

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122729.6 + phase: 0 /pseudo
(247 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
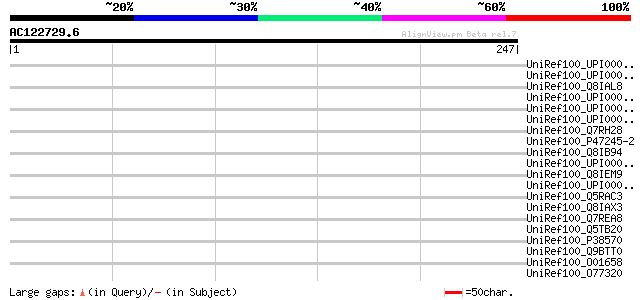
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI000029B1C4 UPI000029B1C4 UniRef100 entry 45 0.001
UniRef100_UPI000026C442 UPI000026C442 UniRef100 entry 44 0.003
UniRef100_Q8IAL8 Hypothetical protein MAL8P1.154 [Plasmodium fal... 44 0.003
UniRef100_UPI000046B7AB UPI000046B7AB UniRef100 entry 41 0.032
UniRef100_UPI00004998B5 UPI00004998B5 UniRef100 entry 40 0.042
UniRef100_UPI000046D02D UPI000046D02D UniRef100 entry 40 0.042
UniRef100_Q7RH28 Mature-parasite-infected erythrocyte surface an... 40 0.042
UniRef100_P47245-2 Splice isoform 2 of P47245 [Rattus norvegicus] 40 0.042
UniRef100_Q8IB94 Ubiquitin-protein ligase 1, putative [Plasmodiu... 40 0.054
UniRef100_UPI000036AABC UPI000036AABC UniRef100 entry 40 0.071
UniRef100_Q8IEM9 Hypothetical protein MAL13P1.41 [Plasmodium fal... 40 0.071
UniRef100_UPI000046678E UPI000046678E UniRef100 entry 39 0.093
UniRef100_Q5RAC3 Hypothetical protein DKFZp459A0729 [Pongo pygma... 39 0.093
UniRef100_Q8IAX3 Hypothetical protein PF08_0078 [Plasmodium falc... 39 0.093
UniRef100_Q7REA8 Hypothetical protein [Plasmodium yoelii yoelii] 39 0.093
UniRef100_Q5TB20 Acidic (Leucine-rich) nuclear phosphoprotein 32... 39 0.093
UniRef100_P38570 Integrin alpha-E precursor [Homo sapiens] 39 0.093
UniRef100_Q9BTT0 Acidic leucine-rich nuclear phosphoprotein 32 f... 39 0.093
UniRef100_O01658 Hypothetical protein T28F2.4 [Caenorhabditis el... 39 0.12
UniRef100_O77320 Hypothetical protein MAL3P3.3 [Plasmodium falci... 38 0.21
>UniRef100_UPI000029B1C4 UPI000029B1C4 UniRef100 entry
Length = 299
Score = 45.4 bits (106), Expect = 0.001
Identities = 26/75 (34%), Positives = 42/75 (55%), Gaps = 6/75 (8%)
Query: 30 IFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEK 89
+ EEE E+I+E FE+E E Y+EEY + EE ++ +++E L+++ E
Sbjct: 168 MLEEELEEIFEEFEEELEETEEYEEEYFAEETL----EEEAVEEVFEE--LEELEEERLA 221
Query: 90 EEHNEPIYDEEYVLA 104
EE E + DEE +A
Sbjct: 222 EEEVEEVLDEEIAVA 236
>UniRef100_UPI000026C442 UPI000026C442 UniRef100 entry
Length = 223
Score = 44.3 bits (103), Expect = 0.003
Identities = 32/104 (30%), Positives = 51/104 (48%), Gaps = 4/104 (3%)
Query: 31 FEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLD-DIHEAFEK 89
FE++ + + E FED+ E P+ DEE+ D E FD+E + DE D D E F+
Sbjct: 65 FEDDEDPLDEEFEDD--EDPL-DEEFEDEDEDEEFDDEDEDEEFDDEEFEDEDEDEEFDD 121
Query: 90 EEHNEPIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITK 133
E+ +E DEE+ + E + + + ++FL I K
Sbjct: 122 EDEDEEFDDEEFDDEDEDEEFDDEDEDEDEDLVELFLSKLFIVK 165
>UniRef100_Q8IAL8 Hypothetical protein MAL8P1.154 [Plasmodium falciparum]
Length = 2568
Score = 44.3 bits (103), Expect = 0.003
Identities = 26/102 (25%), Positives = 59/102 (57%), Gaps = 9/102 (8%)
Query: 24 NREINNIFEEEREDIYESFED-ETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDD 82
