

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.18 + phase: 0
(391 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
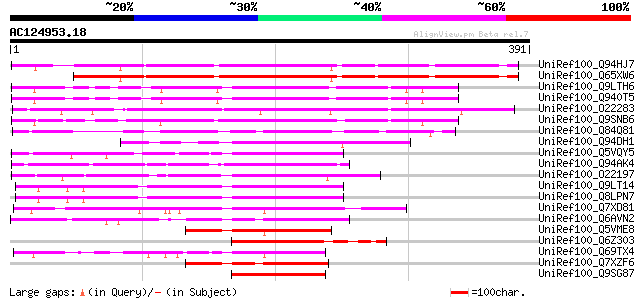
Sequences producing significant alignments: (bits) Value
UniRef100_Q94HJ7 Putative RING-H2 finger protein [Oryza sativa] 305 1e-81
UniRef100_Q65XW6 Putative ring-H2 finger protein [Oryza sativa] 280 4e-74
UniRef100_Q9LTH6 Emb|CAB62332.1 [Arabidopsis thaliana] 280 5e-74
UniRef100_Q940T5 AT5g59550/f2o15_210 [Arabidopsis thaliana] 280 5e-74
UniRef100_O22283 Expressed protein [Arabidopsis thaliana] 279 8e-74
UniRef100_Q9SNB6 Hypothetical protein F12A12.140 [Arabidopsis th... 276 5e-73
UniRef100_Q84Q81 Hypothetical protein OJ1261C08.10 [Oryza sativa] 236 1e-60
UniRef100_Q94DH1 Putative RING-H2 finger protein RHC2a [Oryza sa... 177 3e-43
UniRef100_Q5VQY5 Putative ring finger protein 126 isoform 1 [Ory... 141 3e-32
UniRef100_Q94AK4 Hypothetical protein At3g56580 [Arabidopsis tha... 129 2e-28
UniRef100_O22197 Expressed protein [Arabidopsis thaliana] 128 2e-28
UniRef100_Q9LT14 Arabidopsis thaliana genomic DNA, chromosome 3,... 124 3e-27
UniRef100_Q8LPN7 AT3g19950/MPN9_19 [Arabidopsis thaliana] 124 3e-27
UniRef100_Q7XD81 Putative zinc finger protein [Oryza sativa] 110 5e-23
UniRef100_Q6AVN2 Hypothetical protein OJ1119_H02.11 [Oryza sativa] 107 7e-22
UniRef100_Q5VME8 Putative ring finger protein 126 isoform 1 [Ory... 105 2e-21
UniRef100_Q6Z303 Zinc finger-like [Oryza sativa] 105 2e-21
UniRef100_Q69TX4 Zinc finger-like [Oryza sativa] 103 8e-21
UniRef100_Q7XZF6 Hypothetical protein OSJNBb0033J23.18 [Oryza sa... 100 7e-20
UniRef100_Q9SG87 Putative RING zinc finger protein [Arabidopsis ... 98 5e-19
>UniRef100_Q94HJ7 Putative RING-H2 finger protein [Oryza sativa]
Length = 386
Score = 305 bits (781), Expect = 1e-81
Identities = 177/391 (45%), Positives = 229/391 (58%), Gaps = 42/391 (10%)
Query: 2 SSHWCYRCNKFVRVWRL---GMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMY 58
SS+WCY C++FVR CPDC GFLE+ M P A Y
Sbjct: 19 SSYWCYSCDRFVRAPAPHDDSAVACPDCGGGFLEE---------------MSAPPPRAAY 63
Query: 59 MIGHRNNNYNQNTFRRHRRNNVNG--GDISPFNPIIMIRGGGGSSEGTSREREENNEFEL 116
+ R ++ N RR RR GD SPFNP+I++R ++ G FEL
Sbjct: 64 LRRPRAHHANDLRLRRTRRAAAAAAAGDRSPFNPVIVLRRSPAAA-GDDDSLAAATSFEL 122
Query: 117 FYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVEL 176
FY+DGAGSGLR LP MS+ ++GSGFER+++QL+ +EA G +++ PA K++VE
Sbjct: 123 FYDDGAGSGLRPLPETMSDFLMGSGFERLLDQLTQIEA---GGLARARENPPASKASVES 179
Query: 177 LPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHE 236
+PT+ I SH+ +SHCAVCKEPFELG AREMPC HIYH +CILPWLA++NSCPVCRHE
Sbjct: 180 MPTVTIAASHVGADSHCAVCKEPFELGDEAREMPCSHIYHQDCILPWLALRNSCPVCRHE 239
Query: 237 LPCES--PQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENRMEHPIVYTEV 294
+P ++ P+ +N E+E VGLTIWRLPGGGFAVGRF+ GG E P+VYTE+
Sbjct: 240 MPTDAARPRPSNA---GTEEETVGLTIWRLPGGGFAVGRFA--GGRRPEERELPVVYTEM 294
Query: 295 DGAFNNVGEPRRISWSLTSSRGGIGRSRGGAFRRMLSNLFGCL-RGGGVRNQHSPFTTRE 353
DG FNN G PRRISW SR + A RR+ N+F C R +Q S +R
Sbjct: 295 DGGFNNGGAPRRISWGSRQSRS----TERSAIRRIFRNVFSCFGRSHSSNSQASSSHSRP 350
Query: 354 FPQPMTMRNNSASHTNENPSLRSRRT-WSMD 383
