

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126006.10 - phase: 0 /pseudo
(352 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
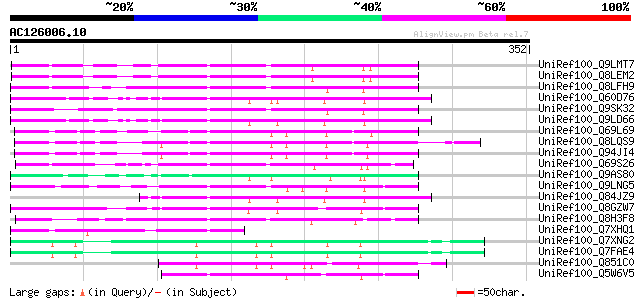
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 104 4e-21
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 103 9e-21
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 84 6e-15
UniRef100_Q60D76 Putative polyprotein [Oryza sativa] 84 8e-15
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 84 8e-15
UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein ... 82 2e-14
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 78 4e-13
UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa] 78 4e-13
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 77 5e-13
UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [... 77 5e-13
UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa] 76 1e-12
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 75 3e-12
UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa] 69 1e-10
UniRef100_Q8GZW7 Putative transposable element [Oryza sativa] 64 8e-09
UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like p... 59 2e-07
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 56 1e-06
UniRef100_Q7XNG2 OSJNBa0096F01.11 protein [Oryza sativa] 55 3e-06
UniRef100_Q7FAE4 OSJNBa0052P16.6 protein [Oryza sativa] 55 3e-06
UniRef100_Q851C0 Putative mutator-like transposase [Oryza sativa] 52 2e-05
UniRef100_Q5W6V5 Putative polyprotein [Oryza sativa] 52 3e-05
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 104 bits (259), Expect = 4e-21
Identities = 77/283 (27%), Positives = 133/283 (46%), Gaps = 32/283 (11%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + +R LG + G+ G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLGKLPK------GELSG 136
Query: 62 GYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSN 121
++ L++ + A ++E+ Y R YL+Y++G +F
Sbjct: 137 NRVTAKWLKESF----------AECPKGATMKEIE----YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLIF 181
+I + YL ED + Y+WG LA+LY ++ +A + +GG +TLL +
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL---QCWS 238
Query: 182 YFFVNVDPNTDYMENYPVAARWK---LQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREI 238
YF +N+D +P+A WK + + YR LD + +V W P+E +I
Sbjct: 239 YFHLNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDLDI 298
Query: 239 --QDFED--VFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
Q F+D + S + G + V H P+R ++Q+G Q IP
Sbjct: 299 VPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP 341
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 103 bits (256), Expect = 9e-21
Identities = 76/283 (26%), Positives = 133/283 (46%), Gaps = 32/283 (11%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + ++ LG + G+ G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLKLLGKLPK------GELSG 136
Query: 62 GYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSN 121
++ L++ + A ++E+ Y R YL+Y++G +F
Sbjct: 137 NRVTAKWLKESF----------AECPKGATMKEIE----YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLIF 181
