

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
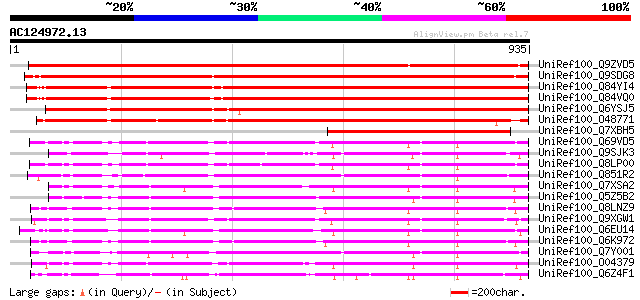
Query= AC124972.13 - phase: 0
(935 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZVD5 Putative Argonaute (AGO1) protein [Arabidopsis ... 1350 0.0
UniRef100_Q9SDG8 ESTs AU068544 [Oryza sativa] 1296 0.0
UniRef100_Q84YI4 ARGONAUTE9 protein [Arabidopsis thaliana] 1231 0.0
UniRef100_Q84VQ0 PAZ (Piwi Argonaut and Zwille) family [Arabidop... 1230 0.0
UniRef100_Q6YSJ5 Putative ARGONAUTE9 protein [Oryza sativa] 1102 0.0
UniRef100_O48771 Argonaute (AGO1)-like protein [Arabidopsis thal... 940 0.0
UniRef100_Q7XBH5 Zwille pinhead-like protein [Oryza sativa] 502 e-140
UniRef100_Q69VD5 ZLL/PNH homologous protein [Oryza sativa] 493 e-137
UniRef100_Q9SJK3 Argonaute-like protein At2g27880 [Arabidopsis t... 485 e-135
UniRef100_Q8LP00 ZLL/PNH homologous protein [Oryza sativa] 484 e-135
UniRef100_Q851R2 Putative argonaute protein [Oryza sativa] 482 e-134
UniRef100_Q7XSA2 OSJNBa0005N02.3 protein [Oryza sativa] 481 e-134
UniRef100_Q5Z5B2 Putative AGO1 homologous protein [Oryza sativa] 479 e-133
UniRef100_Q8LNZ9 AGO1 homologous protein [Oryza sativa] 476 e-132
UniRef100_Q9XGW1 PINHEAD protein [Arabidopsis thaliana] 476 e-132
UniRef100_Q6EU14 Putative argonaute protein [Oryza sativa] 474 e-132
UniRef100_Q6K972 AGO1 homologous protein [Oryza sativa] 474 e-132
UniRef100_Q7Y001 Putative leaf development and shoot apical meri... 469 e-130
UniRef100_O04379 Argonaute protein [Arabidopsis thaliana] 464 e-129
UniRef100_Q6Z4F1 Putative leaf development protein Argonaute [Or... 444 e-123
>UniRef100_Q9ZVD5 Putative Argonaute (AGO1) protein [Arabidopsis thaliana]
Length = 924
Score = 1350 bits (3495), Expect = 0.0
Identities = 644/902 (71%), Positives = 777/902 (85%), Gaps = 4/902 (0%)
Query: 35 ECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNT 94
E LPPPPP++P ++EP++++ ++ +KK P +VPMAR+G G++G K+PLLTNHFKV+V N
Sbjct: 24 EALPPPPPVIPPNVEPVRVKTELAEKKGPVRVPMARKGFGTRGQKIPLLTNHFKVDVANL 83
Query: 95 DGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLA 154
G+FF YSVALFY+DGRPVE KG GRKILD+V +TY S+L+GK+ AYDGEKTLFT G+L
Sbjct: 84 QGHFFHYSVALFYDDGRPVEQKGVGRKILDKVHQTYHSDLDGKEFAYDGEKTLFTYGALP 143
Query: 155 QNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQ 214
NK++F+VVLE+V++ R NGN SP+G+ SP+D DRKRL++ +RSK ++VEIS+A+KIPLQ
Sbjct: 144 SNKMDFSVVLEEVSATRANGNGSPNGNESPSDGDRKRLRRPNRSKNFRVEISYAAKIPLQ 203
Query: 215 AIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCR 274
A+ANA++G E+EN QEAIRVLDIILRQHAA+QGCLLVRQ+FFHNDP N VGG +LGCR
Sbjct: 204 ALANAMRGQESENSQEAIRVLDIILRQHAARQGCLLVRQSFFHNDPTNCEPVGGNILGCR 263
Query: 275 GLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTLKNLRI 334
G HSSFRTTQ G+SLN+DV+TTMI+ PGPVVDFLIANQN RDP+S+DW+KAKRTLKNLR+
Sbjct: 264 GFHSSFRTTQGGMSLNMDVTTTMIIKPGPVVDFLIANQNARDPYSIDWSKAKRTLKNLRV 323
Query: 335 TTSPTNQEYKITGLSEMPCKDQLFTLKKRGA-VPGEDDTEEITVYDYFVNRRKISLQYSA 393
SP+ QE+KITGLS+ PC++Q F LKKR GE +T E+TV DYF + R I LQYSA
Sbjct: 324 KVSPSGQEFKITGLSDKPCREQTFELKKRNPNENGEFETTEVTVADYFRDTRHIDLQYSA 383
Query: 394 DLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDAL 453
DLPCINVGKPKRPT++P+ELC+LV LQRYTKAL+T QRS+LVEKSRQKPQERM VL+ AL
Sbjct: 384 DLPCINVGKPKRPTYIPLELCALVPLQRYTKALTTFQRSALVEKSRQKPQERMTVLSKAL 443
Query: 454 KTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQP 513
K S+Y +EP+LR+CGISI+S FTQV+GRVL AP+LK G G + PRNGRWNFNNK+ V+P
Sbjct: 444 KVSNYDAEPLLRSCGISISSNFTQVEGRVLPAPKLKMGCGSETFPRNGRWNFNNKEFVEP 503
Query: 514 VKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKM 573
KI++W VVNFSARC+VR +V DLIK GG KGI + PF FEE QFRRAPP++RVE M
Sbjct: 504 TKIQRWVVVNFSARCNVRQVVDDLIKIGGSKGIEIASPFQVFEEGNQFRRAPPMIRVENM 563
Query: 574 FEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTR-VNDQYLTNV 632
F+ +QSKLPG P+F+LC+L ++KNSDLYGPWKKKNL EFGIVTQC+APTR NDQYLTN+
Sbjct: 564 FKDIQSKLPGVPQFILCVLPDKKNSDLYGPWKKKNLTEFGIVTQCMAPTRQPNDQYLTNL 623
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQW 692
LLKINAKLGG+NS+L VE +P+ ++SK PT+ILGMDVSHGSPGQ+++PSIAAVVSSR+W
Sbjct: 624 LLKINAKLGGLNSMLSVERTPAFTVISKVPTIILGMDVSHGSPGQSDVPSIAAVVSSREW 683
Query: 693 PLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRD 752
PLISKYRA VRTQ +K EMI++L K + TED+GII+ELL+DFY SS RKP++IIIFRD
Sbjct: 684 PLISKYRASVRTQPSKAEMIESLVKK-NGTEDDGIIKELLVDFYTSSNKRKPEHIIIFRD 742
Query: 753 GVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGTV 812
GVSESQFNQVLNIEL QIIEACK LD WNPKFL++VAQKNHHTKFFQP SP+NVPPGT+
Sbjct: 743 GVSESQFNQVLNIELDQIIEACKLLDANWNPKFLLLVAQKNHHTKFFQPTSPENVPPGTI 802
Query: 813 VDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTT 872
+DNKICHP+N DFY+CAHAGMIGT+RPTHYHVL DEIGFS D+LQELVHSLSYVYQRST+
Sbjct: 803 IDNKICHPKNNDFYLCAHAGMIGTTRPTHYHVLYDEIGFSADELQELVHSLSYVYQRSTS 862
Query: 873 AISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSM 932
AISVVAPICYAHLAA+Q+G FMKFED+SETSSSHGG S + QLP+L D+V NSM
Sbjct: 863 AISVVAPICYAHLAAAQLGTFMKFEDQSETSSSHGGITAPGPIS-VAQLPRLKDNVANSM 921
Query: 933 FF 934
FF
Sbjct: 922 FF 923
>UniRef100_Q9SDG8 ESTs AU068544 [Oryza sativa]
Length = 904
Score = 1296 bits (3355), Expect = 0.0
Identities = 640/908 (70%), Positives = 747/908 (81%), Gaps = 6/908 (0%)
Query: 27 EMGSQENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNH 86
E S E EE LPPPPP+ P + EPIK + K P + MAR G G KG + LLTNH
Sbjct: 2 ESNSGEIEE-LPPPPPL-PPNAEPIKTD-DTKKLSKPKRALMARSGCGKKGQPIQLLTNH 58
Query: 87 FKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKT 146
FKV++ D +F Y V L YED RPV+GKG GRK+LD++Q+TY SEL KD AYDGEK+
Sbjct: 59 FKVSLKAADEFFHHYYVNLKYEDDRPVDGKGIGRKVLDKLQQTYASELANKDFAYDGEKS 118
Query: 147 LFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEIS 206
LFTIG+L Q EFTVVLED + +++ N G+ SP + DRKR+++ +++KT+KVE++
Sbjct: 119 LFTIGALPQVNNEFTVVLEDFNTGKSSANGGSPGNDSPGN-DRKRVRRPYQTKTFKVELN 177
Query: 207 FASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDV 266
FA+KIP+ AIA AL+G E+EN QEAIRV+DIILRQH+AKQGCLLVRQ+FFHN+P NF D+
Sbjct: 178 FAAKIPMSAIAQALRGQESENTQEAIRVIDIILRQHSAKQGCLLVRQSFFHNNPSNFVDL 237
Query: 267 GGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAK 326
GGGV+GCRG HSSFR TQSGLSLNIDVSTTMIV PGPVVDFL+ANQ V P +DW KAK
Sbjct: 238 GGGVMGCRGFHSSFRATQSGLSLNIDVSTTMIVKPGPVVDFLLANQKVDHPNKIDWAKAK 297
Query: 327 RTLKNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRK 386
R LKNLRI TSP N EYKI GLSE C +Q+FTLK+R GE + E++VY+YFV R
Sbjct: 298 RALKNLRIKTSPANTEYKIVGLSERNCYEQMFTLKQRNG-DGEPEGVEVSVYEYFVKNRG 356
Query: 387 ISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERM 446
I L+YS D PCINVGKPKRPT+ P+ELCSLV LQRYTKALSTLQRSSLVEKSRQKP+ERM
Sbjct: 357 IELRYSGDFPCINVGKPKRPTYFPIELCSLVPLQRYTKALSTLQRSSLVEKSRQKPEERM 416
Query: 447 RVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFN 506
VL+D LK S+Y SEPML +CGISI GFTQV GRVLQAP+LK GNGED RNGRWNFN
Sbjct: 417 SVLSDVLKRSNYDSEPMLNSCGISIARGFTQVAGRVLQAPKLKAGNGEDLFARNGRWNFN 476
Query: 507 NKKIVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPP 566
NK++++ IEKWAVVNFSARC++R LVRD+IKCGGMKGI VE PFD EE+ RRAP
Sbjct: 477 NKRLIKASSIEKWAVVNFSARCNIRDLVRDIIKCGGMKGIKVEDPFDVIEEDPSMRRAPA 536
Query: 567 LVRVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVND 626
RV+ M + +Q KLPG PKFLLC+L+ERKNSD+YGPWK+K LAEFGI+TQC+APTRVND
Sbjct: 537 ARRVDGMIDKMQKKLPGQPKFLLCVLAERKNSDIYGPWKRKCLAEFGIITQCVAPTRVND 596
Query: 627 QYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAV 686
QY+TNVLLKINAKLGG+NSLL +E SPSIP+VSK PT+ILGMDVSHGSPGQ++IPSIAAV
Sbjct: 597 QYITNVLLKINAKLGGLNSLLQIETSPSIPLVSKVPTIILGMDVSHGSPGQSDIPSIAAV 656
Query: 687 VSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDN 746
VSSR+WPL+SKYRA VR+Q K+EMID LFKP ED+G+IRELL+DFY S+G RKPD
Sbjct: 657 VSSREWPLVSKYRASVRSQSPKLEMIDGLFKPQGAQEDDGLIRELLVDFYTSTGKRKPDQ 716
Query: 747 IIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDN 806
+IIFRDGVSESQF QVLNIEL QIIEACKFLDE W+PKF +IVAQKNHHTKFF PGS +N
Sbjct: 717 VIIFRDGVSESQFTQVLNIELDQIIEACKFLDENWSPKFTLIVAQKNHHTKFFVPGSQNN 776
Query: 807 VPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYV 866
VPPGTVVDN +CHPRN DFYMCAHAGMIGT+RPTHYH+L DEIGFS DDLQELVHSLSYV
Sbjct: 777 VPPGTVVDNAVCHPRNNDFYMCAHAGMIGTTRPTHYHILHDEIGFSADDLQELVHSLSYV 836
Query: 867 YQRSTTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMD 926
YQRSTTAISVVAPICYAHLAA+QV QF+KF++ SETSSSHGG ++P+P+LP+L +
Sbjct: 837 YQRSTTAISVVAPICYAHLAAAQVSQFIKFDEMSETSSSHGGH-TSAGSAPVPELPRLHN 895
Query: 927 SVCNSMFF 934
V +SMFF
Sbjct: 896 KVRSSMFF 903
>UniRef100_Q84YI4 ARGONAUTE9 protein [Arabidopsis thaliana]
Length = 896
