

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
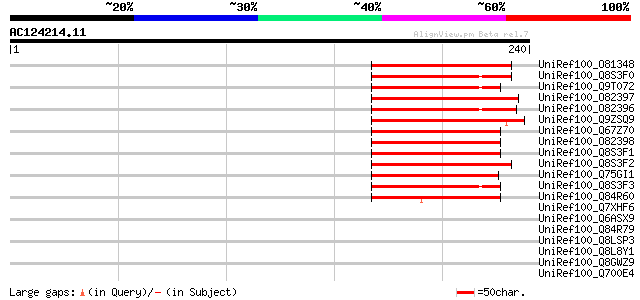
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81348 Symbiotic ammonium transporter [Glycine max] 96 1e-18
UniRef100_Q8S3F0 Putative bHLH transcription factor [Arabidopsis... 66 9e-10
UniRef100_Q9T072 Hypothetical protein AT4g37850 [Arabidopsis tha... 65 1e-09
UniRef100_O82397 Putative bHLH transcription factor [Arabidopsis... 63 6e-09
UniRef100_O82396 Putative bHLH transcription factor [Arabidopsis... 63 6e-09
UniRef100_Q9ZSQ9 Transporter homolog [Mesembryanthemum crystalli... 62 2e-08
UniRef100_Q67Z70 Putative bHLH transcription factor [Arabidopsis... 61 2e-08
UniRef100_O82398 Putative bHLH transcription factor [Arabidopsis... 61 2e-08
UniRef100_Q8S3F1 Putative bHLH transcription factor [Arabidopsis... 61 2e-08
UniRef100_Q8S3F2 Putative bHLH transcription factor [Arabidopsis... 60 6e-08
UniRef100_Q75GI1 Putative symbiotic ammonium transport protein [... 59 8e-08
UniRef100_Q8S3F3 Putative bHLH transcription factor [Arabidopsis... 59 1e-07
UniRef100_Q84R60 Putative ammonium transporter [Oryza sativa] 47 3e-04
UniRef100_Q7XHF6 Putative bHLH transcription factor [Oryza sativa] 45 0.001
UniRef100_Q6ASX9 Hypothetical protein B1130H05.3 [Oryza sativa] 44 0.003
UniRef100_Q84R79 Putative ammonium transporter [Oryza sativa] 44 0.004
UniRef100_Q8LSP3 Putative bHLH transcription factor [Oryza sativa] 44 0.005
UniRef100_Q8L8Y1 Hypothetical protein [Arabidopsis thaliana] 36 0.97
UniRef100_Q8GWZ9 Putative bHLH transcription factor bHLH067 [Ara... 36 0.97
UniRef100_Q700E4 Hypothetical protein [Arabidopsis thaliana] 36 0.97
>UniRef100_O81348 Symbiotic ammonium transporter [Glycine max]
Length = 347
Score = 95.5 bits (236), Expect = 1e-18
Identities = 48/65 (73%), Positives = 55/65 (83%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIEARF +RNVLIR+HCEK KGV+EKTI +IEKL+LKV NSS +TFGS LDITIIAQ
Sbjct: 264 LPEIEARFWERNVLIRIHCEKNKGVIEKTISEIEKLHLKVINSSALTFGSFILDITIIAQ 323
Query: 228 PTLSF 232
+ F
Sbjct: 324 MDMEF 328
>UniRef100_Q8S3F0 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 304
Score = 65.9 bits (159), Expect = 9e-10
Identities = 35/65 (53%), Positives = 46/65 (69%), Gaps = 1/65 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE RF D +VLI++ CEK KG + K + +IEKL++ +TNSS + FG LDITIIA+
Sbjct: 222 LPEIEVRFSDEDVLIKILCEKQKGHLAKIMAEIEKLHILITNSSVLNFGP-TLDITIIAK 280
Query: 228 PTLSF 232
F
Sbjct: 281 KESDF 285
>UniRef100_Q9T072 Hypothetical protein AT4g37850 [Arabidopsis thaliana]
Length = 314
Score = 65.1 bits (157), Expect = 1e-09
Identities = 34/60 (56%), Positives = 45/60 (74%), Gaps = 1/60 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE RF D +VLI++ CEK KG + K + +IEKL++ +TNSS + FG LDITIIA+
Sbjct: 222 LPEIEVRFSDEDVLIKILCEKQKGHLAKIMAEIEKLHILITNSSVLNFGP-TLDITIIAK 280
>UniRef100_O82397 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 284
Score = 63.2 bits (152), Expect = 6e-09
Identities = 28/68 (41%), Positives = 44/68 (64%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIEA+ ++LIR+ CEK KG + ++ IE L++ NS + FG LDIT++AQ
Sbjct: 214 LPEIEAKISQNDILIRILCEKSKGCMINILNTIENFQLRIENSIVLPFGDSTLDITVLAQ 273
Query: 228 PTLSFYVF 235
++ Y++
Sbjct: 274 VIINNYIY 281
>UniRef100_O82396 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 288
Score = 63.2 bits (152), Expect = 6e-09
