

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125481.7 + phase: 0 /pseudo
(489 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
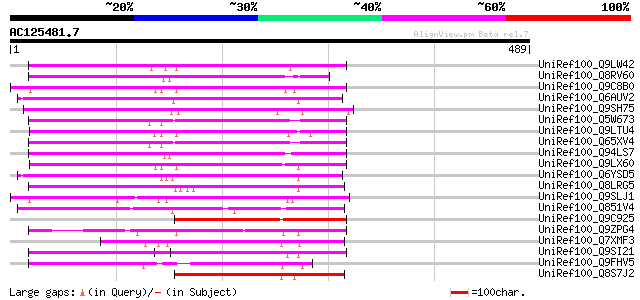
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 269 2e-70
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 261 3e-68
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 252 1e-65
UniRef100_Q6AUV2 Hypothetical protein OSJNBa0091B22.10 [Oryza sa... 248 2e-64
UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana] 247 5e-64
UniRef100_Q5W673 Putative helicase [Oryza sativa] 246 1e-63
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 244 4e-63
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 243 7e-63
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 239 2e-61
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 238 4e-61
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 236 1e-60
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 229 1e-58
UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana] 219 1e-55
UniRef100_Q851V4 Putative helicase [Oryza sativa] 209 1e-52
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 182 2e-44
UniRef100_Q9ZPG4 F5K24.3 protein [Arabidopsis thaliana] 171 6e-41
UniRef100_Q7XMF3 OSJNBa0061G20.6 protein [Oryza sativa] 166 1e-39
UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana] 162 3e-38
UniRef100_Q9FHV5 Helicase [Arabidopsis thaliana] 153 1e-35
UniRef100_Q8S7J2 Hypothetical protein OSJNBa0095C06.16 [Oryza sa... 147 5e-34
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 269 bits (687), Expect = 2e-70
Identities = 147/370 (39%), Positives = 198/370 (52%), Gaps = 70/370 (18%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD +TMIES RLR+I+KNQ +R + +A D G+ D SN G+ +++ PSF GG Y
Sbjct: 630 VDAFTMIESNRLRFIKKNQTKLRSTNKQAVQDASDAGDNDLSNKGKSVIIPPSFTGGPAY 689
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVF----- 132
M N DAM CK +G+PDLF+T CN+ W +I F+KER L A D+PDI+CR++
Sbjct: 690 MQQNYLDAMGTCKHFGFPDLFITFTCNSKWPKITRFVKERKLNAEDKPDIICRIYKMKLE 749
Query: 133 -------------KMKLDKLMDDFEK---------------------------------- 145
K KL +F+K
Sbjct: 750 NLMDDLTKNHIFGKTKLATYTIEFQKRGLPHAHIIVWMDPRYKFPIADHVDKIIFAEIPD 809
Query: 146 EECLAKLMQR------HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRR 198
+E KL Q H PC P SPCM+ G+C+KYY KN+ T +D EGYP Y+RR
Sbjct: 810 KENDPKLYQAVSECMIHGPCRLVNPNSPCMENGKCSKYYPKNHVENTFLDNEGYPIYRRR 869
Query: 199 DFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALM 258
D G + K + DNRYVVPYN LL +Y H+NVE CN+S S+KYLFKYVNK PD+ +
Sbjct: 870 DTGRFIKKNKYPCDNRYVVPYNDFLLRKYRAHINVEWCNQSVSVKYLFKYVNKGPDRVTV 929
Query: 259 QLSVD-----------GDNRDKSKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVL 307
L G+ + + ++++ Y+DCRYVS CEA+WRI + IH++ V
Sbjct: 930 SLEPHRKEVVSEENNVGETNNDPQEQNQVEDYFDCRYVSACEAMWRIKGYPIHYRQTLVT 989
Query: 308 KLLFHLHNEQ 317
KL FH +Q
Sbjct: 990 KLTFHEKGKQ 999
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 261 bits (667), Expect = 3e-68
Identities = 139/333 (41%), Positives = 192/333 (56%), Gaps = 56/333 (16%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD TMIE++R+ Y++ NQKT+R + + + + G D+S G++ VL SF GG Y
Sbjct: 257 VDGLTMIEAERISYLQFNQKTLRAEKYSSVEASAEKGNNDSSMCGKRYVLPSSFVGGARY 316
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVFKMKLD 137
M DAM++C+ YGYP LF+T CN W EI +L R L A DR DIV RV+KMKLD
Sbjct: 317 MQQKYMDAMALCQHYGYPHLFITFTCNPKWPEITRYLNARGLSAEDRADIVTRVYKMKLD 376
Query: 138 KLMDDF----------------------EKEECL-------------------------- 149
L+ D EKE +
Sbjct: 377 HLIWDIKNNNVFGPSRAGLPHAHILLFLEKEAKIPTTADVDKFIWAEIPDKYTHPILHEV 436