+ R+ SH + RSR T W ++
Sbjct: 351 ELNDASDRSAVFSHGS-----RSRSTSWRLE 376
>UniRef100_Q65XW6 Putative ring-H2 finger protein [Oryza sativa]
Length = 333
Score = 280 bits (717), Expect = 4e-74
Identities = 160/341 (46%), Positives = 208/341 (60%), Gaps = 24/341 (7%)
Query: 49 MRFPMAAAMYMIGHRNNNYNQNTFRRHRRNNVNG--GDISPFNPIIMIRGGGGSSEGTSR 106
M P A Y+ R ++ N RR RR GD SPFNP+I++R ++ G
Sbjct: 1 MSAPPPRAAYLRRPRAHHANDLRLRRTRRAAAAAAAGDRSPFNPVIVLRRSPAAA-GDDD 59
Query: 107 EREENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQH 166
FELFY+DGAGSGLR LP MS+ ++GSGFER+++QL+ +EA G +++
Sbjct: 60 SLAAATSFELFYDDGAGSGLRPLPETMSDFLMGSGFERLLDQLTQIEA---GGLARAREN 116
Query: 167 LPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
PA K++VE +PT+ I SH+ +SHCAVCKEPFELG AREMPC HIYH +CILPWLA+
Sbjct: 117 PPASKASVESMPTVTIAASHVGADSHCAVCKEPFELGDEAREMPCSHIYHQDCILPWLAL 176
Query: 227 QNSCPVCRHELPCES--PQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENR 284
+NSCPVCRHE+P ++ P+ +N E+E VGLTIWRLPGGGFAVGRF+ GG
Sbjct: 177 RNSCPVCRHEMPTDAARPRPSNA---GTEEETVGLTIWRLPGGGFAVGRFA--GGRRPEE 231
Query: 285 MEHPIVYTEVDGAFNNVGEPRRISWSLTSSRGGIGRSRGGAFRRMLSNLFGCL-RGGGVR 343
E P+VYTE+DG FNN G PRRISW SR + A RR+ N+F C R
Sbjct: 232 RELPVVYTEMDGGFNNGGAPRRISWGSRQSRS----TERSAIRRIFRNVFSCFGRSHSSN 287
Query: 344 NQHSPFTTREFPQPMTMRNNSASHTNENPSLRSRRT-WSMD 383
+Q S +R + R+ SH + RSR T W ++
Sbjct: 288 SQASSSHSRPELNDASDRSAVFSHGS-----RSRSTSWRLE 323
>UniRef100_Q9LTH6 Emb|CAB62332.1 [Arabidopsis thaliana]
Length = 512
Score = 280 bits (716), Expect = 5e-74
Identities = 161/367 (43%), Positives = 216/367 (57%), Gaps = 64/367 (17%)
Query: 2 SSHWCYRCNKFVRVWR------LGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAA 55
+S+WCY C +FV VW +G CP CD GF+E + S+ +A + P +
Sbjct: 121 ASYWCYSCTRFVSVWADQGTTTVGSVACPHCDGGFIEQINDSSSAAT-----ELTIPAST 175
Query: 56 AMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNE-- 113
+ I NN ++ RR R G FNP+I+++GG G EREE E
Sbjct: 176 EVRSI----NNNRRSVIRRRR-----SGRRPSFNPVIVLQGGAG-------EREEGEEGD 219
Query: 114 -------FELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEAN-----RSGNEG 161
FE +Y+DG+GSGLR LP +SE+++GSGFER++EQLS +EA+ RSGN
Sbjct: 220 AARDRRAFEFYYDDGSGSGLRPLPDSVSEILMGSGFERLLEQLSQIEASATGIGRSGNP- 278
Query: 162 HNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECIL 221
PA KSA+E LP +EI++ H+ E++CAVC E FE AREMPCKH++H++CI+
Sbjct: 279 ------PASKSAIESLPRVEISDCHIGSEANCAVCTEIFETETEAREMPCKHLFHDDCIV 332
Query: 222 PWLAIQNSCPVCRHELPCESPQINNEISNSNEDEN-VGLTIWRLPGGGFAVGRFSGDGGG 280
PWL+I+NSCPVCR ELP E N SN+NE++N VG+TIWRLPGGGFAVGRF+
Sbjct: 333 PWLSIRNSCPVCRFELPSEP----NRRSNNNEEDNAVGMTIWRLPGGGFAVGRFNAAMRD 388
Query: 281 GENRMEHPIVYTEVDGA--FNNVGEPRRISW-------SLTSSRGGIGRSRGGAFRRMLS 331
GE + P+V TE+DG N+ PRRISW S+ G G GG RRM+
Sbjct: 389 GERVL--PVVLTEMDGGGIGNSQDGPRRISWVRSHGTLESDSNGTGSGSGSGGRLRRMVR 446
Query: 332 NLFGCLR 338
+ +R
Sbjct: 447 GMVSLMR 453
>UniRef100_Q940T5 AT5g59550/f2o15_210 [Arabidopsis thaliana]
Length = 407
Score = 280 bits (716), Expect = 5e-74
Identities = 161/367 (43%), Positives = 216/367 (57%), Gaps = 64/367 (17%)
Query: 2 SSHWCYRCNKFVRVWR------LGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAA 55
