 247
K+ISA SI+FE+MPY+L+ +TG IDYDQ+E +A LFRPK+I+AG SAYARL DYARI+K+
Sbjct: 179 KRISATSIYFESMPYKLNPATGLIDYDQMEMTAKLFRPKIIIAGTSAYARLIDYARIKKL 238
Query: 248 CDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVN 307
C A ++ADMAHISGLVAAG IPSPF+YAD+VT+TTHKSLRG R +IF+RKG++ +
Sbjct: 239 CTSVNAYLMADMAHISGLVAAGAIPSPFEYADLVTSTTHKSLRGARSGLIFYRKGIRSKD 298
Query: 308 KQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKF 367
K+GKE+ YD EDK+N +VFP LQGGPHNH I G+AVALKQA +P +K Y QVL N
Sbjct: 299 KKGKEIMYDLEDKVNFSVFPSLQGGPHNHGIAGVAVALKQAQSPMFKDYIAQVLKNAKAM 358
Query: 368 AQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVP 427
A AL KGY LVSGGT+NHLVLV+L+ GIDG+R E+VLE I ANKNT PGD SA+ P
Sbjct: 359 AAALISKGYTLVSGGTDNHLVLVDLRPMGIDGARAERVLELASITANKNTCPGDTSALTP 418
Query: 428 GGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQ 487
GG+R+G PALTSR F E DFV+V E+ D + L +K K KL++F L
Sbjct: 419 GGLRLGAPALTSRQFKEADFVQVVEFMDEGFKIGLDVK--KKTGKLQEFKNFLVQDPETV 476
Query: 488 SEISKLRHDVEEFAKQFPTIGF 509
+ I+ LRH VE FA+ FP GF
Sbjct: 477 ARIADLRHRVEAFARPFPMPGF 498
>UniRef100_Q68EQ3 Shmt2-prov protein [Xenopus tropicalis]
Length = 496
Score = 580 bits (1494), Expect = e-164
Identities = 293/503 (58%), Positives = 373/503 (73%), Gaps = 10/503 (1%)
Query: 10 LSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDPEIADIIEL 69
L+ S+ + ++PL S V + S + + ++V W Q SL E DPE+ D+++
Sbjct: 2 LTFSLRRLAQPLRSCCHVRSQHS----QAWTQAGNQV-WTGQ--ESLAEGDPEMWDLVQK 54
Query: 70 EKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKR 129
EK RQ +GLE+I ENF S + ++A+GS + NKYSEGYPG RYYGG E +D E LCQ+R
Sbjct: 55 EKDRQCRGLEMIALENFCSRAALEALGSCLNNKYSEGYPGKRYYGGAEVVDKIELLCQQR 114
Query: 130 ALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDTKK 189
AL+AF L+P KWGVNVQP SGSP+NF YTA+L+PHDRIM LDLP GGHL+HGY +D K+
Sbjct: 115 ALDAFDLNPEKWGVNVQPYSGSPANFAAYTAVLQPHDRIMGLDLPDGGHLTHGYMSDVKR 174
Query: 190 ISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKVCD 249
ISA SI+FE+MPY+L+ +TG IDYDQLE +A LFRPKLI+AG SAYARL DYAR+RKVCD
Sbjct: 175 ISATSIYFESMPYKLNPATGLIDYDQLEMTARLFRPKLIIAGTSAYARLIDYARMRKVCD 234
Query: 250 KQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNKQ 309
+ KA +LADMAHISGLVAAGVIPSPF++AD+VT+TTHK+LRG R +IF+RKG+K V+K+
Sbjct: 235 EMKAYLLADMAHISGLVAAGVIPSPFEHADIVTSTTHKTLRGARSGLIFYRKGVKSVDKK 294
Query: 310 -GKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFA 368
GK+V Y+ EDK+N +VFP +QGGPHNH I +AVALKQA++P ++ Y QVL N A
Sbjct: 295 TGKDVLYNLEDKVNFSVFPSIQGGPHNHAIAAVAVALKQASSPMFREYAVQVLKNAKSMA 354
Query: 369 QALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPG 428
