

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.5 - phase: 0
(340 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
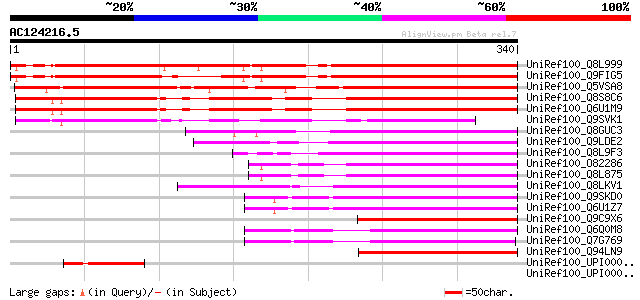
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8L999 Hypothetical protein [Arabidopsis thaliana] 434 e-120
UniRef100_Q9FIG5 Gb|AAF18661.1 [Arabidopsis thaliana] 373 e-102
UniRef100_Q5VSA8 Putative basic pentacysteine 4 [Oryza sativa] 340 3e-92
UniRef100_Q8S8C6 Expressed protein [Arabidopsis thaliana] 294 3e-78
UniRef100_Q6U1M9 Basic pentacysteine 4 [Arabidopsis thaliana] 293 5e-78
UniRef100_Q9SVK1 Hypothetical protein F19H22.10 [Arabidopsis tha... 220 4e-56
UniRef100_Q8GUC3 Barley B recombinant [Hordeum vulgare] 159 1e-37
UniRef100_Q9LDE2 F10B6.5 [Arabidopsis thaliana] 156 7e-37
UniRef100_Q8L9F3 Hypothetical protein [Arabidopsis thaliana] 154 3e-36
UniRef100_O82286 Hypothetical protein At2g35550 [Arabidopsis tha... 153 7e-36
UniRef100_Q8L875 Hypothetical protein At2g35550/T32F12.7 [Arabid... 153 7e-36
UniRef100_Q8LKV1 GAGA-binding protein [Glycine max] 152 2e-35
UniRef100_Q9SKD0 Hypothetical protein At2g01930 [Arabidopsis tha... 151 3e-35
UniRef100_Q6U1Z7 Basic pentacysteine 1 [Arabidopsis thaliana] 151 3e-35
UniRef100_Q9C9X6 Hypothetical protein T23K23.3 [Arabidopsis thal... 148 2e-34
UniRef100_Q6Q0M8 Barley B recombinant like-protein B [Oryza sativa] 146 7e-34
UniRef100_Q7G769 Hypothetical protein OJ1014H12.2 [Oryza sativa] 145 1e-33
UniRef100_Q94LN9 Hypothetical protein OSJNBa0092N12.3 [Oryza sat... 137 6e-31
UniRef100_UPI000031F17A UPI000031F17A UniRef100 entry 50 9e-05
UniRef100_UPI000023D377 UPI000023D377 UniRef100 entry 43 0.011
>UniRef100_Q8L999 Hypothetical protein [Arabidopsis thaliana]
Length = 342
Score = 434 bits (1115), Expect = e-120
Identities = 237/360 (65%), Positives = 279/360 (76%), Gaps = 38/360 (10%)
Query: 1 MDD---RENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEK 57
MDD RENGRHKA + QGQWLMQ HQ PSMKQ+MSI+AERDAAIQERNLA+SEK
Sbjct: 1 MDDGGHRENGRHKA----AVQGQWLMQ---HQ-PSMKQVMSIIAERDAAIQERNLAISEK 52
Query: 58 KAALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANG---GMSSCPPGCQIS 114
KAA+AERDMAFLQRDTAIAERNNA+MERD+A+ LQ+REN++ MS+CPPGCQIS
Sbjct: 53 KAAVAERDMAFLQRDTAIAERNNAIMERDSALTALQYRENSMVTAPAANMSACPPGCQIS 112
Query: 115 RGVKHIHHLPQ----QVNHLPNMGDSSYGTRELHTTDALPAAPVS---LEVGKPPRRAKR 167
RGVKH+HH Q +H+P + +++Y TRE+ D LP +P + LE KP +R KR
Sbjct: 113 RGVKHLHHPHMHHHHQQHHIPQLTENAYETREMEPNDGLPTSPPAGSTLESAKP-KRGKR 171
Query: 168 --PKESKSDSPNKKTPKS-RKVKKEG-DDLNKTMFANNKELEWKSSEEIINGDDDLNKQL 223
PK + + NK+ PK+ RKVKKE DDLNK MF K++ + D+D +K +
Sbjct: 172 VNPKATTQTAANKRGPKNQRKVKKESEDDLNKIMFV-------KTTHDYT--DEDSSKHI 222
