

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
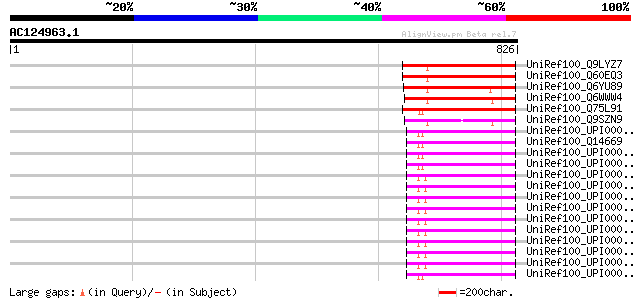
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LYZ7 Hypothetical protein F9G14_190 [Arabidopsis tha... 236 3e-60
UniRef100_Q60EQ3 Hypothetical protein OJ1280_A04.10 [Oryza sativa] 227 1e-57
UniRef100_Q6YU89 Putative HECT ubiquitin-protein ligase 3 [Oryza... 204 1e-50
UniRef100_Q6WWW4 HECT ubiquitin-protein ligase 3 [Arabidopsis th... 191 8e-47
UniRef100_Q75L91 Hypothetical protein OJ1729_E02.3 [Oryza sativa] 187 8e-46
UniRef100_Q9SZN9 Hypothetical protein F20M13.160 [Arabidopsis th... 178 7e-43
UniRef100_UPI000036988D UPI000036988D UniRef100 entry 150 1e-34
UniRef100_Q14669 Thyroid receptor interacting protein 12 [Homo s... 150 1e-34
UniRef100_UPI0000360B01 UPI0000360B01 UniRef100 entry 150 2e-34
UniRef100_UPI0000360B00 UPI0000360B00 UniRef100 entry 150 2e-34
UniRef100_UPI00001D02B5 UPI00001D02B5 UniRef100 entry 149 3e-34
UniRef100_UPI000017E000 UPI000017E000 UniRef100 entry 149 3e-34
UniRef100_UPI000043822D UPI000043822D UniRef100 entry 148 7e-34
UniRef100_UPI000043822C UPI000043822C UniRef100 entry 148 7e-34
UniRef100_UPI00004370EB UPI00004370EB UniRef100 entry 148 7e-34
UniRef100_UPI00004370EA UPI00004370EA UniRef100 entry 148 7e-34
UniRef100_UPI00004370E1 UPI00004370E1 UniRef100 entry 148 7e-34
UniRef100_UPI00004370E0 UPI00004370E0 UniRef100 entry 148 7e-34
UniRef100_UPI00002490A8 UPI00002490A8 UniRef100 entry 148 7e-34
UniRef100_UPI00003AECB5 UPI00003AECB5 UniRef100 entry 147 1e-33
>UniRef100_Q9LYZ7 Hypothetical protein F9G14_190 [Arabidopsis thaliana]
Length = 1502
Score = 236 bits (601), Expect = 3e-60
Identities = 119/206 (57%), Positives = 149/206 (71%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF---------------------- 678
+F +KIEDLCL+F+LPGY L P+S + NL +
Sbjct: 1294 SFHGTKIEDLCLEFALPGYTDYDLAPYSANDMVNLDNLEEYIKGIVNATVCNGIQKQVEA 1353
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFNQVF IEHLR+F+EEELE +LCGE + ++ N++L H KFDHGYT+ SPP+ LL+
Sbjct: 1354 FRSGFNQVFSIEHLRIFNEEELETMLCGECDLFSMNEVLDHIKFDHGYTSSSPPVEYLLQ 1413
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
IL EF+ E++RAF+QFVT SPRLP GGLASL PKLT+V+K + +DTDLPSVMTCANYL
Sbjct: 1414 ILHEFDREQQRAFLQFVTGSPRLPHGGLASLSPKLTIVRKHGSDSSDTDLPSVMTCANYL 1473
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKE+MK+KL+YAITEG+G F
Sbjct: 1474 KLPPYSSKEKMKEKLIYAITEGQGSF 1499
Score = 38.9 bits (89), Expect = 0.63
Identities = 17/34 (50%), Positives = 28/34 (82%), Gaps = 1/34 (2%)
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAI 34
V+++ALQ+ E++L+K+ D F+ FI+EGV+FAI
Sbjct: 503 VIVVALQVAEVLLEKY-RDTFLNSFIKEGVFFAI 535
>UniRef100_Q60EQ3 Hypothetical protein OJ1280_A04.10 [Oryza sativa]
Length = 523
Score = 227 bits (578), Expect = 1e-57
Identities = 114/206 (55%), Positives = 150/206 (72%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFL-LPFSIY*IT--NLYSF------------------- 678
++R +IEDL ++F+LPGY + L L S+ ++ NL +
Sbjct: 315 SYRGCRIEDLAIEFALPGYPEYVLSLENSLDNVSADNLEQYVSFVVDATIRSGIARQLEA 374
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFN+VFP+ L+VFSE+ELE +LCGE ++W F L+ H KFDHGYT+ SPP++NLLE
Sbjct: 375 FKSGFNEVFPLSMLQVFSEDELERLLCGEQDTWDFAKLVDHIKFDHGYTSSSPPVINLLE 434
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
++QEF +RRAF+QF+T SPRLPPGGLA+L+PKLTVV+K + N D DLPSVMTCANYL
Sbjct: 435 VIQEFEGHQRRAFLQFITGSPRLPPGGLAALNPKLTVVRKHNSNEADDDLPSVMTCANYL 494
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSK++M++KLLYAITEG+G F
Sbjct: 495 KLPPYSSKDKMREKLLYAITEGQGSF 520
>UniRef100_Q6YU89 Putative HECT ubiquitin-protein ligase 3 [Oryza sativa]
Length = 1781
Score = 204 bits (518), Expect = 1e-50
Identities = 110/217 (50%), Positives = 136/217 (61%), Gaps = 34/217 (15%)
Query: 642 FRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF----------------------F 679
FR + IEDLCLDF+LPGY L I N+Y+ F
Sbjct: 1562 FRGTPIEDLCLDFTLPGYPDYILKEGEENTIVNIYNLEEYVTLVVDATVKSGIMRQVEAF 1621