Score = 1231 bits (3185), Expect = 0.0
Identities = 607/906 (66%), Positives = 735/906 (80%), Gaps = 17/906 (1%)
Query: 31 QENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMAR-RGLGSKGAKLPLLTNHFKV 89
+ N LPPPPP VPA++ P ++EP VKK + +PMAR RG GSKG K+PLLTNHF V
Sbjct: 5 EPNGSGLPPPPPFVPANLVP-EVEP--VKKNI--LLPMARPRGSGSKGQKIPLLTNHFGV 59
Query: 90 NVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFT 149
GYFF YSVA+ YEDGRPVE KG GRKILD+VQETY S+L K AYDGEKTLFT
Sbjct: 60 KFNKPSGYFFHYSVAINYEDGRPVEAKGIGRKILDKVQETYQSDLGAKYFAYDGEKTLFT 119
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
+G+L NKL+F+VVLE++ S+RN+ ND DRKR ++ +++K + VEIS+A+
Sbjct: 120 VGALPSNKLDFSVVLEEIPSSRNHAG------NDTNDADRKRSRRPNQTKKFMVEISYAA 173
Query: 210 KIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGG 269
KIP+QAIA+AL+G ETEN Q+A+RVLDIILRQ AA+QGCLLVRQ+FFHND KNF +GGG
Sbjct: 174 KIPMQAIASALQGKETENLQDALRVLDIILRQSAARQGCLLVRQSFFHNDVKNFVPIGGG 233
Query: 270 VLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTL 329
V GCRG HSSFRTTQ GLSLNID STTMIV PGP+VDFL+ANQN +DP+ +DWNKA+R L
Sbjct: 234 VSGCRGFHSSFRTTQGGLSLNIDTSTTMIVQPGPIVDFLLANQNKKDPYGMDWNKARRVL 293
Query: 330 KNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISL 389
KNLR+ + +N+EYKI+GLSE CKDQLFT +K GE + EITV +Y+ R I +
Sbjct: 294 KNLRVQITLSNREYKISGLSEHSCKDQLFTWRKPND-KGEFEEVEITVLNYY-KERNIEV 351
Query: 390 QYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVL 449
+YS D PCINVGKPKRPT+ P+E C+LVSLQRYTK+L+ QR++LVEKSRQKP ERM L
Sbjct: 352 RYSGDFPCINVGKPKRPTYFPIEFCNLVSLQRYTKSLTNFQRAALVEKSRQKPPERMASL 411
Query: 450 TDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKK 509
T LK S+Y ++P+L++ G+SI + FTQV+GR+L P LK G GE+ +P G+WNF K
Sbjct: 412 TKGLKDSNYNADPVLQDSGVSIITNFTQVEGRILPTPMLKVGKGENLSPIKGKWNFMRKT 471
Query: 510 IVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPF-DCFEENGQFRRAPPLV 568
+ +P + +WAVVNFSARCD L+RDLIKCG KGI+VE PF D EN QFR AP V
Sbjct: 472 LAEPTTVTRWAVVNFSARCDTNTLIRDLIKCGREKGINVEPPFKDVINENPQFRNAPATV 531
Query: 569 RVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQY 628
RVE MFE ++SKLP P FLLC+L+ERKNSD+YGPWKKK+L + GIVTQCIAPTR+NDQY
Sbjct: 532 RVENMFEQIKSKLPKPPLFLLCILAERKNSDVYGPWKKKDLVDLGIVTQCIAPTRLNDQY 591
Query: 629 LTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVS 688
LTNVLLKINAKLGG+NSLL +E SP++P V++ PT+I+GMDVSHGSPGQ++IPSIAAVVS
Sbjct: 592 LTNVLLKINAKLGGLNSLLAMERSPAMPKVTQVPTIIVGMDVSHGSPGQSDIPSIAAVVS 651
Query: 689 SRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNII 748
SRQWPLISKY+ACVRTQ K+EMIDNLFKPV+ +DEG+ RELL+DFY SS NRKP++II
Sbjct: 652 SRQWPLISKYKACVRTQSRKMEMIDNLFKPVNG-KDEGMFRELLLDFYYSSENRKPEHII 710
Query: 749 IFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVP 808
IFRDGVSESQFNQVLNIEL Q+++ACKFLD+ W+PKF VIVAQKNHHTKFFQ PDNVP
Sbjct: 711 IFRDGVSESQFNQVLNIELDQMMQACKFLDDTWHPKFTVIVAQKNHHTKFFQSRGPDNVP 770
Query: 809 PGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQ 868
PGT++D++ICHPRN+DFY+CAHAGMIGT+RPTHYHVL DEIGF+ DDLQELVHSLS+VYQ
Sbjct: 771 PGTIIDSQICHPRNFDFYLCAHAGMIGTTRPTHYHVLYDEIGFATDDLQELVHSLSHVYQ 830
Query: 869 RSTTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSV 928
RSTTAISVVAP+CYAHLAA+Q+G MK+E+ SETSSSHGG P+P +P+L ++V
Sbjct: 831 RSTTAISVVAPVCYAHLAAAQMGTVMKYEELSETSSSHGGITTP-GTVPVPPMPQLHNNV 889
Query: 929 CNSMFF 934
SMFF
Sbjct: 890 STSMFF 895
>UniRef100_Q84VQ0 PAZ (Piwi Argonaut and Zwille) family [Arabidopsis thaliana]
Length = 892
Score = 1230 bits (3182), Expect = 0.0
Identities = 609/906 (67%), Positives = 733/906 (80%), Gaps = 21/906 (2%)
Query: 31 QENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMAR-RGLGSKGAKLPLLTNHFKV 89
+ N LPPPPP VPA++ P ++EP VKK + +PMAR RG GSKG K+PLLTNHF V
Sbjct: 5 EPNGSGLPPPPPFVPANLVP-EVEP--VKKNI--LLPMARPRGSGSKGQKIPLLTNHFGV 59
Query: 90 NVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFT 149
GYFF YSVA+ YEDGRPVE KG GRKILD+VQETY S+L K AYDGEKTLFT
Sbjct: 60 KFNKPSGYFFHYSVAINYEDGRPVEAKGIGRKILDKVQETYQSDLGAKYFAYDGEKTLFT 119
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
+G+L NKL+F+VVLE++ S+RN+ ND DRKR ++ +++K + VEIS+A+
Sbjct: 120 VGALPSNKLDFSVVLEEIPSSRNHAG------NDTNDADRKRSRRPNQTKKFMVEISYAA 173
Query: 210 KIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGG 269
KIP+QAIA+AL+G ETEN Q+A+RVLDIILRQ AA+QGCLLVRQ+FFHND KNF +GGG
Sbjct: 174 KIPMQAIASALQGKETENLQDALRVLDIILRQSAARQGCLLVRQSFFHNDVKNFVPIGGG 233
Query: 270 VLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTL 329
V GCRG HSSFRTTQ GLSLNID STTMIV PGPVVDFL+ANQN +DP+ +DWNKA+R L
Sbjct: 234 VSGCRGFHSSFRTTQGGLSLNIDTSTTMIVQPGPVVDFLLANQNKKDPYGMDWNKARRVL 293
Query: 330 KNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISL 389
KNLR+ + +N+EYKI+GLSE CKDQL K GE + EITV +Y+ R I +
Sbjct: 294 KNLRVQITLSNREYKISGLSEHSCKDQLKPNDK-----GEFEEVEITVLNYY-KERNIEV 347
Query: 390 QYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVL 449
+YS D PCINVGKPKRPT+ P+E C+LVSLQRYTK+L+ QR++LVEKSRQKP ERM L
Sbjct: 348 RYSGDFPCINVGKPKRPTYFPIEFCNLVSLQRYTKSLTNFQRAALVEKSRQKPPERMASL 407
Query: 450 TDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKK 509
T LK S+Y ++P+L++ G+SI + FTQV+GR+L P LK G GE+ +P G+WNF K
Sbjct: 408 TKGLKDSNYNADPVLQDSGVSIITNFTQVEGRILPTPMLKVGKGENLSPIKGKWNFMRKT 467
Query: 510 IVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPF-DCFEENGQFRRAPPLV 568
+ +P + +WAVVNFSARCD L+RDLIKCG KGI+VE PF D EN QFR AP V
Sbjct: 468 LAEPTTVTRWAVVNFSARCDTNTLIRDLIKCGREKGINVEPPFKDVINENPQFRNAPATV 527
Query: 569 RVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQY 628
RVE MFE ++SKLP P FLLC+L+ERKNSD+YGPWKKKNL + GIVTQCIAPTR+NDQY
Sbjct: 528 RVENMFEQIKSKLPKPPLFLLCILAERKNSDVYGPWKKKNLVDLGIVTQCIAPTRLNDQY 587
Query: 629 LTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVS 688
LTNVLLKINAKLGG+NSLL +E SP++P V++ PT+I+GMDVSHGSPGQ++IPSIAAVVS
Sbjct: 588 LTNVLLKINAKLGGLNSLLAMERSPAMPKVTQVPTIIVGMDVSHGSPGQSDIPSIAAVVS 647
Query: 689 SRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNII 748
SRQWPLISKY+ACVRTQ K+EMIDNLFKPV+ +DEG+ RELL+DFY SS NRKP++II
Sbjct: 648 SRQWPLISKYKACVRTQSRKMEMIDNLFKPVNG-KDEGMFRELLLDFYYSSENRKPEHII 706
Query: 749 IFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVP 808
IFRDGVSESQFNQVLNIEL Q+++ACKFLD+ W+PKF VIVAQKNHHTKFFQ PDNVP
Sbjct: 707 IFRDGVSESQFNQVLNIELDQMMQACKFLDDTWHPKFTVIVAQKNHHTKFFQSRGPDNVP 766
Query: 809 PGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQ 868
PGT++D++ICHPRN+DFY+CAHAGMIGT+RPTHYHVL DEIGF+ DDLQELVHSLSYVYQ
Sbjct: 767 PGTIIDSQICHPRNFDFYLCAHAGMIGTTRPTHYHVLYDEIGFATDDLQELVHSLSYVYQ 826
Query: 869 RSTTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSV 928
RSTTAISVVAP+CYAHLAA+Q+G MK+E+ SETSSSHGG A P+P +P+L ++V
Sbjct: 827 RSTTAISVVAPVCYAHLAAAQMGTVMKYEELSETSSSHGGITTP-GAVPVPPMPQLHNNV 885
Query: 929 CNSMFF 934
SMFF
Sbjct: 886 STSMFF 891
>UniRef100_Q6YSJ5 Putative ARGONAUTE9 protein [Oryza sativa]
Length = 889
Score = 1102 bits (2849), Expect = 0.0
Identities = 551/878 (62%), Positives = 684/878 (77%), Gaps = 16/878 (1%)
Query: 65 KVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILD 124
+VP+AR G +G ++ LL+NHF V ++ D F+QYSV++ ED + ++GKG GRK++D
Sbjct: 19 RVPIARPSFGREGKQIKLLSNHFTVKLSGIDAVFYQYSVSIKSEDDKVIDGKGIGRKVMD 78
Query: 125 RVQETYGSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSP 184
+V +TY SEL GK+ AYDGEK LFT+G L QN EFTV+LE+ +S G+ GHGSP
Sbjct: 79 KVLQTYSSELAGKEFAYDGEKCLFTVGPLPQNNFEFTVILEETSSRAAGGSL---GHGSP 135
Query: 185 NDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAA 244
N D+KR K +H +K V IS+A+KIPL+++A AL+G E+++ Q+A+RVLDI+LRQ A
Sbjct: 136 NQGDKKRSKCTHLAKKIVVGISYAAKIPLKSVALALQGSESDHAQDALRVLDIVLRQQQA 195
Query: 245 KQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPV 304
K+GCLLVRQ+FF +D +N D+ GGV GCRGLHSSFRTT GLSLN+DVSTTMIV PGPV
Sbjct: 196 KRGCLLVRQSFFSDDFRNLVDLTGGVSGCRGLHSSFRTTIGGLSLNMDVSTTMIVTPGPV 255
Query: 305 VDFLIANQNVRDPFSLDWNKAKRTLKNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRG 364
DFL+ NQNVRD +DW +AK+ LKNLR+ N E+KI GLS+ PC Q F +K R
Sbjct: 256 FDFLLTNQNVRDIRDIDWPRAKKMLKNLRVKAIHNNMEFKIIGLSDEPCSRQTFPMKVRN 315
Query: 365 AVPGEDDTEEITVYDYFVNRR-KISLQYSADLPCINVGKPKRPTFVPVE------LCSLV 417
E +T EITV +YF +++ +++ Y LPC++VGKPKRP +VP+E LC +V
Sbjct: 316 G-SSEGETVEITVQEYFKSKQVDLTMPY---LPCLDVGKPKRPNYVPIEVAGTNPLCHMV 371
Query: 418 SLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQ 477
SLQRYTKALS+ QR++LVEKSRQKPQERMRV+TDA+K + Y +P+L +CGI I T+
Sbjct: 372 SLQRYTKALSSQQRATLVEKSRQKPQERMRVVTDAVKNNRYDDDPILSSCGIKIEKQLTR 431
Query: 478 VDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDL 537
VDGRVL AP L GN ED P GRWN+NNK++ +PVKIE+WA+VNFSARCD+ + RDL
