

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124971.1 + phase: 0
(397 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
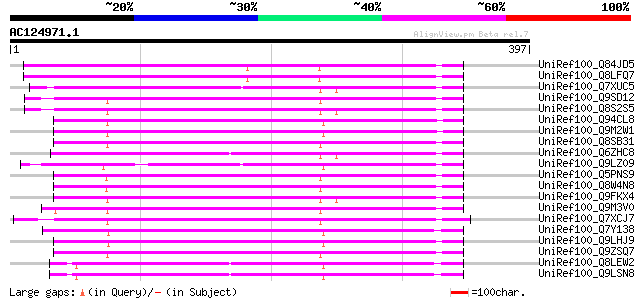
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84JD5 Hypothetical protein At5g06750 [Arabidopsis tha... 220 6e-56
UniRef100_Q8LFQ7 Protein phosphatase 2C-like [Arabidopsis thaliana] 218 2e-55
UniRef100_Q7XUC5 OSJNBa0088A01.23 protein [Oryza sativa] 213 1e-53
UniRef100_Q9SD12 Protein phosphatase 2C-like protein [Arabidopsi... 211 4e-53
UniRef100_Q8S2S5 Protein phosphatase 2c-like protein [Thellungie... 210 5e-53
UniRef100_Q94CL8 Ser/Thr protein phosphatase 2C [Arabidopsis tha... 207 5e-52
UniRef100_Q9M2W1 Protein phosphatase 2C-like protein [Arabidopsi... 207 5e-52
UniRef100_Q8SB31 Hypothetical protein OJ1540_H01.4 [Oryza sativa] 207 5e-52
UniRef100_Q6ZHC8 Hypothetical protein OJ1717_A09.24 [Oryza sativa] 205 2e-51
UniRef100_Q9LZ09 Protein phosphatase-like protein [Arabidopsis t... 203 6e-51
UniRef100_Q5PNS9 At4g38520 [Arabidopsis thaliana] 202 2e-50
UniRef100_Q8W4N8 Hypothetical protein At4g38520; F20M13.80 [Arab... 202 2e-50
UniRef100_Q9FKX4 Protein phosphatase 2C-like protein [Arabidopsi... 200 7e-50
UniRef100_Q9M3V0 Protein phosphatase 2C [Fagus sylvatica] 200 7e-50
UniRef100_Q7XCJ7 Hypothetical protein [Oryza sativa] 197 6e-49
UniRef100_Q7Y138 Hypothetical protein OSJNBa0078D06.30 [Oryza sa... 195 2e-48
UniRef100_Q9LHJ9 Protein phosphatase 2C [Arabidopsis thaliana] 195 2e-48
UniRef100_Q9ZSQ7 Protein phosphatase 2C homolog [Mesembryanthemu... 194 5e-48
UniRef100_Q8LEW2 Protein phosphatase-2c, putative [Arabidopsis t... 192 1e-47
UniRef100_Q9LSN8 Protein phosphatase 2C-like protein [Arabidopsi... 192 2e-47
>UniRef100_Q84JD5 Hypothetical protein At5g06750 [Arabidopsis thaliana]
Length = 393
Score = 220 bits (560), Expect = 6e-56
Identities = 130/347 (37%), Positives = 199/347 (56%), Gaps = 14/347 (4%)
Query: 11 RSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFG 69
R G+ ++++ + + D+L+W + + + + SIA Q+N V+ED QVE G
Sbjct: 18 RYGRMNRDDDDDDDHDGDSSSSGDSLLWSRELERHSFGDFSIAVVQANEVIEDHSQVETG 77
Query: 70 KNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVT 129
++FVGVYDGH G +A+R+I LF L R+ E +SE+ + A E+GF V
Sbjct: 78 NGAVFVGVYDGHGGPEASRYISDHLFSHLMRVSRERSCISEEALRAAFSATEEGFLTLVR 137
Query: 130 NNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVV---QLTVDH 186
+ +VGSCCL G+IW TL +ANVGDSRA+LGS R + + QLT DH
Sbjct: 138 RTCGLKPLIAAVGSCCLVGVIWKGTLLIANVGDSRAVLGSMGSNNNRSNKIVAEQLTSDH 197
Query: 187 HVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS----DP-HPSF 241
+ + R+E+R+ +D ++ G R+K +I+++RSIGDAYLK DP P F
Sbjct: 198 NAALEEVRQELRSLHPDDSHIVVLKHGVWRIKGIIQVSRSIGDAYLKRPEFSLDPSFPRF 257
Query: 242 ETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKR 301
+ V+S +P R + SDKF+IFAS G W+ MTN +A +IV + + GI++R
Sbjct: 258 HLAEELQRPVLSAEPCVYTRVLQTSDKFVIFASDGLWEQMTNQQAVEIVNKHPRPGIARR 317
Query: 302 LVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
LVR A+ A + Y +L+ ++ G RR ++DD+TV+VIF++
Sbjct: 318 LVRRAITIAAKKREMNYDDLKKVERG----VRRFFHDDITVVVIFID 360
>UniRef100_Q8LFQ7 Protein phosphatase 2C-like [Arabidopsis thaliana]
Length = 393
Score = 218 bits (556), Expect = 2e-55
Identities = 130/347 (37%), Positives = 198/347 (56%), Gaps = 14/347 (4%)
Query: 11 RSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFG 69
R G+ ++++ + + D+L+W + + + + SIA Q+N V+ED QVE G
Sbjct: 18 RYGRMNRDDDDDDDHDGDSSSSGDSLLWSRELERHSFGDFSIAVVQANEVIEDHSQVETG 77