+ +IN ++E + E I + ++D E + +P D+E +I +++D+ I+ YD++ ++
Sbjct: 609 HEQINQLYEND-EQINQPYDDNEEINQPYDDDE----EINQLYDDHEEINQPYDDD--EE 661
Query: 83 IHEAFEKEEH-NEPIYDEEYVLAEYGEYLETKKRFQTSTNKD 123
I++ ++ +E N+P D+E + Y + E + + +TN D
Sbjct: 662 INQLYDNDEQINQPYDDDEEINQPYDDDEEQQNKINNNTNDD 703
Score = 41.6 bits (96), Expect = 0.019
Identities = 23/84 (27%), Positives = 51/84 (60%), Gaps = 7/84 (8%)
Query: 26 EINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHE 85
EIN I+E++ E+I + ++D+ IY+++ +I + +D++ I+ +YD++ + I++
Sbjct: 561 EINQIYEDD-EEINQPYDDDEEINQIYEDD---EEINQPYDDDEEINQLYDDH--EQINQ 614
Query: 86 AFEKEEH-NEPIYDEEYVLAEYGE 108
+E +E N+P D E + Y +
Sbjct: 615 LYENDEQINQPYDDNEEINQPYDD 638
>UniRef100_UPI000046B7AB UPI000046B7AB UniRef100 entry
Length = 1445
Score = 40.8 bits (94), Expect = 0.032
Identities = 36/152 (23%), Positives = 61/152 (39%), Gaps = 42/152 (27%)
Query: 21 TIVNREINNIFEEEREDIYESFE-DETLEKPIYDEEYVGADICEVF-------------- 65
T + E ++ ++ + IY FE DE L K Y +++ CE+
Sbjct: 867 TTKSHENKMLYTDDEDFIYNEFERDEFLMKNEYSKDFTNFSNCEILNADTLSYMKRDNNK 926
Query: 66 ---------DEEGNIDPIYDEN---------------GLDDIHEAFEKEEHNEPIYDEEY 101
D+ NID I EN L+++ EE+ E +D+++
Sbjct: 927 MITKKNDIKDKSNNIDSITTENVNGQSASLLKDMTLSSLNNVSNEIASEENEEKEHDQDF 986
Query: 102 VLAEYG-EYLETK--KRFQTSTNKDMFLMSSS 130
+ +EY YLE F +S K MF ++S+
Sbjct: 987 IASEYSFSYLENNDISLFSSSKIKKMFNLNST 1018
>UniRef100_UPI00004998B5 UPI00004998B5 UniRef100 entry
Length = 1416
Score = 40.4 bits (93), Expect = 0.042
Identities = 29/95 (30%), Positives = 47/95 (48%), Gaps = 8/95 (8%)
Query: 16 NYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIY 75
NYK ++ + N E+E E+ YE ++E E YDE+ ++ E D+E + Y
Sbjct: 790 NYKDTVVIQEDNTNYDEDENEEDYEYGDEEDEEVEYYDED---GNLIEPGDDE---EWEY 843
Query: 76 DENGLDD--IHEAFEKEEHNEPIYDEEYVLAEYGE 108
+E+ +++ E E + NEP D E E GE
Sbjct: 844 EEDEIEEEKPEETEENNKGNEPKEDNESNETENGE 878
>UniRef100_UPI000046D02D UPI000046D02D UniRef100 entry
Length = 324
Score = 40.4 bits (93), Expect = 0.042
Identities = 31/83 (37%), Positives = 41/83 (49%), Gaps = 14/83 (16%)
Query: 31 FEEEREDIYESFEDE-TLEKPIY------DEEYVGADICEVFDEEGNIDPIYDENGL--- 80
+ E+ ED E EDE + Y DEEY D E +EE D Y+E+ L
Sbjct: 218 YSEDDEDYSEEIEDEENSSEESYECISEEDEEYDAEDENEENEEEDMSDNSYEESDLEDG 277
Query: 81 --DDIHEAFEKEEHNEPIYDEEY 101
D+ E +EKEE ++ YDEEY
Sbjct: 278 YEDEFSEEYEKEEGSD--YDEEY 298
>UniRef100_Q7RH28 Mature-parasite-infected erythrocyte surface antigen [Plasmodium
yoelii yoelii]
Length = 1003
Score = 40.4 bits (93), Expect = 0.042
Identities = 32/103 (31%), Positives = 49/103 (47%), Gaps = 6/103 (5%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
EE+ E I E +DE +E+ DE + EV +EE D + +E DD E E+EE
Sbjct: 275 EEDAEVIEEEEDDEVIEEEEDDEVIEEEEDDEVIEEEEEDDEVIEEEEEDD--EVIEEEE 332
Query: 92 HNEPIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITKV 134
+E I +E+ E E E + + NK+ LM ++
Sbjct: 333 DDEIIGEED----EEDEMGEKESTQSVTNNKNEELMKDKENEI 371
Score = 34.3 bits (77), Expect = 3.0
Identities = 24/79 (30%), Positives = 42/79 (52%), Gaps = 2/79 (2%)