+S+WCY C +FV VW +G CP CD GF+E + S+ +A + P +
Sbjct: 16 ASYWCYSCTRFVSVWADQGTTTVGSVACPHCDGGFIEQINDSSSAAT-----ELTIPAST 70
Query: 56 AMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNE-- 113
+ I NN ++ RR R G FNP+I+++GG G EREE E
Sbjct: 71 EVRSI----NNNRRSVIRRRR-----SGRRPSFNPVIVLQGGAG-------EREEGEEGD 114
Query: 114 -------FELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEAN-----RSGNEG 161
FE +Y+DG+GSGLR LP +SE+++GSGFER++EQLS +EA+ RSGN
Sbjct: 115 AARDRRAFEFYYDDGSGSGLRPLPDSVSEILMGSGFERLLEQLSQIEASATGIGRSGNP- 173
Query: 162 HNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECIL 221
PA KSA+E LP +EI++ H+ E++CAVC E FE AREMPCKH++H++CI+
Sbjct: 174 ------PASKSAIESLPRVEISDCHIGSEANCAVCTEIFETETEAREMPCKHLFHDDCIV 227
Query: 222 PWLAIQNSCPVCRHELPCESPQINNEISNSNEDEN-VGLTIWRLPGGGFAVGRFSGDGGG 280
PWL+I+NSCPVCR ELP E N SN+NE++N VG+TIWRLPGGGFAVGRF+
Sbjct: 228 PWLSIRNSCPVCRFELPSEP----NRRSNNNEEDNAVGMTIWRLPGGGFAVGRFNAAMRD 283
Query: 281 GENRMEHPIVYTEVDGA--FNNVGEPRRISW-------SLTSSRGGIGRSRGGAFRRMLS 331
GE + P+V TE+DG N+ PRRISW S+ G G GG RRM+
Sbjct: 284 GERVL--PVVLTEMDGGGIGNSQDGPRRISWVRSHGTLESDSNGTGSGSGSGGRLRRMVR 341
Query: 332 NLFGCLR 338
+ +R
Sbjct: 342 GMVSLMR 348
>UniRef100_O22283 Expressed protein [Arabidopsis thaliana]
Length = 401
Score = 279 bits (714), Expect = 8e-74
Identities = 171/407 (42%), Positives = 224/407 (55%), Gaps = 41/407 (10%)
Query: 3 SHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQS---------THSANTVGGRRMRFPM 53
S+WCY C++FV W CPDCD GFLE +++ T + T RFP
Sbjct: 5 SYWCYSCSRFV--WVSDSISCPDCDGGFLELIQEPLDFTPSDSFTTTTTTQHRSPTRFPP 62
Query: 54 AAAMYMIG----HRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGS-SEGTSRER 108
++ H +N+ R R N SP NP+I++RG + S E
Sbjct: 63 PSSSSSTPSASMHADNSPTPTIVTRTRSNR------SP-NPVIVLRGSAAAPSSDVVSEG 115
Query: 109 EENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLP 168
+ + F+++Y+DG SGLR LPP M+E +LGSGF+R+++Q+S +E N + N + +H P
Sbjct: 116 LDRSAFQMYYDDGTDSGLRPLPPSMTEFLLGSGFDRLLDQISQIELNTNRNL-RSCEHPP 174
Query: 169 ALKSAVELLPTIEINESHM--NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
A KSA+E LP IEI+ +H+ + +SHCAVCKE F L SAREMPC HIYH +CILPWLAI
Sbjct: 175 ASKSAIEALPLIEIDPTHLLSDSQSHCAVCKENFVLKSSAREMPCNHIYHPDCILPWLAI 234
Query: 227 QNSCPVCRHELPCE--------SPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDG 278
+NSCPVCRHELP E + + ED GLTIWRLPGGGFAVGR G
Sbjct: 235 RNSCPVCRHELPAEDLTDGTGAALTAVTATAEEEEDSAAGLTIWRLPGGGFAVGRIPGGW 294
Query: 279 GGGENRMEHPIVYTEVDGA-FNNVGEPRRISW-SLTSSR-GGIGRSRGGAFRRMLSNLFG 335
GG+ M P+VYTEVDG + PRR++W S R GG R RGG F + LFG
Sbjct: 295 RGGDRMM--PVVYTEVDGGRLGDERLPRRVAWGSRRGGRDGGGSRERGGGFAGRIMRLFG 352
Query: 336 CLRG--GGVRNQHSPFTTREFPQPMTMRNNSASHTNENPSLRSRRTW 380
C G G + + + +T R S S + S RR W
Sbjct: 353 CFSGSSGSIAAAAAASSGSGSRIRVTRRTRSFSMFSTASSSSRRRNW 399
>UniRef100_Q9SNB6 Hypothetical protein F12A12.140 [Arabidopsis thaliana]
Length = 395
Score = 276 bits (707), Expect = 5e-73
Identities = 155/355 (43%), Positives = 211/355 (58%), Gaps = 42/355 (11%)
Query: 2 SSHWCYRCNKFVRVWR-----LGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAA 56
+S+WCY C +F+ VW G+ +CP C+ GF+E++E S++S TV P +