+I + YL ED + Y+WG LA+LY ++ +A + +GG +TLL +
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL---QCWS 238
Query: 182 YFFVNVDPNTDYMENYPVAARWK---LQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREI 238
YF +N+D +P+A WK + + YR LD + +V W P+E +I
Sbjct: 239 YFHLNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDLDI 298
Query: 239 --QDFED--VFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
Q F+D + S + G + V H P+R ++Q+G Q IP
Sbjct: 299 VPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP 341
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 84.0 bits (206), Expect = 6e-15
Identities = 76/283 (26%), Positives = 118/283 (40%), Gaps = 28/283 (9%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P+GEMT+TLD+VA + L I+G + G K+ + R LG A K
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIV-GSKVDDEVAMDMCGRLLGKLPSAANK------ 144
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
++ R+ + N L T + V R YLLYLIG +F
Sbjct: 145 --EVNCSRV---------KLNWLKRTFSECPEDASFDVVKCHTRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLI 180
++ + YL ED + Y+WG LA LY L +A + G +TLL +
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL---QCW 249
Query: 181 FYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIM--YRSLLDRIQLDDVCWRPYEEHREI 238
YF +++ +P+A WK + + + YR LD + + W PYE +
Sbjct: 250 SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERFENL 309
Query: 239 ----QDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
+ + S + ++ H P+R LRQ+G Q IP
Sbjct: 310 IPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>UniRef100_Q60D76 Putative polyprotein [Oryza sativa]
Length = 1754
Score = 83.6 bits (205), Expect = 8e-15
Identities = 88/308 (28%), Positives = 133/308 (42%), Gaps = 39/308 (12%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P GE+TVTL+DVA + LPI G+ ++ + +LG+ A GQ
Sbjct: 974 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRER--VEEYLGLEPPVAPD--GQRQ 1029
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
P ++L RAN + P E +E A V+ YC R Y+LY+ G +LF D
Sbjct: 1030 TKTSGVP------LSWL-RANF---GQCPAEADE-ATVQRYC-RAYVLYIFGSILFPDSG 1077
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACR--PGDKALGGSVTLLTI-- 176
++L + D + +SWG LA+LY +L DACR GD L G V LL +
Sbjct: 1078 GDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLLQVWM 1136
Query: 177 --RKLI---FYFFVNVDPNTDYMENYPVAARWKLQKGHEEG------IMYRSLLDRIQLD 225
R L+ + P+ D VA W+ G Y + +D +Q +
Sbjct: 1137 WMRLLVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYCHYTNEMDYLQPE 1196
Query: 226 DVCWRPYEEHREIQ-------DFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIPR 278
V W PY+ ++ ED + V + RV+RQ+G +QTIP
Sbjct: 1197 HVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRLQTIPH 1256
Query: 279 HPTDVVEL 286
+ ++L
Sbjct: 1257 RFSTSIDL 1264
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 83.6 bits (205), Expect = 8e-15
Identities = 75/283 (26%), Positives = 115/283 (40%), Gaps = 28/283 (9%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P+GEMT+TLD+VA + L I+G + K V + A +CG+