Query: 224 AI-SKADWKPQDLA-LNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVY 281
I SK+DWK Q++ LNQV YD++TMP P CSCTGVLRQCYKWGNGGWQS+CCTTTLS+Y
Sbjct: 223 LIGSKSDWKSQEMVGLNQVVYDETTMPPPVCSCTGVLRQCYKWGNGGWQSSCCTTTLSMY 282
Query: 282 PLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEG-HDLSHPVDLKDHWAKHGTNRYITIK 340
PLPA+PNKRHARVGGRKMSGSAFNKLLSRLAAEG HDLS+PVDLKDHWAKHGTNRYITIK
Sbjct: 283 PLPALPNKRHARVGGRKMSGSAFNKLLSRLAAEGHHDLSNPVDLKDHWAKHGTNRYITIK 342
>UniRef100_Q9FIG5 Gb|AAF18661.1 [Arabidopsis thaliana]
Length = 301
Score = 373 bits (957), Expect = e-102
Identities = 216/353 (61%), Positives = 250/353 (70%), Gaps = 65/353 (18%)
Query: 1 MDD---RENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEK 57
MDD RENGRHKA + QGQWLMQ HQ PSMKQ+MSI+AERDAAIQERNLA+SEK
Sbjct: 1 MDDGGHRENGRHKA----AVQGQWLMQ---HQ-PSMKQVMSIIAERDAAIQERNLAISEK 52
Query: 58 KAALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGV 117
KAA+AERDMAFLQRDTAIAERNNA+MERD+A+ LQ+REN++ P I
Sbjct: 53 KAAVAERDMAFLQRDTAIAERNNAIMERDSALTALQYRENSMVTA------PAANI---- 102
Query: 118 KHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVS---LEVGKPPRRAKR--PKESK 172
E+ D LP +P + LE KP +R KR PK +
Sbjct: 103 ------------------------EMEPNDGLPTSPPAGSTLESAKP-KRGKRVNPKATT 137
Query: 173 SDSPNKKTPKS-RKVKKEG-DDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAI-SKAD 229
+ NK+ PK+ RKVKKE DDLNK MF K++ + D+D +K + I SK+D
Sbjct: 138 QTAANKRGPKNQRKVKKESEDDLNKIMFV-------KTTHDYT--DEDSSKHILIGSKSD 188
Query: 230 WKPQDLA-LNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPN 288
WK Q++ LNQV YD++TMP P CSCTGVLRQCYKWGNGGWQS+CCTTTLS+YPLPA+PN
Sbjct: 189 WKSQEMVGLNQVVYDETTMPPPVCSCTGVLRQCYKWGNGGWQSSCCTTTLSMYPLPALPN 248
Query: 289 KRHARVGGRKMSGSAFNKLLSRLAAEG-HDLSHPVDLKDHWAKHGTNRYITIK 340
KRHARVGGRKMSGSAFNKLLSRLAAEG HDLS+PVDLKDHWAKHGTNRYITIK
Sbjct: 249 KRHARVGGRKMSGSAFNKLLSRLAAEGHHDLSNPVDLKDHWAKHGTNRYITIK 301
>UniRef100_Q5VSA8 Putative basic pentacysteine 4 [Oryza sativa]
Length = 331
Score = 340 bits (873), Expect = 3e-92
Identities = 189/348 (54%), Positives = 232/348 (66%), Gaps = 34/348 (9%)
Query: 4 RENGRHKADQYKSAQGQWLM---QQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA 60
RENGR + DQYK QW+M Q+H H SM ++++M +RD AI+ER+ AL+EKKAA
Sbjct: 7 RENGRQRPDQYKGLHTQWMMPQTQRHLKDHQSMN-LLALMNDRDNAIRERDHALAEKKAA 65
Query: 61 LAERDMAFLQRDTAIAERNNALMERDNAIATLQF-RENALANGGMSSCPPGCQISRGVKH 119
+AERDMAF QRD A+AERN A++ERDNA+A L+ R N L + P G G K+
Sbjct: 66 IAERDMAFTQRDAAMAERNAAVVERDNALAALELARTNGLNMNNGNGFPQGSL--SGSKN 123
Query: 120 IHHLPQQVNHLPN----MGDSSYG-TRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSD 174
IHH Q++H + + DS Y RE+H ++A P + GK AKRPK++ S
Sbjct: 124 IHH-HDQLSHAQSSPLQLADSPYDHAREMHISEAYPISTAPGSAGK----AKRPKKNSSQ 178
