

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.1 - phase: 0 /pseudo
(250 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
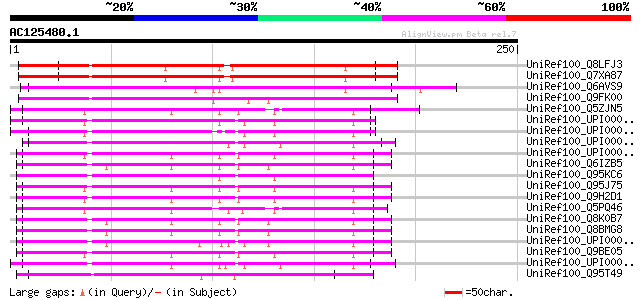
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LFJ3 Contains similarity to peroxisomal membrane car... 210 2e-53
UniRef100_Q7XA87 At5g66380 [Arabidopsis thaliana] 207 2e-52
UniRef100_Q6AVS9 Mitochondrial carrier protein-like protein [Ory... 188 1e-46
UniRef100_Q9FK00 Similarity to peroxisomal membrane carrier prot... 150 3e-35
UniRef100_Q5ZJN5 Hypothetical protein [Gallus gallus] 114 3e-24
UniRef100_UPI00003AC016 UPI00003AC016 UniRef100 entry 113 4e-24
UniRef100_UPI00003AC018 UPI00003AC018 UniRef100 entry 112 7e-24
UniRef100_UPI0000436FA8 UPI0000436FA8 UniRef100 entry 108 1e-22
UniRef100_UPI000036E3DB UPI000036E3DB UniRef100 entry 108 2e-22
UniRef100_Q6IZB5 Mitochondrial folate transporter [Cricetulus gr... 108 2e-22
UniRef100_Q95KC6 Hypothetical protein [Macaca fascicularis] 108 2e-22
UniRef100_Q95J75 Mitochondrial folate transporter/carrier [Macac... 108 2e-22
UniRef100_Q9H2D1 Mitochondrial folate transporter/carrier [Homo ... 108 2e-22
UniRef100_Q5PQ46 Hypothetical protein [Xenopus laevis] 107 2e-22
UniRef100_Q8K0B7 Mitochondrial folate transporter/carrier [Mus m... 107 2e-22
UniRef100_Q8BMG8 Mitochondrial folate transporter/carrier [Mus m... 107 2e-22
UniRef100_UPI00001814D5 UPI00001814D5 UniRef100 entry 107 3e-22
UniRef100_Q9BE05 Hypothetical protein [Macaca fascicularis] 107 3e-22
UniRef100_UPI000027C671 UPI000027C671 UniRef100 entry 107 4e-22
UniRef100_Q95T49 GH22139p [Drosophila melanogaster] 106 5e-22
>UniRef100_Q8LFJ3 Contains similarity to peroxisomal membrane carrier protein
[Arabidopsis thaliana]
Length = 308
Score = 210 bits (535), Expect = 2e-53
Identities = 109/200 (54%), Positives = 135/200 (67%), Gaps = 16/200 (8%)
Query: 5 QWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLK 64
QWE A GFA + +PLDVV+TR QV+DGR S P Y NT HA+FTIAR EGL+
Sbjct: 6 QWENATAGAVAGFATVAAMHPLDVVRTRFQVNDGR-GSSLPTYKNTAHAVFTIARLEGLR 64
Query: 65 GLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-------------SGALMTVCL 111
GLYAGF ++G+T+SW L F+YG K R+AR + ++ +GAL VCL
Sbjct: 65 GLYAGFFPAVIGSTVSWGLYFFFYGRAKQRYARGRDDEKLSPALHLASAAEAGAL--VCL 122
Query: 112 CANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQ 171
C NP ++VKTRLQLQTPLH +PYSGL DAFRTI +EEG A Y+GIVPG L+S AIQ
Sbjct: 123 CTNPIWLVKTRLQLQTPLHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQ 182
Query: 172 FIVYEQLRKTVVNLKTKGSK 191
F YE+LRK +V+LK + K
Sbjct: 183 FTAYEELRKIIVDLKERRRK 202
Score = 47.4 bits (111), Expect = 3e-04
Identities = 48/175 (27%), Positives = 70/175 (39%), Gaps = 21/175 (12%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL----GATIS 80
P+ +V+TRLQ+ Q Y+ A TI ++EG + LY G GL+ GA
Sbjct: 126 PIWLVKTRLQLQTP--LHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQF 183
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMT--------------VCLCANPAFVVKTRLQLQ 126
+ IV + R K E + L+ L P V++ RLQ +
Sbjct: 184 TAYEELRKIIVDLKERRRKSESTDNLLNSADYAALGGSSKVAAVLLTYPFQVIRARLQQR 243
Query: 127 TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRK 180
+ Y R R EG FYRG+ + ++I FIVYE + K
Sbjct: 244 PSTNGIPRYIDSLHVVRETARYEGLRGFYRGLTANLLKNVPASSITFIVYENVLK 298
Score = 42.7 bits (99), Expect = 0.009
Identities = 25/82 (30%), Positives = 38/82 (45%), Gaps = 2/82 (2%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
+Y ++ A + YP V++ RLQ + PRY ++ H + AR EGL+G
Sbjct: 214 DYAALGGSSKVAAVLLTYPFQVIRARLQQRPST--NGIPRYIDSLHVVRETARYEGLRGF 271