Query: 70 KNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVT 129
++FVGVYDGH G +A+R+I LF L R+ E +SE+ + A E+GF V
Sbjct: 78 NGAVFVGVYDGHGGPEASRYISDHLFSHLMRVSRERSCISEEALRAAFSATEEGFLTLVR 137
Query: 130 NNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVV---QLTVDH 186
+ +VGSCCL G+IW TL +ANVGDSRA+LGS R + + QLT DH
Sbjct: 138 RTCGLKPLIAAVGSCCLVGVIWKGTLLIANVGDSRAVLGSMGSNNNRSNKIVAEQLTSDH 197
Query: 187 HVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS----DP-HPSF 241
+ + R+E+R+ +D ++ G R+K +I+++RSIGDAYLK DP P F
Sbjct: 198 NAALEEVRQELRSLHPDDSHIVVLKHGVWRIKGIIQVSRSIGDAYLKRPEFSLDPSFPRF 257
Query: 242 ETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKR 301
+ V S +P R + SDKF+IFAS G W+ MTN +A +IV + + GI++R
Sbjct: 258 HLAEELQRPVSSAEPCVYTRVLQTSDKFVIFASDGLWEQMTNQQAVEIVNKHPRPGIARR 317
Query: 302 LVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
LVR A+ A + Y +L+ ++ G RR ++DD+TV+VIF++
Sbjct: 318 LVRRAITIAAKKREMNYDDLKKVERG----VRRFFHDDITVVVIFID 360
>UniRef100_Q7XUC5 OSJNBa0088A01.23 protein [Oryza sativa]
Length = 388
Score = 213 bits (541), Expect = 1e-53
Identities = 132/339 (38%), Positives = 194/339 (56%), Gaps = 16/339 (4%)
Query: 16 KDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSLF 74
+DEE+ A+ D L+W D + S A Q+N V+ED QVE G + F
Sbjct: 24 RDEEDGSGSGGDAD----DLLLWSRDLVRHAAGEFSFAVVQANDVLEDHSQVETGAAATF 79
Query: 75 VGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDD 134
+GVYDGH G +A+RFI L L RL E +SEDI+ A E+GF V
Sbjct: 80 IGVYDGHGGAEASRFISNHLAAHLVRLAQERGTISEDIVRNAFSATEEGFLSLVRRTHLI 139
Query: 135 DGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAR 194
+ S+GSCCL GIIW TL++AN+GDSRA++G G K QLT DH+ S R
Sbjct: 140 KPSIASIGSCCLVGIIWKGTLYLANLGDSRAVVGCLTGSNKIV-AEQLTRDHNASMEEVR 198
Query: 195 EEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD--PHPSFETFSRYE---A 249
+E+R+ +D ++ G R+K +I+++RSIGDAYLK + PS F E
Sbjct: 199 QELRSLHPDDSQIVVLKNGVWRIKGIIQVSRSIGDAYLKKQEFALDPSMTRFHLSEPLRR 258
Query: 250 NVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEK 309
V++ +P R + D F IFAS G W+ +TN +A +IV+NN ++GI++RLV+AAL++
Sbjct: 259 PVLTSEPSIYTRVLHSQDSFFIFASDGLWEHLTNQQAVEIVHNNPREGIARRLVKAALKE 318
Query: 310 AIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
A + Y +++ L+ G RR ++DD+TV+V+F++
Sbjct: 319 AARKREMKYNDIKKLEKG----VRRFFHDDITVVVVFID 353
>UniRef100_Q9SD12 Protein phosphatase 2C-like protein [Arabidopsis thaliana]
Length = 379
Score = 211 bits (536), Expect = 4e-53
Identities = 125/351 (35%), Positives = 199/351 (56%), Gaps = 28/351 (7%)
Query: 12 SGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGK 70
S GK + K+ D L+W +D ++ S+A Q+N ++ED QVE G
Sbjct: 17 SSSGKSSDSTGKQ---------DGLLWYKDFGQHLVGEFSMAVVQANNLLEDQSQVESGP 67
Query: 71 NSL--------FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEK 122
S F+G+YDGH G + +RF+ LF L R E +S D++++A + E+
Sbjct: 68 LSTLDSGPYGTFIGIYDGHGGPETSRFVNDHLFQHLKRFAAEQASMSVDVIKKAYEATEE 127
Query: 123 GFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQL 182
GF VT ++ +VGSCCL G+I G L++ANVGDSRA+LG + +QL
Sbjct: 128 GFLGVVTKQWPTKPQIAAVGSCCLVGVICGGMLYIANVGDSRAVLGRAMKATGEVIALQL 187
Query: 183 TVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----P 237
+ +H+VS + R+E+ + +D ++ RVK LI+I+RSIGD YLK ++
Sbjct: 188 SAEHNVSIESVRQEMHSLHPDDSHIVMLKHNVWRVKGLIQISRSIGDVYLKKAEFNKEPL 247
Query: 238 HPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDG 297
+ + ++ ++S +P +I DKFLIFAS G W+ M+N EA DIV N+ ++G
Sbjct: 248 YTKYRIREPFKRPILSGEPTITEHEIQPQDKFLIFASDGLWEQMSNQEAVDIVQNHPRNG 307
Query: 298 ISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