Query: 175 SPNKKTPKS--RKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKP 232
+ K P RK KK D WK+ GDD + ++ K +WK
Sbjct: 179 ASPLKRPSGVLRKTKKPSGD-------------WKNVGMSGCGDDSAHA--SVMKNEWKD 223
Query: 233 QDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHA 292
Q+L LNQVA+DDSTMPAPACSCTG LRQCYKWGNGGWQS+CCT +S+YPLP +PNKRHA
Sbjct: 224 QNLGLNQVAFDDSTMPAPACSCTGKLRQCYKWGNGGWQSSCCTMNISMYPLPVMPNKRHA 283
Query: 293 RVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
R+GGRKMSG AF KLLSRLAAEGHDLS PVDLKDHWAKHGTNRYITI+
Sbjct: 284 RMGGRKMSGGAFTKLLSRLAAEGHDLSTPVDLKDHWAKHGTNRYITIR 331
>UniRef100_Q8S8C6 Expressed protein [Arabidopsis thaliana]
Length = 296
Score = 294 bits (752), Expect = 3e-78
Identities = 167/342 (48%), Positives = 210/342 (60%), Gaps = 59/342 (17%)
Query: 5 ENGRHKADQYKSAQGQW-LMQQHQ--HQHPSM---KQIMSIMAERDAAIQERNLALSEKK 58
+N R K D +K AQ W ++ QHQ QH ++ K+IMSI+AERDAA+ ERN A+S KK
Sbjct: 8 DNARFKPDYFKGAQSMWNMIPQHQIKEQHNALVMNKKIMSILAERDAAVHERNQAVSAKK 67
Query: 59 AALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVK 118
A+A RD A QRD A++ER+ AL+ERDNA A LQ EN+L N +S G K
Sbjct: 68 EAVAARDEALQQRDKALSERDKALIERDNAYAALQHHENSL-NFALS----------GGK 116
Query: 119 HIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNK 178
+ GD +G E H + P + + EV KR KE+K
Sbjct: 117 CVD------------GDDCFGIGEPHKLEVFPLSTIPPEVTNTKVVNKRKKENKQGLS-- 162
Query: 179 KTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALN 238
KVKK G+DLN+ + A K+ S+ DW QD+ LN
Sbjct: 163 ------KVKKVGEDLNRRVPAPGKK----------------------SRTDWDSQDVGLN 194
Query: 239 QVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRK 298
V +D++TMP P CSCTG RQCYKWGNGGWQS+CCTTTLS YPLP +PNKRH+R+GGRK
Sbjct: 195 LVTFDETTMPVPMCSCTGSTRQCYKWGNGGWQSSCCTTTLSQYPLPQMPNKRHSRMGGRK 254
Query: 299 MSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
MSG+ F++LLSRL+AEG+DLS PVDLKD+WA+HGTNRYITIK
Sbjct: 255 MSGNVFSRLLSRLSAEGYDLSCPVDLKDYWARHGTNRYITIK 296
>UniRef100_Q6U1M9 Basic pentacysteine 4 [Arabidopsis thaliana]
Length = 296
Score = 293 bits (750), Expect = 5e-78
Identities = 167/342 (48%), Positives = 210/342 (60%), Gaps = 59/342 (17%)
Query: 5 ENGRHKADQYKSAQGQW-LMQQHQ--HQHPSM---KQIMSIMAERDAAIQERNLALSEKK 58
+N R K D +K AQ W ++ QHQ QH ++ K+IMSI+AERDAA+ ERN A+S KK
Sbjct: 8 DNARFKPDYFKVAQSMWNMIPQHQIKEQHNALVMNKKIMSILAERDAAVHERNQAVSAKK 67
Query: 59 AALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVK 118
A+A RD A QRD A++ER+ AL+ERDNA A LQ EN+L N +S G K
Sbjct: 68 EAVAARDEALQQRDKALSERDKALIERDNAYAALQHHENSL-NFALS----------GGK 116
Query: 119 HIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNK 178
+ GD +G E H + P + + EV KR KE+K
Sbjct: 117 CVD------------GDDCFGIGEPHKLEVFPLSTIPPEVTNTKVVNKRKKENKQGLS-- 162