Query: 67 YAGFPAGLLGATISWSLLVFYY 88
Y G A LL + S+ Y
Sbjct: 272 YRGLTANLLKNVPASSITFIVY 293
>UniRef100_Q7XA87 At5g66380 [Arabidopsis thaliana]
Length = 308
Score = 207 bits (527), Expect = 2e-52
Identities = 108/200 (54%), Positives = 134/200 (67%), Gaps = 16/200 (8%)
Query: 5 QWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLK 64
QWE A GFA + + LDVV+TR QV+DGR S P Y NT HA+FTIAR EGL+
Sbjct: 6 QWENATAGAVAGFATVAAMHSLDVVRTRFQVNDGR-GSSLPTYKNTAHAVFTIARLEGLR 64
Query: 65 GLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-------------SGALMTVCL 111
GLYAGF ++G+T+SW L F+YG K R+AR + ++ +GAL VCL
Sbjct: 65 GLYAGFFPAVIGSTVSWGLYFFFYGRAKQRYARGRDDEKLSPALHLASAAEAGAL--VCL 122
Query: 112 CANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQ 171
C NP ++VKTRLQLQTPLH +PYSGL DAFRTI +EEG A Y+GIVPG L+S AIQ
Sbjct: 123 CTNPIWLVKTRLQLQTPLHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQ 182
Query: 172 FIVYEQLRKTVVNLKTKGSK 191
F YE+LRK +V+LK + K
Sbjct: 183 FTAYEELRKIIVDLKERRRK 202
Score = 47.8 bits (112), Expect = 3e-04
Identities = 48/175 (27%), Positives = 70/175 (39%), Gaps = 21/175 (12%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL----GATIS 80
P+ +V+TRLQ+ Q Y+ A TI ++EG + LY G GL+ GA
Sbjct: 126 PIWLVKTRLQLQTP--LHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQF 183
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMT--------------VCLCANPAFVVKTRLQLQ 126
+ IV + R K E + L+ L P V++ RLQ +
Sbjct: 184 TAYEELRKIIVDLKERRRKSESTDNLLNSADYAALGGSSKVAAVLLTYPFQVIRARLQQR 243
Query: 127 TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRK 180
+ Y R R EG FYRG+ + ++I FIVYE + K
Sbjct: 244 PSTNGIPRYIDSLHVIRETARYEGLRGFYRGLTANLLKNVPASSITFIVYENVLK 298
Score = 43.1 bits (100), Expect = 0.007
Identities = 26/82 (31%), Positives = 38/82 (45%), Gaps = 2/82 (2%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
+Y ++ A + YP V++ RLQ + PRY ++ H I AR EGL+G
Sbjct: 214 DYAALGGSSKVAAVLLTYPFQVIRARLQQRPST--NGIPRYIDSLHVIRETARYEGLRGF 271
Query: 67 YAGFPAGLLGATISWSLLVFYY 88
Y G A LL + S+ Y
Sbjct: 272 YRGLTANLLKNVPASSITFIVY 293
>UniRef100_Q6AVS9 Mitochondrial carrier protein-like protein [Oryza sativa]
Length = 318
Score = 188 bits (477), Expect = 1e-46
Identities = 101/227 (44%), Positives = 134/227 (58%), Gaps = 12/227 (5%)
Query: 6 WEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
WE A GFA + +PLDVV+TR QV GR P Y NT HA++TIAR EGL+G
Sbjct: 16 WENAAAGAAAGFATVATLHPLDVVRTRFQVSGGRGCYDLPPYRNTAHAVYTIARSEGLRG 75
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVKDRHARSK----------VEKSGALMTVCLCANP 115
LYAGF +LG+T+SW L F+Y K R+ + K V + A VCL NP
Sbjct: 76 LYAGFYPAVLGSTVSWGLYFFFYNRAKQRYLQGKDDQLRPVHHLVSAAEAGALVCLFTNP 135
Query: 116 AFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVY 175
++VKTRLQLQTP HH YSG DA RTI +EEG+ A YRGI PG L++ AIQF Y
Sbjct: 136 IWLVKTRLQLQTPSHHTSRYSGFSDALRTILKEEGWLALYRGIGPGLLLVTHGAIQFTAY 195
Query: 176 EQLRKTVVNLKTKGSKIQHQKPDQIL--VCYWTLDLFTSEISLVLDY 220
E+LRK ++ K++ ++ ++ D L + Y L + +++L Y
Sbjct: 196 EELRKALIFAKSRQTRTDNRSCDDSLNSIDYAALGAGSKVTAILLTY 242
Score = 55.5 bits (132), Expect = 1e-06
Identities = 52/200 (26%), Positives = 79/200 (39%), Gaps = 23/200 (11%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
V++ G F P+ +V+TRLQ+ + RY+ + A+ TI ++EG LY G
Sbjct: 120 VSAAEAGALVCLFTNPIWLVKTRLQLQTPSHHTS--RYSGFSDALRTILKEEGWLALYRG 177
Query: 70 FPAGLLGATISWSLLVFYYGI------VKDRHARSKVEK--------------SGALMTV 109
GLL T Y + K R R+ +G+ +T
Sbjct: 178 IGPGLLLVTHGAIQFTAYEELRKALIFAKSRQTRTDNRSCDDSLNSIDYAALGAGSKVTA 237
