

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
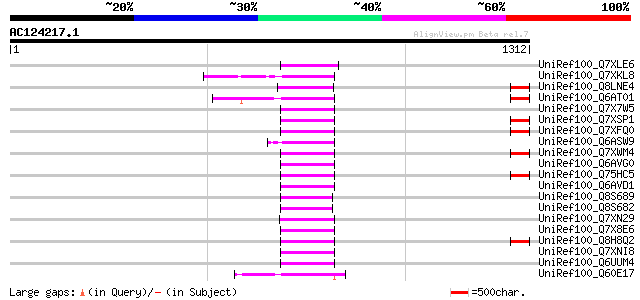
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XLE6 OSJNBa0013A04.23 protein [Oryza sativa] 104 2e-20
UniRef100_Q7XKL8 OSJNBa0054D14.8 protein [Oryza sativa] 102 6e-20
UniRef100_Q8LNE4 Putative retroelement [Oryza sativa] 102 1e-19
UniRef100_Q6AT01 Putative polyprotein [Oryza sativa] 102 1e-19
UniRef100_Q7X7W5 OSJNBb0046P18.13 protein [Oryza sativa] 101 1e-19
UniRef100_Q7XSP1 OSJNBa0070M12.15 protein [Oryza sativa] 101 2e-19
UniRef100_Q7XFQ0 Putative retroelement [Oryza sativa] 100 2e-19
UniRef100_Q6ASW9 Reverse transcriptase (RNA-dependent DNA polyme... 100 2e-19
UniRef100_Q7XWM4 OSJNBb0052B05.5 protein [Oryza sativa] 100 3e-19
UniRef100_Q6AVG0 Putative retrotransposon gag protein [Oryza sat... 100 3e-19
UniRef100_Q75HC5 Putative reverse transcriptase [Oryza sativa] 100 4e-19
UniRef100_Q6AVD1 Putative polyprotein [Oryza sativa] 100 4e-19
UniRef100_Q8S689 Putative retrotransposon polyprotein [Oryza sat... 100 4e-19
UniRef100_Q8S682 Putative retrotransposon polyprotein [Oryza sat... 100 4e-19
UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa] 100 5e-19
UniRef100_Q7X8E6 OSJNBb0012A12.6 protein [Oryza sativa] 100 5e-19
UniRef100_Q8H8Q2 Putative gag-pol polyprotein [Oryza sativa] 100 5e-19
UniRef100_Q7XNI8 OSJNBb0032D24.5 protein [Oryza sativa] 100 5e-19
UniRef100_Q6UUM4 Hypothetical protein [Oryza sativa] 99 7e-19
UniRef100_Q60E17 Putative polyprotein [Oryza sativa] 99 7e-19
>UniRef100_Q7XLE6 OSJNBa0013A04.23 protein [Oryza sativa]
Length = 1235
Score = 104 bits (259), Expect = 2e-20
Identities = 53/146 (36%), Positives = 87/146 (59%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL++K VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 188 ERAIFEKPEVVENRHLKPLYVKGFVNGKPMSKMMVDGGAAVNLMPYATFRKLGKNADDLI 247
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
NV L+ + GV+ V+L +G+ T F V+ K +Y++LLG +WIH +P
Sbjct: 248 KTNVVLNDFGGNPSENKGVLNVELTIGSKTDPTTFFVIDGKGSYSMLLGRDWIHANCCIP 307
Query: 806 STMH*RLALWREDGVLEILKLSKVTL 831
STMH L W+ D + +L S++ +
Sbjct: 308 STMHQSLIQWQGDKIEIVLADSQLKM 333
Score = 48.1 bits (113), Expect = 0.002
Identities = 21/47 (44%), Positives = 34/47 (71%)
Query: 1266 GFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
GF+++K+GIE++Q AI+ + KK+L + +GK+NF+ RFISN
Sbjct: 514 GFLVHKRGIEVSQRSINAINKIQRLEDKKQLLSLIGKVNFVRRFISN 560
>UniRef100_Q7XKL8 OSJNBa0054D14.8 protein [Oryza sativa]
Length = 1702
Score = 102 bits (255), Expect = 6e-20
Identities = 94/337 (27%), Positives = 148/337 (43%), Gaps = 34/337 (10%)
Query: 491 PPTRHQNRGQVRTFTPSSKSPTEKWVFSSGKKSSYTSPPAKWV----KRFTGTDNQNKAP 546
P R + V PSS+S V +G P + +RF ++ A
Sbjct: 586 PFGRSDHGSHVGLAGPSSRSDRRPHVGLTGASGRSDRPSTSGLTRLQRRFDRRFAKSSAG 645
Query: 547 VSKKFAYNNNYKGKNPMTKT*WRRFQRQKKVDGLKYVTNNDKGKNKQMETFKMIRKPATE 606
S N P T+ RR QR + + + K ++ F +R P T+
Sbjct: 646 ASSSLGRLNRGHYLPPGTEPKPRRLQRLRAQE-----LREREAKEQRDRRFNELRPPPTK 700
Query: 607 RIFPPLSAVKENLHVEDEEMTSNFTDFEA---DFDVICVVLILPIE*DVTSEVTEDEEDF 663