I++RLV+ AL++A + Y +L+ ++ G RRH++DD+TV++IFL+
Sbjct: 308 IARRLVKMALQEAAKKREMRYSDLKKIERG----VRRHFHDDITVVIIFLD 354
>UniRef100_Q8S2S5 Protein phosphatase 2c-like protein [Thellungiella halophila]
Length = 378
Score = 210 bits (535), Expect = 5e-53
Identities = 128/351 (36%), Positives = 199/351 (56%), Gaps = 28/351 (7%)
Query: 12 SGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGK 70
S GK + K+ D L+W +D ++ S+A Q+N ++ED QVE G
Sbjct: 16 SSSGKSSDSTGKQ---------DGLLWYKDSGQHLLGEFSMAVVQANNLLEDQSQVESGP 66
Query: 71 NSL--------FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEK 122
S F+G+YDGH G + +RF+ LF L R E+ +S D++ +A + E+
Sbjct: 67 LSTLDSGPFGTFIGIYDGHGGPETSRFVNDHLFQHLKRFAAEHASMSVDVIRKAYEATEE 126
Query: 123 GFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQL 182
GF VT ++ +VGSCCL G+I G L++ANVGDSRA+LG + +QL
Sbjct: 127 GFLGVVTKQWPVKPQIAAVGSCCLVGVICGGRLYIANVGDSRAVLGRAMNATGEVIALQL 186
Query: 183 TVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD--PHPS 240
+ +H+VS + R+E+R+ +D ++ RVK LI+I+RSIGD YLK ++ P
Sbjct: 187 SAEHNVSIESVRQEMRSLHPDDSHIVVLKHNVWRVKGLIQISRSIGDIYLKKAEFNKEPL 246
Query: 241 FETFSRYE---ANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDG 297
+ + E ++S +P +I D+FLIFAS G W+ M+N EA DIV N+ ++G
Sbjct: 247 YTKYRLREPIKRPILSGEPSITEHEIQPQDQFLIFASDGLWEQMSNQEAVDIVQNHPRNG 306
Query: 298 ISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
I++RLV+ AL+ A + Y +L+ ++ G RRH++DD+TV+VIFL+
Sbjct: 307 IARRLVKMALQAAAKKREMRYSDLKKIERG----VRRHFHDDITVVVIFLD 353
>UniRef100_Q94CL8 Ser/Thr protein phosphatase 2C [Arabidopsis thaliana]
Length = 384
Score = 207 bits (526), Expect = 5e-52
Identities = 123/329 (37%), Positives = 190/329 (57%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D + S+A Q+N ++ED Q+E G SL FVGVYDGH G
Sbjct: 35 DGLLWYKDSGNHITGEFSMAVVQANNLLEDHSQLESGPISLHESGPEATFVGVYDGHGGP 94
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+AARF+ LF + R +E + +S D++ + E+ F V ++ SVG+C
Sbjct: 95 EAARFVNDRLFYNIKRYTSEQRGMSPDVITRGFVATEEEFLGLVQEQWKTKPQIASVGAC 154
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL GI+ L+VAN GDSR +LG FK VQL+ +H+ S + REE+R +D
Sbjct: 155 CLVGIVCNGLLYVANAGDSRVVLGKVANPFKELKAVQLSTEHNASIESVREELRLLHPDD 214
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGDAYLK ++ + P F R+E ++ +P
Sbjct: 215 PNIVVLKHKVWRVKGIIQVSRSIGDAYLKRAEFNQEPLLPKFRVPERFEKPIMRAEPTIT 274
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
I D+FLIFAS G W+ ++N EA DIV + ++G++++LV+AAL++A + Y
Sbjct: 275 VHKIHPEDQFLIFASDGLWEHLSNQEAVDIVNSCPRNGVARKLVKAALQEAAKKREMRYS 334
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ ++ G RRH++DD+TVIV+FL+
Sbjct: 335 DLEKIERGI----RRHFHDDITVIVVFLH 359
>UniRef100_Q9M2W1 Protein phosphatase 2C-like protein [Arabidopsis thaliana]
Length = 409
Score = 207 bits (526), Expect = 5e-52
Identities = 123/329 (37%), Positives = 190/329 (57%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D + S+A Q+N ++ED Q+E G SL FVGVYDGH G
Sbjct: 60 DGLLWYKDSGNHITGEFSMAVVQANNLLEDHSQLESGPISLHESGPEATFVGVYDGHGGP 119
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+AARF+ LF + R +E + +S D++ + E+ F V ++ SVG+C
Sbjct: 120 EAARFVNDRLFYNIKRYTSEQRGMSPDVITRGFVATEEEFLGLVQEQWKTKPQIASVGAC 179
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL GI+ L+VAN GDSR +LG FK VQL+ +H+ S + REE+R +D
Sbjct: 180 CLVGIVCNGLLYVANAGDSRVVLGKVANPFKELKAVQLSTEHNASIESVREELRLLHPDD 239
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGDAYLK ++ + P F R+E ++ +P
Sbjct: 240 PNIVVLKHKVWRVKGIIQVSRSIGDAYLKRAEFNQEPLLPKFRVPERFEKPIMRAEPTIT 299
