

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
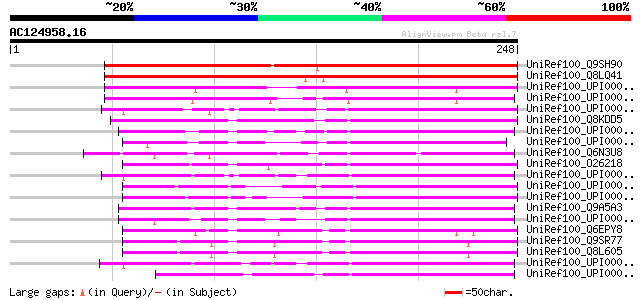
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SH90 Hypothetical protein At2g37970; T8P21.12 [Arabi... 299 5e-80
UniRef100_Q8LQ41 P0529H11.19 protein [Oryza sativa] 273 2e-72
UniRef100_UPI00002EBB25 UPI00002EBB25 UniRef100 entry 127 3e-28
UniRef100_UPI0000298E21 UPI0000298E21 UniRef100 entry 122 1e-26
UniRef100_UPI000029E73E UPI000029E73E UniRef100 entry 119 9e-26
UniRef100_Q8KDD5 Hypothetical protein CT1119 [Chlorobium tepidum] 119 9e-26
UniRef100_UPI0000344277 UPI0000344277 UniRef100 entry 112 1e-23
UniRef100_UPI00002A4BD5 UPI00002A4BD5 UniRef100 entry 105 1e-21
UniRef100_Q6N3U8 Hypothetical protein [Rhodopseudomonas palustris] 105 1e-21
UniRef100_O26218 Hypothetical protein MTH115 [Methanobacterium t... 104 2e-21
UniRef100_UPI00002E5468 UPI00002E5468 UniRef100 entry 103 4e-21
UniRef100_UPI00002BD7B5 UPI00002BD7B5 UniRef100 entry 103 4e-21
UniRef100_UPI00002E4A7D UPI00002E4A7D UniRef100 entry 102 7e-21
UniRef100_Q9A5A3 Hypothetical protein CC2549 [Caulobacter cresce... 102 9e-21
UniRef100_UPI00002772EA UPI00002772EA UniRef100 entry 100 3e-20
UniRef100_Q6EPY8 SOUL heme-binding protein-like [Oryza sativa] 100 4e-20
UniRef100_Q9SR77 T22K18.4 protein [Arabidopsis thaliana] 99 1e-19
UniRef100_Q8L605 Hypothetical protein At3g10130 [Arabidopsis tha... 99 1e-19
UniRef100_UPI00002C69CE UPI00002C69CE UniRef100 entry 98 2e-19
UniRef100_UPI000034075C UPI000034075C UniRef100 entry 97 3e-19
>UniRef100_Q9SH90 Hypothetical protein At2g37970; T8P21.12 [Arabidopsis thaliana]
Length = 215
Score = 299 bits (765), Expect = 5e-80
Identities = 151/216 (69%), Positives = 172/216 (78%), Gaps = 15/216 (6%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIG 106
MGMVFGKI VETPKY VTK+ YEIR Y P+VAAEVTYD S+FKG+KDGGF +LA YIG
Sbjct: 1 MGMVFGKIAVETPKYTVTKSGDGYEIREYPPAVAAEVTYDASEFKGDKDGGFQLLAKYIG 60
Query: 107 ALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGE--------------RNK 152
G P+N KPEKIAMTAPVITK EKIAMTAPV+TK SE+ E R K
Sbjct: 61 VFGKPENEKPEKIAMTAPVITK-EGEKIAMTAPVITKESEKIEMTSPVVTKEGGGEGRKK 119
Query: 153 MVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLS 212
+VTMQF+LPS Y+KAEEAP+PTDERVVI+EEG RKYGV+KFSG+AS+ VV EKV+KL
Sbjct: 120 LVTMQFLLPSMYKKAEEAPRPTDERVVIKEEGGRKYGVIKFSGIASESVVSEKVKKLSSH 179
Query: 213 LERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGFK+ GDF+L RYNPPWTLP FRTNEVMIP+E
Sbjct: 180 LEKDGFKITGDFVLARYNPPWTLPPFRTNEVMIPVE 215
>UniRef100_Q8LQ41 P0529H11.19 protein [Oryza sativa]
Length = 216
Score = 273 bits (699), Expect = 2e-72
Identities = 138/216 (63%), Positives = 163/216 (74%), Gaps = 14/216 (6%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIG 106
MGMV GKI VETPK+EV T YE+R Y P V AEVTYDP++ KG++DGGF VL NYIG