Sbjct: 29 TSYWCYSCTRFISVWEDQDANAGV-LCPYCNGGFIEEIEDSSNS--TVAAIPASTPEVRS 85
Query: 57 MYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENN---- 112
+ +++ RR R N FNP+I++ GGGG G E EE +
Sbjct: 86 V-------EETHRSIIRRRRSNRRTS-----FNPVIVLHGGGGGGAGERVENEEGDGATR 133
Query: 113 ---EFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPA 169
+E +Y+DG+GSGLR LP +SE+++GSGFER++EQLS +EA SGN + PA
Sbjct: 134 ERRAYEFYYDDGSGSGLRPLPDSVSEILMGSGFERLLEQLSQIEA--SGNGIGRSGNPPA 191
Query: 170 LKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNS 229
KSA+E LP +EI++ H E++CAVC E FE GI REMPCKHI+H +CI+PWL+I+NS
Sbjct: 192 SKSAIESLPRVEISDCHTKAEANCAVCTEVFEAGIEGREMPCKHIFHGDCIVPWLSIRNS 251
Query: 230 CPVCRHELPCESPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENRMEHPI 289
CPVCR ELP + Q +NE E+ VG+TIWRLPGGGFAVGRF+ GE + P+
Sbjct: 252 CPVCRFELPSDPIQRSNE-----EEHAVGMTIWRLPGGGFAVGRFNAGVREGERIL--PV 304
Query: 290 VYTEVDGAFNNVGE-PRRISW-----SLTSSRGGIGRSRGGAFRRMLSNLFGCLR 338
V TE+DG E PRRISW + SR G GG RR + + +R
Sbjct: 305 VLTEMDGGGLGSNEGPRRISWVRAHETPEMSRNGGRSGNGGRLRRAVRGMVSFMR 359
>UniRef100_Q84Q81 Hypothetical protein OJ1261C08.10 [Oryza sativa]
Length = 279
Score = 236 bits (601), Expect = 1e-60
Identities = 137/336 (40%), Positives = 178/336 (52%), Gaps = 82/336 (24%)
Query: 3 SHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMIGH 62
S+WCY C++FVRV +CP+CD GFLE Q GRR
Sbjct: 7 SYWCYHCSRFVRV-SPSTVVCPECDGGFLEQFPQPPPRGGGGSGRR-------------- 51
Query: 63 RNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYEDGA 122
NP+I++RGG S FEL+Y+DG+
Sbjct: 52 -----------------------GAMNPVIVLRGGSLSG------------FELYYDDGS 76
Query: 123 GSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEI 182
G GLR LP +S L++GSGF R+++Q S +EA PA K+AVE +P++ +
Sbjct: 77 GDGLRPLPGDVSHLLMGSGFHRLLDQFSRLEAAAP--------RPPASKAAVESMPSVTV 128
Query: 183 NESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESP 242
S +HCAVC+E FE G SAREMPCKH+YH +CILPWL+++NSCPVCR ELP +
Sbjct: 129 AGSG----AHCAVCQEAFEPGASAREMPCKHVYHQDCILPWLSLRNSCPVCRRELPAAAA 184
Query: 243 QINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENRMEHPIVYTEVDGAFNNVG 302
+ + GLTIWRLP GGFAVGRF+G R + P+VYTE+DG F+N
Sbjct: 185 P--------ESEADAGLTIWRLPRGGFAVGRFAGGP-----REQLPVVYTELDGGFSNGV 231
Query: 303 EPRRISWSLTSSR--GGIGRSRGGAFRRMLSNLFGC 336
PRR++W GG GR RR+ NLFGC
Sbjct: 232 GPRRVTWPEGDGHVDGGEGR-----IRRVFRNLFGC 262
>UniRef100_Q94DH1 Putative RING-H2 finger protein RHC2a [Oryza sativa]
Length = 279
Score = 177 bits (450), Expect = 3e-43
Identities = 101/229 (44%), Positives = 132/229 (57%), Gaps = 39/229 (17%)
Query: 84 DISPFNP-IIMIRGGGGSSEGTSREREENNEFELFYEDGAGSGLRALPPRMSELILGSGF 142
D SPFNP +I++R + T+ F+L Y+DGA S LR L F
Sbjct: 50 DHSPFNPPVIVLRRSASPDDATT--------FDLLYDDGASSALRPL------------F 89
Query: 143 ERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFEL 202
+R++ ++ N + PA K+AV+ +PTI I H+ +SHCAVCKEPF L
Sbjct: 90 DRLLLRIPSASDNPNP---------PASKAAVDSMPTILIGACHLAADSHCAVCKEPFHL 140
Query: 203 GISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEIS------NSNED-E 255
AREMPC HIYH+ CILPWLA+ NSCPVCRH +P + N + +S+ED
Sbjct: 141 AAEAREMPCAHIYHHNCILPWLALHNSCPVCRHRMPTDDHDSTNAAAAQAAAGSSDEDAT 200