Query: 680 LVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEI 739
GFNQVF I L++FS EEL+ ++CG W + L+ + KFDHGYTA+SP I+NLLEI
Sbjct: 1622 RSGFNQVFDISSLKIFSPEELDYLICGRREIWEPDSLVDNIKFDHGYTAKSPAIVNLLEI 1681
Query: 740 LQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYN------------HTDTD 787
+ EF E++ AF QFVT +PRLPPGGLA+L+PKLT+V+K + D D
Sbjct: 1682 MAEFTPEQQHAFCQFVTGAPRLPPGGLAALNPKLTIVRKHPSSAVNTSNIAGVTESADDD 1741
Query: 788 LPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
LPSVMTCANYLKLPPYS+KE M++KLLYAI EGRG F
Sbjct: 1742 LPSVMTCANYLKLPPYSTKEVMRKKLLYAILEGRGSF 1778
>UniRef100_Q6WWW4 HECT ubiquitin-protein ligase 3 [Arabidopsis thaliana]
Length = 1888
Score = 191 bits (485), Expect = 8e-47
Identities = 107/214 (50%), Positives = 132/214 (61%), Gaps = 32/214 (14%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 1672 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 1731
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+NLLEI+ E
Sbjct: 1732 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIINLLEIMGE 1791
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 1792 LTADQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 1851
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 1852 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 1885
Score = 35.0 bits (79), Expect = 9.2
Identities = 17/45 (37%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAISHFHHLGKHLH 45
VL+ ALQ+ E++++K + F K+F+ EGV A+ +GK H
Sbjct: 620 VLVPALQVAEILMEKL-PETFSKVFVREGVVHAVDQLVLVGKPSH 663
>UniRef100_Q75L91 Hypothetical protein OJ1729_E02.3 [Oryza sativa]
Length = 1321
Score = 187 bits (476), Expect = 8e-46
Identities = 101/206 (49%), Positives = 132/206 (64%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLP----------------FSIY*------ITNLYSF 678
+++N ++EDLCLDF+LPG + L+P SI I+N
Sbjct: 1113 SYKNVRLEDLCLDFTLPGNPEYELVPGGSEKMVTLDNLEEYVSSIVDATLKSGISNQIEA 1172
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F G N+VF ++ LR+FSE+E+E ILCGE +SW N L H FD+GY A S +++ LE
Sbjct: 1173 FKAGINKVFALKTLRLFSEDEMERILCGEQDSWASNKLEDHINFDYGYDANSASVISFLE 1232
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
IL+EF E++RAF+ F T +P+LP GGLASLDPKLTVV+K D +LPSV TC ++
Sbjct: 1233 ILREFGREDQRAFLHFTTGAPQLPLGGLASLDPKLTVVRKQCDGKVDNELPSVNTCRHFF 1292
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKE M+QKL YAI EG G F
Sbjct: 1293 KLPPYSSKEIMRQKLKYAIKEGLGSF 1318
>UniRef100_Q9SZN9 Hypothetical protein F20M13.160 [Arabidopsis thaliana]
Length = 757
Score = 178 bits (451), Expect = 7e-43
Identities = 102/214 (47%), Positives = 129/214 (59%), Gaps = 35/214 (16%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 544 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 603
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+N +
Sbjct: 604 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIIN---VCYP 660
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 661 SSSDQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 720
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 721 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 754
>UniRef100_UPI000036988D UPI000036988D UniRef100 entry
Length = 1993
Score = 150 bits (380), Expect = 1e-34
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1788 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1847
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1848 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1907
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1908 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1967
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1968 DYSSIEIMREKLLIAAREGQQSF 1990
>UniRef100_Q14669 Thyroid receptor interacting protein 12 [Homo sapiens]
Length = 1992
Score = 150 bits (380), Expect = 1e-34
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1787 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1846
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1847 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1906
