

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122165.8 - phase: 1 /pseudo
(144 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
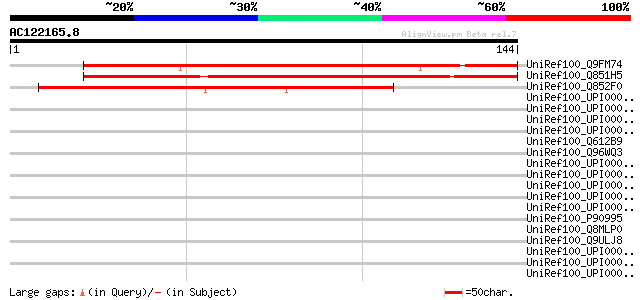
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FM74 Arabidopsis thaliana genomic DNA, chromosome 5,... 144 3e-34
UniRef100_Q851H5 Hypothetical protein OSJNBb0042N11.13 [Oryza sa... 144 3e-34
UniRef100_Q852F0 Hypothetical protein OJ1212_C05.17 [Oryza sativa] 95 4e-19
UniRef100_UPI000035F0E4 UPI000035F0E4 UniRef100 entry 38 0.054
UniRef100_UPI00002DA95D UPI00002DA95D UniRef100 entry 37 0.092
UniRef100_UPI000046BF91 UPI000046BF91 UniRef100 entry 37 0.12
UniRef100_UPI0000219509 UPI0000219509 UniRef100 entry 36 0.20
UniRef100_Q612B9 Hypothetical protein CBG16810 [Caenorhabditis b... 35 0.27
UniRef100_Q96WQ3 Ray38p [Candida maltosa] 35 0.27
UniRef100_UPI00003654B3 UPI00003654B3 UniRef100 entry 35 0.35
UniRef100_UPI0000361765 UPI0000361765 UniRef100 entry 35 0.35
UniRef100_UPI0000361764 UPI0000361764 UniRef100 entry 35 0.35
UniRef100_UPI0000361763 UPI0000361763 UniRef100 entry 35 0.35
UniRef100_UPI0000258295 UPI0000258295 UniRef100 entry 35 0.35
UniRef100_P90995 Hypothetical protein B0432.11 [Caenorhabditis e... 35 0.35
UniRef100_Q8MLP0 CG30165-PA [Drosophila melanogaster] 35 0.35
UniRef100_Q9ULJ8 Neurabin-I [Homo sapiens] 35 0.35
UniRef100_UPI0000365FFE UPI0000365FFE UniRef100 entry 35 0.46
UniRef100_UPI0000368249 UPI0000368249 UniRef100 entry 35 0.46
UniRef100_UPI0000452A0A UPI0000452A0A UniRef100 entry 35 0.46
>UniRef100_Q9FM74 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MDF20
[Arabidopsis thaliana]
Length = 149
Score = 144 bits (364), Expect = 3e-34
Identities = 76/127 (59%), Positives = 94/127 (73%), Gaps = 5/127 (3%)
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVE-SPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIP 80
DTDGFVIPSL I++ + NN + VE S S+ K+K EE IYLGPHGAPPSQ +
Sbjct: 20 DTDGFVIPSLEIEEDKVNNNNDSEVETSKPSSPKSKAEENIYLGPHGAPPSQLQDGGSNT 79
Query: 81 SNRKQRFKQKLKEADKRISGTGRENKVDNLRELVG---GEKTSVGMAKGSSPKDWLDPHC 137
S+RKQRFKQ LKEAD+++SG+GRENK+ NLRELVG G + M KG S +DWLDPHC
Sbjct: 80 SSRKQRFKQMLKEADQKMSGSGRENKMANLRELVGGGAGAEKGTNMGKGVS-RDWLDPHC 138
Query: 138 HEAEFER 144
HE++FE+
Sbjct: 139 HESQFEK 145
>UniRef100_Q851H5 Hypothetical protein OSJNBb0042N11.13 [Oryza sativa]
Length = 157
Score = 144 bits (364), Expect = 3e-34
Identities = 71/123 (57%), Positives = 89/123 (71%), Gaps = 3/123 (2%)
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS 81
D DGFVIP+L Q+ D +++ A + P + KEEKIYLGPHGAPPSQ KQQE+
Sbjct: 35 DADGFVIPNLTTQEDDVTEHTAPKPKDPEPLKE--KEEKIYLGPHGAPPSQAKQQEINIV 92
Query: 82 NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE 141
RKQRF+ KLKEAD + +G +ENKV+ LREL+G S G+ K SSP+DWLDPHCHE+E
Sbjct: 93 GRKQRFRNKLKEADNKFTGNAQENKVETLRELMGARTHSKGVPK-SSPRDWLDPHCHESE 151
Query: 142 FER 144
F+R
Sbjct: 152 FDR 154