Sbjct: 1 MGMVLGKITVETPKHEVLHTGAGYEVRKYPPCVVAEVTYDPAEMKGDRDGGFTVLGNYIG 60
Query: 107 ALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTK-------------SSEEGERNK- 152
ALGNPQNTKPEKI MTAPVIT G E IAMTAPV+T ++EE + K
Sbjct: 61 ALGNPQNTKPEKIDMTAPVITSGEPESIAMTAPVITSGEPEPVAMTAPVITAEERSQGKG 120
Query: 153 MVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLS 212
+TMQF+LPS Y K EEAP+PTDERVV+R+ GERKYGVV+FSG+ D+VVKEK E L+ +
Sbjct: 121 QMTMQFLLPSKYSKVEEAPRPTDERVVLRQVGERKYGVVRFSGLTGDKVVKEKAEWLKAA 180
Query: 213 LERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGF V G F+L RYNPP+TLP RTNEVM+P+E
Sbjct: 181 LEKDGFTVKGPFVLARYNPPFTLPPLRTNEVMVPVE 216
>UniRef100_UPI00002EBB25 UPI00002EBB25 UniRef100 entry
Length = 194
Score = 127 bits (319), Expect = 3e-28
Identities = 79/207 (38%), Positives = 109/207 (52%), Gaps = 19/207 (9%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQ--FKGNKDGGFMVLANY 104
MG + GKI E PK+EV T YE+R YAP V AE ++ S+ F G++ G FM LA +
Sbjct: 1 MGSIIGKITEELPKHEVVGKTAAYELRRYAPCVVAETSFASSKGMFGGDQGGSFMRLAKF 60
Query: 105 IGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSS- 163
IG + PQN + E IAMTAPV+ + + + M F LP+S
Sbjct: 61 IGVMAKPQNEQAEPIAMTAPVL--------------MQRDDSRRGADGAYKMAFFLPASR 106
Query: 164 YEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG--FKVI 221
+ KA +APKPT+ V + + R FSG ++ EK +LR LERDG K
Sbjct: 107 FAKAADAPKPTNPDVTLVDLPARVVAAKTFSGNMRAGLIAEKERELREELERDGVRLKPG 166
Query: 222 GDFLLGRYNPPWTLPMFRTNEVMIPIE 248
+ + +NPPWT +TNEVM+ +E
Sbjct: 167 AEVMAAGFNPPWTPSFLKTNEVMLEVE 193
>UniRef100_UPI0000298E21 UPI0000298E21 UniRef100 entry
Length = 196
Score = 122 bits (305), Expect = 1e-26
Identities = 83/209 (39%), Positives = 105/209 (49%), Gaps = 23/209 (11%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPS--QFKGNKDGGFMVLANY 104
MG FG+I E P Y +T +YEIR Y P E +Y S G++ FM LA Y
Sbjct: 1 MGSAFGRITEEQPSYATIRTADEYEIRRYEPCCVIETSYASSAGMVSGDQGTSFMALARY 60
Query: 105 IGALGNPQNTKPEKIAMTAPVI--TKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPS 162
IG L P N + EKI MTAPV T G+A S + R M+F+LP
Sbjct: 61 IGVLSEPANERREKIDMTAPVFMTTDGAA------------SDTDAIR---YVMRFVLPK 105
Query: 163 SY--EKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG--F 218
S + A+ AP T ERV +R+ R V +FSG ++ V+ E LR +L RDG F
Sbjct: 106 SMFADGAKSAPASTQERVRVRDLDARVVAVRRFSGRMREDAVRAHAEILREALRRDGVAF 165
Query: 219 KVIGDFLLGRYNPPWTLPMFRTNEVMIPI 247
YNPPWT +FRTNEVMI +
Sbjct: 166 AENAASEYAGYNPPWTPGLFRTNEVMIEV 194
>UniRef100_UPI000029E73E UPI000029E73E UniRef100 entry
Length = 206
Score = 119 bits (297), Expect = 9e-26
Identities = 83/208 (39%), Positives = 117/208 (55%), Gaps = 20/208 (9%)
Query: 46 IMGMVFGKI--GVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDG---GFMV 100