Sbjct: 432 VDGRVLSAPTLVVGNSEDCIPNRGRWNYNNKRLFEPVKIERWAIVNFSARCDMSRISRDL 491
Query: 538 IKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKFLLCLLSERKN 597
I CG KGI +E+PF +E+ Q RR P+VRVE MFE V++ LPG P+FLLC+L ERKN
Sbjct: 492 INCGRTKGIIIERPFTLVDEDSQSRRCTPVVRVESMFEKVKANLPGPPEFLLCVLPERKN 551
Query: 598 SDLYGPWKKKNLAEFGIVTQCIAPT-RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIP 656
DLYGPWKKKNL E GI+TQCI P+ ++NDQY TNVLLKINAKLGGMNS L +EH IP
Sbjct: 552 CDLYGPWKKKNLHEMGIITQCIVPSVKMNDQYYTNVLLKINAKLGGMNSKLSLEHRHMIP 611
Query: 657 IVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLF 716
IV++ PTLILGMDVSHGSPG+ ++PSIAAVV SR WPLIS+YRA VRTQ KVEMID+LF
Sbjct: 612 IVNQTPTLILGMDVSHGSPGRADVPSIAAVVGSRCWPLISRYRASVRTQSPKVEMIDSLF 671
Query: 717 KPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKF 776
KP+ D +D+GIIRELL+DFY +S RKP IIIFRDGVSESQF+QVLN+EL+QII+A ++
Sbjct: 672 KPLDDGKDDGIIRELLLDFYKTSQQRKPKQIIIFRDGVSESQFSQVLNVELNQIIKAYQY 731
Query: 777 LDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGT 836
+D+ PKF VI+AQKNHHTK FQ +PDNVPPGTVVD+ I HPR YDFYM AHAG IGT
Sbjct: 732 MDQGPIPKFTVIIAQKNHHTKLFQENTPDNVPPGTVVDSGIVHPRQYDFYMYAHAGPIGT 791
Query: 837 SRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKF 896
SRPTHYHVLLDEIGF PDD+Q+LV SLSYVYQRSTTAISVVAPICYAHLAA+Q+GQFMKF
Sbjct: 792 SRPTHYHVLLDEIGFLPDDVQKLVLSLSYVYQRSTTAISVVAPICYAHLAAAQMGQFMKF 851
Query: 897 EDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
E+ +ETSS GG + + +P+LP+L VC+SMFF
Sbjct: 852 EEFAETSSGSGGVPSS-SGAVVPELPRLHADVCSSMFF 888
>UniRef100_O48771 Argonaute (AGO1)-like protein [Arabidopsis thaliana]
Length = 887
Score = 940 bits (2430), Expect = 0.0
Identities = 496/900 (55%), Positives = 632/900 (70%), Gaps = 35/900 (3%)
Query: 48 IEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFY 107
+ PI IEP+ + RRG+G+ G + L TNHF V+V D F+QY+V++
Sbjct: 9 LSPISIEPEQPSHR--DYDITTRRGVGTTGNPIELCTNHFNVSVRQPDVVFYQYTVSITT 66
Query: 108 EDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDV 167
E+G V+G G RK++D++ +TY S+L+GK LAYDGEKTL+T+G L QN+ +F V++E
Sbjct: 67 ENGDAVDGTGISRKLMDQLFKTYSSDLDGKRLAYDGEKTLYTVGPLPQNEFDFLVIVEGS 126
Query: 168 TSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHET-- 225
S R+ G + DG GS + T KR K+S ++YKV+I +A++IPL+ + +G T
Sbjct: 127 FSKRDCGVS--DG-GSSSGTC-KRSKRSFLPRSYKVQIHYAAEIPLKTVLGTQRGAYTPD 182
Query: 226 ENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQS 285
++ Q+A+RVLDI+LRQ AA++GCLLVRQ FFH+D VGGGV+G RGLHSSFR T
Sbjct: 183 KSAQDALRVLDIVLRQQAAERGCLLVRQAFFHSDGHPMK-VGGGVIGIRGLHSSFRPTHG 241
Query: 286 GLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNK-AKRTLKNLRITTSPTNQEYK 344
GLSLNIDVSTTMI+ PGPV++FL ANQ+V P +DW K A + LK++R+ + N E+K
Sbjct: 242 GLSLNIDVSTTMILEPGPVIEFLKANQSVETPRQIDWIKVAAKMLKHMRVKATHRNMEFK 301
Query: 345 ITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPK 404
I GLS PC QLF++K + E EITVYDYF + SA PC++VGKP
Sbjct: 302 IIGLSSKPCNQQLFSMKIKDG-EREVPIREITVYDYFKQTYTEPIS-SAYFPCLDVGKPD 359
Query: 405 RPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPML 464
RP ++P+E C+LVSLQRYTK LS QR LVE SRQKP ER++ L DA+ T Y +P L
Sbjct: 360 RPNYLPLEFCNLVSLQRYTKPLSGRQRVLLVESSRQKPLERIKTLNDAMHTYCYDKDPFL 419
Query: 465 RNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNF 524
CGISI TQV+GRVL+ P LKFG EDF P NGRWNFNNK +++P I+ WA+VNF
Sbjct: 420 AGCGISIEKEMTQVEGRVLKPPMLKFGKNEDFQPCNGRWNFNNKMLLEPRAIKSWAIVNF 479
Query: 525 SARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGA 584
S CD + R+LI CG KGI +++PF EE+ Q+++A P+ RVEKM ++ K P
Sbjct: 480 SFPCDSSHISRELISCGMRKGIEIDRPFALVEEDPQYKKAGPVERVEKMIATMKLKFPDP 539
Query: 585 PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQYLTNVLLKINAKLGGMN 644
P F+LC+L ERK SD+YGPWKK L E GI TQCI P +++DQYLTNVLLKIN+KLGG+N
Sbjct: 540 PHFILCILPERKTSDIYGPWKKICLTEEGIHTQCICPIKISDQYLTNVLLKINSKLGGIN 599
Query: 645 SLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRT 704
SLLG+E+S +IP+++K PTLILGMDVSHG PG+ ++PS+AAVV S+ WPLIS+YRA VRT
Sbjct: 600 SLLGIEYSYNIPLINKIPTLILGMDVSHGPPGRADVPSVAAVVGSKCWPLISRYRAAVRT 659
Query: 705 QGAKVEMIDNLFKPVSDTE--DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQV 762
Q ++EMID+LF+P+ +TE D GI+ EL ++FY +S RKP IIIFRDGVSESQF QV
Sbjct: 660 QSPRLEMIDSLFQPIENTEKGDNGIMNELFVEFYRTSRARKPKQIIIFRDGVSESQFEQV 719
Query: 763 LNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGTVVDNKICHPRN 822
L IE+ QII+A + L E PKF VIVAQKNHHTK FQ P+NVP GTVVD KI HP N
Sbjct: 720 LKIEVDQIIKAYQRLGESDVPKFTVIVAQKNHHTKLFQAKGPENVPAGTVVDTKIVHPTN 779
Query: 823 YDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAIS------- 875
YDFYMCAHAG IGTSRP HYHVLLDEIGFSPDDLQ L+HSLSY S +S
Sbjct: 780 YDFYMCAHAGKIGTSRPAHYHVLLDEIGFSPDDLQNLIHSLSYKLLNSIFNVSSLLCVFV 839
Query: 876 -VVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
VAP+ YAHLAA+QV QF KFE SE +P+LP+L ++V +MFF
Sbjct: 840 LSVAPVRYAHLAAAQVAQFTKFEGISEDGK-------------VPELPRLHENVEGNMFF 886
>UniRef100_Q7XBH5 Zwille pinhead-like protein [Oryza sativa]
Length = 361
Score = 502 bits (1293), Expect = e-140
Identities = 246/330 (74%), Positives = 284/330 (85%), Gaps = 1/330 (0%)
Query: 573 MFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPT-RVNDQYLTN 631
MFE V++ LPG P+FLLC+L ERKN DLYGPWKKKNL E GI+TQCI P+ ++NDQY TN
Sbjct: 1 MFEKVKANLPGPPEFLLCVLPERKNCDLYGPWKKKNLHEMGIITQCIVPSVKMNDQYYTN 60
Query: 632 VLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQ 691
VLLKINAKLGGMNS L +EH IPIV++ PTLILGMDVSHGSPG+ ++PSIAAV SR
Sbjct: 61 VLLKINAKLGGMNSKLSLEHRHMIPIVNQTPTLILGMDVSHGSPGRADVPSIAAVAGSRC 120
Query: 692 WPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFR 751
WPLIS+YRA VRTQ KVEMID+LFKP+ D +D+GIIRELL+DFY +S RKP IIIFR
Sbjct: 121 WPLISRYRASVRTQSPKVEMIDSLFKPLDDGKDDGIIRELLLDFYKTSQQRKPKQIIIFR 180
Query: 752 DGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGT 811
DGVSESQF+QVLN+EL+QII+A +++D+ PKF VI+AQKNHHTK FQ +PDNVPPGT
Sbjct: 181 DGVSESQFSQVLNVELNQIIKAYQYMDQGPIPKFTVIIAQKNHHTKLFQENTPDNVPPGT 240
Query: 812 VVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRST 871
VVD+ I HPR YDFYM AHAG IGTSRPTHYHVLLDEIGF PDD+Q+LV SLSYVYQRST
Sbjct: 241 VVDSGIVHPRQYDFYMYAHAGPIGTSRPTHYHVLLDEIGFLPDDVQKLVLSLSYVYQRST 300
Query: 872 TAISVVAPICYAHLAASQVGQFMKFEDKSE 901
TAISVVAPICYAHLAA+Q+GQFMKFE+ +E
Sbjct: 301 TAISVVAPICYAHLAAAQMGQFMKFEEFAE 330
>UniRef100_Q69VD5 ZLL/PNH homologous protein [Oryza sativa]
Length = 979
Score = 493 bits (1269), Expect = e-137
Identities = 325/932 (34%), Positives = 490/932 (51%), Gaps = 84/932 (9%)
Query: 37 LPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDG 96
L P +V A I P + K L R G G+ GA+ + NHF + + D
Sbjct: 97 LAAAPAVVVAPAARAVIGPPVASKGLSF---CRRPGFGTVGARCVVKANHFLAELPDKD- 152
Query: 97 YFFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFTIGSLAQ 155
QY V + E V + R I+ + Y S+L G+ AYDG K L+T G+L
Sbjct: 153 -LTQYDVKITPE----VSSRSVNRAIMSELVRLYHDSDLGGRLPAYDGRKNLYTAGTLPF 207
Query: 156 NKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQA 215
+ EF V L D DG G P R + Y+V I FA++ L
Sbjct: 208 DAREFVVRLTD----------DDDGTGVPP-----------REREYRVAIKFAARADLHH 246
Query: 216 IANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRG 275
+ + G + + QEA++VLDI+LR+ A ++ + + ++F+ D + +G G+ G
Sbjct: 247 LRQFIAGRQADAPQEALQVLDIVLRELANRR-YVSIGRSFYSPDIRKPQRLGDGLQSWCG 305
Query: 276 LHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF---LIANQNVRDPFSLDWN--KAKRTLK 330
+ S R TQ GLSLNID+S+T + P PV++F ++ + P S D N K K+ L+
Sbjct: 306 FYQSIRPTQMGLSLNIDMSSTAFIEPLPVIEFVAQILGKDVISRPLS-DANRIKIKKALR 364
Query: 331 NLRITTSP---TNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKI 387
+++ + ++Y+I+GL+ P + +F P +D +V +YF
Sbjct: 365 GVKVEVTHRGNVRRKYRISGLTTQPTHELIF--------PIDDQMNMKSVVEYFKEMYGF 416
Query: 388 SLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMR 447
++Q+ LPC+ VG K+ ++P+E C +V QRYTK L+ Q +SL++ + ++P+E+
Sbjct: 417 TIQHP-HLPCLQVGNQKKANYLPMEACKIVEGQRYTKRLNEKQITSLLKVTCRRPREQEM 475
Query: 448 VLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPRNGRWN 504
+ ++ + Y +P + GI+I+ T V+ RVL AP LK+ G ++ P+ G+WN
Sbjct: 476 DILQTVQQNGYEQDPYAKEFGINISEKLTSVEARVLPAPWLKYHDTGKEKECLPQVGQWN 535
Query: 505 FNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMKGIHVEQPFDCFEENGQF 561
NKK++ K+ WA +NFS RG ++L + + G+ E
Sbjct: 536 MVNKKVINGCKVNHWACINFSRSVQETTARGFCQELAQMCQISGMEFNS-----EPVIPI 590
Query: 562 RRAPPLVRVEKMFEHVQS----KLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQ 617
A P +VEK +HV + KL G LL + N LYG K+ + G+++Q
Sbjct: 591 YSARP-DQVEKALKHVYNMSLNKLKGKELELLLAILPDNNGSLYGDIKRICETDLGLISQ 649