AL KGY LVSGGT+NHLVLV+L+ KGIDG+R E+VLE V I ANKNT PGD SA+ PG
Sbjct: 355 AALLSKGYTLVSGGTDNHLVLVDLRPKGIDGARAERVLELVSITANKNTCPGDKSALTPG 414
Query: 429 GIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQS 488
G+R+G PALTSR F E DF KV + D + + L +K K KL+DF L +
Sbjct: 415 GLRLGAPALTSRNFKEADFEKVVHFIDEGIRIGLDVK--RKTNKLQDFKNFLLEDHETVN 472
Query: 489 EISKLRHDVEEFAKQFPTIGFEK 511
I+ LR VE+FA+ FP GF++
Sbjct: 473 RIADLRKQVEQFARSFPMPGFDE 495
>UniRef100_Q8AVC0 Shmt1-prov protein [Xenopus laevis]
Length = 485
Score = 580 bits (1494), Expect = e-164
Identities = 285/461 (61%), Positives = 354/461 (75%), Gaps = 2/461 (0%)
Query: 50 KQLNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPG 109
K + L+ DPE+ +II EK RQ GLELI SENF S +V+QA+GS + NKYSEGYPG
Sbjct: 21 KMVLEPLDTNDPEVYEIIRKEKHRQRYGLELIASENFASCAVLQALGSCLNNKYSEGYPG 80
Query: 110 ARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIM 169
RYYGG E++D E LCQKRALE + L+P KWGVNVQP SGSP+NF +YTAL++PH RIM
Sbjct: 81 QRYYGGTEFVDEMERLCQKRALEVYGLEPQKWGVNVQPYSGSPANFAIYTALVEPHGRIM 140
Query: 170 ALDLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIV 229
LDLP GGHL+HG+ TD KKISA SIFFE+MPY++ TGYIDYD+LE++A LF PK+I+
Sbjct: 141 GLDLPDGGHLTHGFMTDKKKISATSIFFESMPYKVHPETGYIDYDRLEENARLFHPKMII 200
Query: 230 AGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSL 289
AG S Y+R DYAR+R++ D+ AV++ADMAHISGLVAAGV+PSPF++ DVV+TTTHK+L
Sbjct: 201 AGVSCYSRNLDYARMRRIADENNAVLMADMAHISGLVAAGVVPSPFEHCDVVSTTTHKTL 260
Query: 290 RGPRGAMIFFRKGLKEVN-KQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQA 348
RG R MIF+RKG++ V+ K GKE Y+YE INQAVFPGLQGGPHNH I G+AVALKQA
Sbjct: 261 RGCRSGMIFYRKGVRSVDPKTGKETLYNYESLINQAVFPGLQGGPHNHAIAGVAVALKQA 320
Query: 349 TTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEA 408
+PE+K YQ+QV+SNC + A+ E GY +V+GG++NHL+LVNL++K DG R EKVLEA
Sbjct: 321 LSPEFKLYQKQVVSNCKALSLAIEELGYHVVTGGSDNHLILVNLRDKKTDGGRAEKVLEA 380
Query: 409 VHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKI-KAE 467
IA NKNT PGD SA+ P G+R+GTPALTSRGF EEDF KVA++ + L L+I K+
Sbjct: 381 CSIACNKNTCPGDKSALRPSGLRLGTPALTSRGFNEEDFKKVAQFIHRGIELTLEIQKSM 440
Query: 468 SKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIG 508
+ G LKDF E L S +I LR +VE+FA FP G
Sbjct: 441 NPGATLKDFKEKLASQDVHTPKILALRAEVEKFAGTFPIPG 481
>UniRef100_Q6AXB3 MGC79128 protein [Xenopus laevis]
Length = 496
Score = 579 bits (1492), Expect = e-164
Identities = 285/458 (62%), Positives = 352/458 (76%), Gaps = 3/458 (0%)
Query: 55 SLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYG 114
S+ E DPE+ D+++ EK RQ +GLELI SENF S + ++A+GS + NKYSEGYPG RYYG