I G++ G + VE P Y+V ++ Q+ EIR Y P + AEV F KD GF +
Sbjct: 11 IAGVLAGPVMSNVEKPDYKVVQSEQNIEIRQYEPMIIAEVEV----FGKRKDAINDGFRL 66
Query: 101 LANYIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFIL 160
LA+YI GN NT + I+MTAPV K + +KIAMTAPV + +E + + F++
Sbjct: 67 LADYI--FGN--NTLEKSISMTAPVQQKEN-QKIAMTAPVQQQLTENSWK-----ISFVM 116
Query: 161 PSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKV 220
PS Y K + P P + RV ++E +K+ V++FSG S E + E +L +ER ++
Sbjct: 117 PSKY-KMDSLPVPNNNRVRLKEIFTKKFVVIEFSGTNSTENLNEHENELMNYIERKQIQI 175
Query: 221 IGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
IG YN PWTLP R NEVMI I+
Sbjct: 176 IGSPKYAFYNAPWTLPFLRRNEVMIEIK 203
>UniRef100_Q8KDD5 Hypothetical protein CT1119 [Chlorobium tepidum]
Length = 215
Score = 119 bits (297), Expect = 9e-26
Identities = 72/199 (36%), Positives = 105/199 (52%), Gaps = 10/199 (5%)
Query: 50 VFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALG 109
V GK P YE+ K +E+R Y P V AE D + GF LA YI
Sbjct: 18 VLGKREAAEPPYELLKHDGAFEVRRYGPMVIAETILDEKSYSAASGKGFNRLAGYIFG-- 75
Query: 110 NPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
+N I+MTAPV+ + S+EKI+MTAPV+ + + G +M F+LP + +
Sbjct: 76 --KNRSKTSISMTAPVLQERSSEKISMTAPVLQQPQKGGW-----SMAFVLPEGFT-LQS 127
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRY 229
AP+P D V +RE VV FSG+ S +++ +L+ L++ G++ + + L Y
Sbjct: 128 APEPLDPEVKLRELPPSTIAVVTFSGLHSAANLEKYSRQLQAWLKKQGYRALSEPKLASY 187
Query: 230 NPPWTLPMFRTNEVMIPIE 248
+PPWT+P R NEV I IE
Sbjct: 188 DPPWTIPFLRRNEVQIRIE 206
>UniRef100_UPI0000344277 UPI0000344277 UniRef100 entry
Length = 203
Score = 112 bits (279), Expect = 1e-23
Identities = 73/194 (37%), Positives = 110/194 (56%), Gaps = 19/194 (9%)
Query: 54 IGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+ +E PKY++ K+ ++YEIR Y +A EV Y N+D GF L NYI +N
Sbjct: 23 MALEEPKYQIIKSNKNYEIRKYKDRLAVEVEYS------NEDSGFRYLFNYISG----EN 72
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
T EKI MT PV + KI MTAPV ++ + NKM+ MQF LPS + + APKP
Sbjct: 73 TSLEKINMTVPVT---QSVKINMTAPV----TQTTKNNKML-MQFFLPSKFS-LDTAPKP 123
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
T+++V + Y V+K+SG +++ ++ ++L+ L +D + +N P+
Sbjct: 124 TNKKVRLVILKSCYYAVIKYSGRVTNKNFQKHYKELKKYLIKDNINFTEPAIRATFNGPF 183
Query: 234 TLPMFRTNEVMIPI 247
TLP+FR NEVM+ I
Sbjct: 184 TLPVFRRNEVMMKI 197
>UniRef100_UPI00002A4BD5 UPI00002A4BD5 UniRef100 entry
Length = 164
Score = 105 bits (261), Expect = 1e-21
Identities = 71/190 (37%), Positives = 96/190 (50%), Gaps = 33/190 (17%)
Query: 56 VETPKYEVTKT--TQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P Y V + + EIR YAP V A D N D GF VLA YI N
Sbjct: 6 IEEPAYTVEQAWEAEQIEIRAYAPRVMAVTGMDE-----NADSGFRVLAGYIFG----GN 56
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+ EKIAMTAPV + S GE+ M F++P+ + E+ P+P