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK----------------VGDEVAMDMCGRLL 135
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
G S + N L T + V R YLLYLIG +F
Sbjct: 136 GKLPSAANKE--VNCSRVKLNWLKRTFSECPEDASFDVVKCHTRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLI 180
++ + YL ED + Y+WG LA LY L +A + G +TLL +
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL---QCW 249
Query: 181 FYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIM--YRSLLDRIQLDDVCWRPYEEHREI 238
YF +++ +P+A WK + + + YR LD + + W PYE +
Sbjct: 250 SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERFENL 309
Query: 239 ----QDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
+ + S + ++ H P+R LRQ+G Q IP
Sbjct: 310 IPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein [Oryza sativa]
Length = 1230
Score = 82.0 bits (201), Expect = 2e-14
Identities = 86/308 (27%), Positives = 131/308 (41%), Gaps = 39/308 (12%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P GE+TVTL+DVA + LPI G+ ++ + +LG+ A GQ
Sbjct: 592 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRER--VEEYLGLEPPVAPD--GQRQ 647
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
P ++L RAN + P E +E A V+ YC R Y+LY+ G +LF D
Sbjct: 648 TKTSGVP------LSWL-RANF---GQCPAEADE-ATVQRYC-RAYVLYIFGSILFPDSG 695
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACR--PGDKALGGSVTLLTIRK 178
++L + D + +SWG LA+LY +L DACR GD L G V LL +
Sbjct: 696 GDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLLQVWM 754
Query: 179 LI-------FYFFVNVDPNTDYMENYPVAARWKLQKGHEEG------IMYRSLLDRIQLD 225
+ + P+ D VA W+ G Y + +D +Q +
Sbjct: 755 WMRLPVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYYHYTNEMDYLQPE 814
Query: 226 DVCWRPYEEHREIQ-------DFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIPR 278
V W PY+ ++ ED + V + RV+RQ+G +QTIP
Sbjct: 815 HVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRLQTIPH 874
Query: 279 HPTDVVEL 286
+ ++L
Sbjct: 875 RFSTSIDL 882
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 77.8 bits (190), Expect = 4e-13
Identities = 89/293 (30%), Positives = 128/293 (43%), Gaps = 43/293 (14%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGY 63
GEMTVTL DV+ L LPI+GR + H+ A +++ LG + G+ +
Sbjct: 153 GEMTVTLQDVSMLLALPIDGRPVC---STTDHDYAQMVIDCLG-HDPRGPSMPGK---SF 205
Query: 64 ISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSNKR 123
+ Y L+ + L T+D VR Y+L L+ +LF D + R
Sbjct: 206 LHYKWLKKHF------YELPEGTDDQTVERH--------VRAYILSLLCGVLFPDGTG-R 250
Query: 124 IELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELAD-ACRPGDKALGGSVTLLTI----RK 178
+ LIYL + D + + YSWG LA+LY L A K +GGS+ LL + R
Sbjct: 251 MSLIYLPLIAD-LSLVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 309
Query: 179 LIFYFFVN--VDPNTDYMEN-YPVAARWKLQKGHEEGI-----MYRSLLDRIQLDDVCWR 230
+ V V P TD ++ PV RW + E YR L+ + D V W
Sbjct: 310 HVGRPLVRSPVCPETDIPQDVLPVGFRWVGARTQSENATRCLKQYRDELNLQRADQVKWE 369
Query: 231 PYEEHREIQDFEDV------FWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 370 PY-LHIESASLPLLCTKNADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 421
>UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa]
Length = 718
Score = 77.8 bits (190), Expect = 4e-13
Identities = 99/337 (29%), Positives = 141/337 (41%), Gaps = 66/337 (19%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGY 63
GEM VTL DVA L LPI+GR + H+ A +++ LG + G+ +
Sbjct: 184 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLG-HDPRGPSMPGK---SF 236
Query: 64 ISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTY--CVRCYLLYLIGCLLFGDRSN 121
+ Y L+ + EL E A +T VR Y+L L+ +LF D +
Sbjct: 237 LHYKWLKKHF----------------YELPEGADDQTVERHVRAYILSLLCGVLFPDGTG 280
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELAD-ACRPGDKALGGSVTLLTI---- 176
R+ LIYL + D + + YSWG LA+LY L A K +GGS+ LL +
Sbjct: 281 -RMSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWE 338
Query: 177 RKLIFYFFVN--VDPNTDYMENY-PVAARWKLQKGHEEGI-----MYRSLLDRIQLDDVC 228
R + V + P TD ++ PV RW + E YR L+ + D V
Sbjct: 339 RSHVGRPLVRSPLCPETDIPQDVPPVGFRWVGARTQSENATRCLKQYRDELNLQRADQVK 398
Query: 229 WRPYEEHREIQDF------EDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIPRHPTD 282
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 399 WEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP----- 452
Query: 283 VVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHGGW 319
FR TL+A +SR+ H W
Sbjct: 453 -------------PPFRP-TLQALHRISRRGREHENW 475
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 77.4 bits (189), Expect = 5e-13
Identities = 90/295 (30%), Positives = 129/295 (43%), Gaps = 47/295 (15%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGY 63
GEM VTL DVA L LPI+GR + H+ A +++ LG + G+ +
Sbjct: 170 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLG-HDPRGPSMPGK---SF 222
Query: 64 ISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTY--CVRCYLLYLIGCLLFGDRSN 121
+ Y L+ + EL E A +T VR Y+L L+ +LF D +
Sbjct: 223 LHYKWLKKHF----------------YELPEGADDQTVERHVRAYILSLLCGVLFPDGTG 266
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELAD-ACRPGDKALGGSVTLLTI---- 176
R+ LIYL + D + + YSWG LA+LY L A K +GGS+ LL +
Sbjct: 267 -RMSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWE 324
Query: 177 RKLIFYFFVN--VDPNTDYMENY-PVAARWKLQKGHEEGI-----MYRSLLDRIQLDDVC 228
R + V + P TD ++ PV RW + E YR L+ + D V
Sbjct: 325 RSHVGRPLVRSPLCPETDIPQDLPPVGFRWVGARTQSENATRCLKQYRDELNLQRADQVK 384
Query: 229 WRPYEEHREIQDF------EDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 385 WEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 438
>UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [Oryza sativa]
Length = 619
Score = 77.4 bits (189), Expect = 5e-13
Identities = 76/283 (26%), Positives = 125/283 (43%), Gaps = 38/283 (13%)
Query: 5 EMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYI 64
EM +L DV+ + +P+ G +++ + + + + LG S E++CG
Sbjct: 96 EMAPSLRDVSYILGIPVTGHVVT-AEPIGDEAVRRMCLHFLGESPGNGEQLCGL------ 148
Query: 65 SYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSNKRI 124
T+L R E+P + E+A Y R YLLYL+G LF D +