Sbjct: 40 SMAEGDPEMWDLVQKEKDRQCRGLELIASENFCSRAALEALGSCLNNKYSEGYPGKRYYG 99
Query: 115 GNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLP 174
G E +D E LCQ+RAL+AF LDP KWGVNVQP SGSP+NF YTA+L+PHDRIM LDLP
Sbjct: 100 GAEVVDQIELLCQQRALDAFDLDPEKWGVNVQPYSGSPANFAAYTAVLQPHDRIMGLDLP 159
Query: 175 HGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASA 234
GGHL+HGY +D K+ISA SI+FE+MPY+L+ +TG I+YDQLE +A LFRPKLI+AG SA
Sbjct: 160 DGGHLTHGYMSDVKRISATSIYFESMPYKLNPATGLINYDQLEMTARLFRPKLIIAGTSA 219
Query: 235 YARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRG 294
YARL DYA++RKVCD+ KA +LADMAHISGLVAAGVIPSPF +AD+VT+TTHK+LRG R
Sbjct: 220 YARLIDYAKMRKVCDEVKAYLLADMAHISGLVAAGVIPSPFQHADIVTSTTHKTLRGARS 279
Query: 295 AMIFFRKGLKEVNKQ-GKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEY 353
+IFFRKG+K V+K+ GKE+ Y+ EDKIN +VFP +QGGPHNH I +AVALKQA++P +
Sbjct: 280 GLIFFRKGVKSVDKKTGKEIPYNLEDKINFSVFPSIQGGPHNHAIAAVAVALKQASSPMF 339
Query: 354 KAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAA 413
+ Y QVL N A AL KGY LVSGGT+NHLVLV+L+ KGIDG+R E+VLE V I A
Sbjct: 340 REYAVQVLKNAKSMAAALLSKGYTLVSGGTDNHLVLVDLRPKGIDGARAERVLELVSITA 399
Query: 414 NKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKL 473
NKNT PGD SA+ PGG+R+G PALTSR F E DF KV ++ D + + L +K K KL
Sbjct: 400 NKNTCPGDKSALTPGGLRLGAPALTSRNFKEADFEKVVDFIDEGIRIGLDVK--RKTNKL 457
Query: 474 KDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEK 511
+DF L I LR VE+FA+ FP GF++
Sbjct: 458 QDFKNFLLEDQETVKRIGDLRKQVEQFARAFPMPGFDE 495
>UniRef100_P34897 Serine hydroxymethyltransferase, mitochondrial precursor [Homo
sapiens]
Length = 504
Score = 577 bits (1488), Expect = e-163
Identities = 286/458 (62%), Positives = 355/458 (77%), Gaps = 3/458 (0%)
Query: 55 SLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYG 114
SL + DPE+ ++++ EK RQ +GLELI SENF S + ++A+GS + NKYSEGYPG RYYG
Sbjct: 48 SLSDSDPEMWELLQREKDRQCRGLELIASENFCSRAALEALGSCLNNKYSEGYPGKRYYG 107
Query: 115 GNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLP 174
G E +D E LCQ+RALEAF LDPA+WGVNVQP SGSP+N VYTALL+PHDRIM LDLP
Sbjct: 108 GAEVVDEIELLCQRRALEAFDLDPAQWGVNVQPYSGSPANLAVYTALLQPHDRIMGLDLP 167
Query: 175 HGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASA 234
GGHL+HGY +D K+ISA SIFFE+MPY+L+ TG IDY+QL +A LFRP+LI+AG SA
Sbjct: 168 DGGHLTHGYMSDVKRISATSIFFESMPYKLNPKTGLIDYNQLALTARLFRPRLIIAGTSA 227
Query: 235 YARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRG 294
YARL DYAR+R+VCD+ KA +LADMAHISGLVAA VIPSPF +AD+VTTTTHK+LRG R
Sbjct: 228 YARLIDYARMREVCDEVKAHLLADMAHISGLVAAKVIPSPFKHADIVTTTTHKTLRGARS 287
Query: 295 AMIFFRKGLKEVN-KQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEY 353