Query: 150 AKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRRDFGIHVDKQR 208
K M H PCG A SPCM++G+C+K + K++ +T I+ +G+P YKRRD G V+K
Sbjct: 437 VKEMMIHGPCGLANKNSPCMEKGKCSKKFPKDFTSSTHINKDGFPIYKRRDDGRFVEKNG 496
Query: 209 VLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQLSVDGDNRD 268
+LLDNR+V+PYNP LL++Y H+NVE ++S +KYLFKY+NK DK ++ ++
Sbjct: 497 ILLDNRFVIPYNPTLLLKYQAHINVEWVSQSKIVKYLFKYINKGNDKVTARI------KE 550
Query: 269 KSKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHH 301
+ P +EIK++YDCRY+SPCEA WRIF FD+HH
Sbjct: 551 GTGP-NEIKRWYDCRYISPCEAAWRIFGFDLHH 582
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 252 bits (644), Expect = 1e-65
Identities = 150/393 (38%), Positives = 200/393 (50%), Gaps = 76/393 (19%)
Query: 1 MQDRVVDYGNVVNSRRSV-----DCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGE 55
+Q+R V+ ++ S+R D YT IES RL YI+ NQ +RC + EA G
Sbjct: 628 IQEREVECQTLLRSKRLFQQCLCDAYTTIESNRLNYIKFNQSKLRCENYTSVKEAAAAGA 687
Query: 56 TDASNVGQKIVLQPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLK 115
T G ++++ SF GG YM + DAM+ICK YG+PDLF+T CN W EI +
Sbjct: 688 TTMEEEGNQLLIPASFTGGPRYMVQSYYDAMAICKHYGFPDLFITFTCNPKWPEITRYYD 747
Query: 116 ERNLKASDRPDIVCRVFKMKL--------------------------------------- 136
+R L DRP+I+ R+FK+KL
Sbjct: 748 KRGLNPEDRPNIIARIFKIKLDSLMNDLTVKKMLGKTVASMYTVEFQKRGLPHAHILLFM 807
Query: 137 ------------DKLMD----DFEKEECLAKLMQR---HVPCGGARPFSPCMDRGRCTKY 177
DKL+ D EKE L ++++ H PCG A SPCM G C+K
Sbjct: 808 HAKSKLPTADDIDKLISAEIPDKEKEPDLYEVIKNSMIHGPCGSANVNSPCMVDGECSKL 867
Query: 178 YRKNYKGTTTI-DEGYPRYKRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECC 236
Y K ++ T I +GYP Y+RR +V+K + DNRYVVPYN L +RY H+NVE C
Sbjct: 868 YPKKHQDITKIGSDGYPIYRRRKTDDYVEKGGIKCDNRYVVPYNKKLSLRYNAHINVEWC 927
Query: 237 NKSNSIKYLFKYVNKVPDKALM-----QLSVDGDNR-------DKSKPVDEIKQYYDCRY 284
N++ SIKYLFKY+NK PDK + Q + GD+ K +EIK ++DCRY
Sbjct: 928 NQNGSIKYLFKYINKGPDKVVFIVEPTQQTTAGDSETPQQEQGSAEKKKNEIKDWFDCRY 987
Query: 285 VSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNEQ 317
VS EAVWRIF + I H V KL FH+ +Q
Sbjct: 988 VSASEAVWRIFKYPIQHISTPVQKLSFHVEGKQ 1020
>UniRef100_Q6AUV2 Hypothetical protein OSJNBa0091B22.10 [Oryza sativa]
Length = 1500
Score = 248 bits (634), Expect = 2e-64
Identities = 131/322 (40%), Positives = 195/322 (59%), Gaps = 17/322 (5%)
Query: 8 YGNVVNSRRSVDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVL 67
YG + + + VD ++ IE+ RL++ ++K +R ++G+ +A+D G TD ++G++++L
Sbjct: 222 YGRL-SDQIDVDIFSTIEANRLQFFIDHRKELRGESVDGIVDAIDKGVTDGDSIGKRVIL 280
Query: 68 QPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLK-ERNLKASDRPD 126
SF GGR YM N QDAM+IC YG PDLFVT CN+ W++I + ++ E + SDR D
Sbjct: 281 PASFIGGRRYMVMNYQDAMAICHVYGSPDLFVTYTCNSKWQDIAEAIRFEPGQQPSDRAD 340
Query: 127 IVCRVFKMKLDKLMDDFEKEECLAKLM---------QRHVPCGGARPFSPCMDRGRCTKY 177
I+ RVF MK+++ + D + K++ H PCG A PCM +C+K
Sbjct: 341 IIVRVFNMKVNEFITDIREGRTFGKVLAGYELVSEFMMHGPCGDANKSCPCMKNDKCSKN 400
Query: 178 YRKNYKGTTTIDE-GYPRYKRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECC 236
Y K+++ T IDE + Y++R+ G + K + LDNR VVPYN LL +Y H+NVE C
Sbjct: 401 YPKDFQDETIIDEAAFTIYRQRNDGRCIIKNGIPLDNRSVVPYNMKLLKKYEAHINVEWC 460
Query: 237 NKSNSIKYLFKYVNKVPDKALMQLSVDGDNRDKS-----KPVDEIKQYYDCRYVSPCEAV 291
NKSN IKYLFKY+ K D+A + + S P DEI +Y D R++S CEA+
Sbjct: 461 NKSNLIKYLFKYITKGQDRAKIYFETTAKTTNASPNHDLAPRDEILEYMDARFLSTCEAL 520
Query: 292 WRIFAFDIHHK*PHVLKLLFHL 313
R+F FDIH++ P V +L+ HL
Sbjct: 521 HRLFEFDIHYRVPLVERLVVHL 542
>UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana]
Length = 1241
Score = 247 bits (631), Expect = 5e-64
