

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
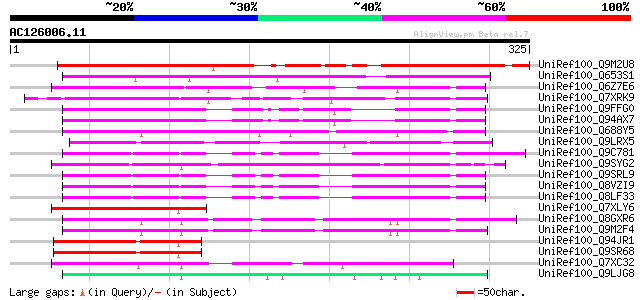
Query= AC126006.11 + phase: 0
(325 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M2U8 Hypothetical protein T12E18_80 [Arabidopsis tha... 303 3e-81
UniRef100_Q653S1 Hypothetical protein OJ1065_E04.5 [Oryza sativa] 160 3e-38
UniRef100_Q6Z7E6 Putative 6b-interacting protein 1 [Oryza sativa] 105 1e-21
UniRef100_Q7XRK9 OSJNBa0027P08.13 protein [Oryza sativa] 105 1e-21
UniRef100_Q9FFG0 Gb|AAF01512.1 [Arabidopsis thaliana] 101 2e-20
UniRef100_Q94AX7 AT5g05550/MOP10_9 [Arabidopsis thaliana] 101 2e-20
UniRef100_Q688Y5 Putative 6b-interacting protein 1 [Oryza sativa] 100 4e-20
UniRef100_Q9LRX5 Gb|AAC27475.1 [Arabidopsis thaliana] 96 1e-18
UniRef100_Q9C781 Hypothetical protein F11B9.6 [Arabidopsis thali... 91 6e-17
UniRef100_Q9SYG2 F15I1.14 protein [Arabidopsis thaliana] 90 7e-17
UniRef100_Q9SRL9 F9F8.9 protein [Arabidopsis thaliana] 90 9e-17
UniRef100_Q8VZI9 AT3g11100/F11B9_105 [Arabidopsis thaliana] 90 9e-17
UniRef100_Q8LF33 Hypothetical protein [Arabidopsis thaliana] 89 1e-16
UniRef100_Q7XLY6 OSJNBa0042I15.4 protein [Oryza sativa] 88 4e-16
UniRef100_Q8GXR6 Hypothetical protein At3g58630/F14P22_220 [Arab... 84 4e-15
UniRef100_Q9M2F4 Hypothetical protein F14P22.220 [Arabidopsis th... 84 5e-15
UniRef100_Q94JR1 AT3g10030/T22K18_15 [Arabidopsis thaliana] 84 7e-15
UniRef100_Q9SR68 Putative uridylate kinase [Arabidopsis thaliana] 84 7e-15
UniRef100_Q7XC32 Hypothetical protein [Oryza sativa] 82 3e-14
UniRef100_Q9LJG8 Similarity to DNA-binding protein [Arabidopsis ... 80 6e-14
>UniRef100_Q9M2U8 Hypothetical protein T12E18_80 [Arabidopsis thaliana]
Length = 296
Score = 303 bits (777), Expect = 3e-81
Identities = 178/298 (59%), Positives = 211/298 (70%), Gaps = 35/298 (11%)
Query: 31 ERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQC 90
+RLKRDEWSEGAV+TLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSR+ TKSPKTQTQC
Sbjct: 31 DRLKRDEWSEGAVSTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRATHTKSPKTQTQC 90
Query: 91 KNKIESMKKRYRSESASSDVSSWPLYSRLDLLLRGT---GQLSTTTLNNNQAGLVLLEPS 147
KNKIESMKKRYRSESA++D SSWPLY RLD LLRGT Q N L+LLEP
Sbjct: 91 KNKIESMKKRYRSESATADGSSWPLYPRLDHLLRGTQPQPQPQAVLPLNCSVPLLLLEPP 150
Query: 148 SLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSS 207
++H PP I S+GSNGV K+ KED + ++ ++TDSS
Sbjct: 151 LPAVAH----------PPQI-----SYGSNGVGKIPKEDGFKPENKPE--KDAEMDTDSS 193
Query: 208 TPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEI 267
TP + K + R KK + +R K+E IA SIRW+AEVV+RSE+ RMET+KEI
Sbjct: 194 TPVV---KTKVRGKKVK--------RRYKEEKEEIAGSIRWLAEVVMRSERARMETMKEI 242
Query: 268 EKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFANVNNKDKDKDKDVDVDSSLRIGRS 325
E+MR EAEAKRGE+++KRTEI+ANTQLEIARIFA + ++K VDSSLRIGR+
Sbjct: 243 ERMRAEAEAKRGELDLKRTEIMANTQLEIARIFAAAASSGQNK----GVDSSLRIGRN 296
