

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124967.9 - phase: 0
(122 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
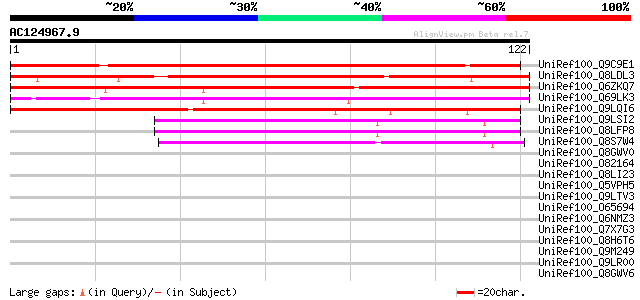
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C9E1 Hypothetical protein T10D10.10 [Arabidopsis tha... 116 1e-25
UniRef100_Q8LDL3 Hypothetical protein [Arabidopsis thaliana] 110 6e-24
UniRef100_Q6ZKQ7 Auxin-induced protein-related-like protein [Ory... 110 1e-23
UniRef100_Q69LK3 Auxin-induced protein-like [Oryza sativa] 103 9e-22
UniRef100_Q9LQI6 F28G4.20 protein [Arabidopsis thaliana] 102 2e-21
UniRef100_Q9LSI2 Gb|AAF27147.1 [Arabidopsis thaliana] 61 7e-09
UniRef100_Q8LFP8 Hypothetical protein [Arabidopsis thaliana] 61 7e-09
UniRef100_Q8S7W4 Hypothetical protein OSJNBa0091P11.19 [Oryza sa... 51 7e-06
UniRef100_Q8GWV0 Hypothetical protein At2g35290 [Arabidopsis tha... 42 0.002
UniRef100_O82164 Hypothetical protein At2g35290 [Arabidopsis tha... 42 0.002
UniRef100_Q8LI23 Hypothetical protein OJ1339_F05.143 [Oryza sativa] 42 0.003
UniRef100_Q5VPH5 Putative auxin-induced protein TGSAUR22 [Oryza ... 42 0.003
UniRef100_Q9LTV3 Auxin-regulated protein-like [Arabidopsis thali... 39 0.035
UniRef100_O65694 Hypothetical protein T4L20.330 [Arabidopsis tha... 38 0.046
UniRef100_Q6NMZ3 At4g34750 [Arabidopsis thaliana] 38 0.060
UniRef100_Q7X7G3 OSJNBa0073E02.12 protein [Oryza sativa] 37 0.079
UniRef100_Q8H6T6 Auxin-regulated protein [Phaseolus vulgaris] 36 0.23
UniRef100_Q9M249 Hypothetical protein F7M19_130 [Arabidopsis tha... 34 0.87
UniRef100_Q9LR00 F10A5.21 [Arabidopsis thaliana] 34 0.87
UniRef100_Q8GWV6 Hypothetical protein At5g20810/T1M15_210 [Arabi... 33 1.1
>UniRef100_Q9C9E1 Hypothetical protein T10D10.10 [Arabidopsis thaliana]
Length = 119
Score = 116 bits (290), Expect = 1e-25
Identities = 61/120 (50%), Positives = 82/120 (67%), Gaps = 3/120 (2%)
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVT 60
MAK GKL KLKSA+KKW SF K HS S+ A D S ++LH V+VG++RR Y +
Sbjct: 1 MAKVGKLTKLKSAMKKWPSFA--KNHHHSTSSAAVSDELSEDNNLHVVYVGQTRRPYMLR 58
Query: 61 SDVVDNPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQPESLDELVDFY 120
D++ +P+F+ELV+RS + D VVACEVVLFEHLLWML++ + S++EL +FY
Sbjct: 59 PDIISHPLFQELVDRSSSRSIEQDREIVVACEVVLFEHLLWMLKSGQ-EGGSVEELAEFY 117
>UniRef100_Q8LDL3 Hypothetical protein [Arabidopsis thaliana]
Length = 127
Score = 110 bits (276), Expect = 6e-24
Identities = 67/131 (51%), Positives = 87/131 (66%), Gaps = 13/131 (9%)
Query: 1 MAKGG-KLMKLKSALKKWNSFGNGK-----QSRHSISAVADEDSSSSRSDLHTVFVGKSR 54
MAKGG KLMKLKS LKK NSF Q+ H S+ S+ DL TV+VG++R
Sbjct: 1 MAKGGNKLMKLKSVLKKLNSFNTKPNQPQAQTNHGRSSAV---SAFPSEDLQTVYVGRTR 57
Query: 55 RLYRVTSDVVDNPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENAD---PQPE 111
R Y V+SDVV +P+F++L ++ +++V +CEVVLFEHLLWMLENAD +PE