Query: 256 NVG-LTIWRLPGGGFAVGRFSGDGGGGENRMEHPIVYTEV-DGAFNNVG 302
VG LTIWRLPGGG+AVGRF+ GG E P++YT++ DG FN G
Sbjct: 201 TVGTLTIWRLPGGGYAVGRFAAAGGTRAGERELPVLYTQMDDGGFNGGG 249
>UniRef100_Q5VQY5 Putative ring finger protein 126 isoform 1 [Oryza sativa]
Length = 329
Score = 141 bits (356), Expect = 3e-32
Identities = 89/263 (33%), Positives = 130/263 (48%), Gaps = 30/263 (11%)
Query: 2 SSHWCYRCNKFVRVWRLGMPI-CPDCDSGFLEDVEQSTHSANTVG--------GRRMRFP 52
++HWCY C + +RV G I CP+C+ GF++++ + S NT G R F
Sbjct: 5 ATHWCYACRRPIRV--SGQDITCPNCNDGFIQEISEIGGSLNTYGIFDPSFDERRDRSFG 62
Query: 53 MAAAMY-MIGHRNNNYNQNT---FRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSRER 108
M AM ++ R +N F R + + G P+++ S R
Sbjct: 63 MVEAMSDLMRQRMAEMGRNRVLDFHGTRGASSHQGRQPTVRPMLIF-----GSNAPDRVS 117
Query: 109 EENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLP 168
+ E ++ G G A P S ++G E + EQL + NR G P
Sbjct: 118 SSSEEADILLRQGRRIG--ADRPNFSRFLVGPSLEALFEQLL-LHNNRQGPP-------P 167
Query: 169 ALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQN 228
A +SA++ +P ++IN H+ + HC VC + FE+G AREMPCKH+YH ECI+PWL N
Sbjct: 168 APQSAIDSMPVVKINLRHLRDDPHCPVCTDKFEVGTEAREMPCKHLYHAECIIPWLVQHN 227
Query: 229 SCPVCRHELPCESPQINNEISNS 251
SCPVCRH LP S + + S+S
Sbjct: 228 SCPVCRHPLPSSSHRSGSTRSSS 250
>UniRef100_Q94AK4 Hypothetical protein At3g56580 [Arabidopsis thaliana]
Length = 320
Score = 129 bits (323), Expect = 2e-28
Identities = 83/258 (32%), Positives = 129/258 (49%), Gaps = 18/258 (6%)
Query: 2 SSHWCYRCNKFVRVW-RLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMI 60
++HWC+RC + VW R +C C GF+E+++ A+ R F + A
Sbjct: 6 NTHWCHRCQR--AVWLRARDAVCSYCGGGFVEEIDIGPSRAHRDVERDPTFDLMEAFSAF 63
Query: 61 GHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYED 120
+ + ++ R + F+ + + GG + +N+ E F +
Sbjct: 64 --MRSRLAERSYDREISGRLGSAGSESFSNLAPLLIFGGQAP-FRLAGGDNSSVEAFV-N 119
Query: 121 GAGSGLR-ALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPT 179
GA G+ A + G G E ++EQLS SG H++ PA KS+++ LPT
Sbjct: 120 GAAPGIGIARGTNAGDYFFGPGLEELIEQLS------SGT--HHRGPPPAPKSSIDALPT 171
Query: 180 IEINESHM-NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP 238
I+I + H+ + +SHC VCK+ FEL A++MPC HIYH++CI+PWL NSCPVCR ELP
Sbjct: 172 IKITQKHLKSSDSHCPVCKDEFELKSEAKQMPCHHIYHSDCIVPWLVQHNSCPVCRKELP 231
Query: 239 CESPQINNEISNSNEDEN 256
+ + S+ N N
Sbjct: 232 SRGSSSSTQ-SSQNRSTN 248
>UniRef100_O22197 Expressed protein [Arabidopsis thaliana]
Length = 328
Score = 128 bits (322), Expect = 2e-28
Identities = 92/307 (29%), Positives = 137/307 (43%), Gaps = 45/307 (14%)
Query: 2 SSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQST-------HSANTVGGRRMRFPMA 54
++HWC+RC + VR+ P+C C GF+E+++ + S V R F +
Sbjct: 6 NTHWCHRCQRAVRLHGQE-PVCFYCGGGFVEELDMAQASPFDMFRSHRGVVERDQTFDLM 64
Query: 55 AAMYMIGHRNNNYNQNTFRRHRRNNVNGGDIS-PFNPIIMIRGGGGSSEGTSREREENNE 113
A + + RN ++ R R ++ G + P ++I GG T +N
Sbjct: 65 DA-FSVFMRNRLAERSHDREIRGRTISSGPENFPGLAPLLIFGGQVPYRLTG-----DNA 118
Query: 114 FELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSA 173
E + +G G+ + G G E + EQLS R PA +SA
Sbjct: 119 VEALF-NGGSPGIGITRGNTGDYFFGPGLEELFEQLSAGTTRRGPP--------PAPRSA 169
