

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.2 + phase: 0
(53 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
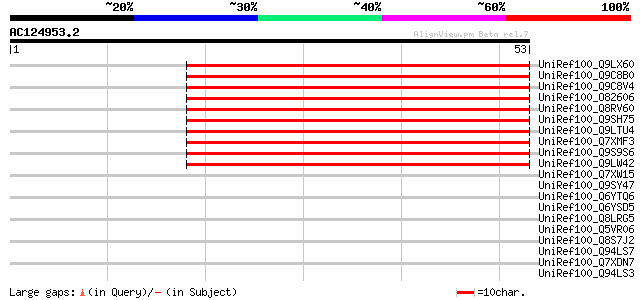
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 52 5e-06
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 51 9e-06
UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis tha... 51 9e-06
UniRef100_O82606 T2L5.8 protein [Arabidopsis thaliana] 49 3e-05
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 49 5e-05
UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana] 48 6e-05
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 48 6e-05
UniRef100_Q7XMF3 OSJNBa0061G20.6 protein [Oryza sativa] 48 8e-05
UniRef100_Q9S9S6 F28J9.3 [Arabidopsis thaliana] 47 1e-04
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 44 9e-04
UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa] 44 0.001
UniRef100_Q9SY47 Hypothetical protein T5L23.19 [Arabidopsis thal... 42 0.004
UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa] 42 0.006
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 42 0.006
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 42 0.006
UniRef100_Q5VR06 Helicase-like protein [Oryza sativa] 41 0.007
UniRef100_Q8S7J2 Hypothetical protein OSJNBa0095C06.16 [Oryza sa... 40 0.021
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 37 0.14
UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa] 37 0.14
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 37 0.14
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 51.6 bits (122), Expect = 5e-06
Identities = 24/35 (68%), Positives = 30/35 (85%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D++G+ TTNVV+KEVFQN+
Sbjct: 1717 RVTSKKGLKILILDKDGNMQKQTTNVVFKEVFQNI 1751
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 50.8 bits (120), Expect = 9e-06
Identities = 24/35 (68%), Positives = 29/35 (82%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D++G TTNVV+KEVFQN+
Sbjct: 1643 RVTSKKGLKILILDKDGKLQKQTTNVVFKEVFQNI 1677
>UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis thaliana]
Length = 729
Score = 50.8 bits (120), Expect = 9e-06
Identities = 24/35 (68%), Positives = 29/35 (82%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D++G TTNVV+KEVFQN+
Sbjct: 694 RVTSKKGLKILILDKDGKLQKQTTNVVFKEVFQNI 728
>UniRef100_O82606 T2L5.8 protein [Arabidopsis thaliana]
Length = 1073
Score = 49.3 bits (116), Expect = 3e-05
Identities = 23/35 (65%), Positives = 28/35 (79%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLK LI D++G TTNVV+KEVFQN+
Sbjct: 1038 RVTSKKGLKFLILDKDGKLQKQTTNVVFKEVFQNI 1072
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 48.5 bits (114), Expect = 5e-05
Identities = 24/35 (68%), Positives = 28/35 (79%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV SR+GLKILI DE G+ TTNVV+KEVFQN+
Sbjct: 1267 RVKSRRGLKILIIDEEGNRGKTTTNVVFKEVFQNL 1301
>UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana]
Length = 1241
Score = 48.1 bits (113), Expect = 6e-05
Identities = 21/35 (60%), Positives = 28/35 (80%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLK+LI D+ G+ T NVV+KE+FQN+
Sbjct: 1205 RVTSKKGLKVLIVDKEGNTQSQTMNVVFKEIFQNL 1239
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 48.1 bits (113), Expect = 6e-05
Identities = 23/35 (65%), Positives = 27/35 (76%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+ GLKILI D+ G TTNVV+KEVFQN+
Sbjct: 1394 RVTSKTGLKILILDKEGKIQKQTTNVVFKEVFQNI 1428
>UniRef100_Q7XMF3 OSJNBa0061G20.6 protein [Oryza sativa]
Length = 1410
Score = 47.8 bits (112), Expect = 8e-05
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
R TSRKGLKILI D++G+C T NVVY E+ Q +
Sbjct: 595 RATSRKGLKILIVDDDGNCSSETRNVVYHEILQAI 629
>UniRef100_Q9S9S6 F28J9.3 [Arabidopsis thaliana]
Length = 436
Score = 47.0 bits (110), Expect = 1e-04