Sbjct: 58 RPYHVSSDVVSHPLFQQLAAVDGGCGSEDGSISV-SCEVVLFEHLLWMLENADADESRPE 116
Query: 112 SLDELVDFYAC 122
S+ ELV+FYAC
Sbjct: 117 SVYELVEFYAC 127
>UniRef100_Q6ZKQ7 Auxin-induced protein-related-like protein [Oryza sativa]
Length = 133
Score = 110 bits (274), Expect = 1e-23
Identities = 61/134 (45%), Positives = 82/134 (60%), Gaps = 13/134 (9%)
Query: 1 MAKGGKLMKLKSALKKWNSFG-------NGKQSRHSISAVADEDSSSSRSD-----LHTV 48
M KGG L KL+ +++W+S + + +A A +S +D LH V
Sbjct: 1 MGKGGGLSKLRCMIRRWHSSSRIARAPPSAGELEEGSAAAAGRAASFHGADEVPKGLHPV 60
Query: 49 FVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADP 108
+VGKSRR Y + ++V +P+F+ LV+R+ VV CEVVLFEHLLWMLENADP
Sbjct: 61 YVGKSRRRYLIAEELVGHPLFQNLVDRTGGGGG-GGAATVVGCEVVLFEHLLWMLENADP 119
Query: 109 QPESLDELVDFYAC 122
QPESLDELV++YAC
Sbjct: 120 QPESLDELVEYYAC 133
>UniRef100_Q69LK3 Auxin-induced protein-like [Oryza sativa]
Length = 165
Score = 103 bits (257), Expect = 9e-22
Identities = 69/168 (41%), Positives = 86/168 (51%), Gaps = 49/168 (29%)
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSD---------------- 44
MAKGG L KLK +K+W+S + + SR A S+S+RS
Sbjct: 1 MAKGG-LSKLKCMIKRWHS--SSRISRTPSGCSASAGSTSARSSHGGGRVGGEEWGRSVV 57
Query: 45 ----------------------------LHTVFVGKSRRLYRVTSDVVDNPVFRELVERS 76
LH V+VGKSRR Y + +D+V +P+F+ LV+RS
Sbjct: 58 ASGGGGGGGGGGRGGSVSFHGADGVPPGLHPVYVGKSRRRYLIAADLVGHPMFQNLVDRS 117
Query: 77 RE--TEQQNDTVNVVACEVVLFEHLLWMLENADPQPESLDELVDFYAC 122
VV CEVVLFEHLLWMLENADPQPESLDELV++YAC
Sbjct: 118 GGGGVGGGGGGGTVVGCEVVLFEHLLWMLENADPQPESLDELVEYYAC 165
>UniRef100_Q9LQI6 F28G4.20 protein [Arabidopsis thaliana]
Length = 131
Score = 102 bits (254), Expect = 2e-21
Identities = 62/130 (47%), Positives = 84/130 (63%), Gaps = 11/130 (8%)
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVT 60
MA GKL KLKSA+KKW S S ++ A + S DLH V+VGKSRR Y ++
Sbjct: 1 MAIFGKLTKLKSAIKKWPSLTKNHHSTMCTASTAVSEVSKCE-DLHVVYVGKSRRPYMLS 59
Query: 61 SDVVDNPVFRELVER-SRETEQQNDTVNV-VACEVVLFEHLLWMLENA--------DPQP 110
S V+ +P+F+EL++R SR E+++D V VACEVVLFEHLLWML+N+ D +
Sbjct: 60 SHVIAHPLFQELLDRSSRFIEERHDQETVLVACEVVLFEHLLWMLKNSSSDHGDEDDRER 119
Query: 111 ESLDELVDFY 120
S++EL +FY
Sbjct: 120 GSVEELAEFY 129
>UniRef100_Q9LSI2 Gb|AAF27147.1 [Arabidopsis thaliana]
Length = 139
Score = 60.8 bits (146), Expect = 7e-09
Identities = 34/92 (36%), Positives = 52/92 (55%), Gaps = 6/92 (6%)
Query: 35 DEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDT----VNVVA 90
D D S + S TV VG++++ Y ++ + +P+ LVE+ + E+ ++ + V
Sbjct: 45 DVDQSPTPSMYQTVLVGRTKKPYLISKKHLKHPLLNALVEKQQRYEEDDNEDGSCIITVK 104
Query: 91 CEVVLFEHLLWMLENADPQP--ESLDELVDFY 120
CEVVLF+HLLWMLE D ESLD+ Y
Sbjct: 105 CEVVLFDHLLWMLEYGDEVHILESLDDFAHLY 136
>UniRef100_Q8LFP8 Hypothetical protein [Arabidopsis thaliana]
Length = 135
Score = 60.8 bits (146), Expect = 7e-09
Identities = 34/92 (36%), Positives = 52/92 (55%), Gaps = 6/92 (6%)