++E++ VE++E S D + D D I VV + P+E DE +
Sbjct: 701 MWRRKAVDIEESIVVEEKEEQSVAKDESSPKEDMD-INVVCMFPME-----FCAMDEVEV 754
Query: 664 IKEMVVHKTMCYYVMNNGCVEGQHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGG 723
+ ++ K A+FE+PD + H+KPL+ K +++ ++++ VDGG
Sbjct: 755 AQFLIGPKV---------------AVFEKPDESNR-HMKPLYRKGRIDGKPVSRMQVDGG 798
Query: 724 VVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVV 783
VVVNL+P S KK+G + LK N+ L+ + + G+ V+L G T F +V
Sbjct: 799 VVVNLMPYSLFKKLGCGDDKLKKTNMILNGFNGEPTEAKGIFSVELTAGNKTLPTGFFIV 858
Query: 784 ASKANYNLLLGTEWIHDVGDVPSTMH*RLALWREDGV 820
+ NYN++LG WIH VPST+H L W D V
Sbjct: 859 DVQGNYNVILGHCWIHANCCVPSTLHQCLIQWDGDDV 895
Score = 42.0 bits (97), Expect = 0.12
Identities = 22/48 (45%), Positives = 33/48 (67%), Gaps = 1/48 (2%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF++++ G+EI+ K + I K + T KE+Q LGK+N+L RFI N
Sbjct: 1241 LGFMVHETGVEIDPKKIEKIRDFK-TPTCKEVQKLLGKVNYLRRFIYN 1287
>UniRef100_Q8LNE4 Putative retroelement [Oryza sativa]
Length = 1170
Score = 102 bits (253), Expect = 1e-19
Identities = 54/141 (38%), Positives = 79/141 (55%)
Query: 678 MNNGCVEGQHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKI 737
MN C + A+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+
Sbjct: 424 MNQDCGQEDEAIFEKPEGTENRHLKPLYINGYVNGKTMSKMMVDGGAAVNLMPYATFRKL 483
Query: 738 GLSESNLKPHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEW 797
G + L N+ L + GV+ V+L VG+ T F V+ +Y+LLLG +W
Sbjct: 484 GRNTEALIKTNMVLKDFGGNPSETRGVLNVELTVGSKTIPTTFFVINGNGSYSLLLGRDW 543
Query: 798 IHDVGDVPSTMH*RLALWRED 818
IH +PSTMH L W+ D
Sbjct: 544 IHANCCIPSTMHQCLIQWQGD 564
Score = 53.1 bits (126), Expect = 5e-05
Identities = 24/48 (50%), Positives = 33/48 (68%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++++GIE+ Q AI +P K ELQ +GKINF+ RFISN
Sbjct: 869 LGFLVHERGIEVTQRSVNAIKKIQPPENKTELQEMIGKINFVRRFISN 916
>UniRef100_Q6AT01 Putative polyprotein [Oryza sativa]
Length = 1739
Score = 102 bits (253), Expect = 1e-19
Identities = 87/321 (27%), Positives = 139/321 (43%), Gaps = 29/321 (9%)
Query: 514 KWVFSSGKKSSYTSPPAKWVKRFTGT--DNQNKAPVSKKFAYNNNYKGKNPMTKT*WRRF 571
+WV SSGK Y K K F + Q + P K+ + ++ R +
Sbjct: 506 QWVKSSGKVEKYDFDITKADKIFDLLLREKQIQLPAGHTIPSTEELGKKSDIVESSSRSY 565
Query: 572 QRQKKVDGLKYVT-------NNDKGKNKQMETFKMIRKPATERIFPPLSAVKENLHVEDE 624
R ++ + N D G+ +E + I + VK + V D+
Sbjct: 566 NRGNRLRQTRVSVHQRLGPVNQDHGEEDSVEVEQEINHRLRKAKPRQEWRVKNQVPVADD 625
Query: 625 EMTSNFTDFEADFDVIC----VVLILPIE*DVT-SEVTEDEEDFIKEMVVHKTMCYYVMN 679
T V+ +V LP+E + ++V E EE+ K ++
Sbjct: 626 VATDEAKRLAKGKSVVTAPINMVFTLPVEFGIDQADVDEVEEESAKLVL----------- 674
Query: 680 NGCVEGQHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGL 739
+ A+F++P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G
Sbjct: 675 ----SPEQAVFQKPEGTENRHLKPLYINGYVNGKLMSKMMVDGGAAVNLMPYATFRKLGR 730
Query: 740 SESNLKPHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIH 799
+ +L N+ L + GV+ V L VG+ T FLV+ K +Y+L LG +WIH
Sbjct: 731 NAEDLIKTNMVLKDFGGNPSETKGVLNVGLTVGSKTIPTTFLVIDGKGSYSLFLGRDWIH 790
Query: 800 DVGDVPSTMH*RLALWREDGV 820
+PSTMH L W+ D V