Sbjct: 149 -------IRLTWLYR-KFHQLPENPT-INEIA----YSTRAYLLYLVGSTLFPDTMRGFV 195
Query: 125 ELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRP-GDKALGGSVTLLTIRKLIFYF 183
YL + D + +R Y+WG LA+LY L+ A P GS TLL I+ +
Sbjct: 196 SPRYLPLLAD-FRKIREYAWGSAALAHLYRGLSVAVTPNATTQFLGSATLL--MAWIYEY 252
Query: 184 FVNVDPNTDYMEN-YPVAARWKL---QKGHEEGIM-YRSLLDRIQLDDVCWRPYEEHRE- 237
P P A RW +G + +M +R + D +Q DV W PY++
Sbjct: 253 LPLTQPQQKNQNTLLPRACRWNFGGATRGQRKKVMEWRKVFDELQFSDVNWNPYKDMNPA 312
Query: 238 -IQDF----EDVFWYSGWIMC-GVRRVYRHLPERVLRQYGYMQ 274
I ++ +++ + W++ ++ VY +P+R RQ+G Q
Sbjct: 313 IIPEYCIAADNICYSRTWLISFNIKEVY--VPDRFSRQFGREQ 353
>UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa]
Length = 792
Score = 76.3 bits (186), Expect = 1e-12
Identities = 86/302 (28%), Positives = 122/302 (39%), Gaps = 49/302 (16%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
MP GE+T+TL DVA + LPI G ++ P++E L+ R+LG +
Sbjct: 278 MPCGEITITLQDVAMILGLPIAGHAVTVNPTEPQNE---LVERYLGKAPPPDR------- 327
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
P LR + RA ED E ++ + R Y+L LI LLF D S
Sbjct: 328 ----PRPGLRVSWV----RAEFNNCPEDADE----ETIKQH-ARAYILSLISGLLFPDAS 374
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACR--PGDKALGGSVTLLTI-- 176
+ Y + + +YSWG TLAYLY + DACR L G + LL
Sbjct: 375 GD-LYTFYPFPLIADLENIGSYSWGSATLAYLYRAMCDACRRQSDGSNLTGCLLLLQFWS 433
Query: 177 --------RKLIFYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIMYR------SLLDRI 222
L+ + NV+ D + V RW + R + D +
Sbjct: 434 WEHFPIGRPDLVKLKYPNVEELEDERDRPTVGLRWVVGVCARRAAPARCYEHFTNEFDLL 493
Query: 223 QLDDVCWRPYEEHR--EIQ-----DFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQT 275
D V W PY E R E+Q + W + + V ++PERV+RQ+G Q
Sbjct: 494 TDDQVVWCPYREERMKELQLAPICTQDSHLWLTRAPLLYFFMVEIYMPERVMRQFGLHQV 553
Query: 276 IP 277
P
Sbjct: 554 CP 555
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 75.1 bits (183), Expect = 3e-12
Identities = 82/302 (27%), Positives = 130/302 (42%), Gaps = 53/302 (17%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P GE+TVTL DV L L ++G ++ K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYC-VRCYLLYLIGCLLFGDR 119
G ++S LR+ + L + D V L+ C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRN-------LPADPDEVTLK--------CHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKL 179
S + L +L + D + + SWG TLA LY EL A + + G + LL +L
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLL---QL 253
Query: 180 IFYFFVNV-------DPNTDYMENY------PVAARWKLQKGHEEGI-----MYRSLLDR 221
+ ++V D YM+ P+ RW+ H+E YR D+
Sbjct: 254 WAWERLHVGRPGRLKDVGASYMDGIDGPLPDPLGCRWRASLSHKENPRGGLDFYRDQFDQ 313
Query: 222 IQLDDVCWRPYEEHREIQ------DFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQT 275
+ + V W+PY + E+++ ++C V H P+RVLRQ+G QT
Sbjct: 314 QKDEQVIWQPYTPDLLAKIPLICVSGENIWRTVAPLIC-FDVVEWHRPDRVLRQFGLHQT 372
Query: 276 IP 277
IP
Sbjct: 373 IP 374
>UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa]
Length = 1381
Score = 69.3 bits (168), Expect = 1e-10
Identities = 62/220 (28%), Positives = 94/220 (42%), Gaps = 25/220 (11%)