Identities = 34/67 (50%), Positives = 46/67 (67%), Gaps = 1/67 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE R ++VLI++ CEK KG V K + +IEKL L +TNS+ + FG DI+IIAQ
Sbjct: 223 LPEIEVRVSGKDVLIKILCEKQKGNVIKIMGEIEKLGLSITNSNVLPFGP-TFDISIIAQ 281
Query: 228 PTLSFYV 234
T+ F +
Sbjct: 282 VTIYFLI 288
>UniRef100_Q9ZSQ9 Transporter homolog [Mesembryanthemum crystallinum]
Length = 300
Score = 61.6 bits (148), Expect = 2e-08
Identities = 34/77 (44%), Positives = 48/77 (62%), Gaps = 6/77 (7%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
L EIEA C+ NVLIR+H +K + +V K +++IE L+L N + + FG A+DITI+AQ
Sbjct: 215 LLEIEAGACNNNVLIRIHAQKDQDLVRKVLNEIENLHLTTLNFNTIPFGGYAMDITIVAQ 274
Query: 228 P------TLSFYVFHLR 238
T+ V HLR
Sbjct: 275 MDDDFELTIKDVVMHLR 291
>UniRef100_Q67Z70 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 320
Score = 61.2 bits (147), Expect = 2e-08
Identities = 30/60 (50%), Positives = 41/60 (68%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+P IEAR DR++LIRVHCEK KG + K + +EK L+V NS + FG+ L ITI+ +
Sbjct: 239 MPMIEARVSDRDLLIRVHCEKNKGCMIKILSSLEKFRLEVVNSFTLPFGNSTLVITILTK 298
>UniRef100_O82398 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 322
Score = 61.2 bits (147), Expect = 2e-08
Identities = 30/60 (50%), Positives = 41/60 (68%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+P IEAR DR++LIRVHCEK KG + K + +EK L+V NS + FG+ L ITI+ +
Sbjct: 239 MPMIEARVSDRDLLIRVHCEKNKGCMIKILSSLEKFRLEVVNSFTLPFGNSTLVITILTK 298
>UniRef100_Q8S3F1 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 320
Score = 61.2 bits (147), Expect = 2e-08
Identities = 30/60 (50%), Positives = 41/60 (68%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+P IEAR DR++LIRVHCEK KG + K + +EK L+V NS + FG+ L ITI+ +
Sbjct: 239 MPMIEARVSDRDLLIRVHCEKNKGCMIKILSSLEKFRLEVVNSFTLPFGNSTLVITILTK 298
>UniRef100_Q8S3F2 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 295
Score = 59.7 bits (143), Expect = 6e-08
Identities = 28/65 (43%), Positives = 40/65 (61%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIEA+ ++LIR+ CEK KG + ++ IE L++ NS + FG LDIT++AQ
Sbjct: 214 LPEIEAKISQNDILIRILCEKSKGCMINILNTIENFQLRIENSIVLPFGDSTLDITVLAQ 273
Query: 228 PTLSF 232
F
Sbjct: 274 MDKDF 278
>UniRef100_Q75GI1 Putative symbiotic ammonium transport protein [Oryza sativa]
Length = 359
Score = 59.3 bits (142), Expect = 8e-08
Identities = 26/59 (44%), Positives = 41/59 (69%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIA 226
LPEIEAR +R VL+++HCE KG + + ++E + L + N++ + F S +LDITI+A
Sbjct: 276 LPEIEARVSERTVLVKIHCENRKGALITALSEVETIGLTIMNTNVLPFTSSSLDITIMA 334
>UniRef100_Q8S3F3 Putative bHLH transcription factor [Arabidopsis thaliana]
Length = 304
Score = 58.9 bits (141), Expect = 1e-07
Identities = 32/60 (53%), Positives = 42/60 (69%), Gaps = 1/60 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE R ++VLI++ CEK KG V K + +IEKL L +TNS+ + FG DI+IIAQ
Sbjct: 223 LPEIEVRVSGKDVLIKILCEKQKGNVIKIMGEIEKLGLSITNSNVLPFGP-TFDISIIAQ 281
>UniRef100_Q84R60 Putative ammonium transporter [Oryza sativa]
Length = 353
Score = 47.4 bits (111), Expect = 3e-04
Identities = 24/61 (39%), Positives = 40/61 (65%), Gaps = 1/61 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKI-KGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIA 226
LPEIEA+ NV++R+H E KG + + + +E L+L +T+++ M F +C ITI+A
Sbjct: 292 LPEIEAKISHGNVMVRIHGENNGKGSLVRLLAAVEGLHLGITHTNVMPFSACTAIITIMA 351
Query: 227 Q 227
+
Sbjct: 352 K 352