Query: 618 CIAPT---RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGS 674
C +++ QYL NV LKIN K+GG N++L S IP+VS PT+I G DV+H
Sbjct: 650 CCLTKHVFKISKQYLANVSLKINVKMGGRNTVLLDAISWRIPLVSDIPTIIFGADVTHPE 709
Query: 675 PGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTEDEGIIRE 730
G+ PSIAAVV+S+ WP ++KY V Q + E+I +L+K P T G+IRE
Sbjct: 710 TGEDSSPSIAAVVASQDWPEVTKYAGLVCAQAHRQELIQDLYKTWHDPQRGTVTGGMIRE 769
Query: 731 LLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVA 790
LLI F ++G +KP II +RDGVSE QF QVL EL I +AC L+ + P +V
Sbjct: 770 LLISFRKATG-QKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPNYQPPVTFVVV 828
Query: 791 QKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHY 842
QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG+ GTSRP HY
Sbjct: 829 QKRHHTRLFANNHKDRSSTDKSGNILPGTVVDSKICHPSEFDFYLCSHAGIQGTSRPAHY 888
Query: 843 HVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKFEDKSET 902
HVL DE F+ D++Q L ++L Y Y R T ++SVV P YAHLAA + +M+ E
Sbjct: 889 HVLWDENNFTADEMQTLTNNLCYTYARCTRSVSVVPPAYYAHLAAFRARFYMEPEMSENQ 948
Query: 903 SSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
++S +G N + + LP + + V MF+
Sbjct: 949 TTSKSSTG--TNGTSVKPLPAVKEKVKRVMFY 978
>UniRef100_Q9SJK3 Argonaute-like protein At2g27880 [Arabidopsis thaliana]
Length = 997
Score = 485 bits (1249), Expect = e-135
Identities = 325/907 (35%), Positives = 475/907 (51%), Gaps = 102/907 (11%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G G+ G K+ + NHF V V + D Y + S+ V K R ++ + +
Sbjct: 150 RPGRGTLGKKVMVRANHFLVQVADRDLYHYDVSI------NPEVISKTVNRNVMKLLVKN 203
Query: 130 Y-GSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD 188
Y S L GK AYDG K+L+T G L + EF V L + ++ ++G P
Sbjct: 204 YKDSHLGGKSPAYDGRKSLYTAGPLPFDSKEFVVNLAEKRADGSSGKDRP---------- 253
Query: 189 RKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGC 248
+KV + + L + L + E + I+VLD++LR +
Sbjct: 254 ------------FKVAVKNVTSTDLYQLQQFLDRKQREAPYDTIQVLDVVLRDKPSND-Y 300
Query: 249 LLVRQNFFHNDPKNFTDVGGGVLG-----CRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
+ V ++FFH G G LG RG S R TQ GLSLNIDVS P
Sbjct: 301 VSVGRSFFHTSLGKDARDGRGELGDGIEYWRGYFQSLRLTQMGLSLNIDVSARSFYEPIV 360
Query: 304 VVDFLIANQNVRD---PF-SLDWNKAKRTLKNLRITTSPTN--QEYKITGLSEMPCKDQL 357
V DF+ N+RD P D K K+ L+ L++ N + KI+G+S +P ++
Sbjct: 361 VTDFISKFLNIRDLNRPLRDSDRLKVKKVLRTLKVKLLHWNCTKSAKISGISSLPIRELR 420
Query: 358 FTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLV 417
FTL +D E TV YF + ++Y A LP I G RP ++P+ELC +
Sbjct: 421 FTL---------EDKSEKTVVQYFAEKYNYRVKYQA-LPAIQTGSDTRPVYLPMELCQID 470
Query: 418 SLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQ 477
QRYTK L+ Q ++L++ + Q+P +R + + + ++Y + + + G+S+T+
Sbjct: 471 EGQRYTKRLNEKQVTALLKATCQRPPDRENSIKNLVVKNNYNDD-LSKEFGMSVTTQLAS 529
Query: 478 VDGRVLQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLV 534
++ RVL P LK+ G + NPR G+WN +KK+V K+ W V+FS R D RGL
Sbjct: 530 IEARVLPPPMLKYHDSGKEKMVNPRLGQWNMIDKKMVNGAKVTSWTCVSFSTRID-RGLP 588
Query: 535 RDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKL-------PGAPKF 587
++ C + G+ C + +F+ P + + EH++ L PG +
Sbjct: 589 QEF--CKQLIGM-------CVSKGMEFKPQPAIPFISCPPEHIEEALLDIHKRAPGL-QL 638
Query: 588 LLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVND---QYLTNVLLKINAKLGGMN 644
L+ +L + S YG K+ E GIV+QC P +VN QY+ NV LKIN K GG N
Sbjct: 639 LIVILPDVTGS--YGKIKRICETELGIVSQCCQPRQVNKLNKQYMENVALKINVKTGGRN 696
Query: 645 SLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRT 704
++L +IP+++ PT+I+G DV+H PG+ PSIAAVV+S WP I+KYR V
Sbjct: 697 TVLNDAIRRNIPLITDRPTIIMGADVTHPQPGEDSSPSIAAVVASMDWPEINKYRGLVSA 756
Query: 705 QGAKVEMIDNLFKPVSDTE----DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFN 760
Q + E+I +L+K V D + G+IRE I F ++G + P II +RDGVSE QF+
Sbjct: 757 QAHREEIIQDLYKLVQDPQRGLVHSGLIREHFIAFRRATG-QIPQRIIFYRDGVSEGQFS 815
Query: 761 QVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF--QPGSPD------NVPPGTV 812
QVL E++ I +AC L E + P+ ++ QK HHT+ F Q G+ D N+ PGTV
Sbjct: 816 QVLLHEMTAIRKACNSLQENYVPRVTFVIVQKRHHTRLFPEQHGNRDMTDKSGNIQPGTV 875
Query: 813 VDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTT 872
VD KICHP +DFY+ +HAG+ GTSRP HYHVLLDE GF+ D LQ L ++L Y Y R T
Sbjct: 876 VDTKICHPNEFDFYLNSHAGIQGTSRPAHYHVLLDENGFTADQLQMLTNNLCYTYARCTK 935
Query: 873 AISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASP-----IPQLPKLMDS 927
++S+V P YAHLAA + +M E+ S GGS R +++ I QLP + D+
Sbjct: 936 SVSIVPPAYYAHLAAFRARYYM------ESEMSDGGSSRSRSSTTGVGQVISQLPAIKDN 989
Query: 928 VCNSMFF 934
V MF+
Sbjct: 990 VKEVMFY 996
>UniRef100_Q8LP00 ZLL/PNH homologous protein [Oryza sativa]
Length = 978
Score = 484 bits (1245), Expect = e-135
Identities = 324/932 (34%), Positives = 489/932 (51%), Gaps = 85/932 (9%)
Query: 37 LPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDG 96
L P +V A I P + K L R G G+ GA+ + NHF + + D
Sbjct: 97 LAAAPAVVVAPAARAVIGPPVASKGLSF---CRRPGFGTVGARCVVKANHFLAELPDKD- 152
Query: 97 YFFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFTIGSLAQ 155
QY V + E V + R I+ + Y S+L G+ AYDG K L+T G+L
Sbjct: 153 -LTQYDVKITPE----VSSRSVNRAIMSELVRLYHDSDLGGRLPAYDGRKNLYTAGTLPF 207
Query: 156 NKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQA 215
+ EF V L D DG G P R + Y+V I FA++ L
Sbjct: 208 DAREFVVRLTD----------DDDGTGVPP-----------REREYRVAIKFAARADLHH 246
Query: 216 IANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRG 275
+ + G + + QEA++VLDI+LR+ A ++ + + ++F+ D + +G G+ G
Sbjct: 247 LRQFIAGRQADAPQEALQVLDIVLRELANRR-YVSIGRSFYSPDIRKPQRLGDGLQSWCG 305
Query: 276 LHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF---LIANQNVRDPFSLDWN--KAKRTLK 330
+ S R TQ GLSLNID+S+T + P PV++F ++ + P S D N K K+ L+
Sbjct: 306 FYQSIRPTQMGLSLNIDMSSTAFIEPLPVIEFVAQILGKDVISRPLS-DANRIKIKKALR 364
Query: 331 NLRITTSP---TNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKI 387
+++ + ++Y+I+GL+ P + +F P +D +V YF
Sbjct: 365 GVKVEVTHRGNVRRKYRISGLTTQPTHELIF--------PIDDQMNMKSVVQYFKEMYGF 416
Query: 388 SLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMR 447
++Q+ LPC+ VG K+ ++P+E C +V QRYTK L+ Q +SL++ + ++P+E+
Sbjct: 417 TIQHP-HLPCLQVGNQKKANYLPMEACKIVEGQRYTKRLNEKQITSLLKVTCRRPREQEM 475
Query: 448 VLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNG---EDFNPRNGRWN 504
+ + + + Y +P + GI+I+ T V+ RVL AP LK+ + ++ P+ G+WN
Sbjct: 476 DILQS-QQNGYEQDPYAKEFGINISEKLTSVEARVLPAPWLKYHDTVKEKECLPQVGQWN 534
Query: 505 FNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMKGIHVEQPFDCFEENGQF 561
NKK++ K+ WA +NFS RG ++L + + G+ E
Sbjct: 535 MVNKKVINGCKVNHWACINFSRSVQETTARGFCQELAQMCQISGMEFNS-----EPVIPI 589
Query: 562 RRAPPLVRVEKMFEHVQS----KLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQ 617
A P +VEK +HV + KL G LL + N LYG K+ + G+++Q
Sbjct: 590 YSARP-DQVEKALKHVYNMSLNKLKGKELELLLAILPDNNGSLYGDIKRICETDLGLISQ 648
Query: 618 CIAPT---RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGS 674
C +++ QYL NV LKIN K+GG N++L S IP+VS PT+I G DV+H
Sbjct: 649 CCLTKHVFKISKQYLANVSLKINVKMGGRNTVLLDAISWRIPLVSDIPTIIFGADVTHPE 708
Query: 675 PGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTEDEGIIRE 730
G+ PSIAAVV+S+ WP ++KY V Q + E+I +L+K P T G+IRE
Sbjct: 709 TGEDSSPSIAAVVASQDWPEVTKYAGLVCAQAHRQELIQDLYKTWHDPQRGTVTGGMIRE 768
Query: 731 LLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVA 790
LLI F ++G +KP II +RDGVSE QF QVL EL I +AC L+ + P +V
Sbjct: 769 LLISFRKATG-QKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPIYQPPVTFVVV 827
Query: 791 QKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHY 842
QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG+ GTSRP HY
Sbjct: 828 QKRHHTRLFANNHKDRSSTDKSGNILPGTVVDSKICHPSEFDFYLCSHAGIQGTSRPAHY 887
Query: 843 HVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKFEDKSET 902
HVL DE F+ D++Q L ++L Y Y R T ++SVV P YAHLAA + +M+ E
Sbjct: 888 HVLWDENNFTADEMQTLTNNLCYTYARCTRSVSVVPPAYYAHLAAFRARFYMEPEMSENQ 947
Query: 903 SSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
++S +G N + + LP + + V MF+
Sbjct: 948 TTSKSSTG--TNGTSVKPLPAVKEKVKRVMFY 977
>UniRef100_Q851R2 Putative argonaute protein [Oryza sativa]
Length = 1058
Score = 482 bits (1241), Expect = e-134
Identities = 330/940 (35%), Positives = 490/940 (52%), Gaps = 80/940 (8%)
Query: 33 NEECLPPPPPIVPADIEP-----IKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHF 87
+E L PP + A P ++ + + AR G G+ G K+ + NHF
Sbjct: 160 SETALAPPAAVASAAAAPAGEASVESDKDLAPVSKKGLAHPARPGFGAAGKKVMIRANHF 219
Query: 88 KVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYG-SELNGKDLAYDGEKT 146