+IF+RKG+K V+ K G+E+ Y +ED+IN AVFP LQGGPHNH I +AVALKQA TP +
Sbjct: 288 GLIFYRKGVKAVDPKTGREIPYTFEDRINFAVFPSLQGGPHNHAIAAVAVALKQACTPMF 347
Query: 354 KAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAA 413
+ Y QVL N A AL E+GY LVSGGT+NHLVLV+L+ KG+DG+R E+VLE V I A
Sbjct: 348 REYSLQVLKNARAMADALLERGYSLVSGGTDNHLVLVDLRPKGLDGARAERVLELVSITA 407
Query: 414 NKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKL 473
NKNT PGD SA+ PGG+R+G PALTSR F E+DF +V ++ D VN+ L++K SK KL
Sbjct: 408 NKNTCPGDRSAITPGGLRLGAPALTSRQFREDDFRRVVDFIDEGVNIGLEVK--SKTAKL 465
Query: 474 KDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEK 511
+DF L S ++ LR VE+FA+ FP GF++
Sbjct: 466 QDFKSFLLKDSETSQRLANLRQRVEQFARAFPMPGFDE 503
>UniRef100_O13425 Serine hydroxymethyltransferase, mitochondrial precursor [Candida
albicans]
Length = 493
Score = 575 bits (1482), Expect = e-162
Identities = 278/457 (60%), Positives = 361/457 (78%)
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
++ S++++DPE+ADI+ E+ RQ + LIPSENFTS +VM +GS M NKYSEGYPG R
Sbjct: 35 ISKSVQDVDPEMADILNQERTRQKNSITLIPSENFTSKAVMDLLGSEMQNKYSEGYPGER 94
Query: 112 YYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMAL 171
YYGGNE ID AE LCQKRALEAF LDP++WGVNVQPLSG+P+N + Y+A+L+ DRIM L
Sbjct: 95 YYGGNEIIDKAEALCQKRALEAFGLDPSQWGVNVQPLSGAPANLYAYSAILEVGDRIMGL 154
Query: 172 DLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
DLPHGGHLSHGYQT T KIS +S +F+TMPYRL+E TG IDYD LEK+A LFRPK+IVAG
Sbjct: 155 DLPHGGHLSHGYQTKTTKISYISKYFQTMPYRLNEETGIIDYDTLEKNAQLFRPKVIVAG 214
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
ASAY+R+ DY R+R++ + A +L+DMAHISGLV+A V SPF Y+D+VTTTTHKSLRG
Sbjct: 215 ASAYSRVIDYKRMRQLSIRLGAYLLSDMAHISGLVSAVVTDSPFPYSDIVTTTTHKSLRG 274
Query: 292 PRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTP 351
PRGAMIFFRKG+++V +GKE+ Y+ E KIN VFPG QGGPHNHTI+ LAVALKQ T P
Sbjct: 275 PRGAMIFFRKGIRKVTTKGKEIPYELERKINFLVFPGHQGGPHNHTISALAVALKQCTEP 334
Query: 352 EYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHI 411
EY YQ++V+SN FA AL KG++LVS GT+ HL+LV+L+++ IDG+RVE VLE +I
Sbjct: 335 EYVKYQQEVVSNAKHFADALVSKGFKLVSDGTDTHLILVDLRSRNIDGARVEAVLERANI 394
Query: 412 AANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGT 471
AANKNTVPGDVSA+ P G+R+GTPA+T+RGF E+F KVAE+ D +VN+A+++KA+ +G
Sbjct: 395 AANKNTVPGDVSALFPSGLRVGTPAMTTRGFGPEEFDKVAEFIDQAVNIAIELKAQEQGK 454
Query: 472 KLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIG 508
K+ + + + + ++ +L +V + ++P G
Sbjct: 455 VPKELLASFKKLADESDKVKQLDKEVVSWVSKYPVPG 491
>UniRef100_UPI0000365291 UPI0000365291 UniRef100 entry
Length = 480