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
I D+FLIFAS G W+ ++N EA DIV + ++G++++LV+AAL++A + Y
Sbjct: 300 VHKIHPEDQFLIFASDGLWEHLSNQEAVDIVNSCPRNGVARKLVKAALQEAAKKREMRYS 359
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ ++ G RRH++DD+TVIV+FL+
Sbjct: 360 DLEKIERGI----RRHFHDDITVIVVFLH 384
>UniRef100_Q8SB31 Hypothetical protein OJ1540_H01.4 [Oryza sativa]
Length = 392
Score = 207 bits (526), Expect = 5e-52
Identities = 122/329 (37%), Positives = 190/329 (57%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D ++ + S+A Q+N ++ED Q+E G S FVGVYDGH G
Sbjct: 32 DGLLWYKDTGQHVNGEFSMAVVQANNLLEDQCQIESGPLSFLDSGPYGTFVGVYDGHGGP 91
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ A +I LF L R +E +S D++++A + E GF VT ++ +VGSC
Sbjct: 92 ETACYINDHLFHHLKRFASEQNSISADVLKKAYEATEDGFFSVVTKQWPVKPQIAAVGSC 151
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL G+I G L+VANVGDSR +LG VQL+ +H+VS + R+E+++ D
Sbjct: 152 CLVGVICGGILYVANVGDSRVVLGRHVKATGEVLAVQLSAEHNVSIESVRKELQSMHPED 211
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTD 259
++ RVK LI++ RSIGDAYLK S+ + F + ++S +P
Sbjct: 212 RHIVVLKHNVWRVKGLIQVCRSIGDAYLKRSEFNREPLYAKFRLREPFHKPILSSEPSIS 271
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
+ + D+FLIFAS G W+ +TN EA DIV+++ ++G ++RL++AAL++A + Y
Sbjct: 272 VQPLQPHDQFLIFASDGLWEHLTNQEAVDIVHSSPRNGSARRLIKAALQEAAKKREMRYS 331
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+FL+
Sbjct: 332 DLKKIDRG----VRRHFHDDITVIVVFLD 356
>UniRef100_Q6ZHC8 Hypothetical protein OJ1717_A09.24 [Oryza sativa]
Length = 387
Score = 205 bits (521), Expect = 2e-51
Identities = 125/323 (38%), Positives = 185/323 (56%), Gaps = 12/323 (3%)
Query: 32 VHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDAARFI 90
V D L+W D + S A Q+N +ED QVE G + FVGVYDGH G DAARFI
Sbjct: 35 VADGLLWSRDLGRHAAGEFSFAVVQANEALEDHSQVETGSAATFVGVYDGHGGADAARFI 94
Query: 91 RVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGII 150
LF L RL E++ VSE+++ A E+GF V + +VGSCCL GII
Sbjct: 95 SDHLFAHLIRLARESETVSEEVVRGAFSATEEGFLTLVRRTQFLKPMIAAVGSCCLVGII 154
Query: 151 WGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCK 210
W L+VAN+GDSRA++G G + Q+T DH+ R+E+ + +D ++
Sbjct: 155 WRGVLYVANLGDSRAVVG-YLGRTNKITAEQITRDHNACKEEVRQELISRHPDDSQIVVL 213
Query: 211 NRGSLRVKSLIEITRSIGDAYLKWSD--PHPSFETFSRYE---ANVISEKPFTDRRDIDE 265
G R+K +I+++R+IGDAYLK + PS F E V++ +P R +
Sbjct: 214 KHGVWRIKGIIQVSRTIGDAYLKRREFALDPSITRFRLSEPLRRPVLTAEPSICTRVLSL 273
Query: 266 SDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYCNLQNLK 324
D+F+IFAS G W+ +TN +A DIVY N + GI+KRLV AL++A + + +L+ ++
Sbjct: 274 QDQFVIFASDGLWEHLTNQQAVDIVYKNPRAGIAKRLVNTALKEAARKREMRFVDLKKVE 333
Query: 325 AGNGLLGRRHYYDDVTVIVIFLN 347
G RR ++DD+TV+V++++
Sbjct: 334 KG----VRRFFHDDITVVVVYID 352
>UniRef100_Q9LZ09 Protein phosphatase-like protein [Arabidopsis thaliana]
Length = 361
Score = 203 bits (517), Expect = 6e-51
Identities = 130/354 (36%), Positives = 197/354 (54%), Gaps = 37/354 (10%)
Query: 9 CGRSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVE 67
C R G G E+ K D L W +D + + S+A Q+N+VMED Q+E
Sbjct: 5 CWRIGAGM-------ERSKINPTKVDGLTWYKDLGLHTFGEFSMAMIQANSVMEDQCQIE 57
Query: 68 FGK--------NSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDF 119
G FVGVYDGH G +A+RFI +FP K +SE ++ +A
Sbjct: 58 SGPLTFNNPTVQGTFVGVYDGHGGPEASRFIADNIFP---------KEISEQVISKAFAE 108
Query: 120 IEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHV 179
+K F + VT + ++ SVGSCCL G+I +++AN GDSRA+LG S+ R
Sbjct: 109 TDKDFLKTVTKQWPTNPQMASVGSCCLAGVICNGLVYIANTGDSRAVLGRSERGGVR--A 166