Sbjct: 57 AEEEKIAMTAPV-----------------QQSMAGEKE----MAFMMPAEFA-LEDLPEP 94
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
D+RV RE V++FSG AS E E ++LR L ++G +++G+ L +YNPPW
Sbjct: 95 EDQRVSFREAPAYTAAVIQFSGWASAEKADENWQQLRRFLIKEGIEIVGEPTLNQYNPPW 154
Query: 234 TLPMFRTNEV 243
TLP R NE+
Sbjct: 155 TLPFLRRNEI 164
>UniRef100_Q6N3U8 Hypothetical protein [Rhodopseudomonas palustris]
Length = 209
Score = 105 bits (261), Expect = 1e-21
Identities = 77/213 (36%), Positives = 117/213 (54%), Gaps = 15/213 (7%)
Query: 37 HFTLSHSRSIMGMVFGKIGVETPKYEVTKTTQD-YEIRIYAPSVAAEVTYDPSQFKGNKD 95
++T+ + S++G+V ++ E P Y V D EIR YAP VAAEV + +GN D
Sbjct: 7 YYTVLVAESVLGIVGLRL-YEEPAYTVLDRPSDTIEIRRYAPRVAAEVDLER---RGNAD 62
Query: 96 G-GFMVLANYIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMV 154
G F +L NYI + E++AMT PV A KIAMTAPV T + + +M
Sbjct: 63 GQAFTLLFNYIAGANRGGSGTSERVAMTVPVDVARPA-KIAMTAPVETATQD-----RMT 116
Query: 155 TMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLE 214
M+F LP+++ A+ APKP+DERV I E+ ++FSG D ++E+ ++L +L
Sbjct: 117 RMRFFLPATFT-ADTAPKPSDERVQIVTVPEQTIATLRFSGTGRD--LREREQQLIAALA 173
Query: 215 RDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPI 247
++ +G Y+ P+TLP R NE + +
Sbjct: 174 NTPWQPVGAPYGLFYDAPFTLPFVRRNEAAVEV 206
>UniRef100_O26218 Hypothetical protein MTH115 [Methanobacterium thermoautotrophicum]
Length = 189
Score = 104 bits (260), Expect = 2e-21
Identities = 63/195 (32%), Positives = 109/195 (55%), Gaps = 9/195 (4%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
VE+P+Y V +EIR Y + A+V + S F+ GF +LANYI N +
Sbjct: 2 VESPEYTVELKDGKFEIRRYPGYILAQVDVEAS-FRDAMVIGFSILANYIFG----GNRR 56
Query: 116 PEKIAMTAPV--ITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
E++ MT+PV + GS+E+I M PV + ++ + K + F +PSSY E P+P
Sbjct: 57 KEELPMTSPVTGVNLGSSERIPMKVPVTEEVPDDADSGKY-RISFTMPSSYT-LETLPEP 114
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
D+R+ REE ++++ +FSG + ++ +++ +L+ LER+ + +F++ +YN P
Sbjct: 115 LDDRIRFREEKDQRFAAYRFSGRVNSDMAAQRIAELKEWLERNSIEPRSNFIIAQYNHPA 174
Query: 234 TLPMFRTNEVMIPIE 248
R NEV++ I+
Sbjct: 175 VPGFLRKNEVLVKID 189
>UniRef100_UPI00002E5468 UPI00002E5468 UniRef100 entry
Length = 202
Score = 103 bits (257), Expect = 4e-21
Identities = 73/204 (35%), Positives = 112/204 (54%), Gaps = 14/204 (6%)
Query: 46 IMGMVFGKI--GVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLAN 103
I G++ G + VE P Y++ ++ Q+ EIR Y P + AEV + + + GF ++A+
Sbjct: 11 ICGVLAGPVMSNVEKPDYKIIESQQNIEIREYDPMIIAEVEVGGKR-EDAINNGFRIIAD 69
Query: 104 YIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSS 163
YI GN N + I+MTAPV K + KIAMTAPV ++ + K + F++PS
Sbjct: 70 YI--FGN--NEVKKTISMTAPVQQKKN-HKIAMTAPV-----QQQLKGKSWRISFVMPSK 119
Query: 164 YEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD 223