Score = 575 bits (1481), Expect = e-162
Identities = 289/473 (61%), Positives = 352/473 (74%), Gaps = 6/473 (1%)
Query: 37 AVYDKENSRVSWPKQLNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVG 96
A + E+ R W Q SL + DPE+ D+++ EK RQ +GLELI SENF S + ++A+G
Sbjct: 11 ATHTMEDDR-PWTGQ--ESLAQDDPEMWDLVQKEKDRQRRGLELIASENFCSRAALEALG 67
Query: 97 SIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFH 156
S + NKYSEGYPG RYYGG E +D E LCQKRAL+AF LDP WGVNVQP SGSP+NF
Sbjct: 68 SCLNNKYSEGYPGRRYYGGAEVVDQIELLCQKRALQAFDLDPTLWGVNVQPYSGSPANFA 127
Query: 157 VYTALLKPHDRIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQL 216
VYTA+LKPHDRIM LDLP GGHL+HGY +D K+ISA SI+FE+MPY+L+ +TG IDYDQ+
Sbjct: 128 VYTAVLKPHDRIMGLDLPDGGHLTHGYMSDVKRISATSIYFESMPYKLNPATGLIDYDQM 187
Query: 217 EKSATLFRPKLIVAGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFD 276
E +A LFRPK+I+AG SAYARL DYARI+K+C A MLADMAHISGLVAAG IPSPF+
Sbjct: 188 EMTAKLFRPKIIIAGTSAYARLIDYARIKKLCSNINAYMLADMAHISGLVAAGAIPSPFE 247
Query: 277 YADVVTTTTHKSLRGPRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNH 336
YAD+VT+TTHKSLRG R +IF+ KG++ +K+G+E+ YD EDK+N +VFP LQGGPHNH
Sbjct: 248 YADLVTSTTHKSLRGARSGLIFY-KGIRSKDKKGREIMYDLEDKVNFSVFPSLQGGPHNH 306
Query: 337 TITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKG 396
I G+AVALKQA +P +K Y QVL N A AL KGY LVSGGT+NHLVLV+L+ G
Sbjct: 307 GIAGVAVALKQAQSPMFKDYIAQVLKNAKAMATALISKGYTLVSGGTDNHLVLVDLRPMG 366
Query: 397 IDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDA 456
IDG+R E+VLE I ANKNT PGD SA+ PGG+R+G PALTSR F E DFVKV E+ D
Sbjct: 367 IDGARAERVLELASITANKNTCPGDTSALTPGGLRLGAPALTSRQFKEADFVKVVEFMDE 426
Query: 457 SVNLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGF 509
+ L +K K KL++F L + I+ LRH VE FA+ FP GF
Sbjct: 427 GFKIGLDVK--KKTGKLQEFKNVLVQDPETVARIADLRHRVEAFARPFPMPGF 477
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.132 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 840,709,142
Number of Sequences: 2790947
Number of extensions: 35278462
Number of successful extensions: 99732
Number of sequences better than 10.0: 1076
Number of HSP's better than 10.0 without gapping: 1006
Number of HSP's successfully gapped in prelim test: 70
Number of HSP's that attempted gapping in prelim test: 96242
Number of HSP's gapped (non-prelim): 1116
length of query: 518
length of database: 848,049,833
effective HSP length: 132
effective length of query: 386
effective length of database: 479,644,829
effective search space: 185142903994
effective search space used: 185142903994
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC126010.6