Query: 35 DEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDT----VNVVA 90
D D S + S TV VG++++ Y ++ + +P+ LVE+ + E+ ++ + V
Sbjct: 41 DVDQSPTPSMYQTVLVGRTKKPYLISKKHLKHPLLNALVEKQQRYEEDDNEDGSCIITVK 100
Query: 91 CEVVLFEHLLWMLENADPQP--ESLDELVDFY 120
CEVVLF+HLLWMLE D ESLD+ Y
Sbjct: 101 CEVVLFDHLLWMLEYGDQVHILESLDDFAHLY 132
>UniRef100_Q8S7W4 Hypothetical protein OSJNBa0091P11.19 [Oryza sativa]
Length = 119
Score = 50.8 bits (120), Expect = 7e-06
Identities = 31/91 (34%), Positives = 48/91 (52%), Gaps = 6/91 (6%)
Query: 36 EDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDTVNVVACEVVL 95
+D S + TV VGK RR++ V V+D FR L+E + E+ + V + +L
Sbjct: 27 DDGPSPAAVTTTVVVGKERRVFSVDQLVLDTYPFRLLLETAVRKEESKAAL-FVDVDAIL 85
Query: 96 FEHLLWMLENADPQPES-----LDELVDFYA 121
FEH+LW+ + D S L E++DFY+
Sbjct: 86 FEHILWLAGHHDRSSSSLLHLDLKEIIDFYS 116
>UniRef100_Q8GWV0 Hypothetical protein At2g35290 [Arabidopsis thaliana]
Length = 121
Score = 42.4 bits (98), Expect = 0.002
Identities = 29/81 (35%), Positives = 43/81 (52%), Gaps = 7/81 (8%)
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRELV--ERSRETEQQNDTVNVV---ACEVVLFEHLLWM 102
V VGK ++ + V V++ FR L+ + R + N T VV + +LFEHLLW+
Sbjct: 38 VVVGKEKKEFMVEPYVLEEYPFRVLIGSAKDRAKNRLNRTGRVVWLDHVDSILFEHLLWL 97
Query: 103 LENADPQPESLD--ELVDFYA 121
L N LD E+++FYA
Sbjct: 98 LRNDASTFSDLDVLEIIEFYA 118
>UniRef100_O82164 Hypothetical protein At2g35290 [Arabidopsis thaliana]
Length = 91
Score = 42.4 bits (98), Expect = 0.002
Identities = 29/81 (35%), Positives = 43/81 (52%), Gaps = 7/81 (8%)
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRELV--ERSRETEQQNDTVNVV---ACEVVLFEHLLWM 102
V VGK ++ + V V++ FR L+ + R + N T VV + +LFEHLLW+
Sbjct: 8 VVVGKEKKEFMVEPYVLEEYPFRVLIGSAKDRAKNRLNRTGRVVWLDHVDSILFEHLLWL 67
Query: 103 LENADPQPESLD--ELVDFYA 121
L N LD E+++FYA
Sbjct: 68 LRNDASTFSDLDVLEIIEFYA 88
>UniRef100_Q8LI23 Hypothetical protein OJ1339_F05.143 [Oryza sativa]
Length = 205
Score = 42.0 bits (97), Expect = 0.003
Identities = 39/126 (30%), Positives = 58/126 (45%), Gaps = 25/126 (19%)
Query: 10 LKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKS---RRLYRVTSDVVD- 65
L+SA ++ + G G R DSSS+ D+ V VG + R ++ V V++
Sbjct: 78 LRSASRRGGAGGGGGYRRL--------DSSSAAGDVIRVEVGTTKGERSVFHVDQAVLEA 129
Query: 66 NPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWM----------LENADPQPESLDE 115
PV R L R T VA +V+LFEHLLW+ ++ + L E
Sbjct: 130 GPVRRLLAAAGRRTR---GGAVAVAVDVLLFEHLLWLQAAGDKGMLGYDDDESAAADLSE 186
Query: 116 LVDFYA 121
+VDFY+
Sbjct: 187 IVDFYS 192
>UniRef100_Q5VPH5 Putative auxin-induced protein TGSAUR22 [Oryza sativa]
Length = 119
Score = 42.0 bits (97), Expect = 0.003
Identities = 33/106 (31%), Positives = 51/106 (47%), Gaps = 9/106 (8%)
Query: 3 KGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSR-SDL----HTVFVGKSRRLY 57