Query: 110 CLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQA 168
L P V++ RLQ + Y + + R EG FYRGI + A
Sbjct: 238 ILLTYPYQVIRARLQQRPGSDGTPKYKDSWHVVKETARHEGVRGFYRGITSNLLKNLPAA 297
Query: 169 AIQFIVYEQLRKTVVNLKTK 188
++ F+VYE + K K K
Sbjct: 298 SLTFVVYENVIKLFKAAKEK 317
>UniRef100_Q9FK00 Similarity to peroxisomal membrane carrier protein [Arabidopsis
thaliana]
Length = 310
Score = 150 bits (379), Expect = 3e-35
Identities = 89/200 (44%), Positives = 115/200 (57%), Gaps = 14/200 (7%)
Query: 5 QWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLK 64
QWE A GFA + + LDVV+TR QV+DGR S P Y NT HA+FTIAR EGL+
Sbjct: 6 QWENATAGAVAGFATVAAMHSLDVVRTRFQVNDGR-GSSLPTYKNTAHAVFTIARLEGLR 64
Query: 65 GLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSK-VEKSGALMTVCLCANPA------- 116
GLYAGF ++G+T+SW L F+YG K R+AR + EK + + A
Sbjct: 65 GLYAGFFPAVIGSTVSWGLYFFFYGRAKQRYARGRDDEKLSPALHLASAAEAGALGMMLS 124
Query: 117 --FVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQ 171
F+ K+ L Q T + S + A RTI +EEG A Y+GIVPG L+S AIQ
Sbjct: 125 GLFMHKSYLACQNKVTASDTSSSNSTILRAIRTIVKEEGPRALYKGIVPGLVLVSHGAIQ 184
Query: 172 FIVYEQLRKTVVNLKTKGSK 191
F YE+LRK +V+LK + K
Sbjct: 185 FTAYEELRKIIVDLKERRRK 204
Score = 43.1 bits (100), Expect = 0.007
Identities = 26/82 (31%), Positives = 38/82 (45%), Gaps = 2/82 (2%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
+Y ++ A + YP V++ RLQ + PRY ++ H I AR EGL+G
Sbjct: 216 DYAALGGSSKVAAVLLTYPFQVIRARLQQRPST--NGIPRYIDSLHVIRETARYEGLRGF 273
Query: 67 YAGFPAGLLGATISWSLLVFYY 88
Y G A LL + S+ Y
Sbjct: 274 YRGLTANLLKNVPASSITFIVY 295
Score = 42.0 bits (97), Expect = 0.015
Identities = 41/147 (27%), Positives = 58/147 (38%), Gaps = 19/147 (12%)
Query: 53 AIFTIARKEGLKGLYAGFPAGLL----GATISWSLLVFYYGIVKDRHARSKVEKSGALMT 108
AI TI ++EG + LY G GL+ GA + IV + R K E + L+
Sbjct: 154 AIRTIVKEEGPRALYKGIVPGLVLVSHGAIQFTAYEELRKIIVDLKERRRKSESTDNLLN 213
Query: 109 --------------VCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAF 154
L P V++ RLQ + + Y R R EG F
Sbjct: 214 SADYAALGGSSKVAAVLLTYPFQVIRARLQQRPSTNGIPRYIDSLHVIRETARYEGLRGF 273
Query: 155 YRGIVPGFFL-ISQAAIQFIVYEQLRK 180
YRG+ + ++I FIVYE + K
Sbjct: 274 YRGLTANLLKNVPASSITFIVYENVLK 300
>UniRef100_Q5ZJN5 Hypothetical protein [Gallus gallus]
Length = 322
Score = 114 bits (284), Expect = 3e-24
Identities = 70/216 (32%), Positives = 104/216 (47%), Gaps = 17/216 (7%)
Query: 1 MNELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARK 60
+ +Q E A L+ G +PLD+V+ R V DG P+YN H + T+ ++
Sbjct: 25 LRHVQLENLAAGLSGGVVSTLVLHPLDLVKIRFAVSDG--LELRPKYNGILHCMTTVWKR 82
Query: 61 EGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTV 109
EGL+GLY G ++GA SW L F+Y +K K+E MT+
Sbjct: 83 EGLRGLYQGVTPNMVGAGASWGLYFFFYNAIKAYKKEGKLESLTATEHLVSAAEAGAMTL 142
Query: 110 CLCANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS 166
C+ NP +V KTRL LQ R Y+G+ DA I + EG Y+G VPG F S
Sbjct: 143 CI-TNPIWVTKTRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRGLYKGFVPGLFGTS 201
Query: 167 QAAIQFIVYEQLRKTVVNLKTKGSKIQHQKPDQILV 202
A+QF+ YE L++ + + S + + I++
Sbjct: 202 HGALQFMAYEDLKQRYNKYRNRVSDTKLNTAEYIMM 237
Score = 71.2 bits (173), Expect = 2e-11
Identities = 56/187 (29%), Positives = 91/187 (47%), Gaps = 20/187 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
E+ V++ G + P+ V +TRL + +D + +Y + A+ I + EG++G
Sbjct: 129 EHLVSAAEAGAMTLCITNPIWVTKTRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRG 188
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ L Y ++R + +K+ + +M +
Sbjct: 189 LYKGFVPGLFGTSHGALQFMAYEDLKQRYNKYRNRVSDTKLNTAEYIMMAAVSKIFAVTA 248