>UniRef100_Q852F0 Hypothetical protein OJ1212_C05.17 [Oryza sativa]
Length = 172
Score = 94.7 bits (234), Expect = 4e-19
Identities = 53/103 (51%), Positives = 66/103 (63%), Gaps = 2/103 (1%)
Query: 9 VLKCMFFGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAK-TKKEEKIYLGPHG 67
VL+ MF + V D DGFVIPSL +++SD AA V P K TK EKIYLGPHG
Sbjct: 50 VLRKMFLHVRNVTDNDGFVIPSLSVEESDLGDWEAAQVSRPQPPPKATKDTEKIYLGPHG 109
Query: 68 APPSQPKQQE-VIPSNRKQRFKQKLKEADKRISGTGRENKVDN 109
APPS+ K+QE + R K K+KEAD+++ GTGR+NK N
Sbjct: 110 APPSRAKKQEDTAAAATGYRDKSKVKEADQKVLGTGRDNKGGN 152
>UniRef100_UPI000035F0E4 UPI000035F0E4 UniRef100 entry
Length = 1763
Score = 37.7 bits (86), Expect = 0.054
Identities = 35/122 (28%), Positives = 58/122 (46%), Gaps = 21/122 (17%)
Query: 24 DGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKI--------YLGPHGA------- 68
DGF P++G S +K SV N+ KT KE KI Y H +
Sbjct: 1349 DGFT-PNVGSDGSSLNKTADDSVLPENTEKKTHKEAKIENFVSVLDYQRNHPSVDRLASK 1407
Query: 69 PPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSS 128
PP+ P + PS+ ++ + ++E D ISG + +K ++ +L GG ++ ++ SS
Sbjct: 1408 PPTHPS---LSPSSLRKFMSKAVQELDIEISGGHQSDKPED--DLSGGSTPTLPLSCESS 1462
Query: 129 PK 130
P+
Sbjct: 1463 PR 1464
>UniRef100_UPI00002DA95D UPI00002DA95D UniRef100 entry
Length = 1372
Score = 37.0 bits (84), Expect = 0.092
Identities = 24/100 (24%), Positives = 42/100 (42%), Gaps = 5/100 (5%)
Query: 31 LGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQK 90
LG D D++ + +++E ++ + K+Y S P Q + S R
Sbjct: 717 LGSDDDDETNDEESALELRQFSSVAHRFSKVYSSTEHLSTSTPSNQSLSSSERSHS---- 772
Query: 91 LKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPK 130
+E ++R G G + D R L GGE+ + G P+
Sbjct: 773 -EEKEERWEGRGTLSPGDGYRHLSGGERRADGHRGSLRPR 811
>UniRef100_UPI000046BF91 UPI000046BF91 UniRef100 entry
Length = 509
Score = 36.6 bits (83), Expect = 0.12
Identities = 25/86 (29%), Positives = 40/86 (46%), Gaps = 1/86 (1%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA 94
DS+ KN V+S KKE I LG Q K++ I ++ + KQKL +
Sbjct: 178 DSENEKNKTKEVKSTFPIENNKKE-SILLGNQNKEDGQGKRKNDISLDKGKNKKQKLGKE 236
Query: 95 DKRISGTGRENKVDNLRELVGGEKTS 120
+K +G + K +N + + +TS
Sbjct: 237 NKNQNGINNKQKKNNGIDSMANNETS 262
>UniRef100_UPI0000219509 UPI0000219509 UniRef100 entry
Length = 1592
Score = 35.8 bits (81), Expect = 0.20
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 10/87 (11%)
Query: 45 SVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRE 104
S +P S+ K + + L PH PSQ QE++ K++ K+KLK + T RE
Sbjct: 1495 SQSNPGSSTKQSRPNRTGLNPHAQTPSQ-ATQEILDRKAKKKKKKKLKLKTE----TERE 1549
Query: 105 NKVDNLRELVGGEKTSVGMAKGSSPKD 131
++ V G GMA S+ +D
Sbjct: 1550 PRLPEWANKVSG-----GMASESTNRD 1571
>UniRef100_Q612B9 Hypothetical protein CBG16810 [Caenorhabditis briggsae]
Length = 823
Score = 35.4 bits (80), Expect = 0.27
Identities = 28/116 (24%), Positives = 52/116 (44%), Gaps = 15/116 (12%)
Query: 40 KNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA--DKR 97
