

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123571.7 - phase: 0
(247 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
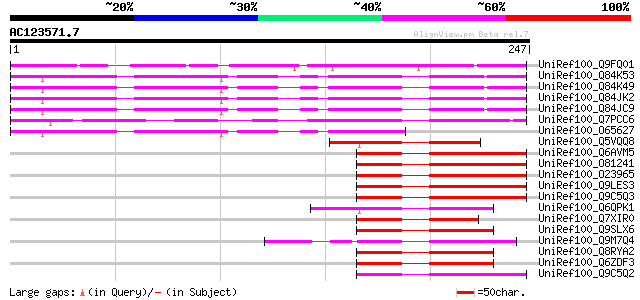
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FQ01 Basic leucine zipper transcription factor [Popu... 120 2e-26
UniRef100_Q84K53 BZIP transcription factor [Arabidopsis thaliana] 116 6e-25
UniRef100_Q84K49 BZIP transcription factor [Arabidopsis thaliana] 116 6e-25
UniRef100_Q84JK2 BZIP transcription factor [Arabidopsis thaliana] 116 6e-25
UniRef100_Q84JC9 BZIP transcription factor [Arabidopsis thaliana] 116 6e-25
UniRef100_Q7PCC6 Putative basic leucine zipper transcription fac... 100 3e-20
UniRef100_O65627 Putative bZIP transcription factor [Arabidopsis... 85 1e-15
UniRef100_Q5VQQ8 Putative bZIP protein DPBF3 [Oryza sativa] 70 5e-11
UniRef100_Q6AVM5 Putative ABA-responsive element-binding protein... 70 6e-11
UniRef100_O81241 Dc3 promoter-binding factor-3 [Helianthus annuus] 69 8e-11
UniRef100_O23965 Dc3 promoter-binding factor-2 [Helianthus annuus] 68 2e-10
UniRef100_Q9LES3 Promoter-binding factor-like protein [Arabidops... 67 3e-10
UniRef100_Q9C5Q3 BZIP protein DPBF3 [Arabidopsis thaliana] 67 3e-10
UniRef100_Q6QPK1 AREB-like protein [Lycopersicon esculentum] 65 2e-09
UniRef100_Q7XIR0 Putative bZIP protein DPBF3 [Oryza sativa] 64 3e-09
UniRef100_Q9SLX6 TRAB1 [Oryza sativa] 64 5e-09
UniRef100_Q9M7Q4 Abscisic acid responsive elements-binding facto... 64 5e-09
UniRef100_Q8RYA2 Raba1 [Oryza sativa] 64 5e-09
UniRef100_Q6ZDF3 TRAB1 [Oryza sativa] 64 5e-09
UniRef100_Q9C5Q2 BZIP protein DPBF4 [Arabidopsis thaliana] 63 6e-09
>UniRef100_Q9FQ01 Basic leucine zipper transcription factor [Populus balsamifera
subsp. trichocarpa x Populus deltoides]
Length = 301
Score = 120 bits (302), Expect = 2e-26
Identities = 97/266 (36%), Positives = 140/266 (52%), Gaps = 41/266 (15%)
Query: 1 MEEVWKDINLSSLNDQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKSLDYSSNNSS 60
MEEVW DINL+SL+D + +T + +F G ++ QDFL RP D S+
Sbjct: 54 MEEVWDDINLASLHDHSNTNTSSNTNHHSFNG-MVFQDFLARPSNKD----------TST 102
Query: 61 SSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYDPLIRHNKHNNSQLLLQQQQH 120
+ + + ++ + + S PPP T LSLNS D + + + +N+ + Q
Sbjct: 103 RAASKEPSSGGGNSFLKNSLGPPP-ATMLSLNSGSDHFH-----YLESSNTVPVRPNPQM 156
Query: 121 NIGVSNVSPCFVNA--SPCDQNVGVPASSSFTCF-GKRFGEAPDISPGERRNKRMIKNRE 177
+ + + F ++ SP D + +SS+F KR E D+S G+RR+KRMIKNRE
Sbjct: 157 HSHANGGTISFDSSLDSPFD---ALGSSSAFLSICKKRPQENGDVSGGDRRHKRMIKNRE 213
Query: 178 SAARSRARKQEKITSF-----------------LFSKFSAYTNELEQKVQLLQEENARLR 220
SAARSRARKQE + F L AYT ELE++ L +ENA+LR
Sbjct: 214 SAARSRARKQESSSPFENLFLVKFNDYRMLMFYLLLILQAYTVELEREAAHLAQENAKLR 273
Query: 221 RQQQELWEAESGGQQKKKSSLYRTSS 246
R QQE + A + Q KK++LYRTS+
Sbjct: 274 R-QQERFLAAAPAQLPKKNTLYRTST 298
>UniRef100_Q84K53 BZIP transcription factor [Arabidopsis thaliana]
Length = 285
Score = 116 bits (290), Expect = 6e-25
Identities = 96/259 (37%), Positives = 124/259 (47%), Gaps = 52/259 (20%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLNGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLCSSPFPPPATVLSLNSGAGFDFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELW 227
R+KRMIKNRESAARSRARKQ AYTNELE +V LQ ENARL+RQQ +L