Query: 33 EEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEH 92
EE +++ E E+E E + +EE V + EV EE + + E ++++ E +E+ E
Sbjct: 183 EEGKEVIEEEEEEYEEVEVEEEEEVEDEYEEVEVEE-EYEEVEVEEEVEEVEEEYEEVEE 241
Query: 93 NEPIYDEEYVLAEYGEYLE 111
E + +EEY E E +E
Sbjct: 242 EEEV-EEEYEEVEEEEEVE 259
Score = 33.1 bits (74), Expect = 6.6
Identities = 25/69 (36%), Positives = 37/69 (53%), Gaps = 9/69 (13%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
EEE E++ E +E E E+ +EEY E +EE ++ ++E DD E E+EE
Sbjct: 226 EEEVEEVEEEYE-EVEEEEEVEEEY------EEVEEEEEVEEEHEEIEEDD--EVIEEEE 276
Query: 92 HNEPIYDEE 100
E I +EE
Sbjct: 277 DAEVIEEEE 285
>UniRef100_P47245-2 Splice isoform 2 of P47245 [Rattus norvegicus]
Length = 1229
Score = 40.4 bits (93), Expect = 0.042
Identities = 34/103 (33%), Positives = 52/103 (50%), Gaps = 14/103 (13%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
EEE E+ E E+E + D+E GA+I + DE G DD E F+ +E
Sbjct: 144 EEEEEEEGEEEEEEEEDDDDDDDEDSGAEIQDD-----------DEEGFDD-EEEFDDDE 191
Query: 92 HNEPIYD-EEYVLAEYGEYLET-KKRFQTSTNKDMFLMSSSIT 132
H++ D EE L E E +E KK + ++++FL+ S +T
Sbjct: 192 HDDDDLDNEENELEELEERVEARKKTTEKQQSQNLFLLWSKLT 234
>UniRef100_Q8IB94 Ubiquitin-protein ligase 1, putative [Plasmodium falciparum]
Length = 8591
Score = 40.0 bits (92), Expect = 0.054
Identities = 23/74 (31%), Positives = 39/74 (52%)
Query: 28 NNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAF 87
N +EE +DI +E+E + + +EE V + E DEE I+ DE D+ ++
Sbjct: 4923 NGESDEEDDDIRYEYEEEMDDDYMEEEEIVNQEDGEEEDEEEAIEGEDDEEAQDEENDEE 4982
Query: 88 EKEEHNEPIYDEEY 101
E NE ++++EY
Sbjct: 4983 EGNNENETLFNDEY 4996
>UniRef100_UPI000036AABC UPI000036AABC UniRef100 entry
Length = 1128
Score = 39.7 bits (91), Expect = 0.071
Identities = 33/134 (24%), Positives = 62/134 (45%), Gaps = 13/134 (9%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
E+E E+ E EDE +EE G +I V D G+IDP + D I
Sbjct: 130 EKEEEEDKEEEEDE-------EEEEAGTEIAIVLDGSGSIDPPDFQRAKDFISNMM--RN 180
Query: 92 HNEPIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITKVVKMLHQTI*KKS*SSE* 151
E ++ + L +YG ++T+ F ++D+ + + + ++ ++ K + + +
Sbjct: 181 FYEKCFECNFALVQYGGVIQTE--FDLRDSQDVMASLARVQNITQV--GSVTKTASAMQH 236
Query: 152 LITKIHTSCNGSTR 165
++ I TS +GS R
Sbjct: 237 VLDSIFTSSHGSRR 250
>UniRef100_Q8IEM9 Hypothetical protein MAL13P1.41 [Plasmodium falciparum]
Length = 885
Score = 39.7 bits (91), Expect = 0.071
Identities = 30/128 (23%), Positives = 58/128 (44%), Gaps = 26/128 (20%)
Query: 13 DCVNYKAVTIVNREINNIFE---EEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEG 69
D YK + N++INN+F E+ + + + + K I+DE+ + I + +
Sbjct: 497 DMFTYKLKDMDNKKINNLFNINANEKRTLIDIIYNNNVNKNIHDEKRIKKRIAYILYKNN 556
Query: 70 NIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQT------STNKD 123
NI HN+ I DEE+ L + +Y+ KK++ + TN+
Sbjct: 557 NI----------------YNSVHNDIIIDEEF-LNTFFKYVVNKKKYNSVCSFYFMTNES 599
Query: 124 MFLMSSSI 131
++L+ +S+
Sbjct: 600 LYLILNSL 607
>UniRef100_UPI000046678E UPI000046678E UniRef100 entry