Query: 179 KTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALN 238
KVKK G+DLN+ + A K+ S+ DW QD+ LN
Sbjct: 163 ------KVKKVGEDLNRRVPAPGKK----------------------SRTDWDSQDVGLN 194
Query: 239 QVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRK 298
V +D++TMP P CSCTG RQCYKWGNGGWQS+CCTTTLS YPLP +PNKRH+R+GGRK
Sbjct: 195 LVTFDETTMPVPMCSCTGSTRQCYKWGNGGWQSSCCTTTLSQYPLPQMPNKRHSRMGGRK 254
Query: 299 MSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
MSG+ F++LLSRL+AEG+DLS PVDLKD+WA+HGTNRYITIK
Sbjct: 255 MSGNVFSRLLSRLSAEGYDLSCPVDLKDYWARHGTNRYITIK 296
>UniRef100_Q9SVK1 Hypothetical protein F19H22.10 [Arabidopsis thaliana]
Length = 445
Score = 220 bits (561), Expect = 4e-56
Identities = 130/311 (41%), Positives = 178/311 (56%), Gaps = 66/311 (21%)
Query: 5 ENGRHKADQYKSAQGQWLMQQHQHQHPSM---KQIMSIMAERDAAIQERNLALSEKKAAL 61
ENGR+K D YK Q +M + + QH ++ K+I+SI+AERDAA++ERN A++ K AL
Sbjct: 22 ENGRYKPDYYKGTQSVNVMPKKE-QHNALVMNKKIISILAERDAAVKERNEAVAATKEAL 80
Query: 62 AERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVKHIH 121
A RD A QRD A++ER+NA+ME ++A+ L++REN L + SC RG
Sbjct: 81 ASRDEALEQRDKALSERDNAIMETESALNALRYRENNL--NYILSCA-----KRG----- 128
Query: 122 HLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTP 181
G + T E H + P + + P E+ + P K+
Sbjct: 129 ------------GSQRFITEESHLPNPSPISTI-------------PPEAANTRPTKRKK 163
Query: 182 KSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQVA 241
+S++ KK G+DLN+ + + K+ S+ DW D+ V
Sbjct: 164 ESKQGKKMGEDLNRPVASPGKK----------------------SRKDWDSNDVL---VT 198
Query: 242 YDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSG 301
+D+ TMP P C+CTG RQCYKWGNGGWQS+CCTTTLS YPLP +PNKRH+RVGGRKMSG
Sbjct: 199 FDEMTMPVPMCTCTGTARQCYKWGNGGWQSSCCTTTLSEYPLPQMPNKRHSRVGGRKMSG 258
Query: 302 SAFNKLLSRLA 312
S F++LLSRLA
Sbjct: 259 SVFSRLLSRLA 269
>UniRef100_Q8GUC3 Barley B recombinant [Hordeum vulgare]
Length = 350
Score = 159 bits (401), Expect = 1e-37
Identities = 87/233 (37%), Positives = 121/233 (51%), Gaps = 33/233 (14%)
Query: 119 HIHHLPQQVNHLPNMGDSSYGTRELHTTDAL----PAAPVSLEVGKPPRR------AKRP 168
HI H V+H P R HT + P P ++ P +K P
Sbjct: 140 HIGHPGHAVHHHPTGFGMMPDARGAHTLQMMQPQEPPVPDEEKITPPLVEDHSVVGSKPP 199
Query: 169 KESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAI-SK 227
+ + K PK +K KK+ G+D K A S+
Sbjct: 200 VKKRQQGRQPKVPKPKKPKKDATP----------------------GEDGAPKARAPRSR 237
Query: 228 ADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVP 287
KP ++ +N + +D S +P P CSCTG +QCY+WG GGWQSACCTT++S YPLP
Sbjct: 238 GPLKPVEMVINGIDFDISRIPTPVCSCTGAPQQCYRWGAGGWQSACCTTSISTYPLPMNT 297
Query: 288 NKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
+R AR+ GRKMS AF K+L +LA EG++L++P+DLK WAKHGTN+++TI+