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1907 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1966
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1967 DYSSIEIMREKLLIAAREGQQSF 1989
>UniRef100_UPI0000360B01 UPI0000360B01 UniRef100 entry
Length = 1915
Score = 150 bits (378), Expect = 2e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +IY ++ + F GF
Sbjct: 1710 VEDLGLDFTLPGFPNIELKKGGKDIPVTIYNLEEYLRLVVYWTLNEGVSRQFESFREGFE 1769
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EELE +LCG + +W L++ + DHGYT S + L E++ F
Sbjct: 1770 SVFPLHHLQYFYPEELEQLLCGSKSETWDVKTLMECCRPDHGYTHDSRAVRFLFEVMSGF 1829
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1830 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1889
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1890 DYSSIETMREKLLIAAREGQQSF 1912
>UniRef100_UPI0000360B00 UPI0000360B00 UniRef100 entry
Length = 1982
Score = 150 bits (378), Expect = 2e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +IY ++ + F GF
Sbjct: 1777 VEDLGLDFTLPGFPNIELKKGGKDIPVTIYNLEEYLRLVVYWTLNEGVSRQFESFREGFE 1836
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EELE +LCG + +W L++ + DHGYT S + L E++ F
Sbjct: 1837 SVFPLHHLQYFYPEELEQLLCGSKSETWDVKTLMECCRPDHGYTHDSRAVRFLFEVMSGF 1896
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1897 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1956
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1957 DYSSIETMREKLLIAAREGQQSF 1979
>UniRef100_UPI00001D02B5 UPI00001D02B5 UniRef100 entry
Length = 2056
Score = 149 bits (376), Expect = 3e-34
Identities = 89/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 1851 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 1910
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1911 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1970
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1971 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 2030
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M+ KLL A EG+ F
Sbjct: 2031 DYSSIEIMRDKLLIAAREGQQSF 2053
>UniRef100_UPI000017E000 UPI000017E000 UniRef100 entry
Length = 588
Score = 149 bits (376), Expect = 3e-34
Identities = 89/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 383 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 442
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 443 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 502
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 503 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 562
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M+ KLL A EG+ F
Sbjct: 563 DYSSIEIMRDKLLIAAREGQQSF 585
>UniRef100_UPI000043822D UPI000043822D UniRef100 entry
Length = 1449
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1244 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1303
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1304 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1363
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1364 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1423
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1424 DYSSIEIMREKLLIAAREGQQSF 1446
>UniRef100_UPI000043822C UPI000043822C UniRef100 entry
Length = 1655
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1450 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1509
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1510 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1569
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1570 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1629
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1630 DYSSIEIMREKLLIAAREGQQSF 1652
>UniRef100_UPI00004370EB UPI00004370EB UniRef100 entry
Length = 1510
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1305 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1364