Identities = 146/384 (38%), Positives = 195/384 (50%), Gaps = 73/384 (19%)
Query: 14 SRRSVDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAG 73
+ + ++ YT IES RL YI+ NQ +RC + + A + G T+ S G+ I + SF G
Sbjct: 268 ANKCINGYTTIESNRLAYIKFNQSNLRCKNYDTVKAAREAGNTNMSEQGKSIRIPQSFTG 327
Query: 74 GRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVFK 133
G +M N+ DAM+ICK YG+P+LF+T CN EI + KER L A DRPD+VC +FK
Sbjct: 328 GPRHMLNSYYDAMAICKMYGFPNLFITFTCNPKLPEITSYCKERKLNAEDRPDVVCWIFK 387
Query: 134 MKLDKLMDDFEKEECLAK--------LMQRHV---------------------------- 157
MKLD LM D ++ L K L H+
Sbjct: 388 MKLDSLMKDLTEDHLLGKTVMFQKRGLPHAHILLFMDKSCKFPTSDDIDNILSAEIPDKA 447
Query: 158 ----------------PCGGARPFSPCMDRGRCTKYYRKNYKGTTTI-DEGYPRYKRRDF 200
PCG A SPC+ G+C+K++ K TT+ +GYP Y+RR+
Sbjct: 448 KDPKLYEVVNDCMIHGPCGAANKESPCIVDGKCSKFFPKKLVEQTTVGKDGYPIYRRRES 507
Query: 201 GIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNK--------- 251
V+K + DN YVVPYN L +RY H+NVE C +S SIKYLFKY+NK
Sbjct: 508 EHFVEKGGIKCDNTYVVPYNRMLSLRYRAHINVEWCKQSGSIKYLFKYINKGQDRVAIVV 567
Query: 252 -VPDKALMQLSVDGDNRDKSKPVD---EIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVL 307
DK + G + +D EIK Y+DCRYVS EAVWRIF F I ++ V+
Sbjct: 568 EPKDKTSNMVLFSGSQKLLVAVIDDDKEIKDYFDCRYVSASEAVWRIFKFPIQYRTTPVM 627
Query: 308 KLLFHLHNEQ-------GNIDHVS 324
KL +HL +Q NID +S
Sbjct: 628 KLSYHLPGKQPLCFEDTQNIDELS 651
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 246 bits (627), Expect = 1e-63
Identities = 146/361 (40%), Positives = 191/361 (52%), Gaps = 71/361 (19%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD I RL +IRK+Q +R + GL +A++ G+T A VG++I+L SF GG Y
Sbjct: 550 VDALACIIQYRLDWIRKHQGNLRTELYAGLQDAIERGDTRADQVGKRILLPSSFTGGPRY 609
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNL-KASDRPDIVCR------ 130
N QDAM+IC+ GYPDLFVT CN W EIQ+ L E + K SDRPDIV R
Sbjct: 610 KAQNYQDAMAICRWAGYPDLFVTFTCNAAWPEIQNMLDEIGVQKPSDRPDIVDRVFHIKL 669
Query: 131 ------------------------------------VFKMKLDKLMD------------- 141
+F K DK D
Sbjct: 670 RELMTDIKDKQYFGKTLAIIYTIEFQKRGLPHAHILIFLDKKDKCPDASEIDRIISAEIP 729
Query: 142 ----DFEKEECLAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYK 196
D E E + M H PCG A+ SPCM +C + + K + TT+DE G+P Y+
Sbjct: 730 DKEEDREGFEAVENFMM-HGPCGEAKSNSPCMIENKCIRNFPKKFHSETTVDEDGFPTYR 788
Query: 197 RRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKA 256
RRD G +++K V LDNRYVVPYN LL++Y H+NVE CN+S SIKYLFKY++K D+A
Sbjct: 789 RRDNGRYIEKGNVKLDNRYVVPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHKGDDQA 848
Query: 257 LMQLSVDGDNRDKSKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNE 316
+ D DEIK+Y +C Y+S +A WRIF F++H++ P V +L FHL NE
Sbjct: 849 TALIESDH---------DEIKKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENE 899
Query: 317 Q 317
Q
Sbjct: 900 Q 900
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 244 bits (623), Expect = 4e-63
Identities = 143/381 (37%), Positives = 194/381 (50%), Gaps = 84/381 (22%)
Query: 19 DCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHYM 78
D YT IES RL YI+ NQ +RC N + E+ G T S G ++++ SF GG YM
Sbjct: 309 DAYTTIESNRLAYIKFNQSKLRCENFNSIKESASSGSTTMSEEGNQVLIPSSFTGGPRYM 368
Query: 79 FNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVFKMK--- 135
DAM+ICK +G+PDLF+T CN W EI + ++R L A DRPDIV R+FK+K
Sbjct: 369 LQTYYDAMAICKHFGFPDLFITFTCNPKWPEITRYCEKRGLTADDRPDIVARIFKIKLDS 428
Query: 136 ------------------------------------------------LDKLMD----DF 143
+DK++ +
Sbjct: 429 LMKDLTERHLLGKTVASMYTVEFQKRGLPHAHILLFMAANSKLPTADDIDKIISAEIPNK 488
Query: 144 EKEECLAKLMQR---HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRRD 199
