

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123575.7 - phase: 0 /pseudo
(378 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
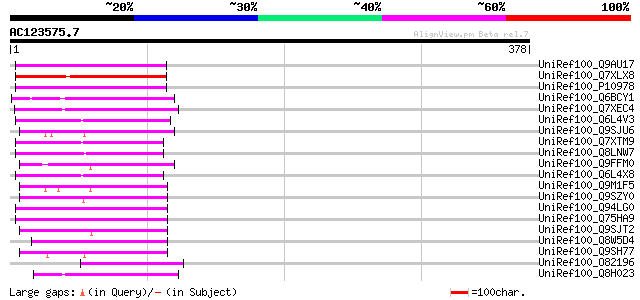
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense] 78 5e-13
UniRef100_Q7XLX8 OSJNBa0042I15.12 protein [Oryza sativa] 77 1e-12
UniRef100_P10978 Retrovirus-related Pol polyprotein from transpo... 76 1e-12
UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas] 73 1e-11
UniRef100_Q7XEC4 Putative polyprotein [Oryza sativa] 71 6e-11
UniRef100_Q6L4V3 Putative polyprotein [Oryza sativa] 67 6e-10
UniRef100_Q9SJU6 Putative retroelement pol polyprotein [Arabidop... 67 1e-09
UniRef100_Q7XTM9 OSJNBa0033G05.13 protein [Oryza sativa] 66 1e-09
UniRef100_Q8LNW7 Putative polyprotein [Oryza sativa] 65 2e-09
UniRef100_Q9FFM0 Copia-like retrotransposable element [Arabidops... 64 7e-09
UniRef100_Q6L4X8 Putative polyprotein [Oryza sativa] 64 7e-09
UniRef100_Q9M1F5 Copia-like polyprotein [Arabidopsis thaliana] 62 3e-08
UniRef100_Q9SZY0 Putative retrotransposon [Arabidopsis thaliana] 62 3e-08
UniRef100_Q94LG0 Putative retroelement pol polyprotein [Oryza sa... 61 6e-08
UniRef100_Q75HA9 Putative polyprotein [Oryza sativa] 60 8e-08
UniRef100_Q9SJT2 Putative retroelement pol polyprotein [Arabidop... 60 8e-08
UniRef100_Q8W5D4 Putative retrotransposon-related protein [Oryza... 60 8e-08
UniRef100_Q9SH77 Putative retroelement pol polyprotein [Arabidop... 57 8e-07
UniRef100_O82196 Copia-like retroelement pol polyprotein [Arabid... 57 1e-06
UniRef100_Q8H023 Putative retrovirus-related pol polyprotein [Or... 56 2e-06
>UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense]
Length = 1328
Score = 77.8 bits (190), Expect = 5e-13
Identities = 42/111 (37%), Positives = 67/111 (59%), Gaps = 1/111 (0%)
Query: 5 KWNIEKFTGSND-FGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAI 63
K+ + KF G F +W+ +M+ +LIQQ +AL GK++ P + + E+++KA SAI
Sbjct: 5 KYEVAKFNGDKPVFSMWQRRMKDLLIQQGLHKALGGKSKKPESMKLEDWEELDEKAASAI 64
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVE 114
L L D V+ + E +W KL++LYM+K+ T++ LK+QLY M E
Sbjct: 65 RLHLTDDVVNNIVDEESACGIWTKLENLYMSKTLTNKLYLKKQLYTLHMDE 115
>UniRef100_Q7XLX8 OSJNBa0042I15.12 protein [Oryza sativa]
Length = 432
Score = 76.6 bits (187), Expect = 1e-12
Identities = 40/110 (36%), Positives = 69/110 (62%), Gaps = 2/110 (1%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+++ KF G +FGLW+ +++ +L QQ+ +ALKG PA + + EM +A + I
Sbjct: 43 KFDLVKFDGFGNFGLWQTRVKDLLAQQEVSKALKGVK--PAKMEDDDCEEMQLQAAATIR 100
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVE 114
CL D+++ V ET +W KL++ +M+K+ T+++ LKQ+LY +M E
Sbjct: 101 RCLSDQIMYHVMEETSPKIIWEKLEAQFMSKTLTNKRYLKQKLYGLKMQE 150
>UniRef100_P10978 Retrovirus-related Pol polyprotein from transposon TNT 1-94
[Contains: Protease (EC 3.4.23.-); Reverse transcriptase
(EC 2.7.7.49); Endonuclease] [Nicotiana tabacum]
Length = 1328
Score = 76.3 bits (186), Expect = 1e-12
Identities = 39/110 (35%), Positives = 64/110 (57%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+ + KF G N F W+ +MR +LIQQ + L ++ P + + +++++A SAI
Sbjct: 5 KYEVAKFNGDNGFSTWQRRMRDLLIQQGLHKVLDVDSKKPDTMKAEDWADLDERAASAIR 64
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVE 114
L L D V+ + E +W +L+SLYM+K+ T++ LK+QLY M E
Sbjct: 65 LHLSDDVVNNIIDEDTARGIWTRLESLYMSKTLTNKLYLKKQLYALHMSE 114
>UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas]
Length = 1298