Sbjct: 217 RHKRMIKNRESAARSRARKQ------------AYTNELELEVAHLQAENARLKRQQDQL- 263
Query: 228 EAESGGQQKKKSSLYRTSS 246
+ + QQ KK++L R+S+
Sbjct: 264 KMAAAIQQPKKNTLQRSST 282
>UniRef100_Q84K49 BZIP transcription factor [Arabidopsis thaliana]
Length = 285
Score = 116 bits (290), Expect = 6e-25
Identities = 96/259 (37%), Positives = 124/259 (47%), Gaps = 52/259 (20%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLNGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLCSSPFPPPATVLSLNSGAGFDFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELW 227
R+KRMIKNRESAARSRARKQ AYTNELE +V LQ ENARL+RQQ +L
Sbjct: 217 RHKRMIKNRESAARSRARKQ------------AYTNELELEVAHLQAENARLKRQQDQL- 263
Query: 228 EAESGGQQKKKSSLYRTSS 246
+ + QQ KK++L R+S+
Sbjct: 264 KMAAAIQQPKKNTLQRSST 282
>UniRef100_Q84JK2 BZIP transcription factor [Arabidopsis thaliana]
Length = 285
Score = 116 bits (290), Expect = 6e-25
Identities = 96/259 (37%), Positives = 124/259 (47%), Gaps = 52/259 (20%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLKGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLYSSPFPPPATVLSLNSGAGFEFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELW 227
R+KRMIKNRESAARSRARKQ AYTNELE +V LQ ENARL+RQQ +L
Sbjct: 217 RHKRMIKNRESAARSRARKQ------------AYTNELELEVAHLQAENARLKRQQDQL- 263
Query: 228 EAESGGQQKKKSSLYRTSS 246
+ + QQ KK++L R+S+
Sbjct: 264 KMAAAIQQPKKNTLQRSST 282
>UniRef100_Q84JC9 BZIP transcription factor [Arabidopsis thaliana]
Length = 285
Score = 116 bits (290), Expect = 6e-25
Identities = 96/259 (37%), Positives = 124/259 (47%), Gaps = 52/259 (20%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLKGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLCSSPFPPPATVLSLNSGAGFEFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELW 227
R+KRMIKNRESAARSRARKQ AYTNELE +V LQ ENARL+RQQ +L
Sbjct: 217 RHKRMIKNRESAARSRARKQ------------AYTNELELEVAHLQAENARLKRQQDQL- 263
Query: 228 EAESGGQQKKKSSLYRTSS 246
+ + QQ KK++L R+S+
Sbjct: 264 KMAAAIQQPKKNTLQRSST 282
>UniRef100_Q7PCC6 Putative basic leucine zipper transcription factor [Arabidopsis
thaliana]
Length = 234
Score = 100 bits (249), Expect = 3e-20
Identities = 89/251 (35%), Positives = 114/251 (44%), Gaps = 64/251 (25%)
Query: 1 MEEVWKDINLSSLNDQNT-----RPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKSLDYS 55
MEEVWK+INL SL+ PM+ + + I QDFL PL P S
Sbjct: 40 MEEVWKEINLGSLHYHRQLNIGHEPMLKNQNPNNS----IFQDFLNMPLNQPPPPPPPPS 95
Query: 56 SNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYDPLIRHNKHNNSQLLL 115
S+ +++ S PP T LSLNS F +
Sbjct: 96 SSTIVTALYG-------------SLPLPPPATVLSLNSGVGFEF---------------- 126
Query: 116 QQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKN 175
N+ SN S+ F C GK+ G+ D + G+RR KRMIKN
Sbjct: 127 LDTTENLLASNPR-------------SFEESAKFGCLGKKRGQDSDDTRGDRRYKRMIKN 173
Query: 176 RESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQ 235
RESAARSRARKQ AYTNELE ++ LQ ENARL+ QQ++L AE+ Q
Sbjct: 174 RESAARSRARKQ------------AYTNELELEIAHLQTENARLKIQQEQLKIAEATQNQ 221
Query: 236 KKKSSLYRTSS 246
KK +L R+S+
Sbjct: 222 VKK-TLQRSST 231
>UniRef100_O65627 Putative bZIP transcription factor [Arabidopsis thaliana]
Length = 244
Score = 85.1 bits (209), Expect = 1e-15