K G LK LK+ +S G +Q + +S +E+ +S SD+ V+VG+ RR +
Sbjct: 4 KKGGAAGLKQILKRCSSLGRRQQEQKQVSEWEEEEEASGLPSDVPRGHFAVYVGERRRRF 63
Query: 58 RVTSDVVDNPVFRELVERSRE----TEQQNDTVNVVACEVVLFEHL 99
V ++D P FR L+ R+ E + V+ CE V F L
Sbjct: 64 VVPLALLDRPEFRSLLRRAEEEFGFAGAGAGGLLVLPCEEVAFRSL 109
>UniRef100_Q9LTV3 Auxin-regulated protein-like [Arabidopsis thaliana]
Length = 132
Score = 38.5 bits (88), Expect = 0.035
Identities = 24/92 (26%), Positives = 49/92 (53%), Gaps = 1/92 (1%)
Query: 27 RHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERS-RETEQQNDT 85
R S++ + + +SS V+VG + V+++++++PVF L+ RS +E +
Sbjct: 36 RSSVTRRSKKQTSSVPEGHVPVYVGDEMERFVVSAELLNHPVFIGLLNRSAQEYGYEQKG 95
Query: 86 VNVVACEVVLFEHLLWMLENADPQPESLDELV 117
V + C V++FE ++ L P P + +L+
Sbjct: 96 VLQIPCHVLVFERIMESLRLGLPVPIDVQDLI 127
>UniRef100_O65694 Hypothetical protein T4L20.330 [Arabidopsis thaliana]
Length = 150
Score = 38.1 bits (87), Expect = 0.046
Identities = 28/114 (24%), Positives = 55/114 (47%), Gaps = 14/114 (12%)
Query: 3 KGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSD 62
K G +++++ LK+W Q + I + ++ S V VG++RR Y V +
Sbjct: 6 KIGSVVRIRQMLKQW-------QKKAHIGSSNNDPVSDVPPGHVAVSVGENRRRYVVRAK 58
Query: 63 VVDNPVFRELVERSRETEQQNDTVNV----VACEVVLFEHLLWMLENADPQPES 112
+++P+FR L+ E E++ NV + C+ LFE ++ ++ + S
Sbjct: 59 HLNHPIFRRLL---AEAEEEYGFANVGPLAIPCDESLFEDIIAIVTRCESSSSS 109
>UniRef100_Q6NMZ3 At4g34750 [Arabidopsis thaliana]
Length = 150
Score = 37.7 bits (86), Expect = 0.060
Identities = 28/114 (24%), Positives = 55/114 (47%), Gaps = 14/114 (12%)
Query: 3 KGGKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSD 62
K G +++++ LK+W Q + I + ++ S V VG++RR Y V +
Sbjct: 6 KIGSVVRIRRMLKQW-------QKKAHIGSSNNDPVSDVPPGHVAVSVGENRRRYVVRAK 58
Query: 63 VVDNPVFRELVERSRETEQQNDTVNV----VACEVVLFEHLLWMLENADPQPES 112
+++P+FR L+ E E++ NV + C+ LFE ++ ++ + S
Sbjct: 59 HLNHPIFRRLL---AEAEEEYGFANVGPLAIPCDESLFEDIIAIVTRCESSSSS 109
>UniRef100_Q7X7G3 OSJNBa0073E02.12 protein [Oryza sativa]
Length = 129
Score = 37.4 bits (85), Expect = 0.079
Identities = 29/86 (33%), Positives = 43/86 (49%), Gaps = 8/86 (9%)
Query: 4 GGKLMKLKSALKKWNSFGNGKQSRHSIS---AVADEDSSSS-RSDL---HTV-FVGKSRR 55
G K L++ L W G + + A D D + SD+ HTV +VG+ R
Sbjct: 5 GSKRRLLRAFLHSWKKLGAAAAAAAPAAGEWAPLDGDGEGAIPSDVPRGHTVVYVGEELR 64
Query: 56 LYRVTSDVVDNPVFRELVERSRETEQ 81
Y V +D+P+FREL++R+RE Q
Sbjct: 65 RYVVRVSSLDHPLFRELLDRAREEYQ 90
>UniRef100_Q8H6T6 Auxin-regulated protein [Phaseolus vulgaris]
Length = 156
Score = 35.8 bits (81), Expect = 0.23
Identities = 30/131 (22%), Positives = 57/131 (42%), Gaps = 29/131 (22%)
Query: 6 KLMKLKSALKKWNSFGNGKQSRHSISAVA---------------------DEDSSSSRSD 44
++++LK +KW + G + + S VA DEDS S
Sbjct: 14 QIVRLKEMFQKWQTVTLGSKESNHDSDVARPGGIPPMINKRLTNVLYCDSDEDSCYSPQP 73
Query: 45 LH-------TVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDTVNV-VACEVVLF 96