VNV D F Y V++ E + + R++L+ + + +G + L GK AYDG K+
Sbjct: 220 LVNVA--DNNLFHYDVSINPES----KSRATNREVLNELIKLHGKTSLGGKLPAYDGRKS 273
Query: 147 LFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEIS 206
L+T GSL EF V L D P D++R ++ YK+ I
Sbjct: 274 LYTAGSLPFESEEFVVKLID-----------------PEKKDKERAERE-----YKITIR 311
Query: 207 FASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDV 266
A + L + L G + + QE I+VLD++LR+ + + V ++FF + D+
Sbjct: 312 IAGRTDLYHLQQFLLGRQRDMPQETIQVLDVVLRE-SPSWNYVTVSRSFFSTQFGHRGDI 370
Query: 267 GGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRD---PFS-LDW 322
G G+ RG + S R TQ GLSLNID+S T P V+ F+ N+RD P S D
Sbjct: 371 GEGLECWRGYYQSLRPTQMGLSLNIDISATSFFKPVTVIQFVEEFLNIRDTSRPLSDRDR 430
Query: 323 NKAKRTLKNLRITTSPTNQE---YKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYD 379
K K+ L+ +RI T+ + YKITG++ +P +F P +D+ TV
Sbjct: 431 VKIKKALRGVRIETNHQEDQIRRYKITGITPIPMSQLIF--------PVDDNGTRKTVVQ 482
Query: 380 YFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSR 439
YF +R L+Y A PC+ G RP ++P+E+C +V QRY+K L+ Q ++++ +
Sbjct: 483 YFWDRYNYRLKY-ASWPCLQSGSDSRPVYLPMEVCKIVEGQRYSKKLNDKQVTNILRATC 541
Query: 440 QKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDF 496
Q+PQ+R + + + + + Y + + GI + + V RVL P LK+ G +
Sbjct: 542 QRPQQREQSIHEMVLHNKYTEDRFAQEFGIKVCNDLVSVPARVLPPPMLKYHDSGREKTC 601
Query: 497 NPRNGRWNFNNKKIVQPVKIEKWAVVNFSARC--DVRGLVRDLIKCGGMKGIHVE-QPFD 553
P G+WN NKK++ ++ W ++FS +V+ DLI+ G+ +P
Sbjct: 602 APSVGQWNMINKKMINGGTVDNWTCLSFSRMRPEEVQRFCGDLIQMCNATGMSFNPRPVV 661
Query: 554 CFEENGQFRRAPPLVRVEKMFEHVQSKL-PGAPKFLLCLLSERKNSDLYGPWKKKNLAEF 612
L V + + ++ G + L+ +L E S YG K+ +
Sbjct: 662 DVRSTNPNNIENALRDVHRRTSELLAREGKGGLQLLIVILPEVSGS--YGKIKRVCETDL 719
Query: 613 GIVTQCIAP---TRVNDQYLTNVLLKINAKLGGMNSLLGVEHSPS-IPIVSKAPTLILGM 668
GIV+QC P +R N QYL NV LKIN K+GG N++L + IP VS+ PT+I G
Sbjct: 720 GIVSQCCLPRHASRPNKQYLENVALKINVKVGGRNTVLERAFIRNGIPFVSEVPTIIFGA 779
Query: 669 DVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSD---TEDE 725
DV+H PG+ SIAAVV+S WP I+KYR V Q + E+I++LF D +
Sbjct: 780 DVTHPPPGEDSASSIAAVVASMDWPEITKYRGLVSAQPHRQEIIEDLFSVGKDPVKVVNG 839
Query: 726 GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKF 785
G+IRELLI F +G R+P+ II +RDGVSE QF+ VL E+ I +AC L+E + P
Sbjct: 840 GMIRELLIAFRKKTG-RRPERIIFYRDGVSEGQFSHVLLHEMDAIRKACASLEEGYLPPV 898
Query: 786 LVIVAQKNHHTKFFQP--GSPD------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTS 837
+V QK HHT+ F G D N+ PGTVVD +ICHP +DFY+C+HAG+ GTS
Sbjct: 899 TFVVVQKRHHTRLFPEVHGRRDMTDKSGNILPGTVVDRQICHPTEFDFYLCSHAGIQGTS 958
Query: 838 RPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKFE 897
RPTHYHVL DE F+ D LQ L ++L Y Y R T A+SVV P YAHLAA + +++ E
Sbjct: 959 RPTHYHVLYDENHFTADALQSLTNNLCYTYARCTRAVSVVPPAYYAHLAAFRARYYVEGE 1018
Query: 898 DKSETSSSHGGSGRDI---NASPIPQLPKLMDSVCNSMFF 934
S+ S+ G SG+ + + QLPK+ ++V + MF+
Sbjct: 1019 -SSDGGSTPGSSGQAVAREGPVEVRQLPKIKENVKDVMFY 1057
>UniRef100_Q7XSA2 OSJNBa0005N02.3 protein [Oryza sativa]
Length = 1101
Score = 481 bits (1239), Expect = e-134
Identities = 320/915 (34%), Positives = 466/915 (49%), Gaps = 94/915 (10%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G G+ G + + NHF + + D QY V++ E V +G R ++ +
Sbjct: 230 RPGKGTYGDRCIVKANHFFAELPDKD--LHQYDVSITPE----VTSRGVNRAVMFELVTL 283
Query: 130 YG-SELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD 188
Y S L G+ AYDG K+L+T G L F + L+D D G T
Sbjct: 284 YRYSHLGGRLPAYDGRKSLYTAGPLPFASRTFEITLQD----------EEDSLGGGQGTQ 333
Query: 189 RKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGC 248
R R + ++V I FA++ L +A L G + + QEA++VLDI+LR+ +
Sbjct: 334 R-------RERLFRVVIKFAARADLHHLAMFLAGRQADAPQEALQVLDIVLRELPTTRYS 386
Query: 249 LLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL 308
+ R +F+ + +G G+ RG + S R TQ GLSLNID+S+T + P PV+DF+
Sbjct: 387 PVGR-SFYSPNLGRRQQLGEGLESWRGFYQSIRPTQMGLSLNIDMSSTAFIEPLPVIDFV 445
Query: 309 IANQN----VRDPFSLDWNKAKRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTLK 361
N VR D K K+ L+ +++ + ++Y+I+GL+ ++ F +
Sbjct: 446 AQLLNRDISVRPLSDSDRVKIKKALRGVKVEVTHRGNMRRKYRISGLTSQATRELSFPVD 505
Query: 362 KRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQR 421
RG V TV YF+ S+Q++ LPC+ VG +RP ++P+E+C +V QR
Sbjct: 506 DRGTVK--------TVVQYFLETYGFSIQHTT-LPCLQVGNQQRPNYLPMEVCKIVEGQR 556
Query: 422 YTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGR 481
Y+K L+ Q ++L++ + Q+PQER + + + Y + + GI I V+ R
Sbjct: 557 YSKRLNEKQITALLKVTCQRPQERELDILRTVSHNAYHEDQYAQEFGIKIDERLASVEAR 616
Query: 482 VLQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDLI 538
VL PRLK+ G +D PR G+WN NKK+V ++ WA +NFS +
Sbjct: 617 VLPPPRLKYHDSGREKDVLPRVGQWNMMNKKMVNGGRVNNWACINFSRN----------V 666
Query: 539 KCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLP-------------GAP 585
+ +G E C F P L + EHV+ L G
Sbjct: 667 QDSAARGFCHELAIMCQISGMDFALEPVLPPLTARPEHVERALKARYQDAMNMLRPQGRE 726
Query: 586 KFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGG 642
LL ++ N LYG K+ + G+V+QC V + QYL NV LKIN K+GG
Sbjct: 727 LDLLIVILPDNNGSLYGDLKRICETDLGLVSQCCLTKHVFKMSKQYLANVALKINVKVGG 786
Query: 643 MNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACV 702
N++L + IP+VS PT+I G DV+H PG+ PSIAAVV+S+ WP ++KY V
Sbjct: 787 RNTVLVDALTRRIPLVSDRPTIIFGADVTHPHPGEDSSPSIAAVVASQDWPEVTKYAGLV 846
Query: 703 RTQGAKVEMIDNLFK----PVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQ 758
Q + E+I +LFK P T G+I+ELLI F ++G +KP II +RDGVSE Q
Sbjct: 847 SAQAHRQELIQDLFKVWQDPHRGTVTGGMIKELLISFKRATG-QKPQRIIFYRDGVSEGQ 905
Query: 759 FNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPG 810
F QVL EL I +AC L+ + P +V QK HHT+ F D N+ PG
Sbjct: 906 FYQVLLYELDAIRKACASLEPNYQPPVTFVVVQKRHHTRLFANNHNDQRTVDRSGNILPG 965
Query: 811 TVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRS 870
TVVD+KICHP +DFY+C+HAG+ GTSRP HYHVL DE F+ D+LQ L ++L Y Y R
Sbjct: 966 TVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENKFTADELQTLTNNLCYTYARC 1025
Query: 871 TTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGG-----------SGRDINASPIP 919
T ++S+V P YAHLAA + +M+ E S + G S R +
Sbjct: 1026 TRSVSIVPPAYYAHLAAFRARFYMEPETSDSGSMASGAATSRGLPPGVRSARVAGNVAVR 1085
Query: 920 QLPKLMDSVCNSMFF 934
LP L ++V MF+
Sbjct: 1086 PLPALKENVKRVMFY 1100
>UniRef100_Q5Z5B2 Putative AGO1 homologous protein [Oryza sativa]
Length = 1038
Score = 479 bits (1233), Expect = e-133
Identities = 322/905 (35%), Positives = 468/905 (51%), Gaps = 81/905 (8%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G GS G + + NHF + + D QY V++ E + + +++ + +
Sbjct: 174 RPGSGSIGTRCLVKANHFFAQLPDKD--LHQYDVSITPELTSRIRSRAVMEELVRLHKMS 231
Query: 130 YGSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDR 189
Y L G+ AYDG K+L+T G L EF + L + DG GS
Sbjct: 232 Y---LGGRLPAYDGRKSLYTAGPLPFTSKEFRISLLE----------EDDGSGS------ 272
Query: 190 KRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCL 249
R KTY V I FA++ L + L G + E QEA++VLDI+LR+ +
Sbjct: 273 -----ERRQKTYNVVIKFAARADLHRLEQFLAGRQAEAPQEALQVLDIVLRELPTARYAP 327
Query: 250 LVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLI 309
R +FF D +G G+ RG + S R TQ GLSLNID+S T P PV+DF+I
Sbjct: 328 FGR-SFFSPDLGRRRSLGEGLETWRGFYQSIRPTQMGLSLNIDMSATAFFEPLPVIDFVI 386
Query: 310 A--NQNVRD-PFS-LDWNKAKRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTLKK 362
N ++R P S + K K+ L+ +++ + ++Y+I+GL+ ++ F + +
Sbjct: 387 QLLNTDIRSRPLSDAERVKIKKALRGVKVGVTHRGNMRRKYRISGLTSQATRELTFPVDQ 446
Query: 363 RGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRY 422
G V +V YF ++Q++ LPC+ VG +RP ++P+E+C +V QRY
Sbjct: 447 GGTVK--------SVVQYFQETYGFAIQHTY-LPCLQVGNQQRPNYLPMEVCKIVEGQRY 497
Query: 423 TKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRV 482
+K L+ Q +L+E++ Q+P +R R + + + Y +P + GI I+ V+ R+
Sbjct: 498 SKRLNQNQIRALLEETCQRPHDRERDIIQMVNHNSYHEDPYAKEFGIKISERLALVEARI 557
Query: 483 LQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRD 536
L APRLK+ G +D PR G+WN NKK+V ++ W VNF+ G R+
Sbjct: 558 LPAPRLKYNETGREKDCLPRVGQWNMMNKKMVNGGRVRSWICVNFARNVQESVASGFCRE 617
Query: 537 LIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKF---LLCLL 592
L + G+ +P + R + R K H + G LL L
Sbjct: 618 LARMCQASGMDFALEPV----LPSMYARPDQVERALKARFHDAMNILGPQHKELDLLIGL 673
Query: 593 SERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGV 649
N LYG K+ + G+V+QC +V N Q L N+ LKIN K+GG N++L
Sbjct: 674 LPDNNGSLYGDLKRICEIDLGLVSQCCCTKQVFKMNKQILANLALKINVKVGGRNTVLVD 733
Query: 650 EHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKV 709
S IP+V+ PT+I G DV+H PG+ PSIAAVV+S+ WP ++KY V Q +