Length = 219
Score = 39.3 bits (90), Expect = 0.093
Identities = 29/83 (34%), Positives = 43/83 (50%), Gaps = 4/83 (4%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
EEE++D YE +D+ + Y+EE G D E +++G D +E+ D E E EE
Sbjct: 74 EEEKDDDYEEGDDDGED---YEEEE-GDDYEEGGEDDGEGDDYEEEDDDGDYEEEDEGEE 129
Query: 92 HNEPIYDEEYVLAEYGEYLETKK 114
+ E EEY E GE E ++
Sbjct: 130 YEEDEEGEEYEEEEDGEDYEEEE 152
>UniRef100_Q5RAC3 Hypothetical protein DKFZp459A0729 [Pongo pygmaeus]
Length = 227
Score = 39.3 bits (90), Expect = 0.093
Identities = 27/85 (31%), Positives = 41/85 (47%), Gaps = 1/85 (1%)
Query: 16 NYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIY 75
N K++ + N EI N+ E+ RE I+E + T E+ D E DE+G+ D
Sbjct: 73 NLKSLDLFNCEITNL-EDYRESIFELLQQITYLDGFDQEDNEAPDSEEEDDEDGDEDDEE 131
Query: 76 DENGLDDIHEAFEKEEHNEPIYDEE 100
+E E +E+EE E DE+
Sbjct: 132 EEENEAGPPEGYEEEEEEEEEEDED 156
>UniRef100_Q8IAX3 Hypothetical protein PF08_0078 [Plasmodium falciparum]
Length = 1419
Score = 39.3 bits (90), Expect = 0.093
Identities = 35/109 (32%), Positives = 53/109 (48%), Gaps = 18/109 (16%)
Query: 26 EINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHE 85
E NN E + +++ E +E LE+ YDEE +D E +DEE + D YDE D ++
Sbjct: 139 EDNNYEESDEDELEEDEYEEELEEDEYDEE---SDQDE-YDEESDQDE-YDEESDQDEYD 193
Query: 86 AFEKEEHNEPIYDEEYVLAEYGEYLETK---------KRFQTSTNKDMF 125
EE ++ YDEE + E+L+ + K F N+D F
Sbjct: 194 ----EESDQDGYDEESNEDDLDEHLKERLMKRKSFYSKLFNIDDNEDKF 238
Score = 33.9 bits (76), Expect = 3.9
Identities = 24/72 (33%), Positives = 36/72 (49%), Gaps = 6/72 (8%)
Query: 39 YESFEDETLEKPIYDEEYVGA--DICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPI 96
Y+ E+ + KP YD+ + DI +E+ N Y+E+ D++ E +EE E
Sbjct: 109 YDEDEEGIIAKPYYDDYHRENRNDIQYDINEDNN----YEESDEDELEEDEYEEELEEDE 164
Query: 97 YDEEYVLAEYGE 108
YDEE EY E
Sbjct: 165 YDEESDQDEYDE 176
>UniRef100_Q7REA8 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 1705
Score = 39.3 bits (90), Expect = 0.093
Identities = 36/153 (23%), Positives = 61/153 (39%), Gaps = 43/153 (28%)
Query: 21 TIVNREINNIFEEEREDIYESFE-DETLEKPIYDEEYVGADICEVFDEE----------- 68
T + E ++ ++ + IY FE DE L K Y +++ CE+ + +
Sbjct: 1128 TTKSHEKKMLYTDDEDFIYNEFERDEFLMKNEYSKDFTNFSNCEILNPDTISYMKRDNNK 1187
Query: 69 -------------GNIDPIYDENG---------------LDDIHEAFEKEEHNEPIYDEE 100
NID I EN L+++ EE+ E +D++
Sbjct: 1188 MITKKNDIKDKSNNNIDSITTENANGQSASLLKDMTISSLNNVSNKIVSEENEEKEHDQD 1247
Query: 101 YVLAEYG-EYLETK--KRFQTSTNKDMFLMSSS 130
++ +EY YLE F +S K MF +SS+
Sbjct: 1248 FIASEYSFSYLENNDISLFSSSKIKKMFNLSST 1280
>UniRef100_Q5TB20 Acidic (Leucine-rich) nuclear phosphoprotein 32 family, member E
[Homo sapiens]
Length = 220
Score = 39.3 bits (90), Expect = 0.093
Identities = 27/85 (31%), Positives = 41/85 (47%), Gaps = 1/85 (1%)
Query: 16 NYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIY 75
N K++ + N EI N+ E+ RE I+E + T E+ D E DE+G+ D
Sbjct: 66 NLKSLDLFNCEITNL-EDYRESIFELLQQITYLDGFDQEDNEAPDSEEEDDEDGDEDDEE 124