Sbjct: 791 ANCCIPSTMHQCLIQWQGDKV 811
Score = 50.4 bits (119), Expect = 3e-04
Identities = 23/48 (47%), Positives = 32/48 (65%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++ +GIE+ Q AI +P K ELQ +GKI+F+ RFISN
Sbjct: 1129 LGFLVHDRGIEVTQRSVNAIKKIQPPENKTELQEMIGKIHFVRRFISN 1176
>UniRef100_Q7X7W5 OSJNBb0046P18.13 protein [Oryza sativa]
Length = 766
Score = 101 bits (252), Expect = 1e-19
Identities = 53/135 (39%), Positives = 81/135 (59%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+F++P+ HLKPL+I VN ++K++VDGG VVNL+P + +K+G + +L
Sbjct: 340 EQAVFDKPEETENWHLKPLYINGYVNGKPMSKMMVDGGAVVNLMPYATFRKLGRNAEDLI 399
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+AL + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 400 KMNMALKDFGGNPSETKGVLNVELSVGSKTVPTTFFVIDGKGSYSLLLGRDWIHANCCIP 459
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 460 STMHQCLIQWQGDKI 474
>UniRef100_Q7XSP1 OSJNBa0070M12.15 protein [Oryza sativa]
Length = 1685
Score = 101 bits (251), Expect = 2e-19
Identities = 53/135 (39%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 722 ERAVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYTTFRKLGRTTEDLM 781
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 782 KTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 841
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D V
Sbjct: 842 STMHQCLIQWQGDKV 856
Score = 53.9 bits (128), Expect = 3e-05
Identities = 26/48 (54%), Positives = 33/48 (68%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++++GIEI Q AI KP K ELQ +GKINF+ RFISN
Sbjct: 1174 LGFLVHERGIEIPQRSINAIKKIKPPGNKTELQEMIGKINFVRRFISN 1221
>UniRef100_Q7XFQ0 Putative retroelement [Oryza sativa]
Length = 1025
Score = 100 bits (250), Expect = 2e-19
Identities = 53/135 (39%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN L+K++VDGG VNL+P + +K+G + +L
Sbjct: 103 EQAVFEKPEGTENRHLKPLYINGYVNGKPLSKMMVDGGAAVNLMPYATFRKLGRNAEDLI 162
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 163 KTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 222
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 223 STMHQCLIQWQGDKI 237
Score = 53.1 bits (126), Expect = 5e-05
Identities = 24/48 (50%), Positives = 33/48 (68%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++++GIE+ Q AI +P K ELQ +GKINF+ RFISN
Sbjct: 481 LGFLVHERGIEVTQRSVNAIKKIQPPENKTELQEMIGKINFVKRFISN 528
>UniRef100_Q6ASW9 Reverse transcriptase (RNA-dependent DNA polymerase) domain
containing protein [Oryza sativa]
Length = 1372
Score = 100 bits (250), Expect = 2e-19
Identities = 61/169 (36%), Positives = 90/169 (53%), Gaps = 10/169 (5%)
Query: 652 VTSEVTEDEEDFIKEMVVHKTMCYYVMNNGCVEGQHAMFERPDYHMKIHLKPLFIKAKVN 711
VT E T DE K + K++ +N A+FE+P+ HLKPL+I VN
Sbjct: 395 VTGEATADEA---KRLAKGKSVVTASVNK-------AIFEKPEGTENRHLKPLYINGYVN 444
Query: 712 QIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVEVDLVV 771
++K++VDGG VNL+P + +K+G + +L N+ L + GV+ V+L V
Sbjct: 445 GKPMSKMMVDGGAAVNLMPYATFRKLGRNVEDLIKTNMVLKDFGGNPSETKGVLNVELTV 504
Query: 772 GTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH*RLALWREDGV 820
