

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126009.11 - phase: 0
(176 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
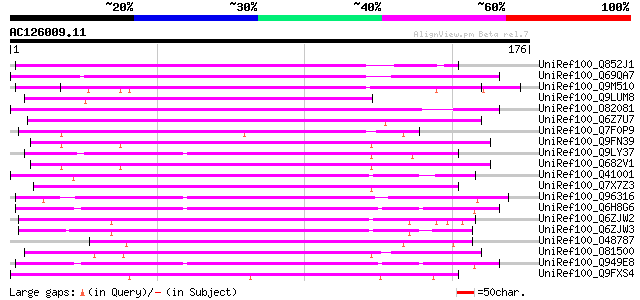
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q852J1 Putative blue copper-binding protein [Oryza sat... 105 5e-22
UniRef100_Q69QA7 Putative blue copper protein [Oryza sativa] 103 1e-21
UniRef100_Q9M510 Dicyanin [Lycopersicon esculentum] 96 4e-19
UniRef100_Q9LUM8 Blue copper-binding protein-like [Arabidopsis t... 96 5e-19
UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana] 96 5e-19
UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa] 93 3e-18
UniRef100_Q7F0P9 Putative blue copper protein [Oryza sativa] 92 7e-18
UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like pr... 90 2e-17
UniRef100_Q9LY37 Stellacyanin (Uclacyanin 3)-like protein [Arabi... 90 3e-17
UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis tha... 89 4e-17
UniRef100_Q41001 Blue copper protein precursor [Pisum sativum] 89 6e-17
UniRef100_Q7X7Z3 OSJNBa0079A21.5 protein [Oryza sativa] 88 8e-17
UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis th... 88 1e-16
UniRef100_Q6H8G6 Putative uclacyanin 3 [Oryza sativa] 87 1e-16
UniRef100_Q6ZJW2 Putative blue copper binding protein [Oryza sat... 87 2e-16
UniRef100_Q6ZJW3 Putative blue copper binding protein [Oryza sat... 86 5e-16
UniRef100_O48787 Putative phytocyanin [Arabidopsis thaliana] 85 7e-16
UniRef100_O81500 F9D12.16 protein [Arabidopsis thaliana] 85 9e-16
UniRef100_Q949E8 Uclacyanin 3-like protein [Oryza sativa] 84 1e-15
UniRef100_Q9FXS4 NtEIG-A1 protein [Nicotiana tabacum] 84 2e-15
>UniRef100_Q852J1 Putative blue copper-binding protein [Oryza sativa]
Length = 172
Score = 105 bits (262), Expect = 5e-22
Identities = 58/150 (38%), Positives = 79/150 (52%), Gaps = 11/150 (7%)
Query: 3 LSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGG 62
L+ A AL + + A A + +VGD GW FDY W A K FR+GDTL F Y G
Sbjct: 7 LAVAAAAVALAVLLPARGAAATEHMVGDGNGWILGFDYAAWAATKQFRVGDTLVFRYKGT 66
Query: 63 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 122
VV V G+DFK+C+ +A +SG+D++ + GRRW+ V DHC KL ITV
Sbjct: 67 NHTVVEVGGADFKACNKTASANEWSSGEDRVALDKEGRRWFFCGVGDHCAKNMKLKITV- 125
Query: 123 PKQDGWSPVPSPSPSPSLDLVTPEAPPSNA 152
+ + +P+P P PPS+A
Sbjct: 126 --------IAAGAPAPGASEAPP--PPSSA 145
>UniRef100_Q69QA7 Putative blue copper protein [Oryza sativa]
Length = 198
Score = 103 bits (258), Expect = 1e-21
Identities = 56/166 (33%), Positives = 80/166 (47%), Gaps = 9/166 (5%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MA S AL L++ A K + VGD GW DY TW ++K F++GDTL FNY
Sbjct: 1 MASSLALVALLLVSCAVVAAAATK-YTVGDTSGWAMGADYTTWASDKKFKMGDTLVFNYA 59
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
GG +V V+ +D+ +C+ +SG + + T G+ ++I + HC NG KL +
Sbjct: 60 GGAHSVDEVSAADYAACTASNALQSDSSGTTTVTLKTAGKHYFICGIAGHCSNGMKLVVD 119
Query: 121 VQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 166