+KE L ++++ H PCG A SPCM G+C+K Y K ++ T + +GYP Y+RR
Sbjct: 489 DKEPELYEVIKNSMMHGPCGSANTSSPCMVDGQCSKLYPKKHQEITKVGADGYPIYRRRL 548
Query: 200 FGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQ 259
+++K V DNRYVVPYN L +RY H+NVE CN++ SIKYLFKY+NK PD+ +
Sbjct: 549 TDDYIEKGGVKCDNRYVVPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYINKGPDRVVFI 608
Query: 260 LS--------------VDGDNRDKSKPVDEIKQYYDC---------RYVSPCEAVWRIFA 296
+ V+ D +K K DEIK ++DC RYVS EA+WRIF
Sbjct: 609 VEPIKEATSSDTTAPVVESDTTEKKK--DEIKDWFDCSSYISFSPARYVSASEAIWRIFK 666
Query: 297 FDIHHK*PHVLKLLFHLHNEQ 317
F I H+ V KL FH +Q
Sbjct: 667 FPIQHRSTPVQKLSFHDKGKQ 687
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 243 bits (621), Expect = 7e-63
Identities = 145/361 (40%), Positives = 190/361 (52%), Gaps = 71/361 (19%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD I RL +IRK+Q +R + GL +A++ G+T A VG++I+L SF G Y
Sbjct: 441 VDALACIIQYRLDWIRKHQGNLRTELYAGLQDAIERGDTRAEQVGKRILLPSSFTGSPRY 500
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNL-KASDRPDIVCR------ 130
N QDAM+IC+ GYPDLFVT CN W EIQ+ L E + K SDRPDIV R
Sbjct: 501 KAQNYQDAMAICRWAGYPDLFVTFTCNAAWPEIQNMLDEIGVQKPSDRPDIVDRVFHIKL 560
Query: 131 ------------------------------------VFKMKLDKLMD------------- 141
+F K DK D
Sbjct: 561 RELMTDIKDKQYFGKTLAIIYTIEFQKRGLPHAHILIFLDKKDKCPDASEIDRIISAEIP 620
Query: 142 ----DFEKEECLAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYK 196
D E E + M H PCG A+ SPCM +C + + K + TT+DE G+P Y+
Sbjct: 621 DKEEDREGFEAVENFMM-HGPCGEAKSNSPCMIENKCIRNFPKKFHSETTVDEDGFPTYR 679
Query: 197 RRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKA 256
RRD G +++K V LDNRYVVPYN LL++Y H+NVE CN+S SIKYLFKY++K D+A
Sbjct: 680 RRDNGRYIEKGNVKLDNRYVVPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHKGDDQA 739
Query: 257 LMQLSVDGDNRDKSKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNE 316
+ D DEIK+Y +C Y+S +A WRIF F++H++ P V +L FHL NE
Sbjct: 740 TALIESDH---------DEIKKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENE 790
Query: 317 Q 317
Q
Sbjct: 791 Q 791
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 239 bits (609), Expect = 2e-61
Identities = 141/360 (39%), Positives = 187/360 (51%), Gaps = 64/360 (17%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD YT IE RL +IR+NQ +R + GL +A+ G+T +G++IVL SF GG
Sbjct: 483 VDAYTCIEQCRLSWIRQNQGILRTELYGGLQDALRTGDTRTEKLGRRIVLPASFTGGPRN 542
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFL-KERNLKASDRPDIVCRVFKMKL 136
N QDAM+IC+ G PDLFVT CN W EIQ L K + K S+RPDIV RVF +KL
Sbjct: 543 KEQNYQDAMAICRWAGNPDLFVTFTCNPKWPEIQCMLEKVGHQKPSERPDIVVRVFMIKL 602
Query: 137 DKLMDDFE---------------------------------KEECL-------------- 149
+LM D + KE+CL
Sbjct: 603 RELMSDIKRNQHFGKTKAIIFTIEFQKKGLPHAHILIFLDKKEKCLKPSQIDKMICAEIP 662
Query: 150 -----------AKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYKR 197
K H PCG P SPCM +C +Y+ K + T IDE +P YKR
Sbjct: 663 DSNKDPETFEAVKNFMMHGPCGETNPKSPCMVDHKCDRYFPKGFSDETIIDEVNFPIYKR 722
Query: 198 RDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKAL 257
RD G + K R+ L+N +VVPYN LL+++ H+NVE N+S SI+YLFK + D+A
Sbjct: 723 RDDGRQIKKGRINLNNGFVVPYNKDLLVKFQAHINVEWFNRSKSIRYLFKSIYNGDDQAT 782
Query: 258 MQLSVDGDNRDKSKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNEQ 317
+ + D +K DEIK+Y C Y++ EA WRIF F +H++ P V +L FH+ NEQ
Sbjct: 783 AVV----EETDTAKDNDEIKRYLGCSYMTATEACWRIFTFPLHYQEPSVQRLFFHVENEQ 838
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 238 bits (606), Expect = 4e-61
Identities = 140/372 (37%), Positives = 188/372 (49%), Gaps = 75/372 (20%)
Query: 19 DCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHYM 78