Sbjct: 298 KRRGARIAGRKMSQGAFKKVLEKLAGEGYNLNNPIDLKTFWAKHGTNKFVTIR 350
>UniRef100_Q9LDE2 F10B6.5 [Arabidopsis thaliana]
Length = 279
Score = 156 bits (395), Expect = 7e-37
Identities = 83/217 (38%), Positives = 119/217 (54%), Gaps = 23/217 (10%)
Query: 124 PQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKS 183
P N LP + LH PV LE + KR +K+ S TPK+
Sbjct: 86 PNYGNVLPETSSAPSMQMNLHHHLQTEENPVKLEEEIVVQTKKRKTNAKAGS----TPKA 141
Query: 184 RKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQVAYD 243
+K +K D+ + N +++ N + K K DL +N V+ D
Sbjct: 142 KKPRKPKDENS-------------------NNNNNNNTNVTRVKPAKKSVDLVINGVSMD 182
Query: 244 DSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSA 303
S +P P C+CTG +QCY+WG GGWQSACCTT +S++PLP +R AR+ GRKMS A
Sbjct: 183 ISGLPVPICTCTGAPQQCYRWGCGGWQSACCTTNISMHPLPMSTKRRGARISGRKMSQGA 242
Query: 304 FNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
F K+L +LA++G + +P+DLK HWA+HGTN+++TI+
Sbjct: 243 FKKVLEKLASDGFNFGNPIDLKSHWARHGTNKFVTIR 279
>UniRef100_Q8L9F3 Hypothetical protein [Arabidopsis thaliana]
Length = 285
Score = 154 bits (390), Expect = 3e-36
Identities = 78/191 (40%), Positives = 108/191 (55%), Gaps = 24/191 (12%)
Query: 150 PAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSS 209
PAAP + GK KR +P PK++K++K K
Sbjct: 119 PAAPCEEQTGK-----KRKMRGSIATPT--VPKAKKMRKP-----------------KEE 154
Query: 210 EEIINGDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGW 269
++ N + +Q K K DL +N V+ D S +P P C+CTG +QCY+WG GGW
Sbjct: 155 RDVTNNNVQQQQQKQRVKPVKKSVDLVINGVSMDISGLPVPVCTCTGTPQQCYRWGCGGW 214
Query: 270 QSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWA 329
QSACCTT +SVYPLP +R AR+ GRKMS AF K+L +L+ EG+ + +DLK HWA
Sbjct: 215 QSACCTTNISVYPLPMSTKRRGARISGRKMSQGAFKKVLEKLSTEGYSFGNAIDLKSHWA 274
Query: 330 KHGTNRYITIK 340
+HGTN+++TI+
Sbjct: 275 RHGTNKFVTIR 285
>UniRef100_O82286 Hypothetical protein At2g35550 [Arabidopsis thaliana]
Length = 226
Score = 153 bits (386), Expect = 7e-36
Identities = 81/182 (44%), Positives = 109/182 (59%), Gaps = 20/182 (10%)
Query: 161 PPRRAKR--PKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDD 218
PP R R + K S N+ K+ K K + K +NK + S E
Sbjct: 63 PPIRDSRNDTETVKQKSVNQSPSKALKPKPQ----RKKRSVSNKSKKTPSIPE------- 111
Query: 219 LNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTL 278
+K + K D+ ++ ++D S +P P CSCTGV R CYKWG GGWQS+CCT ++
Sbjct: 112 -------TKREKKNLDINIDISSFDTSGVPPPVCSCTGVSRVCYKWGMGGWQSSCCTISI 164
Query: 279 SVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYIT 338
S YPLP + AR+ GRKMS A+ KLL+RLA EG+DLSHP+DLK+HWA+HGTN+++T
Sbjct: 165 STYPLPMSTTRPGARLAGRKMSNGAYVKLLARLADEGYDLSHPLDLKNHWARHGTNKFVT 224
Query: 339 IK 340
IK
Sbjct: 225 IK 226