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q H R YSG+ D R R+EG FY+GIVP ++ A I
Sbjct: 249 TYPYQVVRARLQDQ----HNR-YSGVLDVIRRTWRKEGIHGFYKGIVPNVIRVTPACCIT 303
Query: 172 FIVYEQL 178
F+VYE +
Sbjct: 304 FVVYENV 310
>UniRef100_UPI00003AC016 UPI00003AC016 UniRef100 entry
Length = 292
Score = 113 bits (283), Expect = 4e-24
Identities = 68/194 (35%), Positives = 96/194 (49%), Gaps = 17/194 (8%)
Query: 1 MNELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARK 60
+ +Q E A L+ G +PLD+V+ R V DG P+YN H + T+ ++
Sbjct: 3 LRHVQLENLAAGLSGGVVSTLVLHPLDLVKIRFAVSDG--LELRPKYNGILHCMATVWKR 60
Query: 61 EGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTV 109
EGL+GLY G ++GA SW L F+Y +K K+E MT+
Sbjct: 61 EGLRGLYQGVTPNMVGAGASWGLYFFFYNAIKAYKKEGKLESLTATEHLVSAAEAGAMTL 120
Query: 110 CLCANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS 166
C+ NP +V KTRL LQ R Y+G+ DA I + EG Y+G VPG F S
Sbjct: 121 CI-TNPIWVTKTRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRGLYKGFVPGLFGTS 179
Query: 167 QAAIQFIVYEQLRK 180
A+QF+ YE L++
Sbjct: 180 HGALQFMAYEDLKQ 193
Score = 72.0 bits (175), Expect = 1e-11
Identities = 56/182 (30%), Positives = 88/182 (47%), Gaps = 16/182 (8%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
E+ V++ G + P+ V +TRL + +D + +Y + A+ I + EG++G
Sbjct: 107 EHLVSAAEAGAMTLCITNPIWVTKTRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRG 166
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN--------PAF 117
LY GF GL G T +L Y +K R+ + + + + + + P
Sbjct: 167 LYKGFVPGLFG-TSHGALQFMAYEDLKQRYNKYRNRNTAEYIMMAAVSKIFAVTATYPYQ 225
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQFIVYE 176
VV+ RLQ Q H R YSG+ D R R+EG FY+GIVP ++ A I F+VYE
Sbjct: 226 VVRARLQDQ----HNR-YSGVLDVIRRTWRKEGIHGFYKGIVPNVIRVTPACCITFVVYE 280
Query: 177 QL 178
+
Sbjct: 281 NV 282
>UniRef100_UPI00003AC018 UPI00003AC018 UniRef100 entry
Length = 269
Score = 112 bits (281), Expect = 7e-24
Identities = 70/183 (38%), Positives = 98/183 (53%), Gaps = 9/183 (4%)
Query: 1 MNELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARK 60
+ +Q E A L+ G +PLD+V+ R V DG P+YN H + T+ ++
Sbjct: 3 LRHVQLENLAAGLSGGVVSTLVLHPLDLVKIRFAVSDG--LELRPKYNGILHCMATVWKR 60
Query: 61 EGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCANPAFVVK 120
EGL+GLY G ++GA SW L F++G+ K S SGA MT+C+ NP +V K
Sbjct: 61 EGLRGLYQGVTPNMVGAGASWGLYFFFFGMGKTISLTSIC--SGA-MTLCI-TNPIWVTK 116
Query: 121 TRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQ 177
TRL LQ R Y+G+ DA I + EG Y+G VPG F S A+QF+ YE
Sbjct: 117 TRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRGLYKGFVPGLFGTSHGALQFMAYED 176
Query: 178 LRK 180
L++
Sbjct: 177 LKQ 179
Score = 71.6 bits (174), Expect = 2e-11
Identities = 55/179 (30%), Positives = 87/179 (47%), Gaps = 16/179 (8%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYA 68
+ S+ +G + P+ V +TRL + +D + +Y + A+ I + EG++GLY
Sbjct: 96 LTSICSGAMTLCITNPIWVTKTRLVLQYDAGVDPSKRQYTGMSDALIKIYKTEGIRGLYK 155
Query: 69 GFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN--------PAFVVK 120
GF GL G T +L Y +K R+ + + + + + + P VV+
Sbjct: 156 GFVPGLFG-TSHGALQFMAYEDLKQRYNKYRNRNTAEYIMMAAVSKIFAVTATYPYQVVR 214
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQFIVYEQL 178
RLQ Q H R YSG+ D R R+EG FY+GIVP ++ A I F+VYE +
Sbjct: 215 ARLQDQ----HNR-YSGVLDVIRRTWRKEGIHGFYKGIVPNVIRVTPACCITFVVYENV 268
Score = 36.2 bits (82), Expect = 0.80
Identities = 22/72 (30%), Positives = 32/72 (43%), Gaps = 7/72 (9%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
EY + + + +T YP VV+ RLQ Q+ RY+ I RKEG+ G