Query: 180 VQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH- 238
VQL+V+H+ + +AR+E+ + NDP +L RVK +I++TRSIGDAYLK ++ +
Sbjct: 167 VQLSVEHNANLESARQELWSLHPNDPTILVMKHRLWRVKGVIQVTRSIGDAYLKRAEFNR 226
Query: 239 ----PSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNS 294
P F + ++S P + D+F+I AS G W+ ++N EA DIV+N+
Sbjct: 227 EPLLPKFRLPEHFTKPILSADPSVTITRLSPQDEFIILASDGLWEHLSNQEAVDIVHNSP 286
Query: 295 QDGISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+ GI++RL++AAL++A + Y +L + G RRH++DD+TVIV++LN
Sbjct: 287 RQGIARRLLKAALKEAAKKREMRYSDLTEIHPG----VRRHFHDDITVIVVYLN 336
>UniRef100_Q5PNS9 At4g38520 [Arabidopsis thaliana]
Length = 400
Score = 202 bits (513), Expect = 2e-50
Identities = 117/329 (35%), Positives = 186/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNS--------LFVGVYDGHKGL 84
+ L+W D ++ + S+A Q+N+++ED Q+E G S FVGVYDGH G
Sbjct: 32 EGLLWFRDSGQHVFGDFSMAVVQANSLLEDQSQLESGSLSSHDSGPFGTFVGVYDGHGGP 91
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ +RFI +F L R E + +S +++++A E+GF VTN ++ +VGSC
Sbjct: 92 ETSRFINDHMFHHLKRFTAEQQCMSSEVIKKAFQATEEGFLSIVTNQFQTRPQIATVGSC 151
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL +I L+VAN GDSRA+LG H QL+ +H+ S + R E++ +
Sbjct: 152 CLVSVICDGKLYVANAGDSRAVLGQVMRVTGEAHATQLSAEHNASIESVRRELQALHPDH 211
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGD YLK S+ + F S + ++S +P
Sbjct: 212 PDIVVLKHNVWRVKGIIQVSRSIGDVYLKRSEFNREPLYAKFRLRSPFSKPLLSAEPAIT 271
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
++ D+F+I AS G W+ M+N EA DIV N+ ++GI+KRLV+ AL++A + Y
Sbjct: 272 VHTLEPHDQFIICASDGLWEHMSNQEAVDIVQNHPRNGIAKRLVKVALQEAAKKREMRYS 331
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+F +
Sbjct: 332 DLKKIDRG----VRRHFHDDITVIVVFFD 356
>UniRef100_Q8W4N8 Hypothetical protein At4g38520; F20M13.80 [Arabidopsis thaliana]
Length = 400
Score = 202 bits (513), Expect = 2e-50
Identities = 117/329 (35%), Positives = 186/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNS--------LFVGVYDGHKGL 84
+ L+W D ++ + S+A Q+N+++ED Q+E G S FVGVYDGH G
Sbjct: 32 EGLLWFSDSGQHVFGDFSMAVVQANSLLEDQSQLESGSLSSHDSGPFGTFVGVYDGHGGP 91
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ +RFI +F L R E + +S +++++A E+GF VTN ++ +VGSC
Sbjct: 92 ETSRFINDHMFHHLKRFTAEQQCMSSEVIKKAFQATEEGFLSIVTNQFQTRPQIATVGSC 151
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL +I L+VAN GDSRA+LG H QL+ +H+ S + R E++ +
Sbjct: 152 CLVSVICDGKLYVANAGDSRAVLGQVMRVTGEAHATQLSAEHNASIESVRRELQALHPDH 211
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGD YLK S+ + F S + ++S +P
Sbjct: 212 PDIVVLKHNVWRVKGIIQVSRSIGDVYLKRSEFNREPLYAKFRLRSPFSKPLLSAEPAIT 271
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
++ D+F+I AS G W+ M+N EA DIV N+ ++GI+KRLV+ AL++A + Y
Sbjct: 272 VHTLEPHDQFIICASDGLWEHMSNQEAVDIVQNHPRNGIAKRLVKVALQEAAKKREMRYS 331
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+F +
Sbjct: 332 DLKKIDRG----VRRHFHDDITVIVVFFD 356
>UniRef100_Q9FKX4 Protein phosphatase 2C-like protein [Arabidopsis thaliana]
Length = 385
Score = 200 bits (508), Expect = 7e-50
Identities = 119/330 (36%), Positives = 189/330 (57%), Gaps = 20/330 (6%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSL---------FVGVYDGHKG 83
D L+W +D + + S+A Q+N ++ED QVE G + FVGVYDGH G
Sbjct: 32 DGLLWYKDSAHHLFGDFSMAVVQANNLLEDQSQVESGPLTTLSSSGPYGTFVGVYDGHGG 91
Query: 84 LDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGS 143
+ +RF+ LF L R E +S D++ +A + E+GF V + +VGS
Sbjct: 92 PETSRFVNDHLFHHLKRFAAEQDSMSVDVIRKAYEATEEGFLGVVAKQWAVKPHIAAVGS 151