>UniRef100_Q653S1 Hypothetical protein OJ1065_E04.5 [Oryza sativa]
Length = 336
Score = 160 bits (406), Expect = 3e-38
Identities = 108/295 (36%), Positives = 156/295 (52%), Gaps = 39/295 (13%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSR----------SNSTKS 83
KRDEWSE + LLEAYE+KW+LRNRAKLK DW D+A VS+ + +
Sbjct: 22 KRDEWSESGIVRLLEAYEAKWLLRNRAKLKWSDWVDIAHEVSAHCAMENAAATGKPGSST 81
Query: 84 PKTQTQCKNKIESMKKRYRSESASSDVS---------SWPLYSRLDLLLRG-TGQLSTTT 133
KT QCKNKIESMKKRYR+ESA++ + SW ++R+D LL+G G
Sbjct: 82 AKTPNQCKNKIESMKKRYRAESAAAARAGPAAAGAGPSWRFFARMDGLLKGPAGSGQPQA 141
Query: 134 LNNNQAGLVLLEPSSLVMSHAQEEVHQPLVPPP------ITTAQNSHGSNGVDKLIKEDE 187
+N L P+ + + + V Q P ++ N +DK+
Sbjct: 142 ELSNSIDLRAPPPAKVEVDVDADFVSQLADAGPGALSELVSAYANGSIQEKLDKVENSGH 201
Query: 188 IETKSSDHIAN-KNPLETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESI 246
+E ++++ N +P +++ A +K SKKR K IA+SI
Sbjct: 202 VEGRAAESDVNVSSPRIKEANEDAEEVDKVWDMSKKR------------KNTEFDIAKSI 249
Query: 247 RWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFA 301
+A ++ E+ RM+ +E E+MRVEAE K+GEME+KRTEI+A T L+IA++FA
Sbjct: 250 ELLASSFLKIERARMDLYRETERMRVEAEIKKGEMELKRTEIMAKTHLQIAKLFA 304
>UniRef100_Q6Z7E6 Putative 6b-interacting protein 1 [Oryza sativa]
Length = 419
Score = 105 bits (263), Expect = 1e-21
Identities = 83/291 (28%), Positives = 130/291 (44%), Gaps = 44/291 (15%)
Query: 27 GSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKT 86
G G + D WS+GA +TL++A+ ++V R L+ W++VA+ VSSR +K PK+
Sbjct: 53 GGGGGGGREDAWSDGATSTLIDAWGERFVALGRGSLRHPQWQEVAEVVSSRDGYSKQPKS 112
Query: 87 QTQCKNKIESMKKRYRSESASSDVSSWPLYSRLDLLL-------------RGTGQLSTTT 133
QCKN+I+++KK+Y+ E A D SSWP + RLD LL G
Sbjct: 113 DVQCKNRIDTLKKKYKVEKAKPD-SSWPYFHRLDTLLAPVHKPAGAYPAAAAAGAAGAGN 171
Query: 134 LNNNQAGLVLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLI---KEDEIET 190
+N A S+ M+ P V P T S+GV + + + +
Sbjct: 172 SGSNSAAAATAARSTAPMA--------PRVNFPQRTRTQFLPSSGVKRRMPSPPQVSASS 223
Query: 191 KSSDHIANKNPLETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKETMG---IAESIR 247
+SSD + P+ K+RR E N T G +A++IR
Sbjct: 224 ESSDGFPPEPPMAA-------------ANGKRRREVEEEVNGADSGHRTQGLRELAQAIR 270
Query: 248 WVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIAR 298
EV R E + E +E+ R+EA E+E +R + Q+E+++
Sbjct: 271 RFGEVYERVELAKREQELRMERDRLEAAR---ELEDQRVQFFLKMQMELSK 318
>UniRef100_Q7XRK9 OSJNBa0027P08.13 protein [Oryza sativa]
Length = 417
Score = 105 bits (263), Expect = 1e-21
Identities = 87/305 (28%), Positives = 137/305 (44%), Gaps = 46/305 (15%)
Query: 10 QIQQNQESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWED 69
Q QQ+ SP N G G R D WSEGA L++A+ ++V R L+ W++
Sbjct: 47 QKQQHAASP-----NPGGGGGGR--EDAWSEGATAALIDAWGERFVALGRGSLRHPQWQE 99
Query: 70 VAKHVSSRSNSTKSPKTQTQCKNKIESMKKRYRSESASSDVSSWPLYSRLDLLL------ 123
VA VSSR K+PK+ QCKN+I+++KK+Y+ E A SSW + RLD LL