V SP+P+P TP P + PAS L P
Sbjct: 120 V--------AAASPAPAPKAPSTTPTTPSTTPATPASPGTSSGLTP 157
>UniRef100_Q9M510 Dicyanin [Lycopersicon esculentum]
Length = 332
Score = 95.9 bits (237), Expect = 4e-19
Identities = 52/148 (35%), Positives = 82/148 (55%), Gaps = 6/148 (4%)
Query: 18 STMAVAKDFVVGDEKGWTTLFD----YQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSD 73
S +VA+ ++VGD GW+ YQ W NK F++GDTL FN+V G NV V+ +
Sbjct: 165 SPASVAQTYIVGDNLGWSVPTSGPNSYQRWANNKSFKVGDTLVFNFVNGTHNVAMVSKAS 224
Query: 74 FKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPS 133
+ SC+ +++G +I +T G +Y+ + HC GQKL I V D +P PS
Sbjct: 225 YDSCNTTSPINTISNGPARIRLTNSGEHYYMCTFPRHCSLGQKLAINV-TGSDVTAPTPS 283
Query: 134 PSPSPSLDLV-TPEAPPSNAPWPASSVP 160
+ +PS V + ++P ++ P P++S P
Sbjct: 284 TAATPSSPTVPSSDSPVNSPPAPSASAP 311
Score = 87.4 bits (215), Expect = 1e-16
Identities = 57/177 (32%), Positives = 85/177 (47%), Gaps = 14/177 (7%)
Query: 3 LSRALFLFALIASIFSTMAVAKD-FVVGDEKGWTT----LFDYQTWTANKVFRLGDTLTF 57
LS + +++ ++ +A+A+ VVGD GWT Y TW A K F +GD L F
Sbjct: 5 LSSLVVFGSILFALLQHVAMAQQTHVVGDTLGWTVPNGGAASYSTWAAGKSFVVGDILVF 64
Query: 58 NYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKL 117
N+ G +V V+ F SC+ + T+G I +++ G +Y+ + HC GQKL
Sbjct: 65 NFRSGSHSVAEVSKGAFDSCNTSSPISISTNGPTNITLSSAGSHYYLCTFPSHCTLGQKL 124
Query: 118 FITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSV-PRRSLLPKKLFQIF 173
I V SP P P+P+ P A P AP P+ SV P + P + Q +
Sbjct: 125 AINVSGSA---SPAPQPAPA-----TPPTATPVMAPAPSVSVAPATAPSPASVAQTY 173
>UniRef100_Q9LUM8 Blue copper-binding protein-like [Arabidopsis thaliana]
Length = 129
Score = 95.5 bits (236), Expect = 5e-19
Identities = 45/120 (37%), Positives = 66/120 (54%), Gaps = 2/120 (1%)
Query: 6 ALFLFALIASIFSTMAVAK--DFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGK 63
+L + I S+ S M ++ + +VGD GW +Y WT + F +GD L FNY +
Sbjct: 10 SLIILYAIFSLSSLMLKSEGTEHIVGDSNGWELFTNYTNWTQGREFHVGDVLVFNYKSDQ 69
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP 123
NV++VN + + C + + T G D II++ G+ W+I V DHC NGQKL I V P
Sbjct: 70 HNVMQVNSTAYTDCGLDNYTTLFTKGNDSIILSEVGKLWFICGVDDHCVNGQKLSINVAP 129
>UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana]
Length = 261
Score = 95.5 bits (236), Expect = 5e-19
Identities = 53/166 (31%), Positives = 78/166 (46%), Gaps = 10/166 (6%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MA L + +++A+ + VA D +G GWT +TW A + F +GD L F+Y
Sbjct: 1 MASREMLIIISVLATTLIGLTVATDHTIGGPSGWTVGASLRTWAAGQTFAVGDNLVFSYP 60
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
+VV V +F SC +G + +TT G+R++I + HC G KL +
Sbjct: 61 AAFHDVVEVTKPEFDSCQAVKPLITFANGNSLVPLTTPGKRYFICGMPGHCSQGMKLEVN 120
Query: 121 VQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 166
V P P P+ PSL NAP P+S +P + LLP
Sbjct: 121 VVPTATVAPTAPLPNTVPSL----------NAPSPSSVLPIQPLLP 156
>UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa]
Length = 246
Score = 92.8 bits (229), Expect = 3e-18