>UniRef100_Q8L875 Hypothetical protein At2g35550/T32F12.7 [Arabidopsis thaliana]
Length = 271
Score = 153 bits (386), Expect = 7e-36
Identities = 81/182 (44%), Positives = 109/182 (59%), Gaps = 20/182 (10%)
Query: 161 PPRRAKR--PKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDD 218
PP R R + K S N+ K+ K K + K +NK + S E
Sbjct: 108 PPIRDSRNDTETVKQKSVNQSPSKALKPKPQ----RKKRSVSNKSKKTPSIPE------- 156
Query: 219 LNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTL 278
+K + K D+ ++ ++D S +P P CSCTGV R CYKWG GGWQS+CCT ++
Sbjct: 157 -------TKREKKNLDINIDISSFDTSGVPPPVCSCTGVSRVCYKWGMGGWQSSCCTISI 209
Query: 279 SVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYIT 338
S YPLP + AR+ GRKMS A+ KLL+RLA EG+DLSHP+DLK+HWA+HGTN+++T
Sbjct: 210 STYPLPMSTTRPGARLAGRKMSNGAYVKLLARLADEGYDLSHPLDLKNHWARHGTNKFVT 269
Query: 339 IK 340
IK
Sbjct: 270 IK 271
>UniRef100_Q8LKV1 GAGA-binding protein [Glycine max]
Length = 282
Score = 152 bits (383), Expect = 2e-35
Identities = 82/228 (35%), Positives = 126/228 (54%), Gaps = 22/228 (9%)
Query: 113 ISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESK 172
IS+ K+++ +P N+ SS ++ LP +E + ++ +
Sbjct: 77 ISQRDKYMNMIPTNHNYGGIPETSSAHQIQMIQAPELPKEEKPVEEAPVIEKENGTRKKR 136
Query: 173 SDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKP 232
S K+PK++K K+ G K D+ + ++ K
Sbjct: 137 QSSKVPKSPKAKKPKR-GPRAPK---------------------DESAPAVQRARIPKKT 174
Query: 233 QDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHA 292
++ +N + D S++P P CSCTG +QCYKWG+GGWQSACCTT +SVYPLP +R A
Sbjct: 175 TEIVINGIDMDISSIPIPVCSCTGAPQQCYKWGSGGWQSACCTTGMSVYPLPMSTKRRGA 234
Query: 293 RVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
R+ GRKMS AF K+L +LAAEG++ S+P+DL+ +WAKHGTN+++TI+
Sbjct: 235 RIAGRKMSIGAFKKVLEKLAAEGYNFSNPIDLRTYWAKHGTNKFVTIR 282
>UniRef100_Q9SKD0 Hypothetical protein At2g01930 [Arabidopsis thaliana]
Length = 283
Score = 151 bits (381), Expect = 3e-35
Identities = 76/186 (40%), Positives = 113/186 (59%), Gaps = 7/186 (3%)
Query: 158 VGKPPRRAKRPKESKSDSP---NKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIIN 214
+ +P + R +E+ P ++T K RK++ G T+ K + K ++ N
Sbjct: 102 IHQPVLNSSRFEENPIPPPAPCEEQTGKKRKMR--GSIATPTVPKAKKMRKPKEERDVTN 159
Query: 215 GDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACC 274
+++ +Q K K DL +N V+ D S +P P C+CTG +QCY+WG GGWQSACC
Sbjct: 160 --NNVQQQQQRVKPVKKSVDLVINGVSMDISGLPVPVCTCTGTPQQCYRWGCGGWQSACC 217
Query: 275 TTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTN 334
TT +SVYPLP +R AR+ GRKMS AF K+L +L+ EG+ + +DLK HWA+HGTN
Sbjct: 218 TTNISVYPLPMSTKRRGARISGRKMSQGAFKKVLEKLSTEGYSFGNAIDLKSHWARHGTN 277
Query: 335 RYITIK 340
+++TI+
Sbjct: 278 KFVTIR 283