Query: 144 CCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITN 203
CCL G++ L+VANVGDSRA+LG + +QL+ +H+VS + R+E+ + +
Sbjct: 152 CCLIGVVCDGKLYVANVGDSRAVLGKVIKATGEVNALQLSAEHNVSIESVRQEMHSLHPD 211
Query: 204 DPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD--PHPSFETFSRYE---ANVISEKPFT 258
D ++ RVK +I+++RSIGD YLK S+ P + + E ++S +P
Sbjct: 212 DSHIVVLKHNVWRVKGIIQVSRSIGDVYLKKSEFNKEPLYTKYRLREPMKRPILSWEPSI 271
Query: 259 DRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITY 317
D+ D+FLIFAS G W+ ++N EA +IV N+ ++GI++RLV+AAL++A + Y
Sbjct: 272 TVHDLQPDDQFLIFASDGLWEQLSNQEAVEIVQNHPRNGIARRLVKAALQEAAKKREMRY 331
Query: 318 CNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L ++ G RRH++DD+TV+V+FL+
Sbjct: 332 SDLNKIERG----VRRHFHDDITVVVLFLD 357
>UniRef100_Q9M3V0 Protein phosphatase 2C [Fagus sylvatica]
Length = 397
Score = 200 bits (508), Expect = 7e-50
Identities = 118/347 (34%), Positives = 192/347 (55%), Gaps = 28/347 (8%)
Query: 25 KEKAEEEVH---------DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL- 73
+ K++ VH D L+W +D ++ + S Q+N ++ED Q+E G SL
Sbjct: 14 RAKSDRHVHTGSNARGRQDGLLWYKDSGQHFNGEFSYGCGQANNLLEDQSQIESGSLSLN 73
Query: 74 -------FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKE 126
F+GVYDGH G + +R++ LF L R +E +S +++ +A E+GF
Sbjct: 74 ETGPYGTFIGVYDGHGGPETSRYVNNHLFQHLKRFTSEQHSMSVEVIRKAYQATEEGFLS 133
Query: 127 YVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDH 186
VT ++ +VGSCCL G+I G TL++AN+GDSRA+LG +QL+ +H
Sbjct: 134 QVTKQWPLKPQIAAVGSCCLVGVICGGTLYIANLGDSRAVLGRVMKATGEVLSIQLSAEH 193
Query: 187 HVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSF 241
+V+ + R+E+ + DP ++ RVK LI+I+RSIGD YLK ++ + F
Sbjct: 194 NVAIESVRQELHSLHPEDPQIVVLKHNVWRVKGLIQISRSIGDVYLKKAEFNREPLYAKF 253
Query: 242 ETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKR 301
++ ++S P + D+F+IFAS G W+ ++N EA DIV N+ + G +R
Sbjct: 254 RLREPFKKPILSADPAISVHQLQPHDQFVIFASDGLWEHLSNQEAVDIVQNHPRSGSVRR 313
Query: 302 LVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
L++ AL++A + Y +L+ + G RRH++DD+TVIV+FL+
Sbjct: 314 LIKVALQEAAKKREMRYSDLKKIDRG----VRRHFHDDITVIVVFLD 356
>UniRef100_Q7XCJ7 Hypothetical protein [Oryza sativa]
Length = 393
Score = 197 bits (500), Expect = 6e-49
Identities = 123/364 (33%), Positives = 194/364 (52%), Gaps = 30/364 (8%)
Query: 4 PPFLTCGRSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMED 62
P GR KG D + D L+W +D + S+A Q+N ++ED
Sbjct: 15 PASSPAGRPRKGSDAAGRQ-----------DGLLWYKDAGQLVAGEFSMAVVQANNLLED 63
Query: 63 FYQVEFGKNSL--------FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIME 114
QVE G S VGVYDGH G + AR+I LF L +E+K +S D++
Sbjct: 64 HSQVESGPLSTTDPNLQGTLVGVYDGHGGPETARYINDHLFNHLRGFASEHKCMSADVIR 123
Query: 115 QAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFF 174
+A E+GF V++ ++ +VGSCCL G+I L++AN+GDSRA+LG
Sbjct: 124 KAFRATEEGFFSVVSSQWSMRPQLAAVGSCCLVGVICAGNLYIANLGDSRAVLGRLVKGT 183
Query: 175 KRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKW 234
+QL+ +H+ S R E++ +DP ++ RVK +I+ITRSIGD YLK
Sbjct: 184 GEVLAMQLSAEHNASFEEVRRELQAAHPDDPHIVVLKHNVWRVKGIIQITRSIGDVYLKK 243
Query: 235 SD-----PHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADI 289
+ H F + ++S +P + +D+F+IFAS G W+ ++N EA D+
Sbjct: 244 PEFNREPLHSKFRLQETFRRPLLSSEPAIVVHQLQTTDQFIIFASDGLWEHISNQEAVDL 303
Query: 290 VYNNSQDGISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLNK 348
V +N ++GI++RLV+AA+++A + Y +L+ + G RRH++DD+TV+V+F +
Sbjct: 304 VQHNPRNGIARRLVKAAMQQAAKKREMRYSDLKKIDRG----VRRHFHDDITVVVVFFDS 359
Query: 349 RSDT 352
+ T
Sbjct: 360 NAIT 363