Sbjct: 100 VADAVSSREGYAKAPKSDVQCKNRIDTLKKKYKIERA-KPASSWQFFGRLDDLLAPTFNQ 158
Query: 124 ----RGTGQLSTTTLNNNQAGLVLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGV 179
G G + + N P++L + Q PL+P P++ + S
Sbjct: 159 KPGGNGGGGVGASVNGRNPV------PAALRVGFPQRS-RTPLMPAPVSAVKRRAPS--- 208
Query: 180 DKLIKEDEIETKSSDHIANKN-----PLETDSSTPALYSEKDEQRSKKRRMRTENNNNKR 234
E ++SSD + PL S DE R + R
Sbjct: 209 ----PEPSASSESSDGFPPERQPAFPPLPLPPPPNGKRSRADEGRGGGAGGGGDRAQGLR 264
Query: 235 QKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQL 294
+ +A++IR E R E ++E E+E+ R++ + E+E +R + NTQ+
Sbjct: 265 E------LAQAIRRFGEAYERVETAKLEQSAEMERRRLDFAS---ELESQRVQFFLNTQM 315
Query: 295 EIARI 299
E++++
Sbjct: 316 ELSQV 320
>UniRef100_Q9FFG0 Gb|AAF01512.1 [Arabidopsis thaliana]
Length = 248
Score = 101 bits (252), Expect = 2e-20
Identities = 80/267 (29%), Positives = 124/267 (45%), Gaps = 56/267 (20%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ +++V N L+ DW+DVA V+SR KT QCKN+
Sbjct: 20 REDWWSEEATATLVEAWGNRYVKLNHGNLRQNDWKDVADAVNSRHGDNSRKKTDLQCKNR 79
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
++++KK+Y++E A S+W Y+RLD+L+ G V+ + + V+
Sbjct: 80 VDTLKKKYKTEKAKLSPSTWRFYNRLDVLI----------------GPVVKKSAGGVVKS 123
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPL--ETDSSTPAL 211
A + H L P T NS GS+ D ED+ E + +A K+P E D S +
Sbjct: 124 APFKNH--LNP----TGSNSTGSSLEDD--DEDDDEVGDWEFVARKHPRVEEVDLSEGST 175
Query: 212 YSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMR 271
E +A +I EV R E + + + E+EK R
Sbjct: 176 CRE---------------------------LATAILKFGEVYERIEGKKQQMMIELEKQR 208
Query: 272 VEAEAKRGEMEIKRTEIIANTQLEIAR 298
+E E+E+KR ++ QLEI +
Sbjct: 209 MEVTK---EVELKRMNMLMEMQLEIEK 232
>UniRef100_Q94AX7 AT5g05550/MOP10_9 [Arabidopsis thaliana]
Length = 246
Score = 101 bits (252), Expect = 2e-20
Identities = 80/267 (29%), Positives = 124/267 (45%), Gaps = 56/267 (20%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ +++V N L+ DW+DVA V+SR KT QCKN+
Sbjct: 20 REDWWSEEATATLVEAWGNRYVKLNHGNLRQNDWKDVADAVNSRHGDNSRKKTDLQCKNR 79
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
++++KK+Y++E A S+W Y+RLD+L+ G V+ + + V+
Sbjct: 80 VDTLKKKYKTEKAKLSPSTWRFYNRLDVLI----------------GPVVKKSAGGVVKS 123
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPL--ETDSSTPAL 211
A + H L P T NS GS+ D ED+ E + +A K+P E D S +
Sbjct: 124 APFKNH--LNP----TGSNSTGSSLEDD--DEDDDEVGDWEFVARKHPRVEEVDLSEGST 175
Query: 212 YSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMR 271
E +A +I EV R E + + + E+EK R
Sbjct: 176 CRE---------------------------LATAILKFGEVYERIEGKKQQMMIELEKQR 208
Query: 272 VEAEAKRGEMEIKRTEIIANTQLEIAR 298
+E E+E+KR ++ QLEI +
Sbjct: 209 MEVTK---EVELKRMNMLMEMQLEIEK 232
>UniRef100_Q688Y5 Putative 6b-interacting protein 1 [Oryza sativa]
Length = 346
Score = 100 bits (250), Expect = 4e-20