Query: 89 PVELEELARVRTYCVRCYLLYLIGCLLFGDRSNKRIELIYLTTMEDGYAGMRNYSWGGMT 148
P E +E A V+ YC R Y+LY+ G +LF D ++L + D + +SWG
Sbjct: 804 PAEADE-ATVQRYC-RAYVLYIFGSILFPDSGGDMASWMWLPLLAD-WDEAGTFSWGSAA 860
Query: 149 LAYLYGELADACR--PGDKALGGSVTLLTIRKLI-------FYFFVNVDPNTDYMENYPV 199
LA+LY +L DACR GD L G V LL + + + P+ D V
Sbjct: 861 LAWLYRQLCDACRRQGGDANLAGCVWLLQVWMWMRLPVGRPMWRTHQAWPHQDADRCPTV 920
Query: 200 AARWKLQKGHEEG------IMYRSLLDRIQLDDVCWRPYEEHREIQ-------DFEDVFW 246
A W+ G Y + +D +Q + V W PY+ ++ ED
Sbjct: 921 AHLWESVPSPVVGRRNLAYYHYTNEMDYLQPEHVVWMPYQAQEVLELELNPMCHIEDALK 980
Query: 247 YSGWIMCGVRRVYRHLPERVLRQYGYMQTIPRHPTDVVEL 286
+ V + RV+RQ+G +QTIP + ++L
Sbjct: 981 TLRCPLICFYAVEFQMCHRVMRQFGRLQTIPHRFSTSIDL 1020
>UniRef100_Q8GZW7 Putative transposable element [Oryza sativa]
Length = 663
Score = 63.5 bits (153), Expect = 8e-09
Identities = 71/305 (23%), Positives = 125/305 (40%), Gaps = 51/305 (16%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P GEMT+TL DVA + LP+ G ++ + P + + ++ ++ + ++
Sbjct: 92 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWHAQVQQLFGIPLNIEQGQGGKKKQN 151
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
G +S+ + + +D E RV Y R Y+L+L+G +LF D
Sbjct: 152 GIPLSW------------LSQNFSHLDDDAEPW---RVECYA-RAYILHLLGGVLFPDAG 195
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADAC--RPGDKALGGSVTLLTIRK 178
I++ + + + +SWG LA+ Y +L +AC + + G V L+ I
Sbjct: 196 GDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQIWM 254
Query: 179 LI----------------FYFFVNVDPNTDYM-ENYPVA-ARWKLQKGHEEGIMYRSLLD 220
+ +Y +++ Y+ E+ A A W + H Y + +D
Sbjct: 255 WLRLPVGRPKWRQSFTPWYYNEPDMEKTVAYLFESTATAHAHWDVAYKH-----YVNEMD 309
Query: 221 RIQLDDVCWRPYEEHRE--------IQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGY 272
+Q + W PY + D F Y ++C V HLP RV+RQ+G
Sbjct: 310 CLQPQHIEWLPYHTNEASSLTLNLMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGK 368
Query: 273 MQTIP 277
Q P
Sbjct: 369 KQDWP 373
>UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like protein [Oryza
sativa]
Length = 589
Score = 59.3 bits (142), Expect = 2e-07
Identities = 71/288 (24%), Positives = 120/288 (41%), Gaps = 42/288 (14%)
Query: 5 EMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYI 64
EM +L D A + +P+ GR+++ G V + E +C Q G
Sbjct: 83 EMAPSLRDAAYILGIPVTGRVVTTG----------------AVLNKSVEDLCFQYLG--- 123
Query: 65 SYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSNKRI 124
P RD +++ + + L S + Y R YLL+LIG L +R +
Sbjct: 124 QVPDCRDCRGSHV-KLSWLQSKFSRIPERPTNDQTMYGTRAYLLFLIGSALLPERDRGYV 182
Query: 125 ELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADA-CRPGDKALGGSVTLLTIRKLIFYF 183
YL + D + ++ Y+WG LA+LY L+ A K L GS LL I+ +
Sbjct: 183 SPKYLPLLSD-FDKVQEYAWGAAALAHLYKALSIAVAHSARKRLFGSAALL--MGWIYEY 239
Query: 184 FVNVDPNT-DYMEN-YPVAARW---KLQKGHEEGIMYRSLLDRIQLDDVCWRPYE----- 233
+ P+ D E+ +P +W + + + R +Q+ DV W PY+
Sbjct: 240 IPALRPDMYDPPEHIFPRVLKWTGSTISQPAKNVSDIRKAFGLLQVSDVNWEPYKGVDPA 299
Query: 234 ---EHREIQDFEDVFWYSGWIMC-GVRRVYRHLPERVLRQYGYMQTIP 277
+H D ++ + W++ ++ +Y P+R RQ+G Q P