Query: 174 VELLPTIEINESHM-NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPV 232
++ LPTI+I + H+ + +S+C VCK+ FELG A++MPC HIYH++CI+PWL NSCPV
Sbjct: 170 IDALPTIKIAQRHLRSSDSNCPVCKDEFELGSEAKQMPCNHIYHSDCIVPWLVQHNSCPV 229
Query: 233 CRHELP--------------------CESPQINNEISNSNEDENVGLTIWRLPGGGFAVG 272
CR ELP S +N N NE N + W G +
Sbjct: 230 CRQELPSASGPSSSQNRTTPTRNYRSSSSSSSSNSRENGNERRNPFSSFWPFRSSGSSSS 289
Query: 273 RFSGDGG 279
GG
Sbjct: 290 STQNRGG 296
>UniRef100_Q9LT14 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MPN9
[Arabidopsis thaliana]
Length = 386
Score = 124 bits (312), Expect = 3e-27
Identities = 76/266 (28%), Positives = 118/266 (43%), Gaps = 32/266 (12%)
Query: 5 WCYRCNKFVRVWRLGM--PICPDCDSGFLEDVEQSTHSAN---TVGGRRMRFPMA----- 54
+CY+CN+ V + P CP C+ GFLE+ E + + FPMA
Sbjct: 80 FCYQCNQTVTISISSSADPFCPICNQGFLEEYEDPNPNQSLNFNPNSSDSFFPMADPFST 139
Query: 55 --------AAMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSR 106
+A G + + + R+ F+P ++
Sbjct: 140 LLPLIFGSSAAAPSGMDFMSLFGPSMQPQARSTQQNPQSDAFDPFTFLQNH------LQT 193
Query: 107 EREENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQH 166
R FE E+ +P + G G E++++QL+ + NR G
Sbjct: 194 LRSSGTHFEFVIENHPSDPGNRMPGNFGDYFFGPGLEQLIQQLAENDPNRYGTP------ 247
Query: 167 LPALKSAVELLPTIEINESHMNVE-SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA 225
PA KSA++ LPT+++ + + E + CAVC + FE G ++MPCKH++H +C+LPWL
Sbjct: 248 -PASKSAIDALPTVKVTKDMLKSEMNQCAVCMDEFEDGSDVKQMPCKHVFHQDCLLPWLE 306
Query: 226 IQNSCPVCRHELPCESPQINNEISNS 251
+ NSCPVCR ELP + P N S
Sbjct: 307 LHNSCPVCRFELPTDDPDYENRSQGS 332
>UniRef100_Q8LPN7 AT3g19950/MPN9_19 [Arabidopsis thaliana]
Length = 328
Score = 124 bits (312), Expect = 3e-27
Identities = 76/266 (28%), Positives = 118/266 (43%), Gaps = 32/266 (12%)
Query: 5 WCYRCNKFVRVWRLGM--PICPDCDSGFLEDVEQSTHSAN---TVGGRRMRFPMA----- 54
+CY+CN+ V + P CP C+ GFLE+ E + + FPMA
Sbjct: 22 FCYQCNQTVTISISSSADPFCPICNQGFLEEYEDPNPNQSLNFNPNSSDSFFPMADPFST 81
Query: 55 --------AAMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSR 106
+A G + + + R+ F+P ++
Sbjct: 82 LLPLIFGSSAAAPSGMDFMSLFGPSMQPQARSTQQNPQSDAFDPFTFLQNH------LQT 135
Query: 107 EREENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQH 166
R FE E+ +P + G G E++++QL+ + NR G
Sbjct: 136 LRSSGTHFEFVIENHPSDPGNRMPGNFGDYFFGPGLEQLIQQLAENDPNRYGTP------ 189
Query: 167 LPALKSAVELLPTIEINESHMNVE-SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA 225
PA KSA++ LPT+++ + + E + CAVC + FE G ++MPCKH++H +C+LPWL
Sbjct: 190 -PASKSAIDALPTVKVTKDMLKSEMNQCAVCMDEFEDGSDVKQMPCKHVFHQDCLLPWLE 248
Query: 226 IQNSCPVCRHELPCESPQINNEISNS 251
+ NSCPVCR ELP + P N S
Sbjct: 249 LHNSCPVCRFELPTDDPDYENRSQGS 274
>UniRef100_Q7XD81 Putative zinc finger protein [Oryza sativa]
Length = 370
Score = 110 bits (276), Expect = 5e-23
Identities = 90/341 (26%), Positives = 139/341 (40%), Gaps = 70/341 (20%)
Query: 4 HWCYRCNKFVRV----WRLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYM 59
++CY+CN+ V + G CP+C F+E+V N + FP A M
Sbjct: 16 YFCYQCNRTVLLPASAAAAGALSCPECRGDFIEEV-------NVPAPAIIPFPFAFPPMM 68
Query: 60 IGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRG---------------GGGSSEGT 104