Identities = 75/281 (26%), Positives = 135/281 (47%), Gaps = 23/281 (8%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNST---KSPKTQTQC 90
+ D WSEG L+ A+ S++V NR L+ + W++VA V+SR + + P+T QC
Sbjct: 37 REDCWSEGETEALVRAWGSRYVELNRGNLRQKQWQEVADAVNSRRGAAARRRPPRTDVQC 96
Query: 91 KNKIESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLV 150
KN+++++KK+Y++E A S+W + LD L+ T S + + V P +
Sbjct: 97 KNRVDTLKKKYKAERARVMPSTWSFFPELDRLVGPTLSASASKRPSPSPSPVPPPPYFAM 156
Query: 151 MSHAQ------EEVHQPLVPPPITTAQNSH--GSNGVDKLIKEDEIETKSSDHIANKNPL 202
H P PPP+ S+ GS + + E ++ +++
Sbjct: 157 PIHPSAVRKPPSPSPSPSPPPPMALPLPSYRRGSPLPAAALIQQEAAAAAAAAVSDSE-- 214
Query: 203 ETDSSTPALYSEKDEQRSKKRRMRTEN-NNNKRQKKETMG----IAESIRWVAEVVVRSE 257
DS P + + QRS + + + + N+NKR ++E G +A +I AE+ R E
Sbjct: 215 --DSEGPGDNNNHNAQRSPSQSVSSRSGNSNKRSRQEVDGGFRELARAIEAFAEMYERVE 272
Query: 258 QTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIAR 298
+ + EIE+ R++ ++E+KR E + +++AR
Sbjct: 273 SAKQKQALEIERQRIDF---LKQLEVKRMENFVDAHVKLAR 310
>UniRef100_Q9LRX5 Gb|AAC27475.1 [Arabidopsis thaliana]
Length = 310
Score = 96.3 bits (238), Expect = 1e-18
Identities = 78/270 (28%), Positives = 129/270 (46%), Gaps = 34/270 (12%)
Query: 38 WSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNKIESM 97
W++ L+E+Y+ KW R LK WE++A SSRS +T TQC++KIE M
Sbjct: 65 WTQDETLLLIESYKEKWFAIGRGPLKSTHWEEIAVAASSRSG---VERTSTQCRHKIEKM 121
Query: 98 KKRYRSESAS-SDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSHAQE 156
+KR+RSE S +S WP Y++++ L S+ + L L P+S E
Sbjct: 122 RKRFRSERQSMGPISIWPFYNQMEEL------DSSNPAPISARPLTRLPPNSNNRYVDDE 175
Query: 157 EVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKS-SDHIANKNPLETDSST---PALY 212
E +N ++ +EDE ++KS S + + P + L
Sbjct: 176 E------------EDEEEDNNNYEEEEEEDERQSKSRSINYILRRPGTVNRFAGVGGGLL 223
Query: 213 SEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRV 272
S ++RS KR+ R + + +R++K +A IR AE V+ E+ ++E KE ++
Sbjct: 224 SWGQKERSSKRK-RNDGDGGERRRKGMRAVAAEIRAFAERVMVMEKKKIEFAKETVRL-- 280
Query: 273 EAEAKRGEMEIKRTEIIANTQLEIARIFAN 302
R EMEI+R +I ++Q ++ + N
Sbjct: 281 -----RKEMEIRRINLIQSSQTQLLQFINN 305
>UniRef100_Q9C781 Hypothetical protein F11B9.6 [Arabidopsis thaliana]
Length = 348
Score = 90.5 bits (223), Expect = 6e-17
Identities = 77/290 (26%), Positives = 126/290 (42%), Gaps = 47/290 (16%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ ++V NR L+ DW++VA V+S S+ PKT QCKN+
Sbjct: 18 REDWWSEDATATLIEAWGDRYVNLNRGNLRQNDWKEVADAVNS-SHGNGRPKTDVQCKNR 76
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
I+++KK+Y++E A +S+W + RLD L+ G V+ + S V+
Sbjct: 77 IDTLKKKYKTEKA-KPLSNWCFFDRLDFLI----------------GPVMKKSSGAVVKS 119
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS 213
A + P + P T S GS+ D +D+ E D
Sbjct: 120 A---LMNPNLNP---TGSKSTGSSLDDDDDDDDDDEEDDDD------------------- 154
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVE 273
+ R+ R + + + +A SI + E R E + + + E+EK R+E