Identities = 22/35 (62%), Positives = 28/35 (79%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVT++KGLKILI D+ G TTNVV+K+VFQN+
Sbjct: 401 RVTAKKGLKILILDKYGKLHKQTTNVVFKKVFQNI 435
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 44.3 bits (103), Expect = 9e-04
Identities = 20/35 (57%), Positives = 27/35 (77%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV S+ GLK+LITD G + TTNVV+KE+F+N+
Sbjct: 1635 RVKSKGGLKVLITDSKGKQKNETTNVVFKEIFRNL 1669
>UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa]
Length = 2052
Score = 43.9 bits (102), Expect = 0.001
Identities = 19/35 (54%), Positives = 25/35 (71%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
R TSR GLKILI D+N C T+NVVY E+ +++
Sbjct: 1384 RATSRSGLKILIEDDNESCASETSNVVYHEILRSL 1418
>UniRef100_Q9SY47 Hypothetical protein T5L23.19 [Arabidopsis thaliana]
Length = 570
Score = 42.0 bits (97), Expect = 0.004
Identities = 19/35 (54%), Positives = 25/35 (71%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV S+ LK+LITD G TTNV++KE+FQN+
Sbjct: 535 RVKSKARLKVLITDSKGKQKKETTNVIFKEIFQNL 569
>UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa]
Length = 1430
Score = 41.6 bits (96), Expect = 0.006
Identities = 19/31 (61%), Positives = 23/31 (73%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEV 49
RVTSR GLKILI +++G C T N+VY EV
Sbjct: 1357 RVTSRSGLKILIENDDGSCGTQTKNIVYSEV 1387
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 41.6 bits (96), Expect = 0.006
Identities = 19/31 (61%), Positives = 23/31 (73%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEV 49
RVTSR GLKILI +++G C T N+VY EV
Sbjct: 1443 RVTSRSGLKILIENDDGSCGTQTKNIVYSEV 1473
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 41.6 bits (96), Expect = 0.006
Identities = 20/35 (57%), Positives = 24/35 (68%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
R TSR GL+ILI D++G C T NVVY EV + V
Sbjct: 1407 RSTSRDGLRILIKDDDGACSSKTRNVVYHEVLEAV 1441
>UniRef100_Q5VR06 Helicase-like protein [Oryza sativa]
Length = 1427
Score = 41.2 bits (95), Expect = 0.007
Identities = 17/34 (50%), Positives = 25/34 (73%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQN 52
RV++ KGLKILI +E+G C T N+VY+E+ +
Sbjct: 1391 RVSNSKGLKILIENEDGTCATQTKNIVYREILDS 1424
>UniRef100_Q8S7J2 Hypothetical protein OSJNBa0095C06.16 [Oryza sativa]
Length = 1414
Score = 39.7 bits (91), Expect = 0.021
Identities = 19/35 (54%), Positives = 24/35 (68%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
R TSR+G++ILI DE+ C T NVVY EV + V
Sbjct: 714 RSTSREGIRILIEDEDEVCCSQTRNVVYHEVLEAV 748
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 37.0 bits (84), Expect = 0.14
Identities = 18/33 (54%), Positives = 22/33 (66%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQ 51
RVTSR GLKI+I D+ N+VYKE+FQ
Sbjct: 1541 RVTSRDGLKIMIADKECPGEGMVKNIVYKEIFQ 1573
>UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa]
Length = 1477
Score = 37.0 bits (84), Expect = 0.14
Identities = 19/33 (57%), Positives = 21/33 (63%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQ 51
RVTSR GLKILI DE N+VYKE+ Q
Sbjct: 1445 RVTSRDGLKILIADEECPGEGMVKNIVYKEILQ 1477
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 37.0 bits (84), Expect = 0.14
Identities = 19/33 (57%), Positives = 21/33 (63%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQ 51
RVTSR GLKILI DE N+VYKE+ Q
Sbjct: 1469 RVTSRDGLKILIADEECPGEGMVKNIVYKEILQ 1501
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 83,573,531
Number of Sequences: 2790947
Number of extensions: 2217254
Number of successful extensions: 7367
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 7340
Number of HSP's gapped (non-prelim): 27
length of query: 53
length of database: 848,049,833
effective HSP length: 29
effective length of query: 24
effective length of database: 767,112,370
effective search space: 18410696880
effective search space used: 18410696880
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC124953.2