>UniRef100_Q7XHF6 Putative bHLH transcription factor [Oryza sativa]
Length = 361
Score = 45.4 bits (106), Expect = 0.001
Identities = 23/58 (39%), Positives = 36/58 (61%)
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
++EA VL+RV C + KGV+ K + ++EKL L N++ + F +L+ITI AQ
Sbjct: 282 KVEANLQGTTVLLRVVCPEKKGVLIKLLTELEKLGLSTMNTNVVPFADSSLNITITAQ 339
>UniRef100_Q6ASX9 Hypothetical protein B1130H05.3 [Oryza sativa]
Length = 269
Score = 44.3 bits (103), Expect = 0.003
Identities = 23/67 (34%), Positives = 42/67 (62%), Gaps = 4/67 (5%)
Query: 169 PEIEARFCDRN--VLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTF--GSCALDITI 224
PEIE R N V++R+HCE +GV+ + + ++E+++L++ N++ M F + ITI
Sbjct: 200 PEIEVRCSPTNNVVMVRIHCENGEGVIVRILAEVEEIHLRIINANVMPFLDQGATMIITI 259
Query: 225 IAQPTLS 231
A+ + S
Sbjct: 260 AAKASSS 266
>UniRef100_Q84R79 Putative ammonium transporter [Oryza sativa]
Length = 301
Score = 43.9 bits (102), Expect = 0.004
Identities = 22/67 (32%), Positives = 42/67 (61%), Gaps = 4/67 (5%)
Query: 169 PEIEARFCDRN--VLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTF--GSCALDITI 224
PEIE R N V++R+HCE +GV+ + + ++E+++L++ N++ M F + ITI
Sbjct: 211 PEIEVRCSPTNNVVMVRIHCENGEGVIVRILAEVEEIHLRIINANVMPFLDQGATMIITI 270
Query: 225 IAQPTLS 231
A+ ++
Sbjct: 271 AAKAKIN 277
>UniRef100_Q8LSP3 Putative bHLH transcription factor [Oryza sativa]
Length = 451
Score = 43.5 bits (101), Expect = 0.005
Identities = 20/68 (29%), Positives = 40/68 (58%)
Query: 169 PEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQP 228
P +EA VL+++ C++ +G++ + ++EK L + N+S + F L+ITI A+
Sbjct: 383 PTVEASIHGNTVLLKICCKERRGLLVMILSELEKQGLSIINTSVVPFTDSCLNITITAKA 442
Query: 229 TLSFYVFH 236
L+ V++
Sbjct: 443 RLALPVYY 450
>UniRef100_Q8L8Y1 Hypothetical protein [Arabidopsis thaliana]
Length = 358
Score = 35.8 bits (81), Expect = 0.97
Identities = 19/51 (37%), Positives = 29/51 (56%)
Query: 164 DYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMT 214
D +P+IEA +V ++V CEK +G + K I +EKL L V + + T
Sbjct: 269 DQTCIPKIEATVIQNHVSLKVQCEKKQGQLLKGIISLEKLKLTVLHLNITT 319
>UniRef100_Q8GWZ9 Putative bHLH transcription factor bHLH067 [Arabidopsis thaliana]
Length = 307
Score = 35.8 bits (81), Expect = 0.97
Identities = 19/51 (37%), Positives = 29/51 (56%)
Query: 164 DYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMT 214
D +P+IEA +V ++V CEK +G + K I +EKL L V + + T
Sbjct: 218 DQTCIPKIEATVIQNHVSLKVQCEKKQGQLLKGIISLEKLKLTVLHLNITT 268
>UniRef100_Q700E4 Hypothetical protein [Arabidopsis thaliana]
Length = 358
Score = 35.8 bits (81), Expect = 0.97
Identities = 19/51 (37%), Positives = 29/51 (56%)
Query: 164 DYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMT 214
D +P+IEA +V ++V CEK +G + K I +EKL L V + + T
Sbjct: 269 DQTCIPKIEATVIQNHVSLKVQCEKKQGQLLKGIISLEKLKLTVLHLNITT 319
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 287,898,356
Number of Sequences: 2790947
Number of extensions: 8947879
Number of successful extensions: 60772
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 60751
Number of HSP's gapped (non-prelim): 22
length of query: 240
length of database: 848,049,833
effective HSP length: 124
effective length of query: 116
effective length of database: 501,972,405
effective search space: 58228798980
effective search space used: 58228798980
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 73 (32.7 bits)
Medicago: description of AC124214.11