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1365 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1424
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1425 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1484
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1485 DYSSIEIMREKLLIAAREGQQSF 1507
>UniRef100_UPI00004370EA UPI00004370EA UniRef100 entry
Length = 1737
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1532 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1591
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1592 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1651
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1652 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1711
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1712 DYSSIEIMREKLLIAAREGQQSF 1734
>UniRef100_UPI00004370E1 UPI00004370E1 UniRef100 entry
Length = 1593
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1388 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1447
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1448 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1507
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1508 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1567
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1568 DYSSIEIMREKLLIAAREGQQSF 1590
>UniRef100_UPI00004370E0 UPI00004370E0 UniRef100 entry
Length = 1820
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1615 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1674
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1675 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1734
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1735 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1794
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1795 DYSSIEIMREKLLIAAREGQQSF 1817
>UniRef100_UPI00002490A8 UPI00002490A8 UniRef100 entry
Length = 1609
Score = 148 bits (373), Expect = 7e-34
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + + F GF
Sbjct: 1404 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYLRLVVYWTLNEGVLRQFESFREGFE 1463
Query: 685 QVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + SW L++ + DHGYT S + L E+L F
Sbjct: 1464 SVFPLHHLQYFYPEELDQLLCGSKSESWDVKTLMECCRPDHGYTHDSRAVRFLFEVLSSF 1523
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1524 DAEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1583
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1584 DYSSIEIMREKLLIAAREGQQSF 1606
>UniRef100_UPI00003AECB5 UPI00003AECB5 UniRef100 entry
Length = 1826
Score = 147 bits (372), Expect = 1e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L P +I+ + + F GF
Sbjct: 1621 VEDLGLDFTLPGFPNIELKKGGKDTPVTIHNLEEYLRLVIFWALNEGVARQFDSFRDGFE 1680
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EELE +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1681 SVFPLSHLQYFYPEELEQLLCGSKTDTWDAKTLMECCRPDHGYTHDSRAVKYLFEILSSF 1740
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
++E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1741 DSEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1800
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YS+ E M++KLL A EG+ F
Sbjct: 1801 DYSTIEIMREKLLIAAREGQQSF 1823
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,168,434,574
Number of Sequences: 2790947
Number of extensions: 42902291
Number of successful extensions: 194845
Number of sequences better than 10.0: 689
Number of HSP's better than 10.0 without gapping: 539
Number of HSP's successfully gapped in prelim test: 150
Number of HSP's that attempted gapping in prelim test: 192970
Number of HSP's gapped (non-prelim): 902
length of query: 826
length of database: 848,049,833
effective HSP length: 136
effective length of query: 690
effective length of database: 468,481,041
effective search space: 323251918290
effective search space used: 323251918290
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 79 (35.0 bits)
Medicago: description of AC124963.1