Sbjct: 734 AVSRRIPLVTDRPTIIFGADVTHPHPGEDSSPSIAAVVASQDWPEVTKYAGLVSAQSHRQ 793
Query: 710 EMIDNLFKPVSDTEDE----GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNI 765
E+ID+L+ D G++RELLI F S+G +KP II +RDGVSE QF QVL
Sbjct: 794 ELIDDLYNITHDPHRGPICGGMVRELLISFKRSTG-QKPQRIIFYRDGVSEGQFYQVLLH 852
Query: 766 ELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKI 817
EL I +AC L+ + P+ IV QK HHT+ F D N+ PGTVVD+KI
Sbjct: 853 ELDAIRKACASLEANYQPQVTFIVVQKRHHTRLFAHNHNDQNSVDRSGNILPGTVVDSKI 912
Query: 818 CHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVV 877
CHP +DF++C+HAG+ GTSRP HYHVL DE F+ D LQ L ++L Y Y R T ++S+V
Sbjct: 913 CHPTEFDFFLCSHAGIKGTSRPAHYHVLWDENNFTADALQTLTNNLCYTYARCTRSVSIV 972
Query: 878 APICYAHLAASQVGQFMKFE--DKSETSSSHGG------SGRDINASPIPQLPKLMDSVC 929
P YAHLAA + +M+ + D +S GG S R + LP L DSV
Sbjct: 973 PPAYYAHLAAFRARFYMESDSSDSGSMASGRGGGSSTSRSTRAAGGGAVRPLPALKDSVK 1032
Query: 930 NSMFF 934
N MF+
Sbjct: 1033 NVMFY 1037
>UniRef100_Q8LNZ9 AGO1 homologous protein [Oryza sativa]
Length = 909
Score = 476 bits (1225), Expect = e-132
Identities = 333/950 (35%), Positives = 481/950 (50%), Gaps = 108/950 (11%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGY 97
P PP P D I + P+ R G G+ G + + NHF ++ N D +
Sbjct: 14 PTPPQEQPVDAATTTPH-HIPSSSKSIRFPL-RPGKGTIGTRCMVKANHFFAHLPNKDLH 71
Query: 98 FFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFTIGSLAQN 156
+ S+ V + R ++ + Y S L G+ AYDG K+L+T G L
Sbjct: 72 HYDVSIT------PEVTSRIVNRAVIKELVNLYKASYLGGRLPAYDGRKSLYTAGPLPFT 125
Query: 157 KLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAI 216
EF + L D DG GS R +T++V I FA++ L +
Sbjct: 126 SQEFQITLLD----------DDDGSGS-----------ERRQRTFRVVIKFAARADLHRL 164
Query: 217 ANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGL 276
L G E QEA++VLDI+LR+ + + R +FF +G G+ RG
Sbjct: 165 ELFLAGRHAEAPQEALQVLDIVLRELPSARYAPFGR-SFFSPYLGRRQPLGEGLESWRGF 223
Query: 277 HSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL--IANQNV--RDPFSLDWNKAKRTLKNL 332
+ S R TQ GLSLNID+S T + P PV+DF+ + N ++ R F + K K+ L+ +
Sbjct: 224 YQSIRPTQMGLSLNIDMSATAFIEPLPVIDFVAQLLNSDIHSRSLFDAERVKIKKALRGV 283
Query: 333 RITTSPTN---QEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISL 389
++ + ++Y+I+GL+ P ++ F + + G V +V YF ++
Sbjct: 284 KVEVTHRGNMRRKYRISGLTIQPTRELTFPVDEGGTVK--------SVVQYFQETYGFAI 335
Query: 390 QYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVL 449
Q++ LPC+ V +R ++P+E+C +V QRY+K L+ Q +L+E++ Q P++R R +
Sbjct: 336 QHTY-LPCLTV---QRLNYLPMEVCKIVEGQRYSKRLNQNQIRALLEETCQHPRDRERDI 391
Query: 450 TDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPRNGRWNFN 506
+K + Y +P + GI I+ V+ R+L APRLK+ G +D PR G+WN
Sbjct: 392 IKMVKHNAYQDDPYAKEFGIKISDRLASVEARILPAPRLKYNETGREKDCLPRVGQWNMM 451
Query: 507 NKKIVQPVKIEKWAVVNFSARCD---VRGLVRDL-IKCGGMKGIHVEQPFDCFEENGQFR 562
NKK+V K+ W VNF+ VRG +L + C +P
Sbjct: 452 NKKMVNGGKVRSWMCVNFARNVQESVVRGFCHELALMCQASGMDFAPEPI---------- 501
Query: 563 RAPPL-------VRVEKMFEHVQSKLPGAPKFLLCLLSER---KNSDLYGPWKKKNLAEF 612
PPL R K H + G + L LL + N LYG K+ +
Sbjct: 502 -LPPLNAHPDQVERALKARYHDAMNVLGPQRRELDLLIGKLPDNNGSLYGDLKRVCEIDL 560
Query: 613 GIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMD 669
GIV+QC +V N Q L N+ LKIN K+GG N++L S IP+V+ PT+I G D
Sbjct: 561 GIVSQCCCTKQVFKMNKQILANLALKINVKVGGRNTVLVDAVSRRIPLVTDRPTIIFGAD 620
Query: 670 VSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTEDE 725
V+H PG+ PSIAAVV+S+ WP ++KY V Q + E+I++L+K P T
Sbjct: 621 VTHPHPGEDSSPSIAAVVASQDWPEVTKYAGLVSAQAHRQELIEDLYKIWQDPQRGTVSG 680
Query: 726 GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKF 785
G+IRELLI F S+G KP II +RDGVSE QF QVL EL+ I +AC L+ + PK
Sbjct: 681 GMIRELLISFKRSTGE-KPQRIIFYRDGVSEGQFYQVLLYELNAIRKACASLETNYQPKV 739
Query: 786 LVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTS 837
IV QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG+ GTS
Sbjct: 740 TFIVVQKRHHTRLFAHNHNDQNSVDRSGNILPGTVVDSKICHPTEFDFYLCSHAGIKGTS 799
Query: 838 RPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKFE 897
RP HYHVL +E F+ D LQ L ++L Y Y R T ++S+V P YAHLAA + +M+
Sbjct: 800 RPAHYHVLWNENNFTADALQILTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYME-P 858
Query: 898 DKSETSSSHGGSG-------------RDINASPIPQLPKLMDSVCNSMFF 934
D S++SS G G R + + LP L DSV MF+
Sbjct: 859 DTSDSSSVVSGPGVRGPLSGSSTSRTRAPGGAAVKPLPALKDSVKRVMFY 908
>UniRef100_Q9XGW1 PINHEAD protein [Arabidopsis thaliana]
Length = 988
Score = 476 bits (1224), Expect = e-132
Identities = 321/948 (33%), Positives = 485/948 (50%), Gaps = 104/948 (10%)
Query: 39 PPPP----------IVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFK 88
PPPP +I + + Q+ +K P R G G+ G K + NHF
Sbjct: 92 PPPPSQTTSSAVSVATAGEIVAVNHQMQMGVRKNSNFAP--RPGFGTLGTKCIVKANHFL 149
Query: 89 VNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDL-AYDGEKTL 147
++ D QY V + E V K R I+ + Y G+ L AYDG K+L
Sbjct: 150 ADLPTKD--LNQYDVTITPE----VSSKSVNRAIIAELVRLYKESDLGRRLPAYDGRKSL 203
Query: 148 FTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISF 207
+T G L EF+V + D NG R ++YKV I F
Sbjct: 204 YTAGELPFTWKEFSVKIVDEDDGIING--------------------PKRERSYKVAIKF 243
Query: 208 ASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVG 267
++ + + L G + QEA+++LDI+LR+ + K+ C + R +FF D K +G
Sbjct: 244 VARANMHHLGEFLAGKRADCPQEAVQILDIVLRELSVKRFCPVGR-SFFSPDIKTPQRLG 302
Query: 268 GGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF---LIANQNVRDPFS-LDWN 323
G+ G + S R TQ GLSLNID+++ + P PV++F L+ + P S D
Sbjct: 303 EGLESWCGFYQSIRPTQMGLSLNIDMASAAFIEPLPVIEFVAQLLGKDVLSKPLSDSDRV 362
Query: 324 KAKRTLKNLRITTSP---TNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDY 380
K K+ L+ +++ + ++Y++ GL+ P ++ +F P +++ +V +Y
Sbjct: 363 KIKKGLRGVKVEVTHRANVRRKYRVAGLTTQPTRELMF--------PVDENCTMKSVIEY 414
Query: 381 FVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQ 440
F ++Q++ LPC+ VG K+ +++P+E C +V QRYTK L+ Q ++L++ + Q
Sbjct: 415 FQEMYGFTIQHT-HLPCLQVGNQKKASYLPMEACKIVEGQRYTKRLNEKQITALLKVTCQ 473
Query: 441 KPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFN 497
+P++R + ++ + Y +P + G++I+ V+ R+L AP LK+ G +D
Sbjct: 474 RPRDRENDILRTVQHNAYDQDPYAKEFGMNISEKLASVEARILPAPWLKYHENGKEKDCL 533
Query: 498 PRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMKGIHVEQPFDC 554
P+ G+WN NKK++ + + +WA VNFS RG +L + + G+
Sbjct: 534 PQVGQWNMMNKKMINGMTVSRWACVNFSRSVQENVARGFCNELGQMCEVSGMEFNP---- 589
Query: 555 FEENGQFRRAPPLVRVEKMFEHV----QSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLA 610
E A P +VEK +HV +K G LL + N LYG K+
Sbjct: 590 -EPVIPIYSARP-DQVEKALKHVYHTSMNKTKGKELELLLAILPDNNGSLYGDLKRICET 647
Query: 611 EFGIVTQCIAPT---RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILG 667
E G+++QC +++ QYL NV LKIN K+GG N++L S IP+VS PT+I G
Sbjct: 648 ELGLISQCCLTKHVFKISKQYLANVSLKINVKMGGRNTVLVDAISCRIPLVSDIPTIIFG 707
Query: 668 MDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTE 723
DV+H G+ PSIAAVV+S+ WP ++KY V Q + E+I +L+K PV T
Sbjct: 708 ADVTHPENGEESSPSIAAVVASQDWPEVTKYAGLVCAQAHRQELIQDLYKTWQDPVRGTV 767
Query: 724 DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNP 783
G+IR+LLI F ++G +KP II +RDGVSE QF QVL EL I +AC L+ + P
Sbjct: 768 SGGMIRDLLISFRKATG-QKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPNYQP 826
Query: 784 KFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIG 835
IV QK HHT+ F D N+ PGTVVD KICHP +DFY+C+HAG+ G
Sbjct: 827 PVTFIVVQKRHHTRLFANNHRDKNSTDRSGNILPGTVVDTKICHPTEFDFYLCSHAGIQG 886
Query: 836 TSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMK 895
TSRP HYHVL DE F+ D +Q L ++L Y Y R T ++S+V P YAHLAA + +
Sbjct: 887 TSRPAHYHVLWDENNFTADGIQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA----R 942
Query: 896 FEDKSETSSSHGGSGR---------DINASPIPQLPKLMDSVCNSMFF 934
F + E +G G+ D+ P LP L ++V MF+
Sbjct: 943 FYLEPEIMQDNGSPGKKNTKTTTVGDVGVKP---LPALKENVKRVMFY 987
>UniRef100_Q6EU14 Putative argonaute protein [Oryza sativa]
Length = 1082
Score = 474 bits (1221), Expect = e-132
Identities = 329/972 (33%), Positives = 480/972 (48%), Gaps = 113/972 (11%)
Query: 19 LMVMCSVQEMGSQENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGA 78
L V S +E+ Q E + P A I+P K + PM R G G+ G
Sbjct: 167 LPVEASSEEVQHQFQELAIQGQSPTSQA------IQPAPPSSK-SVRFPM-RPGKGTFGD 218
Query: 79 KLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYG-SELNGK 137
+ + NHF + + D QY V++ E V +G R ++ + Y S L G+
Sbjct: 219 RCIVKANHFFAELPDKD--LHQYDVSITPE----VPSRGVNRAVIGEIVTQYRQSHLGGR 272
Query: 138 DLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHR 197
YDG K+L+T G L F V+L+D + G + R
Sbjct: 273 LPVYDGRKSLYTAGPLPFTSRTFDVILQDEEESLAVGQGA-----------------QRR 315