Score = 73.2 bits (178), Expect = 1e-11
Identities = 43/120 (35%), Positives = 70/120 (57%), Gaps = 5/120 (4%)
Query: 2 MDSKWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEK-TEMNDKAI 60
M +K+ IEKF G N F LWK+K++ IL + C+ A+ ++ P T +K +EMN+ A+
Sbjct: 1 MAAKFEIEKFNGKN-FSLWKLKVKAILRKDNCLAAI---SERPVDFTDDKKWSEMNEDAM 56
Query: 61 SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTNNE 120
+ + L + D VL + + +W+ L+ LY KS ++ LK++LY RM ES + E
Sbjct: 57 ADLYLSIADGVLSSIEEKKTANEIWDHLNRLYEAKSLHNKIFLKRKLYTLRMSESTSVTE 116
>UniRef100_Q7XEC4 Putative polyprotein [Oryza sativa]
Length = 382
Score = 70.9 bits (172), Expect = 6e-11
Identities = 42/120 (35%), Positives = 68/120 (56%), Gaps = 2/120 (1%)
Query: 4 SKWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAI 63
SK+ + KF G+ +F LW+++++ +L QQ +AL+ MP + + EM +A + I
Sbjct: 7 SKFEVVKFDGTGNFVLWQMRLKDLLAQQGISKALQ--ETMPEKIDADKWNEMKAQAAATI 64
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTNNEATD 123
L L D V+ +V E +W+KL SL+M+KS T + LKQQLY ++ E + D
Sbjct: 65 RLSLSDSVMYQVMDEKSPKEIWDKLASLHMSKSLTSKLYLKQQLYGLQVQEESDLRKHVD 124
>UniRef100_Q6L4V3 Putative polyprotein [Oryza sativa]
Length = 1243
Score = 67.4 bits (163), Expect = 6e-10
Identities = 38/113 (33%), Positives = 64/113 (56%), Gaps = 1/113 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+++ F LW+VKMR +L QQ +AL G + + EK + + KAIS I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEK-KRDRKAISYIH 63
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKT 117
L L + +L+EV +E +W KL+ + MTK T + LKQ+L+ +++ + ++
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKLQDDES 116
>UniRef100_Q9SJU6 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 838
Score = 66.6 bits (161), Expect = 1e-09
Identities = 45/125 (36%), Positives = 65/125 (52%), Gaps = 12/125 (9%)
Query: 8 IEKFTGSNDFGLWKVKM--RIILI-------QQKCVEALKGKAQMPAHLTPAEKT---EM 55
+EK G D+ LWK K+ I L+ + + +E + A+ + LT E E
Sbjct: 8 VEKLDGEGDYVLWKEKLLAHIELLGLLEGLEEDEAIEEEESTAETDSLLTKTEDKVLKEK 67
Query: 56 NDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVES 115
KA S ++L LG+ VLR+V +E M LD L+M KS +R LKQ+LY Y+M +S
Sbjct: 68 RGKARSTVILSLGNHVLRKVIKEKTAAGMIRVLDKLFMAKSLPNRIYLKQRLYGYKMSDS 127
Query: 116 KTNNE 120
T E
Sbjct: 128 MTIEE 132
>UniRef100_Q7XTM9 OSJNBa0033G05.13 protein [Oryza sativa]
Length = 1181
Score = 66.2 bits (160), Expect = 1e-09
Identities = 37/108 (34%), Positives = 62/108 (57%), Gaps = 1/108 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+++ F LW+VKMR +L QQ+ +AL G + + EK + + KA+S I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQELDDALSGFDKRTQDWSNDEK-KRDRKAMSYIH 63
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
L L + +L+EV +E +W KL+ + MTK T + LKQ+L+ +++
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKL 111
>UniRef100_Q8LNW7 Putative polyprotein [Oryza sativa]
Length = 1280
Score = 65.5 bits (158), Expect = 2e-09
Identities = 37/108 (34%), Positives = 61/108 (56%), Gaps = 1/108 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+++ F LW+VKMR +L QQ +AL G + + EK + + KA+S I
Sbjct: 40 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEKKK-DRKAMSYIH 98
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
L L + +L+EV +E +W KL+ + MTK T + LKQ+L+ +++
Sbjct: 99 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKL 146
>UniRef100_Q9FFM0 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1342
Score = 63.9 bits (154), Expect = 7e-09