+ + +++ + SP + + G S+ GT
Sbjct: 69 PTATSASAAAAAAASPTQSSSSSAATSPSSDLSAFLNSMLGPLNLRTDERMPGTTSAAGT 128
Query: 105 SREREENNEFEL--FYE-----------------DGAGSGL-----RALPPRMSELILGS 140
+ +E + F+ F++ D A GL R + +G
Sbjct: 129 ATPEDEPDGFDAVTFFQNYLQNLMDGGANIQVLLDDASVGLAPGIGRVGGASFGDYFVGP 188
Query: 141 GFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVE--SHCAVCKE 198
G E+++EQL+ + NR G PA KSA+ LP + + ++ + + CAVCKE
Sbjct: 189 GLEQLIEQLTENDPNRYGTP-------PAAKSALSTLPDVVVTDAMVAAADGAECAVCKE 241
Query: 199 PFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEISNSNEDENVG 258
F G A++MPCKHIYH +CI+PWL + NSCP+CR ELP + P ++ +
Sbjct: 242 DFSPGEGAKQMPCKHIYHADCIMPWLDLHNSCPICRFELPTDDPDYEGRKKSNPQ----- 296
Query: 259 LTIWRLPGGGFAVGRFSGDGGGGENRMEHPIVYTEVDGAFN 299
P G G SG E R E V+ FN
Sbjct: 297 ------PTAGVDAGAASGSSTAAEEREESGESARLVERRFN 331
>UniRef100_Q6AVN2 Hypothetical protein OJ1119_H02.11 [Oryza sativa]
Length = 323
Score = 107 bits (266), Expect = 7e-22
Identities = 78/288 (27%), Positives = 123/288 (42%), Gaps = 63/288 (21%)
Query: 1 MSSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMI 60
++ +WC+ C + + + CP C GF+E++ T R P + I
Sbjct: 6 ITRYWCHECEQAIEEAMVDEIKCPSCGGGFVEEMTDEEIERLT---NRQPEPGFSQWNPI 62
Query: 61 GHRNNNYNQNT------------FRRHRRNNV-------------------NGGDISPFN 89
H + + RRHRR + + I+ FN
Sbjct: 63 EHPGETMDSDDEDNDLGREFEGFIRRHRRASTLRRVLDSIHDDLADDQERDSSILINAFN 122
Query: 90 PIIMIRGGG-GSSEGTSREREENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQ 148
+ ++G EG + N+ DG + E +LG+G +++
Sbjct: 123 QALALQGSVLDPDEGQGDQGGSTND------DGL----------LEEYVLGAGLSLLLQH 166
Query: 149 LSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISARE 208
L+ + +R+G PA K AVE LPT++I E C+VC + E+G A++
Sbjct: 167 LAESDPSRNGTP-------PAKKEAVEALPTVKIEEV-----VSCSVCLDDLEVGSQAKQ 214
Query: 209 MPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEISNSNEDEN 256
MPC+H +H+ CILPWL + +SCPVCR ELP E + NE SN E+
Sbjct: 215 MPCEHKFHSSCILPWLELHSSCPVCRFELPSEETKDLNEPSNIGRVED 262
>UniRef100_Q5VME8 Putative ring finger protein 126 isoform 1 [Oryza sativa]
Length = 338
Score = 105 bits (263), Expect = 2e-21
Identities = 47/112 (41%), Positives = 72/112 (63%), Gaps = 9/112 (8%)
Query: 133 MSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVE-- 190
+ + +GSG E++++QL+ + NR G PA KSAV LP + ++ M +
Sbjct: 147 LGDYFVGSGLEQLIQQLAENDPNRYGTP-------PAAKSAVAALPDVAVSADMMAADGG 199
Query: 191 SHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESP 242
+ CAVC + F LG +A+++PCKH++H +CILPWL + +SCPVCR ELP + P
Sbjct: 200 AQCAVCMDDFHLGAAAKQLPCKHVFHKDCILPWLDLHSSCPVCRFELPTDDP 251
>UniRef100_Q6Z303 Zinc finger-like [Oryza sativa]
Length = 340
Score = 105 bits (262), Expect = 2e-21
Identities = 52/117 (44%), Positives = 74/117 (62%), Gaps = 16/117 (13%)
Query: 168 PALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
PA SA++ LPT++I +H++ S C VCKE FELG +AR+MPCKH+YH++CI+PWL +
Sbjct: 174 PAPSSAIDSLPTVQITGAHLSDGSQCPVCKEDFELGEAARQMPCKHVYHSDCIVPWLRLH 233
Query: 228 NSCPVCRHELPCESPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENR 284
NSCPVCR++L +++ + SN + R G G G GGGG+ R
Sbjct: 234 NSCPVCRYQL------LSSAAAGSNANS-------RARRGSANNG---GGGGGGDGR 274