Query: 76 DENGLDDIHEAFEKEEHNEPIYDEE 100
+E E +E+EE E DE+
Sbjct: 125 EEENEAGPPEGYEEEEEEEEEEDED 149
>UniRef100_P38570 Integrin alpha-E precursor [Homo sapiens]
Length = 1179
Score = 39.3 bits (90), Expect = 0.093
Identities = 32/134 (23%), Positives = 62/134 (45%), Gaps = 13/134 (9%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEE 91
E+E E+ E EDE +EE G +I + D G+IDP + D I
Sbjct: 181 EKEEEEDKEEEEDE-------EEEEAGTEIAIILDGSGSIDPPDFQRAKDFISNMM--RN 231
Query: 92 HNEPIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITKVVKMLHQTI*KKS*SSE* 151
E ++ + L +YG ++T+ F ++D+ + + + ++ ++ K + + +
Sbjct: 232 FYEKCFECNFALVQYGGVIQTE--FDLRDSQDVMASLARVQNITQV--GSVTKTASAMQH 287
Query: 152 LITKIHTSCNGSTR 165
++ I TS +GS R
Sbjct: 288 VLDSIFTSSHGSRR 301
>UniRef100_Q9BTT0 Acidic leucine-rich nuclear phosphoprotein 32 family member E [Homo
sapiens]
Length = 268
Score = 39.3 bits (90), Expect = 0.093
Identities = 27/85 (31%), Positives = 41/85 (47%), Gaps = 1/85 (1%)
Query: 16 NYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIY 75
N K++ + N EI N+ E+ RE I+E + T E+ D E DE+G+ D
Sbjct: 114 NLKSLDLFNCEITNL-EDYRESIFELLQQITYLDGFDQEDNEAPDSEEEDDEDGDEDDEE 172
Query: 76 DENGLDDIHEAFEKEEHNEPIYDEE 100
+E E +E+EE E DE+
Sbjct: 173 EEENEAGPPEGYEEEEEEEEEEDED 197
>UniRef100_O01658 Hypothetical protein T28F2.4 [Caenorhabditis elegans]
Length = 748
Score = 38.9 bits (89), Expect = 0.12
Identities = 28/78 (35%), Positives = 39/78 (49%), Gaps = 4/78 (5%)
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICE----VFDEEGNIDPIYDENGLDDIHEAF 87
E E ED + EDE + P E Y+ + E + DEE + DE D+ EA
Sbjct: 178 ELEDEDETDIDEDEMMIDPKDIERYINFESVEDEEDMEDEEIEDEEFEDEEFEDEEEEAD 237
Query: 88 EKEEHNEPIYDEEYVLAE 105
E+EE E + DEE V++E
Sbjct: 238 EQEEEEEDVSDEESVVSE 255
>UniRef100_O77320 Hypothetical protein MAL3P3.3 [Plasmodium falciparum]
Length = 3724
Score = 38.1 bits (87), Expect = 0.21
Identities = 30/82 (36%), Positives = 46/82 (55%), Gaps = 18/82 (21%)
Query: 22 IVNREINNIFEEEREDIYE----SFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDE 77
I + INNI++E +IY+ + DE++ IYDE I ++DE NI+ IYDE
Sbjct: 768 ICDNNINNIYDESINNIYDESINNIYDESINN-IYDE-----SINNIYDE--NINNIYDE 819
Query: 78 NGLDDIHEAFEKEEHNEPIYDE 99
N I+ +++ +N IYDE
Sbjct: 820 N----INNIYDENINN--IYDE 835
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.331 0.144 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 398,766,638
Number of Sequences: 2790947
Number of extensions: 16839885
Number of successful extensions: 93752
Number of sequences better than 10.0: 208
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 187
Number of HSP's that attempted gapping in prelim test: 92151
Number of HSP's gapped (non-prelim): 954
length of query: 247
length of database: 848,049,833
effective HSP length: 124
effective length of query: 123
effective length of database: 501,972,405
effective search space: 61742605815
effective search space used: 61742605815
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC122729.6