Query: 198 SKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFH 257
+ +KV I FA++ L +A L G + + QEA++VLDI+LR+ + + R +F+
Sbjct: 316 ERPFKVVIKFAARADLHHLAMFLAGRQADAPQEALQVLDIVLRELPTARYSPVAR-SFYS 374
Query: 258 NDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQN---- 313
+ +G G+ RG + S R TQ GLSLNID+S+T + P PV+DF+ N
Sbjct: 375 PNLGRRQQLGEGLESWRGFYQSIRPTQMGLSLNIDMSSTAFIEPLPVIDFVAQLLNRDIS 434
Query: 314 VRDPFSLDWNKAKRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTLKKRGAVPGED 370
VR D K K+ L+ +++ + ++Y+I+GL+ ++ F + G V
Sbjct: 435 VRPLSDADRVKIKKALRGVKVEVTHRGNMRRKYRISGLTSQATRELSFPIDNHGTVK--- 491
Query: 371 DTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQ 430
TV YF +++++ LPC+ VG +RP ++P+E+C +V QRY+K L+ Q
Sbjct: 492 -----TVVQYFQETYGFNIKHTT-LPCLQVGNQQRPNYLPMEVCKIVEGQRYSKRLNEKQ 545
Query: 431 RSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF 490
++L++ + Q+PQER + + + Y +P + GI I V+ RVL P LK+
Sbjct: 546 ITALLKVTCQRPQERELDILQTVHHNAYHQDPYAQEFGIRIDERLASVEARVLPPPWLKY 605
Query: 491 ---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMK 544
G +D PR G+WN NKK+V ++ W +NFS R R+L +
Sbjct: 606 HDSGREKDVLPRIGQWNMMNKKMVNGGRVNNWTCINFSRHVQDNAARSFCRELAIMCQIS 665
Query: 545 GIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLP-------------GAPKFLLCL 591
G+ F P + V EHV+ L G LL
Sbjct: 666 GM-------------DFSIDPVVPLVTARPEHVERALKARYQEAMNILKPQGGELDLLIA 712
Query: 592 LSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLG 648
+ N LYG K+ + G+V+QC V + QYL NV LKIN K+GG N++L
Sbjct: 713 ILPDNNGSLYGDLKRICETDLGLVSQCCLTKHVFKMSKQYLANVALKINVKVGGRNTVLV 772
Query: 649 VEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAK 708
+ IP+VS PT+I G DV+H PG+ PSIAAVV+S+ WP ++KY V Q +
Sbjct: 773 DALTRRIPLVSDRPTIIFGADVTHPHPGEDSSPSIAAVVASQDWPEVTKYAGLVSAQAHR 832
Query: 709 VEMIDNLFK----PVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLN 764
E+I +LFK P T G+IRELLI F ++G +KP II +RDGVSE QF QVL
Sbjct: 833 QELIQDLFKVWKDPQRGTVSGGMIRELLISFKRATG-QKPQRIIFYRDGVSEGQFYQVLF 891
Query: 765 IELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNK 816
EL I +AC L+ + P +V QK HHT+ F D N+ PGTVVD+K
Sbjct: 892 YELDAIRKACASLEADYQPPVTFVVVQKRHHTRLFANNHKDQRTVDRSGNILPGTVVDSK 951
Query: 817 ICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISV 876
ICHP +DFY+C+HAG+ GTSRP HYHVL DE F+ D LQ L ++L Y Y R T ++S+
Sbjct: 952 ICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSI 1011
Query: 877 VAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIP--------------QLP 922
V P YAHLAA + +M+ + S + G R P+P LP
Sbjct: 1012 VPPAYYAHLAAFRARFYMEPDTSDSGSMASGAHTR--GGGPLPGARSTKPAGNVAVRPLP 1069
Query: 923 KLMDSVCNSMFF 934
L ++V MF+
Sbjct: 1070 DLKENVKRVMFY 1081
>UniRef100_Q6K972 AGO1 homologous protein [Oryza sativa]
Length = 1011
Score = 474 bits (1219), Expect = e-132
Identities = 334/950 (35%), Positives = 480/950 (50%), Gaps = 108/950 (11%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGY 97
P PP P D I + P+ R G G+ G + + NHF ++ N D +
Sbjct: 116 PTPPQEQPVDAATTTPH-HIPSSSKSIRFPL-RPGKGTIGTRCMVKANHFFAHLPNKDLH 173
Query: 98 FFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFTIGSLAQN 156
+ S+ V + R ++ + Y S L G+ AYDG K+L+T G L
Sbjct: 174 HYDVSIT------PEVTSRIVNRAVIKELVNLYKASYLGGRLPAYDGRKSLYTAGPLPFT 227
Query: 157 KLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAI 216
EF + L D DG GS R +T++V I FA++ L +
Sbjct: 228 SQEFQITLLD----------DDDGSGS-----------ERRQRTFRVVIKFAARADLHRL 266
Query: 217 ANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGL 276
L G E QEA++VLDI+LR+ + + R +FF +G G+ RG
Sbjct: 267 ELFLAGRHAEAPQEALQVLDIVLRELPSARYAPFGR-SFFSPYLGRRQPLGEGLESWRGF 325
Query: 277 HSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL--IANQNVRD-PFS-LDWNKAKRTLKNL 332
+ S R TQ GLSLNID+S T + P PV+DF+ + N ++ P S + K K+ L+ +
Sbjct: 326 YQSIRPTQMGLSLNIDMSATAFIEPLPVIDFVAQLLNSDIHSRPLSDAERVKIKKALRGV 385
Query: 333 RITTSPTN---QEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISL 389
++ + ++Y+I+GL+ P ++ F + + G V +V YF ++
Sbjct: 386 KVEVTHRGNMRRKYRISGLTIQPTRELTFPVDEGGTVK--------SVVQYFQETYGFAI 437
Query: 390 QYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVL 449
Q++ LPC+ V +R ++P+E+C +V QRY+K L+ Q +L+E++ Q P++R R +
Sbjct: 438 QHTY-LPCLTV---QRLNYLPMEVCKIVEGQRYSKRLNQNQIRALLEETCQHPRDRERDI 493
Query: 450 TDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPRNGRWNFN 506
+K + Y +P + GI I+ V+ R+L APRLK+ G +D PR G+WN
Sbjct: 494 IKMVKHNAYQDDPYAKEFGIKISDRLASVEARILPAPRLKYNETGREKDCLPRVGQWNMM 553
Query: 507 NKKIVQPVKIEKWAVVNFSARCD---VRGLVRDL-IKCGGMKGIHVEQPFDCFEENGQFR 562
NKK+V K+ W VNF+ VRG +L + C +P
Sbjct: 554 NKKMVNGGKVRSWMCVNFARNVQESVVRGFCHELALMCQASGMDFAPEPI---------- 603
Query: 563 RAPPL-------VRVEKMFEHVQSKLPGAPKFLLCLLS---ERKNSDLYGPWKKKNLAEF 612
PPL R K H + G + L LL N LYG K+ +
Sbjct: 604 -LPPLNAHPDQVERALKARYHDAMNVLGPQRRELDLLIGILPDNNGSLYGDLKRVCEIDL 662
Query: 613 GIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMD 669
GIV+QC +V N Q L N+ LKIN K+GG N++L S IP+V+ PT+I G D
Sbjct: 663 GIVSQCCCTKQVFKMNKQILANLALKINVKVGGRNTVLVDAVSRRIPLVTDRPTIIFGAD 722
Query: 670 VSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTEDE 725
V+H PG+ PSIAAVV+S+ WP ++KY V Q + E+I++L+K P T
Sbjct: 723 VTHPHPGEDSSPSIAAVVASQDWPEVTKYAGLVSAQAHRQELIEDLYKIWQDPQRGTVSG 782
Query: 726 GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKF 785
G+IRELLI F S+G KP II +RDGVSE QF QVL EL+ I +AC L+ + PK
Sbjct: 783 GMIRELLISFKRSTGE-KPQRIIFYRDGVSEGQFYQVLLYELNAIRKACASLETNYQPKV 841
Query: 786 LVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTS 837
IV QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG+ GTS
Sbjct: 842 TFIVVQKRHHTRLFAHNHNDQNSVDRSGNILPGTVVDSKICHPTEFDFYLCSHAGIKGTS 901
Query: 838 RPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKFE 897
RP HYHVL DE F+ D LQ L ++L Y Y R T ++S+V P YAHLAA + +M+
Sbjct: 902 RPAHYHVLWDENNFTADALQILTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYME-P 960
Query: 898 DKSETSSSHGGSG-------------RDINASPIPQLPKLMDSVCNSMFF 934
D S++SS G G R + + LP L DSV MF+
Sbjct: 961 DTSDSSSVVSGPGVRGPLSGSSTSRTRAPGGAAVKPLPALKDSVKRVMFY 1010
>UniRef100_Q7Y001 Putative leaf development and shoot apical meristem regulating
protein [Oryza sativa]
Length = 1055
Score = 469 bits (1208), Expect = e-130
Identities = 316/938 (33%), Positives = 475/938 (49%), Gaps = 102/938 (10%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGY 97
PPP + P+ + + AR +G+ G + + NHF V V + D Y
Sbjct: 178 PPPATLPPSSSKAVTFP--------------ARPDVGTIGRRCRVRANHFLVQVADKDIY 223
Query: 98 FFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLAQNK 157
Y V + E + R I++++ + L+G+ YDG K+++T G L
Sbjct: 224 --HYDVVITPESTY----RERNRSIINKLVALHKQFLDGRLPVYDGRKSIYTAGPLPFKT 277
Query: 158 LEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIA 217
+F V H +P L+ + R + YKV I ASK L ++
Sbjct: 278 KDFVVK-----------------HINP-------LRGNQREEEYKVTIKQASKTDLYSLK 313
Query: 218 NALKGHETENYQEAIRVLDIILRQHAAKQGC-----LLVRQNFFHNDPKNFTDVGGGVLG 272
L G + E Q+ I+ LDI LR+ + + ++FF + ++G G
Sbjct: 314 QFLVGRQRELPQDTIQALDIALRECPTSVNFTCDRYVSISRSFFSQSFGHGGEIGSGTEC 373
Query: 273 CRGLHSSFRTTQSGLSLNI------DVSTTMIVHPGPVVDFLIANQNVRDPF----SLDW 322
RG + S R TQ GLSLNI ++S T PV+DF + N+RD D
Sbjct: 374 WRGYYQSLRPTQMGLSLNIGMDLPQNISATAFYKAQPVMDFAVQYLNIRDVSRRLSDQDR 433
Query: 323 NKAKRTLKNLRITTSPTNQE---YKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYD 379
K K+ LK ++I + ++ YKITG+ P + +F L D I+V
Sbjct: 434 IKLKKALKGVQIVATHWKEKSIRYKITGIPSAPMNELMFDL----------DGNRISVVQ 483
Query: 380 YFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSR 439
YF + SL++ + PC+ G RP ++P+E+CS++ QRY+K L+ Q ++++ +
Sbjct: 484 YFKKQYNYSLKH-VNWPCLQAGSDSRPKYLPMEVCSILEGQRYSKKLNEHQVTNILRMTC 542
Query: 440 QKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDF 496
++P +R + + + T+ YG++ + GI + + VD RVL PRLK+ G +
Sbjct: 543 ERPAQRESSIIEIVNTNSYGNDDCAKEFGIKVANQLAVVDARVLPTPRLKYHDSGREKVC 602
Query: 497 NPRNGRWNFNNKKIVQPVKIEKWAVVNFSARC---DVRGLVRDLIKCGGMKGIHVEQPFD 553
NP G+WN NK++V I W ++F++R D+R DL+ G+ +
Sbjct: 603 NPSVGQWNMINKRMVNGGCINHWTCLSFASRMHVNDIRMFCEDLVGMCNNIGMQMNTRPC 662
Query: 554 CFEENGQFRRAPPLVRV---EKMFEHVQSKLPGAP-KFLLCLLSERKNSDLYGPWKKKNL 609
GQ R +R + + Q L G + L+ +L+E S YG K+
Sbjct: 663 VDIIQGQQRNIEGAIRNIHRQSSEKLDQQDLTGQQLQLLIVILTEISGS--YGRIKRICE 720
Query: 610 AEFGIVTQCIAPTRVND---QYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLIL 666
E G++TQC AP + QYL N+ LK+N K+GG N++L IPI++ PT++
Sbjct: 721 TEVGVITQCCAPKSLQKGGKQYLENLALKMNVKVGGRNTVLEDALHKKIPILTDRPTIVF 780