H V+VG R + + + + + +F+ L+E++ E + + + + CE+ F
Sbjct: 74 PHDVPKGYLAVYVGPELRRFIIPTSYLSHSLFKVLLEKAAEEFGFDQSGGLTIPCEIETF 133
Query: 97 EHLLWMLENAD 107
++LL +EN D
Sbjct: 134 KYLLNCMENHD 144
>UniRef100_Q9M249 Hypothetical protein F7M19_130 [Arabidopsis thaliana]
Length = 160
Score = 33.9 bits (76), Expect = 0.87
Identities = 33/144 (22%), Positives = 65/144 (44%), Gaps = 31/144 (21%)
Query: 6 KLMKLKSALKKWNSFGNGKQS--------RHS--ISAVADE---DSSSSRSDLHT----- 47
++++LK L+KW + G +S +H+ IS V ++ D + SD T
Sbjct: 14 QIVRLKEILQKWQTVTIGSKSDDGELGARKHTAIISPVINKRLLDLKTCDSDEETTCQSP 73
Query: 48 ------------VFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQ-QNDTVNVVACEVV 94
V+VG R + + ++ + + +F+ L+E++ E + + CEV
Sbjct: 74 EPPPDVPKGYLAVYVGPELRRFIIPTNFLSHSLFKVLLEKAEEEYGFDHSGALTIPCEVE 133
Query: 95 LFEHLLWMLENADPQPESLDELVD 118
F++LL +EN S ++ V+
Sbjct: 134 TFKYLLKCIENHPKDDTSAEDPVE 157
>UniRef100_Q9LR00 F10A5.21 [Arabidopsis thaliana]
Length = 108
Score = 33.9 bits (76), Expect = 0.87
Identities = 34/111 (30%), Positives = 53/111 (47%), Gaps = 14/111 (12%)
Query: 1 MAKGGKLMK---LKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDL----HTVFVGKS 53
M K KL + +K LK+ +S G KQS V ED + S ++ V+VG++
Sbjct: 3 MKKANKLTQTAMIKQILKRCSSLGK-KQSN-----VYGEDENGSPLNVPKGHFVVYVGEN 56
Query: 54 RRLYRVTSDVVDNPVFRELVERSRET-EQQNDTVNVVACEVVLFEHLLWML 103
R Y V + P F+ L++++ E +D + CE V+F L ML
Sbjct: 57 RVRYVVPISFLTRPEFQLLLQQAEEEFGFDHDMGLTIPCEEVVFRSLTSML 107
>UniRef100_Q8GWV6 Hypothetical protein At5g20810/T1M15_210 [Arabidopsis thaliana]
Length = 165
Score = 33.5 bits (75), Expect = 1.1
Identities = 30/131 (22%), Positives = 57/131 (42%), Gaps = 31/131 (23%)
Query: 6 KLMKLKSALKKWNS-----------FGNGKQSRHSISAV------------ADEDSSSSR 42
++++LK L+KW + GKQ+ IS +DE++ S
Sbjct: 14 QIVRLKEILQKWQTVTIGPKSEVPPLAAGKQAVAMISPAINKRLLDVKNGDSDEETCQSP 73
Query: 43 SDLH-------TVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDT-VNVVACEVV 94
H V+VG R + + + + + +F+ L+E++ E + + + CEV
Sbjct: 74 EPPHDVPKGNLAVYVGPELRRFIIPTSYLSHSLFKVLLEKAEEEFGFDQSGALTIPCEVE 133
Query: 95 LFEHLLWMLEN 105
F++LL +EN
Sbjct: 134 TFKYLLKCMEN 144
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.130 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 201,694,157
Number of Sequences: 2790947
Number of extensions: 6830816
Number of successful extensions: 18258
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 40
Number of HSP's that attempted gapping in prelim test: 18225
Number of HSP's gapped (non-prelim): 55
length of query: 122
length of database: 848,049,833
effective HSP length: 98
effective length of query: 24
effective length of database: 574,537,027
effective search space: 13788888648
effective search space used: 13788888648
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC124967.9