>UniRef100_Q7Y138 Hypothetical protein OSJNBa0078D06.30 [Oryza sativa]
Length = 386
Score = 195 bits (496), Expect = 2e-48
Identities = 120/332 (36%), Positives = 184/332 (55%), Gaps = 14/332 (4%)
Query: 26 EKAEEEVHDNLVWCEDRKE-NDYHCSIATSQSNTVMEDFYQVEFGKNSLF---VGVYDGH 81
++A E D L+W D + S+A Q N V+ED +VE G L +GV+DGH
Sbjct: 37 DQAAAEGRDGLLWWRDLARCHAGELSVAVVQGNHVLEDQCRVESGPPPLAATCIGVFDGH 96
Query: 82 KGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSV 141
G DAARF L P L + + V+ D + A E+GF V+ + + +V
Sbjct: 97 AGPDAARFACDHLLPNLREAASGPEGVTADAIRDAFLATEEGFLAVVSRMWEAQPDMATV 156
Query: 142 GSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHI 201
G+CCL G++ RTLFVAN+GDSRA+LG G + QL+ +H+ + R+E+
Sbjct: 157 GTCCLVGVVHQRTLFVANLGDSRAVLGKKVGRAGQITAEQLSSEHNANEEDVRQELMAQH 216
Query: 202 TNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKP 256
+DP ++ G RVK +I+++RS+GDAYLK S + P F + ++S P
Sbjct: 217 PDDPQIVALKHGVWRVKGIIQVSRSLGDAYLKHSQYNTEQIKPKFRLPEPFSRPILSANP 276
Query: 257 FTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDII 315
R + SD F+IFAS G W+ ++N +A +IV+N+ + G ++RL++AAL E A +
Sbjct: 277 SIIARCLQPSDCFIIFASDGLWEHLSNQQAVEIVHNHQRAGSARRLIKAALHEAARKREM 336
Query: 316 TYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
Y +L + RRH++DD+TVIV+F+N
Sbjct: 337 RYSDLMKIDK----KVRRHFHDDITVIVLFIN 364
>UniRef100_Q9LHJ9 Protein phosphatase 2C [Arabidopsis thaliana]
Length = 385
Score = 195 bits (495), Expect = 2e-48
Identities = 118/329 (35%), Positives = 186/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSLF--------VGVYDGHKGL 84
D L+W +D + S++ Q+N ++ED ++E G S+F VGVYDGH G
Sbjct: 34 DGLLWYKDSGNHVAGEFSMSVIQANNLLEDHSKLESGPVSMFDSGPQATFVGVYDGHGGP 93
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+AARF+ LF + + +EN +S +++ +A E+ F V ++ SVG+C
Sbjct: 94 EAARFVNKHLFDNIRKFTSENHGMSANVITKAFLATEEDFLSLVRRQWQIKPQIASVGAC 153
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL GII L++AN GDSR +LG + FK VQL+ +H+ S + REE+R+ ND
Sbjct: 154 CLVGIICSGLLYIANAGDSRVVLGRLEKAFKIVKAVQLSSEHNASLESVREELRSLHPND 213
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGDAYLK ++ + F + ++ +P
Sbjct: 214 PQIVVLKHKVWRVKGIIQVSRSIGDAYLKKAEFNREPLLAKFRVPEVFHKPILRAEPAIT 273
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDIITYC 318
I D+FLIFAS G W+ ++N EA DIV ++GI+++L++ AL E A + Y
Sbjct: 274 VHKIHPEDQFLIFASDGLWEHLSNQEAVDIVNTCPRNGIARKLIKTALREAAKKREMRYS 333
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+FL+
Sbjct: 334 DLKKIDRG----VRRHFHDDITVIVVFLD 358
>UniRef100_Q9ZSQ7 Protein phosphatase 2C homolog [Mesembryanthemum crystallinum]
Length = 396
Score = 194 bits (492), Expect = 5e-48
Identities = 114/329 (34%), Positives = 185/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D ++ + S+A Q+N ++ED Q+E G SL FVGVYDGH G
Sbjct: 31 DGLLWYKDTGQHINGEFSMAVVQANNLLEDQSQIESGCLSLLDSGPYGTFVGVYDGHGGP 90
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ +R+I LF L R E + +S D++++A E+GF V ++ +VGSC
Sbjct: 91 ETSRYINDHLFHHLKRFAAEQQSMSVDVIKKAFQATEEGFFSVVAKQWPMKPQIAAVGSC 150
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL G++ L++AN+GDSRA+LG + +QL+ +H+ S + R+E++ D
Sbjct: 151 CLVGVVCNGILYIANLGDSRAVLGRAVKATGEVLAIQLSAEHNASIESVRQEMQATHPED 210
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTD 259
++ RVK LI+ITRSIGD YLK ++ + F ++ ++S P
Sbjct: 211 KDIVVLKHNVWRVKGLIQITRSIGDVYLKKTEYNREPLYSKFRLREPFKKPILSSDPAIS 270
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
++ D+ IFAS G W+ +TN EA D+V + ++G +KRLV+ AL++A + Y
Sbjct: 271 VHELQPHDQVCIFASDGLWEHLTNQEAVDLVQKSPRNGSAKRLVKVALQEAAKKREMRYS 330