Sbjct: 300 SIPKHCAAPD--NLCFSRTWLVSFNLKEIY--APDRFARQFGQEQHRP 343
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 56.2 bits (134), Expect = 1e-06
Identities = 44/163 (26%), Positives = 75/163 (45%), Gaps = 15/163 (9%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQE----AEKIC 56
+P+GEMT+TL DV+CL LPI GR ++ ++ R LG+ +E +K
Sbjct: 87 LPVGEMTITLQDVSCLWGLPIHGRPIT---GQADGSWVDMIERLLGIPMEEQHMKQKKRK 143
Query: 57 GQEYGGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLF 116
++ +SY R + R ++ E+ + R +L +IG ++F
Sbjct: 144 KEDDMTMVSYSRYSISLSKLRDRFRVMPKNATEREI-------NWYTRALVLDIIGSMVF 196
Query: 117 GDRSNKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADA 159
D S + +YL M + + Y+WG L+ LY +L+ A
Sbjct: 197 TDTSGDGVPAMYLQFMMN-LSEQTEYNWGAAALSMLYRQLSIA 238
>UniRef100_Q7XNG2 OSJNBa0096F01.11 protein [Oryza sativa]
Length = 1365
Score = 55.1 bits (131), Expect = 3e-06
Identities = 82/362 (22%), Positives = 132/362 (35%), Gaps = 64/362 (17%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLS-----HGKKMPKHEGAALLMR--HLGVSQQEAE 53
+P GEM TL DV+ L LP+ G + G K +MR HLG +
Sbjct: 825 LPCGEMAPTLQDVSYLLGLPLAGAPVGPVDGVFGWKEDITARFEQVMRLPHLGPANT--- 881
Query: 54 KICGQEYGGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGC 113
L + T +A LL T D + + + + YLL+L G
Sbjct: 882 ---------------LPPYSTVGPSKAWLLQFTADLLHPDADDYLVRRSLEAYLLWLFGW 926
Query: 114 LLFGDRSNKRIE---LIYLTTMEDGYA-GMRNYSWGGMTLAYLYGELADACRPGDKA--L 167
++F ++ + Y ++ D + +SWG LA Y L +AC D +
Sbjct: 927 VMFTSTHGHAVDFRLVHYARSIADAQPQDVPQWSWGSAVLAATYHALCEACTKTDAGAII 986
Query: 168 GGSVTLLTI---------RKLIFYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIM---- 214
G LL + R ++ V + + E+ P + ++G +
Sbjct: 987 AGCPMLLQLWAAERFAIGRPVVDSAPYGVGRSAQWPEDGPTMGTYWCRRGRRYAHVQVRR 1046
Query: 215 ----YRSLLDRIQLDDVCWRPYEEHR----------EIQDFEDVFWYSGWIMCGVRRVYR 260
+ DR+Q DV W PY E + + +W + M V
Sbjct: 1047 GYPDFVFEFDRLQPSDVIWEPYTEEAVAARAPLGLSSLCTRDQAYWLTILPMVFDIFVEP 1106
Query: 261 HLPERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHGGWL 320
H P+RV+RQ+G Q P + V LP + + R L A+ R + W+
Sbjct: 1107 HCPQRVMRQFGLRQVFPGNVQPTV-LPADHSLT-----RRGQLAGALWAPRVQQYVDDWV 1160
Query: 321 MA 322
+A
Sbjct: 1161 LA 1162
>UniRef100_Q7FAE4 OSJNBa0052P16.6 protein [Oryza sativa]
Length = 1489
Score = 55.1 bits (131), Expect = 3e-06
Identities = 82/362 (22%), Positives = 132/362 (35%), Gaps = 64/362 (17%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLS-----HGKKMPKHEGAALLMR--HLGVSQQEAE 53
+P GEM TL DV+ L LP+ G + G K +MR HLG +
Sbjct: 825 LPCGEMAPTLQDVSYLLGLPLAGAPVGPVDGVFGWKEDITARFEQVMRLPHLGPANT--- 881
Query: 54 KICGQEYGGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGC 113
L + T +A LL T D + + + + YLL+L G
Sbjct: 882 ---------------LPPYSTVGPSKAWLLQFTADLLHPDADDYLVRRSLEAYLLWLFGW 926
Query: 114 LLFGDRSNKRIE---LIYLTTMEDGYA-GMRNYSWGGMTLAYLYGELADACRPGDKA--L 167
++F ++ + Y ++ D + +SWG LA Y L +AC D +
Sbjct: 927 VMFTSTHGHAVDFRLVHYARSIADAQPQDVPQWSWGSAVLAATYHALCEACTKTDAGAII 986
Query: 168 GGSVTLLTI---------RKLIFYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIM---- 214