Sbjct: 155 -AGDWGFVVRKHRKVEDVDSSEGSAFRELARSILKLGEAFERIEGKKQQMMIELEKQRME 213
Query: 274 AEAKRGEMEIKRTEIIANTQLEIARIFANVNNKDKDKDKDVDVDSSLRIG 323
E+E++R ++ QLE+ + D DV +G
Sbjct: 214 VAK---ELELQRMNMLMEMQLELEKSKLGKRRAASDCGGVTDVKQGAMVG 260
>UniRef100_Q9SYG2 F15I1.14 protein [Arabidopsis thaliana]
Length = 383
Score = 90.1 bits (222), Expect = 7e-17
Identities = 80/291 (27%), Positives = 140/291 (47%), Gaps = 23/291 (7%)
Query: 27 GSGNERLKRDE-WSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPK 85
GSG RD+ WSE A L+EA+ ++ + LK Q W++VA+ V+ +S K PK
Sbjct: 82 GSGGGGGGRDDCWSEEATKVLIEAWGDRFSEPGKGTLKQQHWKEVAEIVN-KSRQCKYPK 140
Query: 86 TQTQCKNKIESMKKRYRSESA----SSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGL 141
T QCKN+I+++KK+Y+ E A S W + +L+ L+ GT TT + +++A
Sbjct: 141 TDIQCKNRIDTVKKKYKQEKAKIASGDGPSKWVFFKKLESLIGGT----TTFIASSKASE 196
Query: 142 VLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVD-KLIKEDEIETKS-SDHIANK 199
+L S + Q + Q GS+ + K ET+S SD
Sbjct: 197 KAPMGGALGNSRSSMFKRQTKGNQIVQQQQEKRGSDSMRWHFRKRSASETESESDPEPEA 256
Query: 200 NPLETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQT 259
+P E+ S P L + +R++ + + +A +I E ++E
Sbjct: 257 SPEESAESLPPLQPIQPLSFHMPKRLKVDKSGG--GGSGVGDVARAILGFTEAYEKAETA 314
Query: 260 RMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFANVNNKDKDK 310
+++ + E+EK R++ EME++R + + TQLEI + NN+++++
Sbjct: 315 KLKLMAELEKERMKFAK---EMELQRMQFL-KTQLEITQ-----NNQEEEE 356
>UniRef100_Q9SRL9 F9F8.9 protein [Arabidopsis thaliana]
Length = 256
Score = 89.7 bits (221), Expect = 9e-17
Identities = 73/265 (27%), Positives = 121/265 (45%), Gaps = 47/265 (17%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ ++V NR L+ DW++VA V+S S+ PKT QCKN+
Sbjct: 18 REDWWSEDATATLIEAWGDRYVNLNRGNLRQNDWKEVADAVNS-SHGNGRPKTDVQCKNR 76
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
I+++KK+Y++E A +S+W + RLD L+ G V+ + S V+
Sbjct: 77 IDTLKKKYKTEKA-KPLSNWCFFDRLDFLI----------------GPVMKKSSGAVVKS 119
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS 213
A + P + P T S GS+ D +D+ E D
Sbjct: 120 A---LMNPNLNP---TGSKSTGSSLDDDDDDDDDDEEDDDD------------------- 154
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVE 273
+ R+ R + + + +A SI + E R E + + + E+EK R+E
Sbjct: 155 -AGDWGFVVRKHRKVEDVDSSEGSAFRELARSILKLGEAFERIEGKKQQMMIELEKQRME 213
Query: 274 AEAKRGEMEIKRTEIIANTQLEIAR 298
E+E++R ++ QLE+ +
Sbjct: 214 VAK---ELELQRMNMLMEMQLELEK 235
>UniRef100_Q8VZI9 AT3g11100/F11B9_105 [Arabidopsis thaliana]
Length = 249
Score = 89.7 bits (221), Expect = 9e-17
Identities = 73/265 (27%), Positives = 121/265 (45%), Gaps = 47/265 (17%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ ++V NR L+ DW++VA V+S S+ PKT QCKN+
Sbjct: 18 REDWWSEDATATLIEAWGDRYVNLNRGNLRQNDWKEVADAVNS-SHGNGRPKTDVQCKNR 76
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
I+++KK+Y++E A +S+W + RLD L+ G V+ + S V+