>UniRef100_Q6U1Z7 Basic pentacysteine 1 [Arabidopsis thaliana]
Length = 283
Score = 151 bits (381), Expect = 3e-35
Identities = 76/186 (40%), Positives = 113/186 (59%), Gaps = 7/186 (3%)
Query: 158 VGKPPRRAKRPKESKSDSP---NKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIIN 214
+ +P + R +E+ P ++T K RK++ G T+ K + K ++ N
Sbjct: 102 IHQPVLNSSRFEENPIPPPAPCEEQTGKKRKMR--GSIATPTVPKAKKMRKPKEERDVTN 159
Query: 215 GDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACC 274
+++ +Q K K DL +N V+ D S +P P C+CTG +QCY+WG GGWQSACC
Sbjct: 160 --NNVQQQQQRVKPVKKSVDLVINGVSMDISGLPVPVCTCTGTPQQCYRWGCGGWQSACC 217
Query: 275 TTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTN 334
TT +SVYPLP +R AR+ GRKMS AF K+L +L+ EG+ + +DLK HWA+HGTN
Sbjct: 218 TTNISVYPLPMSTKRRGARISGRKMSQGAFKKVLEKLSTEGYSFGNAIDLKSHWARHGTN 277
Query: 335 RYITIK 340
+++TI+
Sbjct: 278 KFVTIR 283
>UniRef100_Q9C9X6 Hypothetical protein T23K23.3 [Arabidopsis thaliana]
Length = 269
Score = 148 bits (374), Expect = 2e-34
Identities = 60/107 (56%), Positives = 83/107 (77%)
Query: 234 DLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHAR 293
++ +N V+ D +P P CSCTG+ +QCY+WG GGWQSACCTT +S+YPLP +R AR
Sbjct: 163 EMVINGVSMDIGGLPVPVCSCTGMPQQCYRWGCGGWQSACCTTNVSMYPLPVNTKRRGAR 222
Query: 294 VGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
+ GRKMS AF K+L +L+++G D S+P+DLK HWAKHGTN+++TI+
Sbjct: 223 IAGRKMSQGAFRKVLEKLSSDGFDFSNPIDLKSHWAKHGTNKFVTIR 269
>UniRef100_Q6Q0M8 Barley B recombinant like-protein B [Oryza sativa]
Length = 341
Score = 146 bits (369), Expect = 7e-34
Identities = 76/183 (41%), Positives = 101/183 (54%), Gaps = 25/183 (13%)
Query: 158 VGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDD 217
+ +PP KR + + P K PK +E D A + K+ +ING D
Sbjct: 184 IDEPPPPKKRQQGRQPKVPRAKKPKKSAAPRE-DGAPPNAPAPRRRGPRKNIGMVINGID 242
Query: 218 DLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTT 277
D S +P P CSCTG +QCY+WG GGWQSACCTTT
Sbjct: 243 ------------------------LDLSRIPTPICSCTGAPQQCYRWGAGGWQSACCTTT 278
Query: 278 LSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYI 337
+S YPLP +R AR+ GRKMS AF K+L +LA EG++L++P+DLK WAKHGTN+++
Sbjct: 279 ISTYPLPMSTKRRGARIAGRKMSHGAFKKVLEKLAGEGYNLNNPIDLKTFWAKHGTNKFV 338
Query: 338 TIK 340
TI+
Sbjct: 339 TIR 341
>UniRef100_Q7G769 Hypothetical protein OJ1014H12.2 [Oryza sativa]
Length = 344
Score = 145 bits (367), Expect = 1e-33
Identities = 76/182 (41%), Positives = 100/182 (54%), Gaps = 25/182 (13%)
Query: 158 VGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDD 217
+ +PP KR + + P K PK +E D A + K+ +ING D
Sbjct: 184 IDEPPPPKKRQQGRQPKVPRAKKPKKSAAPRE-DGAPPNAPAPRRRGPRKNIGMVINGID 242
Query: 218 DLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTT 277