Sbjct: 191 EYIMMAAVSKIFAVTATYPYQVVRARLQ-------DQHNRYSGVLDVIRRTWRKEGIHGF 243
Query: 67 YAGFPAGLLGAT 78
Y G ++ T
Sbjct: 244 YKGIVPNVIRVT 255
>UniRef100_UPI0000436FA8 UPI0000436FA8 UniRef100 entry
Length = 302
Score = 108 bits (271), Expect = 1e-22
Identities = 68/193 (35%), Positives = 97/193 (50%), Gaps = 15/193 (7%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A L+ G +PLD+V+ R V DG P+Y+ H + +I +EG +GLY G
Sbjct: 23 IAGLSGGVLSTLALHPLDLVKIRFAVSDG--LDVRPKYSGIVHCMKSIWHQEGFRGLYQG 80
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL-----------MTVCLCANPAFV 118
+ GA SW L F+Y +K + ++ + A MT+CL NP +V
Sbjct: 81 VTPNIWGAGASWGLYFFFYNAIKGYNKETRQIELTATEHLLSAAVAGAMTLCL-TNPIWV 139
Query: 119 VKTRLQLQTPLHHA-RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQ 177
KTRL LQ + + Y G+ DA I R EG S YRG VPG F S A+QF+ YE+
Sbjct: 140 TKTRLVLQYSADPSQKQYKGMMDALVKIYRHEGISGLYRGFVPGLFGTSHGALQFMAYEE 199
Query: 178 LRKTVVNLKTKGS 190
L++ + K S
Sbjct: 200 LKRDYNKYRKKQS 212
Score = 62.4 bits (150), Expect = 1e-08
Identities = 54/191 (28%), Positives = 86/191 (44%), Gaps = 20/191 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
E+ +++ G + P+ V +TRL + SQ +Y A+ I R EG+ GL
Sbjct: 118 EHLLSAAVAGAMTLCLTNPIWVTKTRLVLQYSADPSQ-KQYKGMMDALVKIYRHEGISGL 176
Query: 67 YAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL-----MTVCLCAN------- 114
Y GF GL G + + Y + +D + K + L +T+ +
Sbjct: 177 YRGFVPGLFGTSHGALQFMAYEELKRDYNKYRKKQSDAKLNPLEYITMAALSKIFAVATT 236
Query: 115 -PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQF 172
P VV+ RLQ Q H+ Y+GL D R EG FY+G+VP ++ A I F
Sbjct: 237 YPYQVVRARLQDQ---HNT--YNGLTDVVWRTWRNEGLLGFYKGMVPNLIRVTPACCITF 291
Query: 173 IVYEQLRKTVV 183
+VYE + + ++
Sbjct: 292 VVYENVSRVLL 302
>UniRef100_UPI000036E3DB UPI000036E3DB UniRef100 entry
Length = 317
Score = 108 bits (269), Expect = 2e-22
Identities = 64/190 (33%), Positives = 94/190 (48%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E +A ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 23 VRYENLIAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 80
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 81 RGLYQGVTPNIWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 139
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 140 TNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 199
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 200 LQFMAYELLK 209
Score = 70.1 bits (170), Expect = 5e-11
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 124 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRG 183
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 184 LYKGFVPGLFGTSHGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 243
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 244 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGVGGFYKGIAPNLIRVTPACCIT 298
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 299 FVVYENVSHFLLDLREK 315
>UniRef100_Q6IZB5 Mitochondrial folate transporter [Cricetulus griseus]
Length = 316
Score = 108 bits (269), Expect = 2e-22
Identities = 66/190 (34%), Positives = 94/190 (48%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+Y H + TI + EGL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LEVRPKYKGILHCLTTIWKVEGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAEQLEPLEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++DA I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYGGVVNPSQRQYKGMFDALVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 67.4 bits (163), Expect = 3e-10