G T F V+ K +Y+LLLG +WIH +PSTMH L W+ D +
Sbjct: 505 GNKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIPSTMHQCLIQWQGDKI 553
Score = 42.7 bits (99), Expect = 0.073
Identities = 19/44 (43%), Positives = 28/44 (63%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIR 1308
LGF+++++GIE+ Q I +P K ELQ +GKINF+ R
Sbjct: 803 LGFLVHERGIEVTQMSINVIKKIQPPENKTELQEMIGKINFVRR 846
>UniRef100_Q7XWM4 OSJNBb0052B05.5 protein [Oryza sativa]
Length = 1823
Score = 100 bits (249), Expect = 3e-19
Identities = 51/135 (37%), Positives = 80/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN+ ++K++VDGG VNL+P + +K+G + +L
Sbjct: 769 EQAIFEKPEGIENQHLKPLYINGYVNRKPMSKMMVDGGAAVNLMPYATFRKLGKNAEDLI 828
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F ++ K +Y+LLLG +WIH +P
Sbjct: 829 KKNMVLKDFGGNPSETKGVLNVELTVGSKTVPTTFFIIDGKGSYSLLLGRDWIHANCCIP 888
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 889 STMHQCLFQWQGDKI 903
Score = 49.3 bits (116), Expect = 8e-04
Identities = 23/48 (47%), Positives = 32/48 (65%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
+GF+++++GIEI Q AI +P K ELQ + KINF+ RFISN
Sbjct: 1158 VGFLVHERGIEITQRSINAIKMIQPPEDKTELQEMISKINFVRRFISN 1205
>UniRef100_Q6AVG0 Putative retrotransposon gag protein [Oryza sativa]
Length = 1251
Score = 100 bits (249), Expect = 3e-19
Identities = 53/135 (39%), Positives = 78/135 (57%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A FE+P+ HLKPL+I VN ++K++VDGG VNL+P + KK+G + +L
Sbjct: 683 EQAEFEKPEGTGNRHLKPLYINGYVNGKSMSKMMVDGGAAVNLMPYATFKKLGRNAEDLI 742
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 743 KTNMVLKDFGGNPSETRGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 802
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 803 STMHQCLIQWQGDKI 817
>UniRef100_Q75HC5 Putative reverse transcriptase [Oryza sativa]
Length = 1149
Score = 100 bits (248), Expect = 4e-19
Identities = 52/135 (38%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 645 EQAVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNAEDLI 704
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 705 KTNMVLEDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 764
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 765 STMHQCLIQWQGDKI 779
Score = 53.1 bits (126), Expect = 5e-05
Identities = 24/48 (50%), Positives = 33/48 (68%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++++GIE+ Q AI +P K ELQ +GKINF+ RFISN
Sbjct: 1097 LGFLVHERGIEVTQRSVNAIKKIQPPENKTELQEMIGKINFVRRFISN 1144
>UniRef100_Q6AVD1 Putative polyprotein [Oryza sativa]
Length = 1418
Score = 100 bits (248), Expect = 4e-19
Identities = 52/135 (38%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 839 EQAVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNAEDLI 898
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 899 KTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 958
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 959 STMHQCLIQWQGDKI 973
>UniRef100_Q8S689 Putative retrotransposon polyprotein [Oryza sativa]
Length = 1094
Score = 100 bits (248), Expect = 4e-19