D S +P P CSCTG +QCY+WG GGWQSACCTTT
Sbjct: 243 ------------------------LDLSRIPTPICSCTGAPQQCYRWGAGGWQSACCTTT 278
Query: 278 LSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYI 337
+S YPLP +R AR+ GRKMS AF K+L +LA EG++L++P+DLK WAKHGTN+++
Sbjct: 279 ISTYPLPMSTKRRGARIAGRKMSHGAFKKVLEKLAGEGYNLNNPIDLKTFWAKHGTNKFV 338
Query: 338 TI 339
TI
Sbjct: 339 TI 340
>UniRef100_Q94LN9 Hypothetical protein OSJNBa0092N12.3 [Oryza sativa]
Length = 343
Score = 137 bits (344), Expect = 6e-31
Identities = 59/106 (55%), Positives = 79/106 (73%)
Query: 235 LALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARV 294
+ +N + D S +P CSCTG +Q Y+WG GGWQSACCTTT+S YPLP R AR+
Sbjct: 238 MVINGIDLDLSRIPTRICSCTGAPQQRYRWGAGGWQSACCTTTVSTYPLPMSMKPRGARI 297
Query: 295 GGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
GRKMS AF K+L +LA+EG++L++P+DLK WAKHGTN+++TI+
Sbjct: 298 AGRKMSHGAFKKVLEKLASEGYNLNNPIDLKTFWAKHGTNKFVTIR 343
>UniRef100_UPI000031F17A UPI000031F17A UniRef100 entry
Length = 496
Score = 50.1 bits (118), Expect = 9e-05
Identities = 23/54 (42%), Positives = 42/54 (77%), Gaps = 3/54 (5%)
Query: 37 MSIMAERDAAIQERNLALSEKKAALAERDMAFLQRDTAIAERNNALMERDNAIA 90
M+++A+++ + + A ++ +AL+ERD A +RDTA+A+RNNA++ER++AIA
Sbjct: 1 MAVLAQKEGLLAD---ACEQRDSALSERDSAVNERDTALAQRNNAIVERNDAIA 51
>UniRef100_UPI000023D377 UPI000023D377 UniRef100 entry
Length = 349
Score = 43.1 bits (100), Expect = 0.011
Identities = 32/133 (24%), Positives = 58/133 (43%), Gaps = 13/133 (9%)
Query: 104 MSSCPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPR 163
+ S PG +R +HH+P P+ +S ++ T + PVS++V P +
Sbjct: 4 LGSASPGPTAARFRDQLHHIPPPS---PSPNITSMKSKRARETAEVDEKPVSIDVEAPAK 60
Query: 164 RAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLN--K 221
+ KR + KK + +K KKE ++ + K +++ DDD+ +
Sbjct: 61 KRKREDDGADVKKAKKEKRDKKFKKES--------RKERKEKRKDLQDLPEQDDDVEDAQ 112
Query: 222 QLAISKADWKPQD 234
++ AD KP D
Sbjct: 113 DTPMAGADDKPAD 125
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.128 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 590,001,565
Number of Sequences: 2790947
Number of extensions: 24868037
Number of successful extensions: 77195
Number of sequences better than 10.0: 296
Number of HSP's better than 10.0 without gapping: 176
Number of HSP's successfully gapped in prelim test: 125
Number of HSP's that attempted gapping in prelim test: 76389
Number of HSP's gapped (non-prelim): 707
length of query: 340
length of database: 848,049,833
effective HSP length: 128
effective length of query: 212
effective length of database: 490,808,617
effective search space: 104051426804
effective search space used: 104051426804
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC124216.5