Identities = 44/136 (32%), Positives = 64/136 (46%), Gaps = 26/136 (19%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMND---------- 57
+EKF G D+ LWK K+ L + + L+G + + TE++D
Sbjct: 8 VEKFDGDGDYILWKEKL---LAHMEMLGLLEGLGEEEEAVVEDSTTEISDGGNQDPETAT 64
Query: 58 -------------KAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLK 104
KA S I+L LG+ VLR+V ++ M LD L+M KS +R LK
Sbjct: 65 SKLEDKILKEKRGKARSTIILSLGNNVLRKVIKQKTAAGMIKVLDQLFMAKSLPNRIYLK 124
Query: 105 QQLYFYRMVESKTNNE 120
Q+LY Y+M E+ T E
Sbjct: 125 QRLYGYKMSENMTMEE 140
>UniRef100_Q6L4X8 Putative polyprotein [Oryza sativa]
Length = 1211
Score = 63.9 bits (154), Expect = 7e-09
Identities = 36/108 (33%), Positives = 60/108 (55%), Gaps = 1/108 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIV 64
K+++ F LW+VKMR +L QQ +AL G + + EK + + K +S I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEK-KRDRKTMSYIH 63
Query: 65 LCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
L L + +L+EV +E +W KL+ + MTK T + LKQ+L+ +++
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKL 111
>UniRef100_Q9M1F5 Copia-like polyprotein [Arabidopsis thaliana]
Length = 1363
Score = 62.0 bits (149), Expect = 3e-08
Identities = 43/123 (34%), Positives = 63/123 (50%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKM--RIILIQQKCV----EALKGKAQMPAHLTPAEKTEMND---- 57
+EKF G D+ +WK K+ I ++ V E GK + EK E
Sbjct: 8 VEKFDGRGDYTMWKEKLLAHIDMLGLSAVLRESETPMGKERDSEKSDEDEKEEREKMEAF 67
Query: 58 -----KAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
KA S IVL + D+VLR++ +ET +M LD LYM+K+ +R LKQ+LY ++M
Sbjct: 68 EEKKRKARSTIVLSVSDRVLRKIKKETSAAAMLEALDRLYMSKALPNRIYLKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>UniRef100_Q9SZY0 Putative retrotransposon [Arabidopsis thaliana]
Length = 1230
Score = 61.6 bits (148), Expect = 3e-08
Identities = 42/121 (34%), Positives = 59/121 (48%), Gaps = 13/121 (10%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEK-------------TE 54
+EKF G D+ LWK K+ + AL+ + L E+ E
Sbjct: 8 MEKFDGHGDYTLWKEKLMAHMDLLGLTVALRETQSVSDPLESEEEGKESEKGDKEALMEE 67
Query: 55 MNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVE 114
KA S IVL + D+VLR+ +E SM LD LYM+K+ +R LKQ+LY Y+M E
Sbjct: 68 KRQKARSTIVLSVSDQVLRKSKKEKTAPSMLEALDKLYMSKALPNRIYLKQKLYSYKMQE 127
Query: 115 S 115
+
Sbjct: 128 N 128
>UniRef100_Q94LG0 Putative retroelement pol polyprotein [Oryza sativa]
Length = 1326
Score = 60.8 bits (146), Expect = 6e-08
Identities = 36/112 (32%), Positives = 62/112 (55%), Gaps = 1/112 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCV-EALKGKAQMPAHLTPAEKTEMNDKAISAI 63
K+++ F LW+VKMR IL Q + EAL+ + + AE+ + KA+ I
Sbjct: 5 KYDLPLLDYKTRFSLWQVKMRAILAQTSDLDEALESFGKKKSTEWTAEEKRKDRKALLLI 64
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVES 115
L L + +L+EV +E +W KL+S+ M+K T + +K +L+ +++ ES
Sbjct: 65 QLHLSNDILQEVLQEKTAAELWLKLESICMSKDLTSKMHIKMKLFSHKLQES 116
>UniRef100_Q75HA9 Putative polyprotein [Oryza sativa]
Length = 1322
Score = 60.5 bits (145), Expect = 8e-08
Identities = 35/112 (31%), Positives = 62/112 (55%), Gaps = 1/112 (0%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIILIQQKCV-EALKGKAQMPAHLTPAEKTEMNDKAISAI 63
K+++ F LW+VKMR +L Q + EAL+ + AE+ + KA+S I
Sbjct: 5 KYDLPLLDYKTRFSLWQVKMRAVLAQTSDLDEALESFGKKKTTEWTAEEKRKDRKALSLI 64
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVES 115
L L + +L+EV ++ +W KL+S+ M+K T + +K +L+ +++ ES
Sbjct: 65 QLHLSNDILQEVLQKKTAAELWLKLESICMSKDLTSKMHIKMKLFSHKLHES 116