Query: 667 GMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDE- 725
G DV+H SPG+ PSIAAVV+S WP ++KY+ V TQ + E+I NL+ V D
Sbjct: 781 GADVTHPSPGEDASPSIAAVVASMDWPEVTKYKCLVSTQSHREEIISNLYTEVKDPLKGI 840
Query: 726 ---GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWN 782
G+IRELL FY +G +KP II +RDG+SE QF+QVL E+ I +AC L E +
Sbjct: 841 IRGGMIRELLRSFYQETG-QKPSRIIFYRDGISEGQFSQVLLYEMDAIRKACASLQEGYL 899
Query: 783 PKFLVIVAQKNHHTKFFQPGSPD------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGT 836
P +V QK HHT+ F D N+ PGTVVD ICHP +DFY+C+H+G+ GT
Sbjct: 900 PPVTFVVVQKRHHTRLFPENRRDMMDRSGNILPGTVVDTMICHPSEFDFYLCSHSGIKGT 959
Query: 837 SRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMKF 896
SRPTHYHVLLDE GF D LQ L ++LSY Y R T A+S+V P YAHL A + +M+
Sbjct: 960 SRPTHYHVLLDENGFKADTLQTLTYNLSYTYARCTRAVSIVPPAYYAHLGAFRARYYMED 1019
Query: 897 EDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
E + SSS + D + P LP++ ++V MF+
Sbjct: 1020 EHSDQGSSSSVTTRTDRSTKP---LPEIKENVKRFMFY 1054
>UniRef100_O04379 Argonaute protein [Arabidopsis thaliana]
Length = 1048
Score = 464 bits (1193), Expect = e-129
Identities = 319/954 (33%), Positives = 482/954 (50%), Gaps = 108/954 (11%)
Query: 39 PPPPIVPADIEPIKIE---PQIVKKKLPT-----KVPMARRGLGSKGAKLPLLTNHFKVN 90
P P ++ E + +E P + +P+ K PM R G G G + + NHF
Sbjct: 144 PEPTVLAQQFEQLSVEQGAPSQAIQPIPSSSKAFKFPM-RPGKGQSGKRCIVKANHFFAE 202
Query: 91 VTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFT 149
+ + D Y V + E V +G R ++ ++ + Y S L + AYDG K+L+T
Sbjct: 203 LPDKD--LHHYDVTITPE----VTSRGVNRAVMKQLVDNYRDSHLGSRLPAYDGRKSLYT 256
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
G L N EF + L D G G R + +KV I +
Sbjct: 257 AGPLPFNSKEFRINLLD----------EEVGAGG-----------QRREREFKVVIKLVA 295
Query: 210 KIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGG 269
+ L + L+G +++ QEA++VLDI+LR+ + + V ++F+ D +G G
Sbjct: 296 RADLHHLGMFLEGKQSDAPQEALQVLDIVLRELPTSR-YIPVGRSFYSPDIGKKQSLGDG 354
Query: 270 VLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL--IANQNVRD-PFS-LDWNKA 325
+ RG + S R TQ GLSLNID+S+T + PV+ F+ + N+++ P S D K
Sbjct: 355 LESWRGFYQSIRPTQMGLSLNIDMSSTAFIEANPVIQFVCDLLNRDISSRPLSDADRVKI 414
Query: 326 KRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFV 382
K+ L+ +++ + ++Y+I+GL+ + ++ F + +R + +V +YF
Sbjct: 415 KKALRGVKVEVTHRGNMRRKYRISGLTAVATRELTFPVDERNT--------QKSVVEYFH 466
Query: 383 NRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKP 442
+Q++ LPC+ VG RP ++P+E+C +V QRY+K L+ Q ++L++ + Q+P
Sbjct: 467 ETYGFRIQHT-QLPCLQVGNSNRPNYLPMEVCKIVEGQRYSKRLNERQITALLKVTCQRP 525
Query: 443 QERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPR 499
+R + + ++ +DY + + GI I++ V+ R+L P LK+ G P+
Sbjct: 526 IDREKDILQTVQLNDYAKDNYAQEFGIKISTSLASVEARILPPPWLKYHESGREGTCLPQ 585
Query: 500 NGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENG 559
G+WN NKK++ + W +NFS + L R + E C+
Sbjct: 586 VGQWNMMNKKMINGGTVNNWICINFSRQVQ-DNLARTFCQ---------ELAQMCYVSGM 635
Query: 560 QFRRAPPLVRVEKMFEHVQSKLP------------GAPKFLLCLLSERKNSDLYGPWKKK 607
F P L V E V+ L G LL ++ N LYG K+
Sbjct: 636 AFNPEPVLPPVSARPEQVEKVLKTRYHDATSKLSQGKEIDLLIVILPDNNGSLYGDLKRI 695
Query: 608 NLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTL 664
E GIV+QC V + QY+ NV LKIN K+GG N++L S IP+VS PT+
Sbjct: 696 CETELGIVSQCCLTKHVFKMSKQYMANVALKINVKVGGRNTVLVDALSRRIPLVSDRPTI 755
Query: 665 ILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTED 724
I G DV+H PG+ PSIAAVV+S+ WP I+KY V Q + E+I +LFK D +
Sbjct: 756 IFGADVTHPHPGEDSSPSIAAVVASQDWPEITKYAGLVCAQAHRQELIQDLFKEWKDPQK 815
Query: 725 E----GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEK 780
G+I+ELLI F S+G+ KP II +RDGVSE QF QVL EL I +AC L+
Sbjct: 816 GVVTGGMIKELLIAFRRSTGH-KPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEAG 874
Query: 781 WNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAG 832
+ P +V QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG
Sbjct: 875 YQPPVTFVVVQKRHHTRLFAQNHNDRHSVDRSGNILPGTVVDSKICHPTEFDFYLCSHAG 934
Query: 833 MIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQ 892
+ GTSRP HYHVL DE F+ D LQ L ++L Y Y R T ++S+V P YAHLAA +
Sbjct: 935 IQGTSRPAHYHVLWDENNFTADGLQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARF 994
Query: 893 FMKFEDKSETSSSHGGSGR------------DINASPIPQLPKLMDSVCNSMFF 934
+M+ E S + G R ++NA+ P LP L ++V MF+
Sbjct: 995 YMEPETSDSGSMASGSMARGGGMAGRSTRGPNVNAAVRP-LPALKENVKRVMFY 1047
>UniRef100_Q6Z4F1 Putative leaf development protein Argonaute [Oryza sativa]
Length = 1052
Score = 444 bits (1142), Expect = e-123
Identities = 315/950 (33%), Positives = 476/950 (49%), Gaps = 124/950 (13%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGY 97
P PP PA PI + + V P +R G G+ G ++ + NHF V V++ D
Sbjct: 173 PAPPAAPPA---PIPVSSKGV-------APPSRPGFGTVGERIVVRANHFLVRVSDND-M 221
Query: 98 FFQYSVALFYEDGRPVEGKGAGRKILDRVQETYG-SELNGKDLAYDGEKTLFTIGSLAQN 156
+ Y V+L P + + R ++ + + S L G AYDG K L+T G L +
Sbjct: 222 IYLYDVSL----SPPPKTRRINRVVMSELARLHRESHLGGISFAYDGSKALYTAGKLPFD 277
Query: 157 KLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAI 216
++F + +L K R YKV I A + L +
Sbjct: 278 SMDFKI----------------------------KLGKELREIEYKVTIRRAGQADLHHL 309
Query: 217 ANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGL 276
+ G + ++ Q+ I+ LD++LR+ + ++ R F++ D+G G+ +G
Sbjct: 310 HEFIAGRQRDSQQQTIQALDVVLRESPSLNYVIVSRS--FYSTMFGRQDIGDGLECWKGY 367
Query: 277 HSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLI------ANQNVRDP----FSLDWNKAK 326
+ S R TQ GLSLNID+S+T P VV+++ N N DP +D K K
Sbjct: 368 YQSLRPTQMGLSLNIDISSTPFFKPISVVEYVKNCLGTPTNANGPDPRRPLSDIDRLKVK 427
Query: 327 RTLKNLRITTSPTNQ--EYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNR 384
+ L+ +R+ T+ + +YKIT ++ P F++ D T + TV YF R
Sbjct: 428 KALRGVRVETTHQGKSSKYKITTITSEPLSQLNFSM---------DGTTQ-TVIQYFSQR 477
Query: 385 RKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQE 444
K LQY++ PC+ G P P ++P+E+C++V QRY+K L+ Q + L+ + Q PQ+
Sbjct: 478 YKYRLQYTS-WPCLQSGNPSNPIYLPMEVCTIVEGQRYSKKLNDKQVTGLLRATCQPPQK 536
Query: 445 RMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPRNG 501
R + + + ++ ++Y ++ ++ + I+I++ + RVL AP L++ G + NPR G
Sbjct: 537 REQKIIEMVQHNNYPADKVVSDFRINISNQMATMPARVLPAPTLRYHDSGKEKTCNPRVG 596
Query: 502 RWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQF 561
+WN NKK+V ++KW VNFS R + + R CG E + C F
Sbjct: 597 QWNMINKKMVGGAVVQKWTCVNFS-RMHIDAVHR---LCG-------ELVYTCNAIGMVF 645
Query: 562 RRAPPLVRVEKMFEHVQSKLPG----APKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQ 617
P + ++++ L AP+ L ++ + YG K+ E GIV+Q
Sbjct: 646 NEMPEIEVGSAAPNNIEAALSNIHTRAPQLQLLIVILPDVNGYYGRIKRVCETELGIVSQ 705
Query: 618 CIAPTR----VNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHG 673
C+ P R ++ Q+L NV LKIN K GG NS+L P +P + T+I G DV+H
Sbjct: 706 CLKPGRKLLSLDRQFLENVSLKINVKAGGRNSVL---QRPLVPGGLENTTIIFGADVTHP 762
Query: 674 SPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----------PVSDTE 723
+ G+ SIAAVV+S WP I+KY+A V Q + E+I +LF P E
Sbjct: 763 ASGEDSSASIAAVVASMDWPEITKYKALVSAQPPRQEIIQDLFTMTEVAQNADAPAQKAE 822
Query: 724 DE-------GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKF 776
G+ RELL+ FY+ + RKP II +RDGVS+ QF VL E+ I +A
Sbjct: 823 GSKKNFICGGMFRELLMSFYSKNAKRKPQRIIFYRDGVSDGQFLHVLLYEMDAIKKAIAS 882
Query: 777 LDEKWNPKFLVIVAQKNHHTKFFQP--GSPD------NVPPGTVVDNKICHPRNYDFYMC 828
LD + P +V QK HHT+ F G D NV PGTVVD ICHP +DFY+C
Sbjct: 883 LDPAYRPLVTFVVVQKRHHTRLFPEVHGRQDLTDRSGNVRPGTVVDTNICHPSEFDFYLC 942
Query: 829 AHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAAS 888
+HAG+ GTSRPTHYHVL DE FS D LQ L ++L Y Y R T ++SVV P YAHLAA
Sbjct: 943 SHAGIQGTSRPTHYHVLHDENRFSADQLQMLTYNLCYTYARCTRSVSVVPPAYYAHLAAF 1002
Query: 889 QVGQFMKFEDKSETSSSHGGSGRDINASPIP----QLPKLMDSVCNSMFF 934
+ ++ + +SS G G A P +LP++ ++V + MF+
Sbjct: 1003 R-ARYYDEPPAMDGASSVGSGGNQAAAGGQPPAVRRLPQIKENVKDVMFY 1051
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,640,137,921
Number of Sequences: 2790947
Number of extensions: 73823750
Number of successful extensions: 185582
Number of sequences better than 10.0: 309
Number of HSP's better than 10.0 without gapping: 265
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 183853
Number of HSP's gapped (non-prelim): 499
length of query: 935
length of database: 848,049,833
effective HSP length: 137
effective length of query: 798
effective length of database: 465,690,094
effective search space: 371620695012
effective search space used: 371620695012
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC124972.13