D YT IES RL YI+ Q +RC N L +A + G T + G ++++ S GG YM
Sbjct: 641 DAYTTIESNRLSYIKFKQSKLRCENYNSLKKASEAGTTSMNEEGNQVLIPSSLTGGPRYM 700
Query: 79 FNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVFKMKL-- 136
N DAM+ICK YG+PDLF+T CN W EI + R L DRPDIV R+FK+KL
Sbjct: 701 VQNYYDAMAICKHYGFPDLFITFTCNPKWPEITRHCQARGLSVDDRPDIVARIFKIKLDS 760
Query: 137 -------------------------------------------------DKLMD----DF 143
DK++ D
Sbjct: 761 LMKDLTDGKMLGKTVASMHTVEFQKRGLPHAHILLFMDAKSKLPTADDIDKIISAEIPDK 820
Query: 144 EKEECLAKLMQR---HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRRD 199
+KE L ++++ H PCG A SPCM G+C+K Y K ++ T + +GYP Y+RR
Sbjct: 821 DKEPELYEVIKNSMIHGPCGAANMNSPCMVEGKCSKQYPKKHQDITKVGKDGYPIYRRRM 880
Query: 200 FGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQ 259
+++K DN YVVPYN L +RY H+NVE CN+S SIKYLFKY+NK D+ +
Sbjct: 881 TEDYIEKGGFKCDNGYVVPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINKGADRVV-- 938
Query: 260 LSVDGDNRDKS--------------KPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PH 305
V+ N+DK+ K DEIK ++DCRYVS EAVWRI+ F + +
Sbjct: 939 FIVEPVNQDKTTENATSGEPPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPLQDRSTA 998
Query: 306 VLKLLFHLHNEQ 317
V +L FH +Q
Sbjct: 999 VQRLSFHDEGKQ 1010
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 236 bits (602), Expect = 1e-60
Identities = 138/371 (37%), Positives = 201/371 (53%), Gaps = 66/371 (17%)
Query: 8 YGNVVNSRRSVDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVL 67
YG + + + VD ++ IE+ RL+Y +QK +R ++G+ +A+D G TD +VG++++L
Sbjct: 363 YGRL-SDQIDVDIFSTIETNRLQYFIDHQKELRSESVDGIVDAIDKGVTDGDSVGKRVIL 421
Query: 68 QPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLK-ERNLKASDRPD 126
SF GGR YM N QDAM+IC+ YG PDLFVT C++ W+EI D ++ E + SDR D
Sbjct: 422 PASFTGGRRYMVMNYQDAMAICRVYGPPDLFVTYTCHSKWQEIADAIRFEPGQQPSDRAD 481
Query: 127 IVCRVFKMKLDKLMD------------------DFEKE-----ECLAKL----------- 152
I+ RVF MK+++ + +F+K CL L
Sbjct: 482 IIVRVFNMKVNEFITDIREGRTFGKVLAVLYTVEFQKRGLPHIHCLVWLAAATADVSASI 541
Query: 153 ------------------------MQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTI 188
H PC A PCM +C+K+Y K+++ T +
Sbjct: 542 IDGFICAEIPDYDTDRLGYELVSEFMMHGPCSDANKKCPCMKNDKCSKHYPKDFQDETIV 601
Query: 189 DE-GYPRYKRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFK 247
DE G+ Y+RR+ G + K +LLDNR VVPYN LL +Y H+NVE CNKSN IKYLFK
Sbjct: 602 DESGFTIYRRRNDGRSIMKNGILLDNRSVVPYNMALLKKYEAHINVEWCNKSNLIKYLFK 661
Query: 248 YVNKVPDKALMQLSVDGDNRDKS-----KPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK 302
Y+ K D+A + G ++ S P +EI +Y D R++S EA+ R+F FDIH++
Sbjct: 662 YITKGHDRARIYFETTGKTQNASPNHDLAPRNEILEYMDARFLSTYEALHRLFEFDIHYR 721
Query: 303 *PHVLKLLFHL 313
P V +L+ HL
Sbjct: 722 VPPVERLVVHL 732
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 229 bits (584), Expect = 1e-58
Identities = 133/363 (36%), Positives = 190/363 (51%), Gaps = 65/363 (17%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD Y+ +E RL++I +Q +R + G+ +A+D G +VG+K VL SF GGR Y
Sbjct: 336 VDAYSTVEGSRLKFIHDHQPELRSECVQGIVDAIDHGLESGDSVGKKYVLPSSFTGGRRY 395
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLK-ERNLKASDRPDIVCRVFKMKL 136
M N QDAM++C+ +G PDLFVT CN+ W+EI D L E SDR D++ RVF MK+
Sbjct: 396 MVQNYQDAMAVCRVFGSPDLFVTFTCNSKWQEIYDALVFEPGQVPSDRSDMIVRVFSMKV 455
Query: 137 DKLMDDFEKEECLAKLMQ------------RHVPC-----GGARPFS------------- 166
D+ + D ++ ++ H+ C FS
Sbjct: 456 DEFISDIKEGRTFGPVLAVLYTVEFQKRGLPHIHCIVWRAAADAEFSATAVDSLICAEIP 515
Query: 167 -------------------PCMDR---------GRCTKYYRKNYKGTTTIDE-GYPRYKR 197
PC D+ G C+K++ K ++ TT+DE G+ Y+R