Identities = 46/164 (28%), Positives = 77/164 (46%), Gaps = 10/164 (6%)
Query: 7 LFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNV 66
L + AL+ + ++ A A + VGD GWT DY +W +K F++GD+L F Y G V
Sbjct: 9 LTIAALLLAACASSAAATSYTVGDASGWTIGVDYTSWAGSKSFKVGDSLVFKYASGAHTV 68
Query: 67 VRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQD 126
V V+ + + +C+ +SG + + T G+ ++I ++ HC G K+ + V
Sbjct: 69 VEVSAAGYLACAAANALGSDSSGSTTVALKTPGKHYFICTIAGHCAGGMKMEVDVSGSSS 128
Query: 127 ----------GWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVP 160
G PS SP+ P P P+P++ +P
Sbjct: 129 SSSGGGGGGGGGGSTPSSPSSPTPTTPNPSTPTPTTPYPSTPMP 172
>UniRef100_Q7F0P9 Putative blue copper protein [Oryza sativa]
Length = 168
Score = 91.7 bits (226), Expect = 7e-18
Identities = 51/140 (36%), Positives = 73/140 (51%), Gaps = 7/140 (5%)
Query: 4 SRALFLFALIASI--FSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 61
SR + L A+++++ M A D+ VGD GWT + W K F++GD L F Y
Sbjct: 3 SRQVLLLAIVSAVALLPAMVSATDYTVGDGHGWTLEYPSTNWADGKSFQIGDKLVFTYTK 62
Query: 62 GKDNVVRVNGSDFKSCS-VPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
GK V V+G+ F +C+ T SG D + + G+RW+ +V +HCE G KL +
Sbjct: 63 GKHTVTEVDGAAFHACNRQGNTLMTWNSGNDTVALDKAGKRWFFCNVDNHCELGMKLVVD 122
Query: 121 VQPKQDGWSPVP-SPSPSPS 139
V D +P P SP P PS
Sbjct: 123 V---ADPNAPAPASPPPPPS 139
>UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like protein [Arabidopsis
thaliana]
Length = 370
Score = 90.1 bits (222), Expect = 2e-17
Identities = 57/165 (34%), Positives = 82/165 (49%), Gaps = 9/165 (5%)
Query: 8 FLFALIASI--FSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNYVGGK 63
F F ++AS F ++A A F VG W T +Y TW F++ D+L F Y G
Sbjct: 10 FSFLILASFATFFSVADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSLYFKYAKGS 69
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV-- 121
D+V +V +DF C+V +G+ + + G ++IS DHC+ GQKL + V
Sbjct: 70 DSVQQVMKADFDGCNVRNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIVVVLA 129
Query: 122 ---QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 163
QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 130 VRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
>UniRef100_Q9LY37 Stellacyanin (Uclacyanin 3)-like protein [Arabidopsis thaliana]
Length = 187
Score = 89.7 bits (221), Expect = 3e-17
Identities = 55/154 (35%), Positives = 74/154 (47%), Gaps = 10/154 (6%)
Query: 6 ALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDN 65
AL L L+ ++ + AV F VGD GWT +Y +W + K FR+GDTL F Y G +
Sbjct: 8 ALLLLLLLVAVPAVFAVT--FQVGDNDGWTIGVEYTSWVSEKTFRVGDTLEFKY-GPSHS 64
Query: 66 VVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV---- 121
V VN +D+ C + G KI +T G ++ HC G KL + V
Sbjct: 65 VAVVNKADYDGCETSRPTQSFSDGDTKIDLTKVGAIHFLCLTPGHCSLGMKLAVQVLAAV 124
Query: 122 --QPKQDGWSPVPSPS-PSPSLDLVTPEAPPSNA 152
+P +P PSPS PSPS +P P NA
Sbjct: 125 SLEPPPSPSAPSPSPSAPSPSPSAPSPSPSPGNA 158
>UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis thaliana]