Identities = 53/131 (40%), Positives = 76/131 (57%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN + K++VDGG VNL+P + KK+G + NL
Sbjct: 450 EQAVFEKPEGTENRHLKPLYINGYVNGKPMPKMMVDGGAAVNLMPYATFKKLGRNAENLI 509
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 510 KTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 569
Query: 806 STMH*RLALWR 816
STMH L W+
Sbjct: 570 STMHQCLIQWQ 580
>UniRef100_Q8S682 Putative retrotransposon polyprotein [Oryza sativa]
Length = 1094
Score = 100 bits (248), Expect = 4e-19
Identities = 53/131 (40%), Positives = 76/131 (57%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN + K++VDGG VNL+P + KK+G + NL
Sbjct: 450 EQAVFEKPEGTENRHLKPLYINGYVNGKPMPKMMVDGGAAVNLMPYATFKKLGRNAENLI 509
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 510 KTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIP 569
Query: 806 STMH*RLALWR 816
STMH L W+
Sbjct: 570 STMHQCLIQWQ 580
>UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa]
Length = 1563
Score = 99.8 bits (247), Expect = 5e-19
Identities = 50/138 (36%), Positives = 79/138 (57%)
Query: 683 VEGQHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSES 742
+ + A+FE+P+ HLKPL++ +N ++K++VDGG VNL+P + +K+G +
Sbjct: 874 LSSEQAVFEKPEGTENRHLKPLYVNGYINGKPMSKMMVDGGAAVNLMPYATFRKLGRNAE 933
Query: 743 NLKPHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVG 802
+L N+ L + GV+ V+L VG T F V+ K +Y+LLLG +WIH
Sbjct: 934 DLIKTNMVLKDFGSNPSETKGVLNVELTVGNKTIPTTFFVIDGKGSYSLLLGRDWIHANC 993
Query: 803 DVPSTMH*RLALWREDGV 820
+PSTMH L W+ D +
Sbjct: 994 CIPSTMHQCLIQWQGDKI 1011
>UniRef100_Q7X8E6 OSJNBb0012A12.6 protein [Oryza sativa]
Length = 243
Score = 99.8 bits (247), Expect = 5e-19
Identities = 52/135 (38%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 31 EQAVFEKPEGTENWHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNSYDLI 90
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 91 KTNMVLKDFGGNPSETKGVLNVELTVGSKTVPTTFFVIDGKGSYSLLLGRDWIHANCCIP 150
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 151 STMHQCLIQWQGDKI 165
>UniRef100_Q8H8Q2 Putative gag-pol polyprotein [Oryza sativa]
Length = 1152
Score = 99.8 bits (247), Expect = 5e-19
Identities = 53/135 (39%), Positives = 79/135 (58%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ AMFE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 724 EQAMFEKPEGTEDRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNAEDLI 783
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 784 KTNMVLKDFGGNPLETKGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANYCIP 843
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 844 STMHHCLIQWQGDKI 858
Score = 51.2 bits (121), Expect = 2e-04
Identities = 23/48 (47%), Positives = 32/48 (65%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
LGF+++++GIE+ Q AI +P K ELQ +G INF+ RFISN
Sbjct: 1059 LGFLVHERGIEVTQRSVNAIKKIQPPENKTELQEMIGNINFVRRFISN 1106
>UniRef100_Q7XNI8 OSJNBb0032D24.5 protein [Oryza sativa]
Length = 1355
Score = 99.8 bits (247), Expect = 5e-19