Sbjct: 516 DVFSDPLGYALVDEFMIHGPCGDKNKSCVCMKNGHCSKHFPKGFQEETTMDEFGFTVYRR 575
Query: 198 RDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKAL 257
R+ G +V K + LDNR+VVPYN LL +Y H+NVE CNKSN IKYLFKY+ K D+
Sbjct: 576 RNDGRYVVKNGIKLDNRWVVPYNMKLLKKYQAHINVESCNKSNMIKYLFKYITKGGDRTK 635
Query: 258 MQLSVDGDNRDKS-----KPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFH 312
+ G+ +K+ P +EI +Y + R++S CEA WR F FDIH++ P V +L H
Sbjct: 636 LYFETTGNTPNKTVDGTVLPPNEIDEYINARFLSTCEAFWRAFEFDIHYRVPAVERLPIH 695
Query: 313 LHN 315
L N
Sbjct: 696 LPN 698
>UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana]
Length = 1250
Score = 219 bits (558), Expect = 1e-55
Identities = 138/393 (35%), Positives = 200/393 (50%), Gaps = 74/393 (18%)
Query: 1 MQDRVVDYGNVVNSRRS-----VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGE 55
+Q R+ + +V S+R VD YT IE +RLR+ R NQK +R N + +A+ G+
Sbjct: 196 IQTRLTEGMTIVRSKRLLHQYIVDAYTSIEQERLRWYRLNQKKLRADQYNNVKDAVARGD 255
Query: 56 TDASNVGQKIVLQPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVT--------------- 100
TDA ++G++++L S+ G YM DAM+IC+ YG PDLF+T
Sbjct: 256 TDAKSIGKRVILPASYTGSPRYMVEKYHDAMAICRWYGNPDLFITMTTNPKWEEISEHLK 315
Query: 101 ------------IACNTNWREIQDFLKERNLKASDRPDIVCRVFKMKLDK---------- 138
I C ++ + L + N K P + V+ ++ K
Sbjct: 316 TYGNDAANVRPDIECRVFKIKLDELLADFN-KGLFFPKPIAIVYTIEFQKRGLPHAHILL 374
Query: 139 -LMDDFEKE----------------------ECLAKLMQRHVPCGGARPFSPCMDRGRCT 175
L D +K L + H PCG RP SPCM++G C+
Sbjct: 375 WLQGDLKKPTPNDIDKYISAEIPVKDKDPEGHTLVEQHMMHGPCGKDRPSSPCMEKGICS 434
Query: 176 KYYRKNYKGTTTIDE-GYPRYKRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVE 234
K + + + T ++E G+ Y+RR+ +V K + LDNR+VVP+N +L +Y H+NVE
Sbjct: 435 KKFPREFVNHTKMNESGFILYRRRNDQRYVLKGQTRLDNRFVVPHNLEILKKYKAHINVE 494
Query: 235 CCNKSNSIKYLFKYVNKVPDKALMQL----SVDGD---NRDKSKPVDEIKQYYDCRYVSP 287
CNKS++IKYLFKY+ K DKA + SV+G N + KP +EI +Y DCRY+S
Sbjct: 495 WCNKSSAIKYLFKYITKGVDKATFIIQKGNSVNGQGSGNGFEEKPRNEINEYLDCRYLSA 554
Query: 288 CEAVWRIFAFDIHHK*PHVLKLLFHLHNEQGNI 320
CEA+WRIF F+IHH P V +L HL EQ I
Sbjct: 555 CEAMWRIFMFNIHHHNPPVQRLPLHLPGEQSTI 587
>UniRef100_Q851V4 Putative helicase [Oryza sativa]
Length = 1629
Score = 209 bits (533), Expect = 1e-52
Identities = 130/338 (38%), Positives = 179/338 (52%), Gaps = 39/338 (11%)
Query: 8 YGNVVNSRRSVDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVL 67
+G + + +VD Y +ES RL Y+ NQK IR + GL +++ GE+ AS VG++ VL
Sbjct: 594 HGGRLFQQFAVDMYIKVESSRLDYVWNNQKEIRADLYQGLMDSIQAGESRASAVGKRTVL 653
Query: 68 QPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDI 127
SF GG M DAM++ +KYG PD+F+T+ N W EI L E DRPD+
Sbjct: 654 PASFVGGGRNMKRRYMDAMALVQKYGKPDVFLTMTSNPKWDEITREL-EPVQTPQDRPDL 712
Query: 128 VCRVFKMKLDKLMDDFEKEECLAKL-------------------------MQRHVPCGGA 162
V RVF+ KL+ L ++ L K H PCG
Sbjct: 713 VVRVFRAKLEDLKKQLFEKHILGKTNMTASSRLSFQTRKKYPELYNMVVKHMMHGPCGRL 772
Query: 163 RPFSPCMDRGRCTKYYRKNYKGTTTI-DEGYPRYKRRDFGIHVDKQRVL----LDNRYVV 217
CM G+C Y + + TT+ + YP Y+RR+ G K RV+ LDNR+VV
Sbjct: 773 NGRCQCMRDGKCRNNYPREFNPTTSQGKDSYPLYRRRNDG----KSRVVRGHPLDNRWVV 828
Query: 218 PYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQLSVDGDNRDKSKPVDEIK 277
PYNP+LL Y H+NVE C+ ++KYLFKY K DKA S+ + +K+ +DEI+
Sbjct: 829 PYNPYLLRMYNCHMNVEVCSSIKAVKYLFKYHYKGHDKA----SISINEANKNGEIDEIQ 884
Query: 278 QYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHN 315
+Y D R+V P EA+W I+ FDI H P V +L HL N
Sbjct: 885 RYRDARWVIPPEALWCIYGFDICHISPSVRQLQLHLPN 922
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 182 bits (461), Expect = 2e-44
Identities = 85/164 (51%), Positives = 117/164 (70%), Gaps = 4/164 (2%)