Identities = 71/201 (35%), Positives = 90/201 (44%), Gaps = 39/201 (19%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLKGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLYSSPFPPPATVLSLNSGAGFEFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQE 188
R+KRMIKNRESAARSRARKQE
Sbjct: 217 RHKRMIKNRESAARSRARKQE 237
>UniRef100_Q5VQQ8 Putative bZIP protein DPBF3 [Oryza sativa]
Length = 366
Score = 70.1 bits (170), Expect = 5e-11
Identities = 41/74 (55%), Positives = 50/74 (67%), Gaps = 14/74 (18%)
Query: 153 GKRFGEAPDISPG--ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQ 210
G++ G + D++ ERR KRMIKNRESAARSRARKQ AYTNELE KV
Sbjct: 249 GRKRGMSGDVADKLMERRQKRMIKNRESAARSRARKQ------------AYTNELENKVS 296
Query: 211 LLQEENARLRRQQQ 224
L+EEN RL+RQ++
Sbjct: 297 RLEEENVRLKRQKE 310
>UniRef100_Q6AVM5 Putative ABA-responsive element-binding protein 3 [Oryza sativa]
Length = 335
Score = 69.7 bits (169), Expect = 6e-11
Identities = 44/81 (54%), Positives = 50/81 (61%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYTNELE KV L+EEN RL++Q++
Sbjct: 264 ERRQKRMIKNRESAARSRARKQ------------AYTNELENKVLRLEEENERLKKQKEL 311
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S + K L RTSS
Sbjct: 312 DEILNSAPPPEPKYQLRRTSS 332
>UniRef100_O81241 Dc3 promoter-binding factor-3 [Helianthus annuus]
Length = 246
Score = 69.3 bits (168), Expect = 8e-11
Identities = 43/81 (53%), Positives = 49/81 (60%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE K+ L+EEN L+RQ++
Sbjct: 175 ERRQKRMIKNRESAARSRARKQ------------AYTHELENKISRLEEENELLKRQKEV 222
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K K L RTSS
Sbjct: 223 GMVLPSAPPPKPKYQLRRTSS 243
>UniRef100_O23965 Dc3 promoter-binding factor-2 [Helianthus annuus]
Length = 174
Score = 68.2 bits (165), Expect = 2e-10
Identities = 44/81 (54%), Positives = 49/81 (60%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN RLRR+++
Sbjct: 103 ERRQKRMIKNRESAARSRARKQ------------AYTHELENKVSRLEEENERLRREKEV 150
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
K K L RTSS
Sbjct: 151 EKVIPWVPPPKPKYQLRRTSS 171
>UniRef100_Q9LES3 Promoter-binding factor-like protein [Arabidopsis thaliana]
Length = 297
Score = 67.4 bits (163), Expect = 3e-10
Identities = 44/81 (54%), Positives = 49/81 (60%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN RLR+Q++
Sbjct: 226 ERRQKRMIKNRESAARSRARKQ------------AYTHELEIKVSRLEEENERLRKQKEV 273
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 274 EKILPSVPPPDPKRQLRRTSS 294
>UniRef100_Q9C5Q3 BZIP protein DPBF3 [Arabidopsis thaliana]
Length = 297
Score = 67.4 bits (163), Expect = 3e-10
Identities = 44/81 (54%), Positives = 49/81 (60%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN RLR+Q++
Sbjct: 226 ERRQKRMIKNRESAARSRARKQ------------AYTHELEIKVSRLEEENERLRKQKEV 273
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 274 EKILPSVPPPDPKRQLRRTSS 294
>UniRef100_Q6QPK1 AREB-like protein [Lycopersicon esculentum]
Length = 447
Score = 65.1 bits (157), Expect = 2e-09
Identities = 44/107 (41%), Positives = 55/107 (51%), Gaps = 32/107 (29%)
Query: 144 PASSSFTCFGKRFGEAPDISPG--------------------ERRNKRMIKNRESAARSR 183
P SS GK G+ P +SP ERR +RMIKNRESAARSR
Sbjct: 326 PGVSSSDGLGKSNGDTPSVSPVPYVFNGGLRGRKYSTVEKVVERRQRRMIKNRESAARSR 385
Query: 184 ARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAE 230