Identities = 57/197 (28%), Positives = 87/197 (43%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + G + + R Y A+ I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYGGVVNPSQRQYKGMFDALVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLEYNKHINRLPEAQLSTPEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q H Y G+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQ----HVS-YGGVMDVIVKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + + L+ K
Sbjct: 297 FVVYENVSHFLCGLREK 313
>UniRef100_Q95KC6 Hypothetical protein [Macaca fascicularis]
Length = 303
Score = 108 bits (269), Expect = 2e-22
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
>UniRef100_Q95J75 Mitochondrial folate transporter/carrier [Macaca fascicularis]
Length = 315
Score = 108 bits (269), Expect = 2e-22
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.9 bits (172), Expect = 3e-11
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 297 FVVYENVSHFLLDLREK 313
>UniRef100_Q9H2D1 Mitochondrial folate transporter/carrier [Homo sapiens]
Length = 315
Score = 108 bits (269), Expect = 2e-22
Identities = 64/190 (33%), Positives = 94/190 (48%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E +A ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLIAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNIWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.1 bits (170), Expect = 5e-11
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGVGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 297 FVVYENVSHFLLDLREK 313
>UniRef100_Q5PQ46 Hypothetical protein [Xenopus laevis]
Length = 318
Score = 107 bits (268), Expect = 2e-22
Identities = 66/190 (34%), Positives = 93/190 (48%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA L+ G +PLD+V+ R V DG P+Y H + T+ ++EGL
Sbjct: 24 VRYENLVAGLSGGVISTLVLHPLDLVKIRFAVSDG--LELRPKYRGIVHCLATVWQREGL 81
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL-----------MTVCLC 112
+GLY G + GA SW L F+Y VK + E A+ +T+C
Sbjct: 82 RGLYQGVTPNMWGAGASWGLYFFFYNAVKAYKKEGRAEDLSAVEHLLSAAGAGALTLCF- 140
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++ A I R EG Y+G VPG S A
Sbjct: 141 TNPIWVTKTRLVLQYDAGIDSSKRQYRGMFHALGKIYRNEGIPGLYKGFVPGLLGTSHGA 200
Query: 170 IQFIVYEQLR 179
+QF+ YE+L+
Sbjct: 201 LQFMAYEELK 210
Score = 73.2 bits (178), Expect = 6e-12
Identities = 60/198 (30%), Positives = 97/198 (48%), Gaps = 26/198 (13%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
E+ +++ G + F P+ V +TRL + +D + S +Y HA+ I R EG+ G
Sbjct: 125 EHLLSAAGAGALTLCFTNPIWVTKTRLVLQYDAGIDSSKRQYRGMFHALGKIYRNEGIPG 184
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCA------ 113
LY GF GLLG + +++ L Y +R + +K+ G L + + A
Sbjct: 185 LYKGFVPGLLGTSHGALQFMAYEELKMEYNKYLNRPSDTKL---GTLEYITMAALSKIFA 241
Query: 114 ----NPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA- 168
P VV+ RLQ Q H R Y+G+ D R+EG FY+GIVP ++ A
Sbjct: 242 VSTTYPYQVVRARLQDQ----HNR-YTGVLDVISRTWRKEGVQGFYKGIVPNIIRVTPAC 296
Query: 169 AIQFIVYEQLRKTVVNLK 186
I F+VYE++ +++ +
Sbjct: 297 CITFVVYEKVSHFLLDFR 314
>UniRef100_Q8K0B7 Mitochondrial folate transporter/carrier [Mus musculus]
Length = 316
Score = 107 bits (268), Expect = 2e-22
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+Y H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LEVRPKYKGILHCLATIWKVDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAEQLEPLEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++DA I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.1 bits (170), Expect = 5e-11