Sbjct: 77 IDTLKKKYKTEKA-KPLSNWCFFDRLDFLI----------------GPVMKKSSGAVVKS 119
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS 213
A + P + P T S GS+ D +D+ E D
Sbjct: 120 A---LMNPNLNP---TGSKSTGSSLDDDDDDDDDDEEDDDD------------------- 154
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVE 273
+ R+ R + + + +A SI + E R E + + + E+EK R+E
Sbjct: 155 -AGDWGFVVRKHRKVEDVDSSEGSAFRELARSILKLGEAFERIEGKKQQMMIELEKQRME 213
Query: 274 AEAKRGEMEIKRTEIIANTQLEIAR 298
E+E++R ++ QLE+ +
Sbjct: 214 VAK---ELELQRMNMLMEMQLELEK 235
>UniRef100_Q8LF33 Hypothetical protein [Arabidopsis thaliana]
Length = 249
Score = 89.4 bits (220), Expect = 1e-16
Identities = 73/265 (27%), Positives = 121/265 (45%), Gaps = 47/265 (17%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A TL+EA+ ++V NR L+ DW++VA V+S S+ PKT QCKN+
Sbjct: 18 REDWWSEDATATLIEAWGDRYVNLNRGNLRQNDWKEVADAVNS-SHGNGRPKTDVQCKNR 76
Query: 94 IESMKKRYRSESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLVMSH 153
I+++KK+Y++E A +S+W + RLD L+ G V+ + S V+
Sbjct: 77 IDTLKKKYKTEKA-KPLSNWCFFDRLDFLI----------------GPVMKKSSGAVVKS 119
Query: 154 AQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS 213
A + P + P T S GS+ D +D+ E D
Sbjct: 120 A---LMNPNLNP---TGYKSTGSSLDDDDDDDDDDEEDDDD------------------- 154
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVE 273
+ R+ R + + + +A SI + E R E + + + E+EK R+E
Sbjct: 155 -AGDWGFVVRKHRKVEDVDSSEGSAFRELARSILKLGEAFERIEGKKQQIMIELEKQRME 213
Query: 274 AEAKRGEMEIKRTEIIANTQLEIAR 298
E+E++R ++ QLE+ +
Sbjct: 214 VAK---ELELQRMNMLMEMQLELEK 235
>UniRef100_Q7XLY6 OSJNBa0042I15.4 protein [Oryza sativa]
Length = 530
Score = 87.8 bits (216), Expect = 4e-16
Identities = 40/103 (38%), Positives = 66/103 (63%), Gaps = 6/103 (5%)
Query: 27 GSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKT 86
GS R R+EW++GA+++LL++Y ++ NR L+G+DWEDVA V+ + K+
Sbjct: 132 GSSEYRKDREEWTDGAISSLLDSYTDRFEQLNRGNLRGRDWEDVAAAVTDGQGKSSGGKS 191
Query: 87 QTQCKNKIESMKKRYRSE------SASSDVSSWPLYSRLDLLL 123
QCKNKI+++KKRY+ E S +S VS WP + +++ ++
Sbjct: 192 VEQCKNKIDNLKKRYKVECQRLAGSGASAVSHWPWFKKMEQIV 234
>UniRef100_Q8GXR6 Hypothetical protein At3g58630/F14P22_220 [Arabidopsis thaliana]
Length = 321
Score = 84.3 bits (207), Expect = 4e-15
Identities = 74/306 (24%), Positives = 140/306 (45%), Gaps = 38/306 (12%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNST---------KSP 84
+ D WSE A TL++A+ +++V +R L+ + W++VA V+ R +T +
Sbjct: 22 REDCWSEEATFTLIQAWGNRYVDLSRGNLRQKHWQEVANAVNDRHYNTGRNVSAAKSQPY 81
Query: 85 KTQTQCKNKIESMKKRYRSESA-------SSDVSSWPLYSRLDLLLRGTGQLSTTTLNNN 137
+T QCKN+I+++KK+Y+ E A + +S WP +S LD LLR + T+ N +
Sbjct: 82 RTDVQCKNRIDTLKKKYKVEKARVSESNPGAYISPWPFFSALDDLLR---ESFPTSSNPD 138
Query: 138 QAGLVLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIA 197
+ + SL MS ++ V P + S + +D + +
Sbjct: 139 STDNIPHQRLSLPMS-----INPVPVAPRSAIPRRPATSPAIIPHAGDDLLGFR-----G 188