Length = 370
Score = 89.4 bits (220), Expect = 4e-17
Identities = 57/165 (34%), Positives = 82/165 (49%), Gaps = 9/165 (5%)
Query: 8 FLFALIASI--FSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNYVGGK 63
F F ++AS F ++A A F VG W T +Y TW F++ D+L F Y G
Sbjct: 10 FSFLILASFATFFSVADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSLYFKYEKGS 69
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV-- 121
D+V +V +DF C+V +G+ + + G ++IS DHC+ GQKL + V
Sbjct: 70 DSVQQVMKADFDGCNVRNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIVVVLA 129
Query: 122 ---QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 163
QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 130 VRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
>UniRef100_Q41001 Blue copper protein precursor [Pisum sativum]
Length = 189
Score = 88.6 bits (218), Expect = 6e-17
Identities = 57/159 (35%), Positives = 81/159 (50%), Gaps = 7/159 (4%)
Query: 1 MALSRALFLFALIASIFSTM-AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNY 59
MA S AL L L+A I + ++A + VGD GW DY TW ++K F +GD+L FNY
Sbjct: 1 MAFSNALVLCFLLAIINMALPSLATVYTVGDTSGWVIGGDYSTWASDKTFAVGDSLVFNY 60
Query: 60 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 119
G V V SD+KSC+ + ++G I + G+ ++I V H G KL I
Sbjct: 61 GAGAHTVDEVKESDYKSCTSGNSISTDSTGATTIPLKKAGKHYFICGVPGHSTGGMKLSI 120
Query: 120 TVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASS 158
V+ G S PS +PS S + PS+ PA++
Sbjct: 121 KVK-ASSGSSAAPSATPSSS-----GKGSPSSDDTPAAT 153
>UniRef100_Q7X7Z3 OSJNBa0079A21.5 protein [Oryza sativa]
Length = 181
Score = 88.2 bits (217), Expect = 8e-17
Identities = 51/147 (34%), Positives = 69/147 (46%), Gaps = 3/147 (2%)
Query: 9 LFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVR 68
L A++ + + A AKD+ VGD GWTT DY W K F +GDTL F Y +VV
Sbjct: 9 LVAVLLAAAAAPASAKDYTVGDSSGWTTGVDYTAWARGKTFNIGDTLLFQYTSAGHSVVE 68
Query: 69 VNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV---QPKQ 125
V+ +D SCS G + +T G R++I T HC G KL +TV
Sbjct: 69 VSEADHTSCSAANPLRSYKDGTTIVTLTRSGTRYFICGSTGHCGAGMKLTVTVATLSGSA 128
Query: 126 DGWSPVPSPSPSPSLDLVTPEAPPSNA 152
G + + PS S + T S+A
Sbjct: 129 AGGTRLAKPSSSDADPTTTTTTRTSSA 155
>UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis thaliana]
Length = 222
Score = 87.8 bits (216), Expect = 1e-16
Identities = 61/173 (35%), Positives = 81/173 (46%), Gaps = 16/173 (9%)
Query: 3 LSRALFLF-ALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 61
++ AL LF A + ++F A F VGD GWT+ DY W K FR+GDTL F Y G
Sbjct: 5 VAAALLLFLAAVPAVF-----AATFKVGDISGWTSNLDYTVWLTGKTFRVGDTLEFVY-G 58
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
+V V+ + + +C G KI +TT G ++ HC+NG KL + V
Sbjct: 59 LSHSVSVVDKAGYDNCDSSGATQNFADGDTKIDLTTVGTMHFLCPTFGHCKNGMKLAVPV 118
Query: 122 QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPAS-----SVPRRSLLPKKL 169
+P PS SP TP +PPS P+S S P SL P L
Sbjct: 119 LAA----APSPSTPSSPPSTPSTPSSPPSTPSTPSSPPSPPSPPSPSLPPSSL 167