ARKQ AYT ELE +V L+EEN L+++Q+E+ E +
Sbjct: 386 ARKQ------------AYTMELEAEVAKLKEENDELQKKQEEMLEMQ 420
>UniRef100_Q7XIR0 Putative bZIP protein DPBF3 [Oryza sativa]
Length = 269
Score = 64.3 bits (155), Expect = 3e-09
Identities = 36/58 (62%), Positives = 39/58 (67%), Gaps = 12/58 (20%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQ 223
ERR KRMIKNRESAARSRARKQ AYTNELE K+ L+EEN RLR +
Sbjct: 177 ERRQKRMIKNRESAARSRARKQ------------AYTNELENKISRLEEENQRLREHK 222
>UniRef100_Q9SLX6 TRAB1 [Oryza sativa]
Length = 318
Score = 63.5 bits (153), Expect = 5e-09
Identities = 35/65 (53%), Positives = 45/65 (68%), Gaps = 12/65 (18%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +VQ L+E+N L+++Q+E
Sbjct: 231 ERRQRRMIKNRESAARSRARKQ------------AYTMELEAEVQKLKEQNMELQKKQEE 278
Query: 226 LWEAE 230
+ E +
Sbjct: 279 IMEMQ 283
>UniRef100_Q9M7Q4 Abscisic acid responsive elements-binding factor [Arabidopsis
thaliana]
Length = 416
Score = 63.5 bits (153), Expect = 5e-09
Identities = 47/120 (39%), Positives = 63/120 (52%), Gaps = 22/120 (18%)
Query: 122 IGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAAR 181
IG SN ++ SP N GV G++ G + ERR +RMIKNRESAAR
Sbjct: 303 IGKSNGDSSSLSPSPYMFNGGVR--------GRKSGTVEKVV--ERRQRRMIKNRESAAR 352
Query: 182 SRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQKKKSSL 241
SRARKQ AYT ELE +V L+EEN L+R+Q + E + + + ++ L
Sbjct: 353 SRARKQ------------AYTVELEAEVAKLKEENDELQRKQARIMEMQKNQETEMRNLL 400
>UniRef100_Q8RYA2 Raba1 [Oryza sativa]
Length = 90
Score = 63.5 bits (153), Expect = 5e-09
Identities = 35/65 (53%), Positives = 45/65 (68%), Gaps = 12/65 (18%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +VQ L+E+N L+++Q+E
Sbjct: 3 ERRQRRMIKNRESAARSRARKQ------------AYTMELEAEVQKLKEQNMELQKKQEE 50
Query: 226 LWEAE 230
+ E +
Sbjct: 51 IMEMQ 55
>UniRef100_Q6ZDF3 TRAB1 [Oryza sativa]
Length = 318
Score = 63.5 bits (153), Expect = 5e-09
Identities = 35/65 (53%), Positives = 45/65 (68%), Gaps = 12/65 (18%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +VQ L+E+N L+++Q+E
Sbjct: 231 ERRQRRMIKNRESAARSRARKQ------------AYTMELEAEVQKLKEQNMELQKKQEE 278
Query: 226 LWEAE 230
+ E +
Sbjct: 279 IMEMQ 283
>UniRef100_Q9C5Q2 BZIP protein DPBF4 [Arabidopsis thaliana]
Length = 262
Score = 63.2 bits (152), Expect = 6e-09
Identities = 42/81 (51%), Positives = 48/81 (58%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN +LRR ++
Sbjct: 191 ERRQKRMIKNRESAARSRARKQ------------AYTHELEIKVSRLEEENEKLRRLKEV 238
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RT+S
Sbjct: 239 EKILPSEPPPDPKWKLRRTNS 259
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 413,772,204
Number of Sequences: 2790947
Number of extensions: 17089119
Number of successful extensions: 78661
Number of sequences better than 10.0: 768
Number of HSP's better than 10.0 without gapping: 184
Number of HSP's successfully gapped in prelim test: 589
Number of HSP's that attempted gapping in prelim test: 77465
Number of HSP's gapped (non-prelim): 1339
length of query: 247
length of database: 848,049,833
effective HSP length: 124
effective length of query: 123
effective length of database: 501,972,405
effective search space: 61742605815
effective search space used: 61742605815
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC123571.7