Query: 198 NKNPLETDSSTPAL-YSEKDEQRSKKRRMRTENNNNKRQKK--ETMG---IAESIRWVAE 251
N N ++ A SE D + S+ R +N KR++K + G +A++I + +
Sbjct: 189 NLNAFAAAAAAAACPASEDDSEGSRSRSSGRSGSNKKRERKIEKKQGYKEVADAIERLGQ 248
Query: 252 VVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFANVNNKDKDKD 311
+ R E+ + + + E+EK R+ E+E R ++ Q+ + ++ +K
Sbjct: 249 IYERVEEKKRKEMVELEKQRMRFAK---ELECHRMQLFTEMQVRLHKLRRTSGSKGPTSS 305
Query: 312 KDVDVD 317
+D
Sbjct: 306 ASAALD 311
>UniRef100_Q9M2F4 Hypothetical protein F14P22.220 [Arabidopsis thaliana]
Length = 311
Score = 84.0 bits (206), Expect = 5e-15
Identities = 72/288 (25%), Positives = 136/288 (47%), Gaps = 38/288 (13%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNST---------KSP 84
+ D WSE A TL++A+ +++V +R L+ + W++VA V+ R +T +
Sbjct: 22 REDCWSEEATFTLIQAWGNRYVDLSRGNLRQKHWQEVANAVNDRHYNTGRNVSAAKSQPY 81
Query: 85 KTQTQCKNKIESMKKRYRSESA-------SSDVSSWPLYSRLDLLLRGTGQLSTTTLNNN 137
+T QCKN+I+++KK+Y+ E A + +S WP +S LD LLR + T+ N +
Sbjct: 82 RTDVQCKNRIDTLKKKYKVEKARVSESNPGAYISPWPFFSALDDLLR---ESFPTSSNPD 138
Query: 138 QAGLVLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIA 197
+ + SL MS ++ V P + S + +D + +
Sbjct: 139 STDNIPHQRLSLPMS-----INPVPVAPRSAIPRRPATSPAIIPHAGDDLLGFR-----G 188
Query: 198 NKNPLETDSSTPAL-YSEKDEQRSKKRRMRTENNNNKRQKK--ETMG---IAESIRWVAE 251
N N ++ A SE D + S+ R +N KR++K + G +A++I + +
Sbjct: 189 NLNAFAAAAAAAACPASEDDSEGSRSRSSGRSGSNKKRERKIEKKQGYKEVADAIERLGQ 248
Query: 252 VVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARI 299
+ R E+ + + + E+EK R+ E+E R ++ Q+ + ++
Sbjct: 249 IYERVEEKKRKEMVELEKQRMRFAK---ELECHRMQLFTEMQVRLHKL 293
>UniRef100_Q94JR1 AT3g10030/T22K18_15 [Arabidopsis thaliana]
Length = 542
Score = 83.6 bits (205), Expect = 7e-15
Identities = 41/98 (41%), Positives = 60/98 (60%), Gaps = 7/98 (7%)
Query: 28 SGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQ 87
+G R R+EWS+ A+ LL+AY K+ NR L+G+DWE+VA VS R K K+
Sbjct: 151 AGEYRKDREEWSDAAIACLLDAYSDKFTQLNRGNLRGRDWEEVASSVSERCE--KLSKSV 208
Query: 88 TQCKNKIESMKKRYRSE-----SASSDVSSWPLYSRLD 120
QCKNKI+++KKRY+ E S + S WP + +++
Sbjct: 209 EQCKNKIDNLKKRYKLERHRMSSGGTAASHWPWFKKME 246
>UniRef100_Q9SR68 Putative uridylate kinase [Arabidopsis thaliana]
Length = 520
Score = 83.6 bits (205), Expect = 7e-15
Identities = 41/98 (41%), Positives = 60/98 (60%), Gaps = 7/98 (7%)
Query: 28 SGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQ 87
+G R R+EWS+ A+ LL+AY K+ NR L+G+DWE+VA VS R K K+
Sbjct: 151 AGEYRKDREEWSDAAIACLLDAYSDKFTQLNRGNLRGRDWEEVASSVSERCE--KLSKSV 208
Query: 88 TQCKNKIESMKKRYRSE-----SASSDVSSWPLYSRLD 120
QCKNKI+++KKRY+ E S + S WP + +++
Sbjct: 209 EQCKNKIDNLKKRYKLERHRMSSGGTAASHWPWFKKME 246
>UniRef100_Q7XC32 Hypothetical protein [Oryza sativa]
Length = 336
Score = 81.6 bits (200), Expect = 3e-14
Identities = 68/269 (25%), Positives = 116/269 (42%), Gaps = 37/269 (13%)
Query: 27 GSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSN--STKSP 84