Y + P P + RV ++E +K+ V++FSG S E + + +L ++ + + IG
Sbjct: 120 YSM-DSLPVPNNNRVHLKEILAKKFVVIQFSGTNSHENLTKHENQLMNYVKVNKIEFIGS 178
Query: 224 FLLGRYNPPWTLPMFRTNEVMIPI 247
YN PWTLP R NEVMI I
Sbjct: 179 PKYAYYNAPWTLPFLRRNEVMIEI 202
>UniRef100_UPI00002BD7B5 UPI00002BD7B5 UniRef100 entry
Length = 174
Score = 103 bits (257), Expect = 4e-21
Identities = 61/193 (31%), Positives = 103/193 (52%), Gaps = 23/193 (11%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+ETP+Y++ + +EIR+Y P + A +T S ++ + GF +ANYI GN +N +
Sbjct: 5 IETPEYQLLEKKGQFEIRVYEPMIIA-ITNINSNYRQSTSTGFRRIANYIFG-GNDENME 62
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
IAMTAPV++ S E + + F++P + + P P
Sbjct: 63 ------------------IAMTAPVISSSPVNNEG--IYDVAFVMPKKHSLSN-LPNPNY 101
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
+ V+I+EE K V++F G A++E + + +KL L + F + G F++ +YN PW +
Sbjct: 102 DNVIIKEENLGKMAVLRFGGWATEEKIIKNQKKLEKLLIENRFVISGKFIVAQYNSPWVI 161
Query: 236 PMFRTNEVMIPIE 248
P FR NE+M+ I+
Sbjct: 162 PPFRRNEIMVSIK 174
>UniRef100_UPI00002E4A7D UPI00002E4A7D UniRef100 entry
Length = 172
Score = 102 bits (255), Expect = 7e-21
Identities = 67/193 (34%), Positives = 98/193 (50%), Gaps = 23/193 (11%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+ETPKY++ K T YEIR Y P A + S ++ GF +A YI GN +N +
Sbjct: 2 LETPKYKLIKKTGAYEIRDYEPMTIARTVVN-SDYRDATSAGFRRIAGYIFG-GNDENME 59
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
IAMTAPVI S+ G+ + F++PSS+E++ PKP
Sbjct: 60 ---IAMTAPVI-----------------SNSPGDVEDEYEILFVMPSSFERSS-LPKPNS 98
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
V I + + F G A+ E K+ KL L+ +G++V G F++ +YN PW L
Sbjct: 99 STVEIMDRSLNLIACISFGGWATKERAKKYHLKLSNWLKEEGYEVDGKFMVAQYNSPWAL 158
Query: 236 PMFRTNEVMIPIE 248
P FR NE+M+ I+
Sbjct: 159 PPFRHNEIMVRIK 171
>UniRef100_Q9A5A3 Hypothetical protein CC2549 [Caulobacter crescentus]
Length = 208
Score = 102 bits (254), Expect = 9e-21
Identities = 70/194 (36%), Positives = 100/194 (51%), Gaps = 11/194 (5%)
Query: 54 IGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+ VE P ++V D+++R Y V AEVT Q K + GF +LA YI N
Sbjct: 24 MAVEEPVFKVVLHEGDFDVRDYPALVVAEVTVSGDQ-KQAANRGFRLLAGYIFG----GN 78
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+ IAMTAPV + + IAMTAPV T++ G+ ++F +PS Y E P+P
Sbjct: 79 RTRQSIAMTAPVAQAPAGQTIAMTAPV-TQTQSAGQW----VVRFTMPSRYS-LEALPEP 132
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
D +V +R + V++FSG+A + V+ K L+ L + G L +YN PW
Sbjct: 133 NDPQVKLRLIPPSRLAVLRFSGLAGADTVEVKTADLKKRLSAHQLQATGPATLAQYNTPW 192
Query: 234 TLPMFRTNEVMIPI 247
T R NEVMIP+
Sbjct: 193 TPWFMRRNEVMIPV 206
>UniRef100_UPI00002772EA UPI00002772EA UniRef100 entry
Length = 172
Score = 100 bits (250), Expect = 3e-20
Identities = 66/196 (33%), Positives = 96/196 (48%), Gaps = 33/196 (16%)