>UniRef100_Q6H8G6 Putative uclacyanin 3 [Oryza sativa]
Length = 202
Score = 87.4 bits (215), Expect = 1e-16
Identities = 54/165 (32%), Positives = 80/165 (47%), Gaps = 6/165 (3%)
Query: 3 LSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGG 62
L+ A + L+A++ AV D+ VGD GW++ DY TW +K F +GD+L F Y
Sbjct: 7 LAAAGLVVLLLAAVAPAFAV--DYTVGDTSGWSSGVDYDTWAKSKTFSVGDSLVFQY-SM 63
Query: 63 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 122
V V+ +D+ +CS + + KI +T G R++I + HC G KL +TV
Sbjct: 64 MHTVAEVSSADYSACSASNSIQSYSDQNTKIALTKPGTRYFICGTSGHCSGGMKLAVTVS 123
Query: 123 PKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPA-SSVPRRSLLP 166
+P P+ S SP TP P S+ SS P + P
Sbjct: 124 AAA-ATTPTPTASSSPP-STATPATPSSDPGMDTPSSTPDATTTP 166
>UniRef100_Q6ZJW2 Putative blue copper binding protein [Oryza sativa]
Length = 188
Score = 87.0 bits (214), Expect = 2e-16
Identities = 57/161 (35%), Positives = 78/161 (48%), Gaps = 7/161 (4%)
Query: 4 SRALFLFALIASIFSTMAVAKDFVVGDEKG-WTTLFDYQTWTANKVFRLGDTLTFNYVGG 62
+RAL + A+ A++ T + VG G W T +Y W + FR+GD L F Y
Sbjct: 3 ARALLVVAMAAAVLGTAMGVTTYTVGAPAGSWDTRTNYAQWVSAITFRVGDQLVFKYSPA 62
Query: 63 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 122
+VV VN +D+ SCS SG D I + G R++I HC G K+ + V+
Sbjct: 63 AHDVVEVNKADYDSCSSSSPISTFNSGDDTIPLAAIGTRYFICGFPGHCTAGMKVAVKVE 122
Query: 123 PKQDGWSPVPSP-SPSPSLDLV-TPEA-PPSNA--PWPASS 158
G +P PSP +P P V P A PP+N P P SS
Sbjct: 123 -AATGSNPTPSPLAPLPRTPTVMAPNAMPPTNGGRPTPPSS 162
>UniRef100_Q6ZJW3 Putative blue copper binding protein [Oryza sativa]
Length = 187
Score = 85.5 bits (210), Expect = 5e-16
Identities = 54/156 (34%), Positives = 76/156 (48%), Gaps = 8/156 (5%)
Query: 4 SRALFLFALIASIFSTMAVAKDFVVGDEKG-WTTLFDYQTWTANKVFRLGDTLTFNYVGG 62
+RAL + A+ A++ T A+ + VG G W +Y W +N FR GD + F Y
Sbjct: 3 ARALLVVAMAAAVLGT-ALGATYTVGAPSGSWDLRTNYDQWVSNINFRAGDQIVFKYSPA 61
Query: 63 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 122
+VV VN +D+ SCS SG D I +T G R++I HC G K+ + V+
Sbjct: 62 AHDVVEVNKADYDSCSSSSPIATFNSGDDTIPLTATGTRYFICGFNGHCTGGMKVAVKVE 121
Query: 123 PKQDGWSPVPSP-SPSPSLDLVTPEAPPSNAPWPAS 157
G +P PSP +P P TP A NA P +
Sbjct: 122 -AATGSNPAPSPMTPRPR----TPTAMAPNAMPPTA 152
>UniRef100_O48787 Putative phytocyanin [Arabidopsis thaliana]
Length = 206
Score = 85.1 bits (209), Expect = 7e-16
Identities = 54/144 (37%), Positives = 71/144 (48%), Gaps = 14/144 (9%)
Query: 28 VGDEKGWTTLF-DYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVL 86
VG+ KGWT + DY+ W +++VF++GDTL F Y +V V +DF+ C
Sbjct: 31 VGNTKGWTMIGGDYEAWASSRVFQVGDTLVFAYNKDYHDVTEVTHNDFEMCESSKPLRRY 90
Query: 87 TSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVP------------SP 134
+G D I +T G + +I V HC+ GQKL I V P G VP SP
Sbjct: 91 KTGSDSISLTKPGLQHFICGVPGHCKKGQKLQIHVLPASLGHVAVPVPGPVRSQSSSSSP 150
Query: 135 SPSPSLDLVTPEAPP-SNAPWPAS 157
SPSP +D AP P PAS
Sbjct: 151 SPSPLVDPPVNNAPQYQMGPTPAS 174