G LL + R ++ V + + E+ P + ++G +
Sbjct: 987 AGCPMLLQLWAAERFAIGRPVVDSAPYGVGRSAQWPEDGPTMGTYWCRRGRRYAHVQVRR 1046
Query: 215 ----YRSLLDRIQLDDVCWRPYEEHR----------EIQDFEDVFWYSGWIMCGVRRVYR 260
+ DR+Q DV W PY E + + +W + M V
Sbjct: 1047 GYPDFVFEFDRLQPSDVIWEPYTEEAVAARAPLGLSSLCTRDQAYWLTILPMVFDIFVEP 1106
Query: 261 HLPERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHGGWL 320
H P+RV+RQ+G Q P + V LP + + R L A+ R + W+
Sbjct: 1107 HCPQRVMRQFGLRQVFPGNVQPTV-LPADHSLT-----RRGQLAGALWAPRVQQYVDDWV 1160
Query: 321 MA 322
+A
Sbjct: 1161 LA 1162
>UniRef100_Q851C0 Putative mutator-like transposase [Oryza sativa]
Length = 1527
Score = 52.4 bits (124), Expect = 2e-05
Identities = 57/218 (26%), Positives = 89/218 (40%), Gaps = 34/218 (15%)
Query: 102 CVRCYLLYLIGCLLFGDRSNKRIE---LIYLTTMEDGYAG-MRNYSWGGMTLAYLYGELA 157
C+ YLL+L G ++F ++ + Y ++ D G + +SWG LA LY L
Sbjct: 1071 CLEAYLLWLFGWVMFCGGHGHAVDKGLVHYARSIADAAVGEVPQWSWGSALLAALYRGLC 1130
Query: 158 DACRPGDKA--LGGSVTLLTI--RKLIFYFFVNVDPNTDYMENYP--VAARW---KLQKG 208
+AC D + GG L+I + I VD + + Y RW ++++
Sbjct: 1131 EACTKTDPSATFGGCPLFLSIWAAERIAIGRPEVDQHAYEVSLYAERPERRWAHVQVRRS 1190
Query: 209 HEEGIMYRSLLDRIQLDDVCWRPYEEH----------REIQDFEDVFWYSGWIMCGVRRV 258
+ E +M DR+ DV W PY + + +W + M V
Sbjct: 1191 YPEFVME---FDRLLPTDVVWEPYSAAATQPQAPLGLSTLCTRDQAYWMTTVPMVYDICV 1247
Query: 259 YRHLPERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFV 296
H P RV+RQ+G+ Q P +P P V A V
Sbjct: 1248 EPHAPFRVMRQFGFRQPFP--------VPFPTTVPAAV 1277
>UniRef100_Q5W6V5 Putative polyprotein [Oryza sativa]
Length = 1525
Score = 51.6 bits (122), Expect = 3e-05
Identities = 45/185 (24%), Positives = 77/185 (41%), Gaps = 16/185 (8%)
Query: 104 RCYLLYLIGCLLFGDRSNKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPG 163
R Y+L+L+G +LF D I++ + + + +SWG LA+ Y +L +AC
Sbjct: 1087 RAYILHLLGGVLFPDAGGDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQ 1145
Query: 164 DKALGGSVTLLTIRKLIFYFFVN---VDPNTDYMENYPVAARWKLQKGHEEGIMYRSLLD 220
+ S +L I+ ++ N ++ Y+ A ++ Y + +D
Sbjct: 1146 APSSNMSGCVLLIQMWMWLRLPNEPDMEKTVAYLFESTATAHAHRDVAYKH---YVNEMD 1202
Query: 221 RIQLDDVCWRPYEEHRE--------IQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGY 272
+Q + W PY + D F Y ++C V HLP RV+RQ+G
Sbjct: 1203 YLQPQHIEWLPYHTNEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGK 1261
Query: 273 MQTIP 277
Q P
Sbjct: 1262 KQDWP 1266
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.143 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 604,507,108
Number of Sequences: 2790947
Number of extensions: 25210992
Number of successful extensions: 59042
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 87
Number of HSP's that attempted gapping in prelim test: 58790
Number of HSP's gapped (non-prelim): 217
length of query: 352
length of database: 848,049,833
effective HSP length: 128
effective length of query: 224
effective length of database: 490,808,617
effective search space: 109941130208
effective search space used: 109941130208
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC126006.10