Identities = 58/197 (29%), Positives = 89/197 (44%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + G + S R Y A+ I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNKHINRLPEAQLSTAEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q H Y G+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQ----HVS-YGGVTDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + + +L+ K
Sbjct: 297 FVVYENVSHLLYDLREK 313
>UniRef100_Q8BMG8 Mitochondrial folate transporter/carrier [Mus musculus]
Length = 316
Score = 107 bits (268), Expect = 2e-22
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+Y H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LEVRPKYKGILHCLATIWKVDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAEQLEPLEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++DA I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 69.7 bits (169), Expect = 7e-11
Identities = 58/197 (29%), Positives = 89/197 (44%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + G + S R Y A+ I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNKHINRLPEAQLSTAEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q H Y G+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQ----HVS-YGGVTDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + + +L+ K
Sbjct: 297 FVVYENVSHFLYDLREK 313
>UniRef100_UPI00001814D5 UPI00001814D5 UniRef100 entry
Length = 316
Score = 107 bits (267), Expect = 3e-22
Identities = 67/190 (35%), Positives = 95/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+Y H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LEVRPKYKGILHCLATIWKVDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL-----------MTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ AL MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAEQLEALEYLISAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G+ DA I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYGGVVNPSQRQYKGMIDALVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYEVLK 207
Score = 67.4 bits (163), Expect = 3e-10
Identities = 60/198 (30%), Positives = 89/198 (44%), Gaps = 22/198 (11%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKG 65
EY +++ G + PL V +TRL + G + + R Y A+ I + EG++G
Sbjct: 122 EYLISAAEAGAMTLCITNPLWVTKTRLMLQYGGVVNPSQRQYKGMIDALVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVK---DRHARSKVEKS---------GALMTVCLCA 113
LY GF GL G T +L Y ++K ++H E AL + A
Sbjct: 182 LYKGFVPGLFG-TSHGALQFMAYEVLKLKYNKHINKLPEAQLSTAEYISVAALSKIFAVA 240
Query: 114 N--PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AI 170
P VV+ RLQ Q H Y G+ D R+EG FY+GI P ++ A I
Sbjct: 241 ATYPYQVVRARLQDQ----HVS-YGGVTDVITKTWRKEGIGGFYKGIAPNLIRVTPACCI 295
Query: 171 QFIVYEQLRKTVVNLKTK 188
F+VYE + + +L+ K
Sbjct: 296 TFVVYENVSHFLYDLREK 313
>UniRef100_Q9BE05 Hypothetical protein [Macaca fascicularis]
Length = 315
Score = 107 bits (267), Expect = 3e-22
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDPVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S+ A
Sbjct: 138 TNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSRGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.9 bits (172), Expect = 3e-11
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSRGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 297 FVVYENVSHFLLDLREK 313
>UniRef100_UPI000027C671 UPI000027C671 UniRef100 entry
Length = 324
Score = 107 bits (266), Expect = 4e-22
Identities = 67/202 (33%), Positives = 99/202 (48%), Gaps = 15/202 (7%)
Query: 1 MNELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARK 60
+ ++ E VA L G A +PLD+V+ R V DG P+YN H + ++ +
Sbjct: 35 LGHVRVENLVAGLAGGVASTLALHPLDLVKIRFAVSDG--LDLRPKYNGILHCMKSVWNQ 92