>UniRef100_O81500 F9D12.16 protein [Arabidopsis thaliana]
Length = 187
Score = 84.7 bits (208), Expect = 9e-16
Identities = 54/165 (32%), Positives = 79/165 (47%), Gaps = 13/165 (7%)
Query: 6 ALFLFALIASIFSTMAVAKDFV--VGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYVG 61
A + A +A I + +++ V VGD GWTT+ DY+ W + K F +GDT+ F Y
Sbjct: 2 AAIIVAALACIVVMLRLSEAAVYKVGDSAGWTTIANVDYKLWASTKTFHIGDTVLFEYNP 61
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
NV+RV ++SC+ T+G D I +T +G ++ V HC GQKL + V
Sbjct: 62 QFHNVMRVTHPMYRSCNTSKPISTFTTGNDSITLTNHGHHFFFCGVPGHCLAGQKLDLHV 121
Query: 122 ------QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVP 160
P D P S S SP + P +P A+S+P
Sbjct: 122 LLPASSTPLSD---PPTSSSSSPPSTTIPAAGVPGPSPSLAASLP 163
>UniRef100_Q949E8 Uclacyanin 3-like protein [Oryza sativa]
Length = 202
Score = 84.3 bits (207), Expect = 1e-15
Identities = 53/165 (32%), Positives = 79/165 (47%), Gaps = 6/165 (3%)
Query: 3 LSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGG 62
L+ A + L+A++ AV D+ VGD GW++ DY TW +K F +GD+L F Y
Sbjct: 7 LAAAGLVVLLLAAVAPAFAV--DYTVGDTSGWSSGVDYVTWAKSKTFSVGDSLVFQY-SM 63
Query: 63 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 122
V V+ +D+ +CS + + KI +T G R++I + HC G KL + V
Sbjct: 64 MHTVAEVSSADYSACSASNSIQSYSDQNTKIALTKPGTRYFICGTSGHCSGGMKLAVMVS 123
Query: 123 PKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPA-SSVPRRSLLP 166
+P P+ S SP TP P S+ SS P + P
Sbjct: 124 AAA-ATTPTPTASSSPP-STATPATPSSDPGMDTPSSTPDATTTP 166
>UniRef100_Q9FXS4 NtEIG-A1 protein [Nicotiana tabacum]
Length = 184
Score = 83.6 bits (205), Expect = 2e-15
Identities = 53/162 (32%), Positives = 77/162 (46%), Gaps = 10/162 (6%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFD----YQTWTANKVFRLGDTLT 56
M + LF +AS+ VVGD +GWT Y W A K F +GDTL
Sbjct: 3 MIMRMVLFGALALASLVQLTTAQTAHVVGDNEGWTIPSSGASAYTNWAAGKTFMVGDTLV 62
Query: 57 FNYVGGKDNVVRVNGSDFKSCSVP-LTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQ 115
FN++ +V++V + F CS + SG + + + G R+YI + HC+NGQ
Sbjct: 63 FNFMTNTHDVLQVPKASFDGCSSQNAIGSAIVSGPANVTLDSAGERYYICTFGRHCQNGQ 122
Query: 116 KLFITVQPK--QDGWSPVPSPSPSPSLDL---VTPEAPPSNA 152
KL ITV G +P S + PS + + P PPS++
Sbjct: 123 KLAITVSSSTGTPGANPPTSFAAGPSGSVPGGIAPPPPPSSS 164
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 314,645,224
Number of Sequences: 2790947
Number of extensions: 14864066
Number of successful extensions: 85004
Number of sequences better than 10.0: 376
Number of HSP's better than 10.0 without gapping: 199
Number of HSP's successfully gapped in prelim test: 183
Number of HSP's that attempted gapping in prelim test: 83116
Number of HSP's gapped (non-prelim): 1264
length of query: 176
length of database: 848,049,833
effective HSP length: 119
effective length of query: 57
effective length of database: 515,927,140
effective search space: 29407846980
effective search space used: 29407846980
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC126009.11