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TV+V+FL+
Sbjct: 331 DLKKIDRG----VRRHFHDDITVVVVFLD 355
>UniRef100_Q8LEW2 Protein phosphatase-2c, putative [Arabidopsis thaliana]
Length = 384
Score = 192 bits (488), Expect = 1e-47
Identities = 120/328 (36%), Positives = 188/328 (56%), Gaps = 19/328 (5%)
Query: 31 EVHDNLVWCEDRKENDYHC----SIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDA 86
E D L+W D + +C S+A Q+N V+ED QVE G FVGVYDGH G +A
Sbjct: 40 EGKDGLLWFRDLGK---YCGGDFSMAVIQANQVLEDQSQVESGNFGTFVGVYDGHGGPEA 96
Query: 87 ARFIRVCLFPELSRLVTENK-VVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCC 145
AR++ LF + E + VV+ + +E+A E+GF V+ + + +VG+CC
Sbjct: 97 ARYVCDHLFXHFREISAETQGVVTRETIERAFHATEEGFASIVSELWQEIPNLATVGTCC 156
Query: 146 LFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDP 205
L G+I+ TLFVA++GDSR +LG KG +QL+ +H+ ++ R E+++ +DP
Sbjct: 157 LVGVIYQNTLFVASLGDSRVVLG-KKGNCGGLSAIQLSTEHNANNEDIRWELKDLHPDDP 215
Query: 206 FVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTDR 260
++ G RVK +I+++RSIGD Y+K + + F + ++S P
Sbjct: 216 QIVVFRHGVXRVKGIIQVSRSIGDMYMKRPEFNKEPISQKFRIAEPMKRPLMSATPTILS 275
Query: 261 RDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDIITYCN 319
+ +D FLIFAS G W+ +TN +A +IV+N+ + G +KRL++AAL E A + Y +
Sbjct: 276 HPLHPNDSFLIFASDGLWEHLTNEKAVEIVHNHPRAGSAKRLIKAALHEAARKREMRYSD 335
Query: 320 LQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
L+ + RRH++DD+TVIV+FLN
Sbjct: 336 LRKIDK----KVRRHFHDDITVIVVFLN 359
>UniRef100_Q9LSN8 Protein phosphatase 2C-like protein [Arabidopsis thaliana]
Length = 379
Score = 192 bits (487), Expect = 2e-47
Identities = 120/328 (36%), Positives = 188/328 (56%), Gaps = 19/328 (5%)
Query: 31 EVHDNLVWCEDRKENDYHC----SIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDA 86
E D L+W D + +C S+A Q+N V+ED QVE G FVGVYDGH G +A
Sbjct: 35 EGKDGLLWFRDLGK---YCGGDFSMAVIQANQVLEDQSQVESGNFGTFVGVYDGHGGPEA 91
Query: 87 ARFIRVCLFPELSRLVTENK-VVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCC 145
AR++ LF + E + VV+ + +E+A E+GF V+ + + +VG+CC
Sbjct: 92 ARYVCDHLFNHFREISAETQGVVTRETIERAFHATEEGFASIVSELWQEIPNLATVGTCC 151
Query: 146 LFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDP 205
L G+I+ TLFVA++GDSR +LG KG +QL+ +H+ ++ R E+++ +DP
Sbjct: 152 LVGVIYQNTLFVASLGDSRVVLG-KKGNCGGLSAIQLSTEHNANNEDIRWELKDLHPDDP 210
Query: 206 FVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTDR 260
++ G RVK +I+++RSIGD Y+K + + F + ++S P
Sbjct: 211 QIVVFRHGVWRVKGIIQVSRSIGDMYMKRPEFNKEPISQKFRIAEPMKRPLMSATPTILS 270
Query: 261 RDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDIITYCN 319
+ +D FLIFAS G W+ +TN +A +IV+N+ + G +KRL++AAL E A + Y +
Sbjct: 271 HPLHPNDSFLIFASDGLWEHLTNEKAVEIVHNHPRAGSAKRLIKAALHEAARKREMRYSD 330
Query: 320 LQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
L+ + RRH++DD+TVIV+FLN
Sbjct: 331 LRKIDK----KVRRHFHDDITVIVVFLN 354
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 676,001,804
Number of Sequences: 2790947
Number of extensions: 29184552
Number of successful extensions: 166562
Number of sequences better than 10.0: 667
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 557
Number of HSP's that attempted gapping in prelim test: 165062
Number of HSP's gapped (non-prelim): 1036
length of query: 397
length of database: 848,049,833
effective HSP length: 129
effective length of query: 268
effective length of database: 488,017,670
effective search space: 130788735560
effective search space used: 130788735560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC124971.1