>UniRef100_Q9SJT2 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1333
Score = 60.5 bits (145), Expect = 8e-08
Identities = 40/123 (32%), Positives = 62/123 (49%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDK--------- 58
+EKF G D+ +WK K+ L ALK + + + + TE +K
Sbjct: 8 VEKFDGRGDYTMWKEKLMAHLDILGLSVALKEEDDLVEKVAEMQLTEEEEKEEVLRRELL 67
Query: 59 ------AISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
A SAIVL + D+VLR++ +E +M LD LYM+K+ +R KQ+LY ++M
Sbjct: 68 EEKRRKARSAIVLSVTDRVLRKIKKEQSAAAMLGVLDKLYMSKALPNRIYQKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>UniRef100_Q8W5D4 Putative retrotransposon-related protein [Oryza sativa]
Length = 1229
Score = 60.5 bits (145), Expect = 8e-08
Identities = 34/100 (34%), Positives = 58/100 (58%), Gaps = 1/100 (1%)
Query: 17 FGLWKVKMRIILIQQKCV-EALKGKAQMPAHLTPAEKTEMNDKAISAIVLCLGDKVLREV 75
F LW+VKMR +L Q + EAL+ + AE+ + KA+S I L L + +L++V
Sbjct: 14 FSLWQVKMRAVLAQTSDLDEALESFGKKKTTEWTAEEKRKDRKALSLIQLHLSNDILQKV 73
Query: 76 ARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVES 115
+E +W KL+S+ M+K T + +K +L+ +++ ES
Sbjct: 74 LQEKTAAELWFKLESICMSKDLTSKMHIKMKLFSHKLQES 113
>UniRef100_Q9SH77 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1356
Score = 57.0 bits (136), Expect = 8e-07
Identities = 38/123 (30%), Positives = 64/123 (51%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKMRI---ILIQQKCVEALKGKAQMPAHLTPAEKT----------- 53
+EKF G D+ +WK K+ IL ++ + + + L +++
Sbjct: 8 VEKFDGRGDYTMWKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEKFEAL 67
Query: 54 -EMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
E KA SAIVL + D+VLR++ +E+ +M LD LYM+K+ +R KQ+LY ++M
Sbjct: 68 EEKKKKARSAIVLSVTDRVLRKIKKESTAAAMLLALDKLYMSKALPNRIYPKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>UniRef100_O82196 Copia-like retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1137
Score = 56.6 bits (135), Expect = 1e-06
Identities = 28/75 (37%), Positives = 42/75 (55%)
Query: 52 KTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYR 111
+ + ++KA+ I + +GDKVLR + W LD LY+ KS +R L+ ++Y YR
Sbjct: 37 RIDQDEKAMDMIFINVGDKVLRNIENSKTAAEAWATLDKLYLVKSLPNRVYLQLKVYNYR 96
Query: 112 MVESKTNNEATDGIQ 126
M +SKT E D Q
Sbjct: 97 MQDSKTLEENVDEFQ 111
>UniRef100_Q8H023 Putative retrovirus-related pol polyprotein [Oryza sativa]
Length = 556
Score = 55.8 bits (133), Expect = 2e-06
Identities = 37/106 (34%), Positives = 57/106 (52%), Gaps = 2/106 (1%)
Query: 18 GLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIVLCLGDKVLREVAR 77
G +++++ +L QQ +AL+ MP + + EM +A + I L L D V+ +V
Sbjct: 152 GSSQMRLKDLLAQQGISKALE--ETMPEKMDVGKWVEMKAQATAIIRLSLSDFVMYQVMD 209
Query: 78 ETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTNNEATD 123
E +W+KL SLYM+KS T + LKQQLY +M E + D
Sbjct: 210 EKTPKEIWDKLASLYMSKSLTSKLYLKQQLYGLQMQEESDLRKHVD 255
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.356 0.157 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 489,962,635
Number of Sequences: 2790947
Number of extensions: 15770847
Number of successful extensions: 70196
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 70023
Number of HSP's gapped (non-prelim): 168
length of query: 378
length of database: 848,049,833
effective HSP length: 129
effective length of query: 249
effective length of database: 488,017,670
effective search space: 121516399830
effective search space used: 121516399830
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC123575.7