Query: 156 HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRRDFGIHVDKQRVLLDNR 214
H PCG P SPCM+ G+C KY+ K+Y TT +D +G+P Y+RRD GI+V+K DNR
Sbjct: 112 HGPCGVGHPNSPCMENGKCKKYFPKSYSDTTKVDNDGFPVYRRRDTGIYVEKNGFQCDNR 171
Query: 215 YVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQLSVDGDNRDKS-KPV 273
YV+PYN + +RY H+NVE CN+S SIKYLFKYV+K D+ + ++V+ +++D + K
Sbjct: 172 YVIPYNEKVSLRYQAHINVELCNQSGSIKYLFKYVHKGHDR--VTVTVEPNDQDTAKKEK 229
Query: 274 DEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNEQ 317
DE+K Y+DCRYVS CEA+WRIF F IH++ V+KL FH +Q
Sbjct: 230 DEVKDYFDCRYVSACEAMWRIFKFPIHYRTTPVVKLFFHEEGKQ 273
>UniRef100_Q9ZPG4 F5K24.3 protein [Arabidopsis thaliana]
Length = 448
Score = 171 bits (432), Expect = 6e-41
Identities = 114/327 (34%), Positives = 161/327 (48%), Gaps = 60/327 (18%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
+D YT+IES RL YI+ NQ +R SF Y
Sbjct: 56 IDAYTIIESNRLAYIKFNQSKLR----------------------------SSFTSASRY 87
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNL-----------KASDRPD 126
M DAM+ICK YG+PD + ++ R+ L ERNL + R
Sbjct: 88 MLQTYYDAMAICKHYGFPDFLSRLRLDSLMRD----LTERNLLRKTVAFKFKKRGLPRAR 143
Query: 127 IVCRVFKMKLDKLMDDFEKEECLAKLMQR---HVPCGGARPFSPCMDRGRCTKYYRKNYK 183
I+ + + + DD E E+ L +L++ H CG A S CM G+C+K Y K ++
Sbjct: 144 ILLFMEANRKLPIADDIETEQELYELIKNSMIHGLCGSANTNSLCMVDGQCSKLYPKKHQ 203
Query: 184 GTTTID-EGYPRYKRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSI 242
T + +GYP Y+RR ++K V DN YVVPYN ++Y H+NVE CN++ SI
Sbjct: 204 ELTKVGADGYPVYRRRPIDGCIEKGGVKCDNMYVVPYN-RFSLKYQAHINVEWCNQNGSI 262
Query: 243 KYLFKYVNKVPDKA--LMQLSVDGDNRDKS----------KPVDEIKQYYDCRYVSPCEA 290
KYLFKY+NK PD+ +++L + N + + K D IK ++D YVS EA
Sbjct: 263 KYLFKYINKGPDRVVFIVELVKEETNSNTTTLGDETVTTKKKKDGIKDWFDYIYVSASEA 322
Query: 291 VWRIFAFDIHHK*PHVLKLLFHLHNEQ 317
+WRIF F I H+ V KL FH+ +Q
Sbjct: 323 IWRIFKFPIQHRSTPVQKLSFHVEGKQ 349
>UniRef100_Q7XMF3 OSJNBa0061G20.6 protein [Oryza sativa]
Length = 1410
Score = 166 bits (421), Expect = 1e-39
Identities = 109/293 (37%), Positives = 142/293 (48%), Gaps = 63/293 (21%)
Query: 86 MSICKKYGYPDLFVTIACNTNWREIQDFL-KERNLKASDRPD------------------ 126
M+IC+ G PDLFVT CN+ W+EI D L E SDR D
Sbjct: 1 MAICRVLGAPDLFVTFTCNSKWQEIYDALIYEPGQVPSDRSDMIVRVFNINEFISDIREG 60
Query: 127 -----IVCRVFKMKLDK----------------------LMDDFEKEE-----------C 148
I+ ++ ++ K ++D F E
Sbjct: 61 NTFGPILAVLYTVEFQKCGLPHIHCIVWLAAQNANFSAAVVDGFISAELPDVFADPLGYA 120
Query: 149 LAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYKRRDFGIHVDKQ 207
L + H PCG PCM +G C+K++ K ++ T +DE G YKRRD G +V K
Sbjct: 121 LVEEFMIHGPCGVDNKTCPCMKKGSCSKHFPKTFQAETILDEFGLTVYKRRDDGRYVIKN 180
Query: 208 RVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPD--KALMQLSVDGD 265
LDNR VVPYN LL +Y H+NVE CNKSN IKYLFKYV K D K + S
Sbjct: 181 GKKLDNRNVVPYNMKLLKKYQAHINVEWCNKSNMIKYLFKYVTKGSDCTKRYFETSTQTM 240
Query: 266 NRDKSK---PVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHN 315
NR S+ P +EI++Y D R++S CEA WR + FDIH++ P V +L HL N
Sbjct: 241 NRSNSENNAPRNEIQEYIDARFLSTCEAFWRAYEFDIHYRLPSVERLTVHLPN 293
>UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana]
Length = 1219
Score = 162 bits (409), Expect = 3e-38
Identities = 84/197 (42%), Positives = 119/197 (59%), Gaps = 18/197 (9%)
Query: 137 DKLMDDFEKEECLAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRY 195
DKL D E + K M H PCG P PCM+ G+C+K+Y K++ T ID EG+P Y
Sbjct: 305 DKLKDPELFE--VIKEMMVHGPCGVVNPKCPCMENGKCSKFYPKDHVPKTIIDKEGFPIY 362
Query: 196 KRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDK 255
+RR V K+ DNRYV+PYN L +RY H+NVE CN+S S+KY+FKY++K PD+