Identities = 53/135 (39%), Positives = 78/135 (57%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+ A+FE+P+ HLKPL+I VN L+K++VDGG VNL+P + +K+G S +L
Sbjct: 335 EQAVFEKPEGTENRHLKPLYINGYVNGKPLSKMIVDGGAAVNLMPYATFRKLGRSSDDLI 394
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +P
Sbjct: 395 KTNMVLKDFGGNQSETKGVLNVELTVGSKTVPTTFFVIDGKGSYSLLLGRDWIHANCCIP 454
Query: 806 STMH*RLALWREDGV 820
STM L W+ D +
Sbjct: 455 STMQQCLIQWQGDKI 469
>UniRef100_Q6UUM4 Hypothetical protein [Oryza sativa]
Length = 760
Score = 99.4 bits (246), Expect = 7e-19
Identities = 52/135 (38%), Positives = 78/135 (57%)
Query: 686 QHAMFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLK 745
+HA+FE+P+ HLKPL+I VN ++K++VDGG VNL+P + +K+G + +L
Sbjct: 404 EHAVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYTTFRKLGRNAEDLI 463
Query: 746 PHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVP 805
N+ L + V+ V+L VG T F V+ K +Y+LLLG +WIH +P
Sbjct: 464 KTNMVLKDFGSNPSETKEVLNVELTVGNKTIPTTFSVIDGKGSYSLLLGRDWIHANCCIP 523
Query: 806 STMH*RLALWREDGV 820
STMH L W+ D +
Sbjct: 524 STMHQCLIQWQGDKI 538
>UniRef100_Q60E17 Putative polyprotein [Oryza sativa]
Length = 1304
Score = 99.4 bits (246), Expect = 7e-19
Identities = 77/285 (27%), Positives = 128/285 (44%), Gaps = 29/285 (10%)
Query: 569 RRFQRQKKVDGLKYVTNNDKGKNKQMETFKMIRKPATERIFPPLSAVKENLHVEDEEMTS 628
RR QR + N ++ + Q + + +++ T++++ S V ++ +E
Sbjct: 426 RRVQRMR---------NMEQFQENQQDIGRRLKRAKTKQVWRVKSKVVSADEIKADEAER 476
Query: 629 NFTDFEADFDVICVVLILPIE*DVTSEVTEDEEDFIKEMVVHKTMCYYVMNNGCVEGQHA 688
+ +V +LP E + + E+ +M++ + A
Sbjct: 477 LAKGKAIASSSVNMVFVLPAEFGIDQRHADIMEESCAQMILSP--------------EQA 522
Query: 689 MFERPDYHMKIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHN 748
+FE+P+ H KPL +K VN ++K+LVDGG VNL+P KK G S +L
Sbjct: 523 IFEKPEGTENRHFKPLCVKGFVNGKPMSKMLVDGGAAVNLMPYVTFKKSGRSADDLIKTK 582
Query: 749 VALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTM 808
+ L + GV+ V+L VG+ T F V+ K +Y+LLLG +WIH +PSTM
Sbjct: 583 MVLKDFGGNHTETKGVLNVELTVGSKIVPTTFFVIDGKGSYSLLLGRDWIHANCCIPSTM 642
Query: 809 H*RLALWREDG------VLEILKLSKVTLLLRLTTSPKRTSTNNW 847
H L W+ D VL I K K+ + + K TS + +
Sbjct: 643 HHSLIQWQGDKIEVSNIVLVIKKNGKLRVCIDFRDLNKATSKDEY 687
Score = 45.8 bits (107), Expect = 0.009
Identities = 20/44 (45%), Positives = 31/44 (70%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIR 1308
LGF+++++GIE+ Q AI KP KK+LQ+ +GK+NF+ R
Sbjct: 799 LGFLVHERGIEMPQRSINAIQKIKPPEDKKQLQSLIGKMNFIRR 842
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,870,462,295
Number of Sequences: 2790947
Number of extensions: 72314492
Number of successful extensions: 376996
Number of sequences better than 10.0: 594
Number of HSP's better than 10.0 without gapping: 551
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 375767
Number of HSP's gapped (non-prelim): 1242
length of query: 1312
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1172
effective length of database: 457,317,253
effective search space: 535975820516
effective search space used: 535975820516
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 81 (35.8 bits)
Medicago: description of AC124217.1