K++ A +E +S++ + + + +PK++ +P R + FK + +E+ D
Sbjct: 45 KDHLAGIEEMSSSSSDEASDSDDV----MDSEEPKKELKLPKKRSKAFKIESEESSSDSY 100
Query: 98 ISGTGRENKVDNLREL---------VGGEKTSVGMAKGSSPKDWLDPHCHEAEFER 144
SG E++ D+ ++ GG K +KGS KD P C +A R
Sbjct: 101 ESGESEESENDDSEDVGGPGPSSKAFGGSKGKSSKSKGSDQKDAHWPKCSKASIAR 156
>UniRef100_Q96WQ3 Ray38p [Candida maltosa]
Length = 269
Score = 35.4 bits (80), Expect = 0.27
Identities = 32/115 (27%), Positives = 46/115 (39%), Gaps = 6/115 (5%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQ-----QEVIPSNRKQRFKQ 89
D ++KNN V +A K K K P + K+ ++ K++FKQ
Sbjct: 47 DPAKAKNNKKKVTGNEAALKNKTNNKEVAPPQSSAAKHTKKPFDRHSRTGKTDSKKKFKQ 106
Query: 90 KLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAEFER 144
E+DKR E D EL E + A S+PK L + E E +R
Sbjct: 107 GWGESDKRELEGEVEGAEDAEAEL-SAEASENADAADSTPKKTLQEYLAEMELKR 160
>UniRef100_UPI00003654B3 UPI00003654B3 UniRef100 entry
Length = 780
Score = 35.0 bits (79), Expect = 0.35
Identities = 22/78 (28%), Positives = 39/78 (49%), Gaps = 3/78 (3%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA 94
DSD S ++++S ++ S K +KE +PK ++ PS++K+ +QK E
Sbjct: 625 DSDSSSSSSSSSDTRRSKRKNRKERS---EEKKKKKEKPKTRKEKPSDKKRSRQQKAAET 681
Query: 95 DKRISGTGRENKVDNLRE 112
D GR ++ + RE
Sbjct: 682 DSDEEQHGRSDRRGHARE 699
>UniRef100_UPI0000361765 UPI0000361765 UniRef100 entry
Length = 908
Score = 35.0 bits (79), Expect = 0.35
Identities = 25/97 (25%), Positives = 43/97 (43%), Gaps = 6/97 (6%)
Query: 31 LGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQK 90
LG D D++ ++ +S+E ++ + K+Y S P Q + S R
Sbjct: 324 LGSDDDDETNDDESSLELRQFSSVAHRFSKVYSSTEHLSTSTPSNQSLSSSERSHS---- 379
Query: 91 LKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGS 127
+E ++R G G + D R L G++ + G KGS
Sbjct: 380 -EEKEERWEGRGILSPGDGYRHLSSGDRRADGQ-KGS 414
>UniRef100_UPI0000361764 UPI0000361764 UniRef100 entry
Length = 1263
Score = 35.0 bits (79), Expect = 0.35
Identities = 25/97 (25%), Positives = 43/97 (43%), Gaps = 6/97 (6%)
Query: 31 LGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQK 90
LG D D++ ++ +S+E ++ + K+Y S P Q + S R
Sbjct: 667 LGSDDDDETNDDESSLELRQFSSVAHRFSKVYSSTEHLSTSTPSNQSLSSSERSHS---- 722
Query: 91 LKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGS 127
+E ++R G G + D R L G++ + G KGS
Sbjct: 723 -EEKEERWEGRGILSPGDGYRHLSSGDRRADGQ-KGS 757
>UniRef100_UPI0000361763 UPI0000361763 UniRef100 entry
Length = 1259
Score = 35.0 bits (79), Expect = 0.35
Identities = 25/97 (25%), Positives = 43/97 (43%), Gaps = 6/97 (6%)
Query: 31 LGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQK 90
LG D D++ ++ +S+E ++ + K+Y S P Q + S R
Sbjct: 653 LGSDDDDETNDDESSLELRQFSSVAHRFSKVYSSTEHLSTSTPSNQSLSSSERSHS---- 708
Query: 91 LKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGS 127
+E ++R G G + D R L G++ + G KGS
Sbjct: 709 -EEKEERWEGRGILSPGDGYRHLSSGDRRADGQ-KGS 743
>UniRef100_UPI0000258295 UPI0000258295 UniRef100 entry
Length = 217