Sbjct: 363 RRRRIDDFVQKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHKGPDR 422
Query: 256 ALMQL--SVDGDNRDKS-------------KPVDEIKQYYDCRYVSPCEAVWRIFAFDIH 300
+ + S++ N++K K +E++ +++CRYVS CEA WRI + IH
Sbjct: 423 VTVVVGSSLNSKNKEKGKQKVNADTDGSEPKKKNEVEDFFNCRYVSACEAAWRILKYPIH 482
Query: 301 HK*PHVLKLLFHLHNEQ 317
++ V+KL FHL EQ
Sbjct: 483 YRSTSVMKLSFHLPGEQ 499
Score = 128 bits (321), Expect = 4e-28
Identities = 61/134 (45%), Positives = 80/134 (59%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD Y IES RL YI+ NQ ++R N + +A + G+ D G L +F GG Y
Sbjct: 126 VDSYIAIESNRLGYIKLNQSSLRADNYNSVQKASEEGKCDLKYQGLACYLPATFTGGPRY 185
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDRPDIVCRVFKMKLD 137
M N DAM++CK +G+PD F+T CN W E+ F ERNL+ DRP+I+C++FKMKLD
Sbjct: 186 MRNMYLDAMAVCKHFGFPDYFITFTCNPKWPELIRFCGERNLRVDDRPEIICKIFKMKLD 245
Query: 138 KLMDDFEKEECLAK 151
LM D K L K
Sbjct: 246 SLMLDLTKRNILGK 259
>UniRef100_Q9FHV5 Helicase [Arabidopsis thaliana]
Length = 1523
Score = 153 bits (386), Expect = 1e-35
Identities = 91/285 (31%), Positives = 145/285 (49%), Gaps = 32/285 (11%)
Query: 18 VDCYTMIESQRLRYIRKNQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRHY 77
VD YT IES L YI+ NQ ++R N + +A + G++D + G L +F GG Y
Sbjct: 557 VDSYTAIESNWLGYIKLNQTSLRAYNYNSVQKASEEGKSDLKDQGLACYLPATFTGGPRY 616
Query: 78 MFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLKERNLKASDR--PDIVCRVFKMK 135
M N DAM++CK +G+PD F+T CN W E+ F ++ R P ++
Sbjct: 617 MRNMYLDAMALCKHFGFPDYFITFTCNPKWPELIRFCAMYTIEFQKRGLPHAHILIWLDS 676
Query: 136 LDKLMDDFEKEECLAKLMQRHVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDEGYPRY 195
KL K E + K++ +P D+ + + + + G+P Y
Sbjct: 677 KSKL----TKAEHIDKVISAEIP-----------DKLKDPELFEVIKESMVHGPCGFPIY 721
Query: 196 KRRDFGIHVDKQRVLLDNRYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDK 255
+RR V+K+ DNRYV+PYN L +RY H+NVE CN+S S+KY+FKY++K PD+
Sbjct: 722 RRRRTDDFVEKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHKGPDR 781
Query: 256 --ALMQLSVDGDNRDKSKPVD-------------EIKQYYDCRYV 285
+++ S++ N++ K D E++ Y++CR +
Sbjct: 782 VTVVVESSLNSKNKENGKQKDNADTDGSETKKKNEVEDYFNCRRI 826
>UniRef100_Q8S7J2 Hypothetical protein OSJNBa0095C06.16 [Oryza sativa]
Length = 1414
Score = 147 bits (372), Expect = 5e-34
Identities = 76/166 (45%), Positives = 105/166 (62%), Gaps = 6/166 (3%)
Query: 156 HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYKRRDFGIHVDKQRVLLDNR 214
H PCG PCM +G C+K++ K+++ T +DE G+ YKRR+ G +V K + L NR
Sbjct: 58 HGPCGNQNRACPCMKKGECSKHFPKSFQEETMMDEFGFTIYKRRNNGRYVVKNGIKLYNR 117
Query: 215 YVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPD--KALMQLSVDGDNR---DK 269
+VVPYN LL +Y H+NVE CNKSN IKYLFKYV K D KA ++S + N+
Sbjct: 118 WVVPYNLELLKKYQAHINVEWCNKSNMIKYLFKYVTKGADRTKAFFEISGNASNKTSESS 177
Query: 270 SKPVDEIKQYYDCRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHN 315
+ P +EI++ D R++S CE+ WR F DIH++ P V +L HL N
Sbjct: 178 TSPRNEIQECIDARFLSTCESHWRAFELDIHYRMPSVERLTVHLPN 223
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.342 0.150 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 780,031,471
Number of Sequences: 2790947
Number of extensions: 32175129
Number of successful extensions: 90914
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 90581
Number of HSP's gapped (non-prelim): 164
length of query: 489
length of database: 848,049,833
effective HSP length: 131
effective length of query: 358
effective length of database: 482,435,776
effective search space: 172712007808
effective search space used: 172712007808
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 77 (34.3 bits)
Medicago: description of AC125481.7