Query: 54 IGVETPKYEVTKTTQD--YEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNP 111
+ +E P YEV K ++ +IR Y+P V A D K+ GF +LA YI
Sbjct: 1 MAIEEPNYEVVKEWENASIQIRAYSPRVMAVTEIDEG-----KNNGFRILAGYIFG---- 51
Query: 112 QNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAP 171
NT+ +KI MTAPV + M P M F++PS Y + P
Sbjct: 52 GNTREQKIEMTAPV-------QQTMDGPP--------------EMAFMMPSEYS-LPDLP 89
Query: 172 KPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNP 231
P D+RV RE V++F+G A + +E+ +KL+ L ++IG L +YNP
Sbjct: 90 TPNDKRVTFRETPAFTAAVIRFNGWADLQKAEERWQKLKTFLVNQNIQIIGQPTLNQYNP 149
Query: 232 PWTLPMFRTNEVMIPI 247
PWTLP R NE+++P+
Sbjct: 150 PWTLPFLRRNEIIVPV 165
>UniRef100_Q6EPY8 SOUL heme-binding protein-like [Oryza sativa]
Length = 287
Score = 100 bits (248), Expect = 4e-20
Identities = 69/202 (34%), Positives = 106/202 (52%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQ---FKGNKDGGFMVLANYIGALGNPQ 112
+ET + V K +YEIR AE T F G+ F VLA+Y+ +
Sbjct: 90 LETVPFRVLKREAEYEIREVESYYVAETTMPGRSGFDFNGSSQS-FNVLASYLFG----K 144
Query: 113 NTKPEKIAMTAPVITKGS---AEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT E++ MT PV T+ EK+ MT PV+TK S + KM F++PS Y +
Sbjct: 145 NTTSEQMEMTTPVFTRKGEPDGEKMDMTTPVITKKSANENKWKM---SFVMPSKY--GPD 199
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG-FKVIGDFL--L 226
P P D V I+E + V FSG+ +D+ + ++ +LR +L++D F+V D + +
Sbjct: 200 LPLPKDPSVTIKEVPAKIVAVAAFSGLVTDDDISQRESRLRETLQKDSQFRVKDDSVVEI 259
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NE+ + ++
Sbjct: 260 AQYNPPFTLPFTRRNEIALEVK 281
>UniRef100_Q9SR77 T22K18.4 protein [Arabidopsis thaliana]
Length = 309
Score = 98.6 bits (244), Expect = 1e-19
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGG---FMVLANYIGALGNPQ 112
+ET + V T YEIR P AE T P + + G F VLA Y+ +
Sbjct: 114 LETMNFRVLFRTDKYEIRQVEPYFVAE-TIMPGETGFDSYGASKSFNVLAEYLFG----K 168
Query: 113 NTKPEKIAMTAPVITK---GSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT EK+ MT PV+T+ EK+ MT PV+T +++ + +M F++PS Y
Sbjct: 169 NTIKEKMEMTTPVVTRKVQSVGEKMEMTTPVITSKAKDQNQWRM---SFVMPSKY--GSN 223
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD---FLL 226
P P D V I++ + VV FSG +DE ++ + +LR +L+ D + D F +
Sbjct: 224 LPLPKDPSVKIQQVPRKIVAVVAFSGYVTDEEIERRERELRRALQNDKKFRVRDGVSFEV 283
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NEV + +E
Sbjct: 284 AQYNPPFTLPFMRRNEVSLEVE 305
>UniRef100_Q8L605 Hypothetical protein At3g10130 [Arabidopsis thaliana]
Length = 309
Score = 98.6 bits (244), Expect = 1e-19
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGG---FMVLANYIGALGNPQ 112
+ET + V T YEIR P AE T P + + G F VLA Y+ +
Sbjct: 114 LETMNFRVLFRTDKYEIRQVEPYFVAE-TIMPGETGFDSYGASKSFNVLAEYLFG----K 168
Query: 113 NTKPEKIAMTAPVITK---GSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT EK+ MT PV+T+ EK+ MT PV+T +++ + +M F++PS Y