Query: 61 EGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGA-----------LMTV 109
EGL+GLY G + GA SW L +Y +K + + A ++T+
Sbjct: 93 EGLRGLYQGVTPNIWGAGASWGLYFLFYNAIKGYIKEGRQSELSASQHLVSAAQAGILTL 152
Query: 110 CLCANPAFVVKTRLQLQTPLHHA-RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA 168
L NP +V KTRL LQ + + Y G++DA I R EG Y+G VPG F S
Sbjct: 153 TL-TNPIWVTKTRLVLQYGADRSSKQYKGMFDALLKIYRHEGVPGLYKGFVPGLFGTSHG 211
Query: 169 AIQFIVYEQLRKTVVNLKTKGS 190
A+QF+ YE+L++ K + S
Sbjct: 212 ALQFMAYEELKRDYNRYKNRPS 233
Score = 65.9 bits (159), Expect = 9e-10
Identities = 57/186 (30%), Positives = 89/186 (47%), Gaps = 20/186 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
++ V++ G +T P+ V +TRL + G S +Y A+ I R EG+ GL
Sbjct: 139 QHLVSAAQAGILTLTLTNPIWVTKTRLVLQYGADRSS-KQYKGMFDALLKIYRHEGVPGL 197
Query: 67 YAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEK-----SGALMTVCLCAN- 114
Y GF GL G + +++ L Y K+R + ++++ AL + A
Sbjct: 198 YKGFVPGLFGTSHGALQFMAYEELKRDYNRYKNRPSDARLDSLEYITMAALSKIFAVATT 257
Query: 115 -PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQF 172
P VV+ RLQ Q H++ YSG+ D R EG + FY+GI P ++ A I F
Sbjct: 258 YPYQVVRARLQDQ---HNS--YSGVMDVIGRTWRNEGAAGFYKGIFPNIIRVTPACCITF 312
Query: 173 IVYEQL 178
+VYE +
Sbjct: 313 VVYENV 318
>UniRef100_Q95T49 GH22139p [Drosophila melanogaster]
Length = 304
Score = 106 bits (265), Expect = 5e-22
Identities = 62/187 (33%), Positives = 95/187 (50%), Gaps = 12/187 (6%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E+ VA ++ G +PLD+++ R V+DGR + P+Y + A TI R+EG
Sbjct: 21 VKYEHLVAGVSGGVVSTLILHPLDLIKIRFAVNDGRT-ATVPQYRGLSSAFTTIFRQEGF 79
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKD-----------RHARSKVEKSGALMTVCLC 112
+GLY G + G+ SW L +Y +K + + + + + L
Sbjct: 80 RGLYKGVTPNVWGSGSSWGLYFMFYNTIKTFIQGGNTTMPLGPTMNMLAAAESGILTLLL 139
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQF 172
NP +VVKTRL LQ + Y G+ A I +EEG YRG VPG +S AIQF
Sbjct: 140 TNPIWVVKTRLCLQCDAASSAEYRGMIHALGQIYKEEGIRGLYRGFVPGMLGVSHGAIQF 199
Query: 173 IVYEQLR 179
+ YE+L+
Sbjct: 200 MTYEELK 206
Score = 54.3 bits (129), Expect = 3e-06
Identities = 46/164 (28%), Positives = 70/164 (42%), Gaps = 20/164 (12%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A+ +G + P+ VV+TRL + S Y HA+ I ++EG++GLY G
Sbjct: 127 LAAAESGILTLLLTNPIWVVKTRLCLQCDAASSA--EYRGMIHALGQIYKEEGIRGLYRG 184
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTV-------------CLCANPA 116
F G+LG + + Y + + K+ L T P
Sbjct: 185 FVPGMLGVSHGAIQFMTYEELKNAYNEYRKLPIDTKLATTEYLAFAAVSKLIAAAATYPY 244
Query: 117 FVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
VV+ RLQ HH R Y+G +D + R E FY+G+VP
Sbjct: 245 QVVRARLQD----HHHR-YNGTWDCIKQTWRYERMRGFYKGLVP 283
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.143 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 415,204,578
Number of Sequences: 2790947
Number of extensions: 16514685
Number of successful extensions: 53203
Number of sequences better than 10.0: 1659
Number of HSP's better than 10.0 without gapping: 609
Number of HSP's successfully gapped in prelim test: 1053
Number of HSP's that attempted gapping in prelim test: 46212
Number of HSP's gapped (non-prelim): 4249
length of query: 250
length of database: 848,049,833
effective HSP length: 124
effective length of query: 126
effective length of database: 501,972,405
effective search space: 63248523030
effective search space used: 63248523030
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 73 (32.7 bits)
Medicago: description of AC125480.1