>UniRef100_Q69TX4 Zinc finger-like [Oryza sativa]
Length = 331
Score = 103 bits (257), Expect = 8e-21
Identities = 79/252 (31%), Positives = 114/252 (44%), Gaps = 55/252 (21%)
Query: 4 HWCYRCNKFVRVW---RLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMI 60
+WCY C + +RV CP C FL +++ + P A
Sbjct: 20 YWCYVCRRALRVVVPSATSDVYCPRCFGRFLHEIDLPVPRVS---------PPA------ 64
Query: 61 GHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEG----TSREREENNEFEL 116
+ + Q F + P ++ GGGG G T+R R +
Sbjct: 65 ---EDQFFQPPFLPYD---------GPRRWVLYTGGGGGGDYGGADVTARRRRLPSPPPA 112
Query: 117 ---FYEDGAGSG-----LRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLP 168
+DGAG G A+ P E G +++ L+ + +R G P
Sbjct: 113 PGTRRQDGAGDGDPPPPAPAIDP--GEYFAGPDLNALIDALT--QDDRPGPP-------P 161
Query: 169 ALKSAVELLPTIEINESHMNVE--SHCAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
A +SA+E LPT+ I+ H+ + S C VCKE FELG +ARE+PCKH YH++CI+PWL +
Sbjct: 162 APESAIESLPTVHISPDHLPADGGSECPVCKEEFELGEAARELPCKHAYHSDCIVPWLRL 221
Query: 227 QNSCPVCRHELP 238
NSCPVCR E+P
Sbjct: 222 HNSCPVCRQEVP 233
>UniRef100_Q7XZF6 Hypothetical protein OSJNBb0033J23.18 [Oryza sativa]
Length = 369
Score = 100 bits (249), Expect = 7e-20
Identities = 47/108 (43%), Positives = 67/108 (61%), Gaps = 12/108 (11%)
Query: 133 MSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESH 192
+ + LG G + +++ L+ + NRSG PA K AVE LPT+ I E
Sbjct: 207 LGDYFLGPGLDILLQHLAESDLNRSGTP-------PAKKEAVEALPTVNIQEV-----LG 254
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCE 240
C+VC E FE+G A+EMPC+H +H++CILPWL + +SCP+CR +LP E
Sbjct: 255 CSVCLEDFEMGTEAKEMPCQHKFHSQCILPWLELHSSCPICRFQLPTE 302
Score = 33.9 bits (76), Expect = 8.1
Identities = 13/40 (32%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Query: 4 HWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQSTHSANT 43
+WC+ C + + M CP CDSGF+E++ + +T
Sbjct: 9 YWCHHCEEVIEPVEPDMK-CPSCDSGFVEEMGSAGFEPST 47
>UniRef100_Q9SG87 Putative RING zinc finger protein [Arabidopsis thaliana]
Length = 684
Score = 97.8 bits (242), Expect = 5e-19
Identities = 39/71 (54%), Positives = 54/71 (75%)
Query: 168 PALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
PA +A+ L I+I + H+ ++ +C VC++ FE+G AR+MPCKHIYH+ECILPWL +
Sbjct: 96 PASLAAINSLQKIKIRQKHLGLDPYCPVCQDQFEIGSDARKMPCKHIYHSECILPWLVQR 155
Query: 228 NSCPVCRHELP 238
N+CPVCR ELP
Sbjct: 156 NTCPVCRKELP 166
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.136 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 744,985,601
Number of Sequences: 2790947
Number of extensions: 35082113
Number of successful extensions: 114679
Number of sequences better than 10.0: 1817
Number of HSP's better than 10.0 without gapping: 1242
Number of HSP's successfully gapped in prelim test: 576
Number of HSP's that attempted gapping in prelim test: 112725
Number of HSP's gapped (non-prelim): 2170
length of query: 391
length of database: 848,049,833
effective HSP length: 129
effective length of query: 262
effective length of database: 488,017,670
effective search space: 127860629540
effective search space used: 127860629540
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC124953.18