G RL W+ L+EAY +W + L+ DW+DVA V++R T +
Sbjct: 42 GGSGRRLPPPCWTHEETLALIEAYRDRWEGLRKGNLRASDWDDVAGAVTARCGRFPTATH 101
Query: 85 KTQTQCKNKIESMKKRYRSESA----SSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQAG 140
K+ QC++KIE ++KRYR+E A S WP + L L G G
Sbjct: 102 KSGVQCRHKIEKLRKRYRAERARAAGRSKGPKWPFFPLLHDLAGG--------------G 147
Query: 141 LVLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKN 200
P+ ++ ++ P P + + S S+ +EDE E ++D +++
Sbjct: 148 APDPSPNPIIKIKSKGPAAAAASPSPASPSPVSSPSS------EEDEEEEAAADAGRSRS 201
Query: 201 PLETDSS-----------TPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWV 249
S+ A S+ QR + ++ E + + + +A ++R V
Sbjct: 202 LHGLISNGGSGSGLRFTIPKASRSKPVAQREQPTAIKVEKSEEDAEAEAMAEVASALRAV 261
Query: 250 AEVVVRSEQTRMETLKEIEKMRVEAEAKR 278
+ +R E+ R+E +IEK R+E+E KR
Sbjct: 262 GDKFLRMEERRLEISLQIEKERMESEMKR 290
>UniRef100_Q9LJG8 Similarity to DNA-binding protein [Arabidopsis thaliana]
Length = 443
Score = 80.5 bits (197), Expect = 6e-14
Identities = 77/328 (23%), Positives = 128/328 (38%), Gaps = 62/328 (18%)
Query: 34 KRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNK 93
+ D WSE A L++A+ +++ +R LK + W++VA+ VSSR + K PKT QCKN+
Sbjct: 80 REDCWSEAATAVLIDAWGERYLELSRGNLKQKHWKEVAEIVSSREDYGKIPKTDIQCKNR 139
Query: 94 IESMKKRYRSESA----SSDVSSWPLYSRLDLLLRGTGQLSTTTLN-NNQAGLVLLEPSS 148
I+++KK+Y+ E S W + +LD L+ T ++ T T + G + P
Sbjct: 140 IDTVKKKYKQEKVRIANGGGRSRWVFFDKLDRLIGSTAKIPTATSGVSGPVGGLHKIPMG 199
Query: 149 LVMSHAQEEVHQ--PLVPPPITT-----------------AQNSHGSNGVDKLIKEDEIE 189
+ M HQ PP G G + +
Sbjct: 200 IPMGSRSNLYHQQAKAATPPFNNLDRLIGATARVSAASFGGSGGGGGGGSVNVPMGIPMS 259
Query: 190 TKSSDHIANKNPLETDSSTPALYSE-----KDEQRSKKRRMRTENNN------------- 231
++S+ L T + K SK+ R R N +
Sbjct: 260 SRSAPFGQQGRTLPQQGRTLPQQQQQGMMVKRCSESKRWRFRKRNASDSDSESEAAMSDD 319
Query: 232 ----------NKRQKKETM------GIAESIRWVAEVVVR----SEQTRMETLKEIEKMR 271
+KR K E G+ R + ++R EQT L+++ +M
Sbjct: 320 SGDSLPPPPLSKRMKTEEKKKQDGDGVGNKWRELTRAIMRFGEAYEQTENAKLQQVVEME 379
Query: 272 VEAEAKRGEMEIKRTEIIANTQLEIARI 299
E E+E++R + TQLEI+++
Sbjct: 380 KERMKFLKELELQRMQFFVKTQLEISQL 407
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.306 0.123 0.328
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 519,713,370
Number of Sequences: 2790947
Number of extensions: 21201294
Number of successful extensions: 81240
Number of sequences better than 10.0: 1230
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 1178
Number of HSP's that attempted gapping in prelim test: 78975
Number of HSP's gapped (non-prelim): 2687
length of query: 325
length of database: 848,049,833
effective HSP length: 127
effective length of query: 198
effective length of database: 493,599,564
effective search space: 97732713672
effective search space used: 97732713672
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC126006.11