Sbjct: 169 NTIKEKMEMTTPVVTRKVQSVGEKMEMTTPVITSKAKDQNQWRM---SFVMPSKY--GSN 223
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD---FLL 226
P P D V I++ + VV FSG +DE ++ + +LR +L+ D + D F +
Sbjct: 224 LPLPKDPSVKIQQVPRKIVAVVAFSGYVTDEEIERRERELRRALQNDKKFRVRDGVSFEV 283
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NEV + +E
Sbjct: 284 AQYNPPFTLPFMRRNEVSLEVE 305
>UniRef100_UPI00002C69CE UPI00002C69CE UniRef100 entry
Length = 211
Score = 98.2 bits (243), Expect = 2e-19
Identities = 67/205 (32%), Positives = 105/205 (50%), Gaps = 14/205 (6%)
Query: 45 SIMGMVFGKI--GVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLA 102
+I G + G I VE P Y++ + T++ EIR Y P + AEV S+ K + GF +LA
Sbjct: 17 AIGGAMIGPIMSNVEIPDYKILEKTENIEIRRYPPLIIAEVRTMGSR-KDSIGDGFRILA 75
Query: 103 NYIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPS 162
++I +N KI+MT PV+ + K MTAPV ++ K + FI+PS
Sbjct: 76 DFIFGKYKGEN----KISMTTPVMQQ-EGTKNEMTAPV-----QQENTGKEWLVSFIMPS 125
Query: 163 SYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIG 222
+ PKPT++++ I + + Y + FSG + + + +L+ L R + + G
Sbjct: 126 KFSM-NTLPKPTNDKIKIVQMPSKNYAAIVFSGRGTKDNLTANTLELKEYLSRSNYSIRG 184
Query: 223 DFLLGRYNPPWTLPMFRTNEVMIPI 247
+ YNPPWTLP R NEV +
Sbjct: 185 NPKYAFYNPPWTLPFMRRNEVQFEL 209
>UniRef100_UPI000034075C UPI000034075C UniRef100 entry
Length = 180
Score = 97.4 bits (241), Expect = 3e-19
Identities = 59/176 (33%), Positives = 98/176 (55%), Gaps = 9/176 (5%)
Query: 72 IRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPVITKGSA 131
IR + AE T D + + +++ F L YI Q+ I MT PV + +
Sbjct: 11 IRRVPSLLVAETTVDIADYDQSRNVAFKRLFKYISGSNRTQSD----IVMTVPVESSYIS 66
Query: 132 EKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVV 191
E+I+MTAPV + + EG+ + M+F LP+ Y + APKPTD+ V + + + V+
Sbjct: 67 EQISMTAPVESIRTAEGQ----MRMRFFLPAEYSY-QTAPKPTDQNVNLVKIPAQSQAVL 121
Query: 192 KFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPI 247
+F+G A++ +V+ K +L L+ G+ +IGD Y+PPWT+ +FR NE++I +
Sbjct: 122 RFNGQANESIVESKTLELLKILDNSGWGIIGDPSAYFYDPPWTISIFRRNEIVISV 177
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 417,293,214
Number of Sequences: 2790947
Number of extensions: 17519143
Number of successful extensions: 35436
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 118
Number of HSP's successfully gapped in prelim test: 102
Number of HSP's that attempted gapping in prelim test: 34997
Number of HSP's gapped (non-prelim): 236
length of query: 248
length of database: 848,049,833
effective HSP length: 124
effective length of query: 124
effective length of database: 501,972,405
effective search space: 62244578220
effective search space used: 62244578220
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 73 (32.7 bits)
Medicago: description of AC124958.16