Score = 35.0 bits (79), Expect = 0.35
Identities = 26/96 (27%), Positives = 47/96 (48%), Gaps = 6/96 (6%)
Query: 47 ESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENK 106
E P K K+E I P +PP Q + S + ++F ++L +I G+ R +
Sbjct: 85 EKPLRENKVKEEASIKPDPAPSPPFQSTSGVKLASPKVRKFARELGANVTQIQGSQRSGR 144
Query: 107 V--DNLRELVGGEKTSVGMAKGS-SPKDWLDPHCHE 139
V +++++ V K+ V + +G PK + D + HE
Sbjct: 145 VTEEDVKKFV---KSQVSVNRGKVEPKKYKDEYSHE 177
>UniRef100_P90995 Hypothetical protein B0432.11 [Caenorhabditis elegans]
Length = 190
Score = 35.0 bits (79), Expect = 0.35
Identities = 26/88 (29%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Query: 47 ESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSN----RKQRFKQKLKEADKRISGTG 102
++ N+ ++TKK P AP P + I + + + K+ +KE + SGTG
Sbjct: 40 KTKNAGSRTKKSATSTSIPAPAPAPAPTEATTIAKDSLMGKLEPPKEPVKEPESDKSGTG 99
Query: 103 RENKVDNLRELVGGEKTSVGMAKGSSPK 130
N V LR K+ G KG PK
Sbjct: 100 PVNSVKWLRGRKKAWKSRKGSKKGKKPK 127
>UniRef100_Q8MLP0 CG30165-PA [Drosophila melanogaster]
Length = 469
Score = 35.0 bits (79), Expect = 0.35
Identities = 15/66 (22%), Positives = 36/66 (53%)
Query: 32 GIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
G +D + ++ +++ +T EE + L + PS+ +QQ+ P R++ ++++
Sbjct: 131 GKRDQSRERSTSSTARPKPPLRRTATEETLDLADEESTPSRQQQQQPPPQRRQESDRERV 190
Query: 92 KEADKR 97
+E D+R
Sbjct: 191 RERDQR 196
>UniRef100_Q9ULJ8 Neurabin-I [Homo sapiens]
Length = 1098
Score = 35.0 bits (79), Expect = 0.35
Identities = 30/104 (28%), Positives = 50/104 (47%), Gaps = 13/104 (12%)
Query: 28 IPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRF 87
IP IQ S + +++ ++ ++P+S K EE P + PK + IPS +
Sbjct: 292 IPGEEIQQSKEPEDSTSNQQTPDSIDKDGPEE-----PCAESKAMPKSE--IPSPQ---- 340
Query: 88 KQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKD 131
Q L++A+ + GRE +EL GG+ TS + S K+
Sbjct: 341 SQLLEDAEANL--VGREAAKQQRKELAGGDFTSPDASASSCGKE 382
>UniRef100_UPI0000365FFE UPI0000365FFE UniRef100 entry
Length = 1080
Score = 34.7 bits (78), Expect = 0.46
Identities = 28/105 (26%), Positives = 48/105 (45%), Gaps = 6/105 (5%)
Query: 24 DGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNR 83
+G I G++D D+ N+ +S + + K+E K +GAP + KQ E + +
Sbjct: 755 EGSGIAGTGLEDGDRPDENSDECKSDGQSNEEKREAKD--SKNGAPEEEEKQAE---NGK 809
Query: 84 KQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSS 128
K+ + K +E ++ T E D +E G E G +G S
Sbjct: 810 KEEERGKEQEGEREGDKTDSE-MGDADKEKDGKEGAEEGQREGES 853
>UniRef100_UPI0000368249 UPI0000368249 UniRef100 entry
Length = 931
Score = 34.7 bits (78), Expect = 0.46
Identities = 27/104 (25%), Positives = 47/104 (44%), Gaps = 7/104 (6%)
Query: 25 GFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGP---HGAPPSQPKQQEVIPS 81
GF I G+ D+ + N+ S +P AA K+ L P G P+ VI
Sbjct: 773 GFPIIFHGVMGKDEREGNSPSFFNPEEAATVTSYLKLLLAPSSKKGKARLSPRSVGVISP 832
Query: 82 NRKQ--RFKQKLKEADKRISGTG--RENKVDNLRELVGGEKTSV 121
RKQ + + + + D+ + G ++ KV ++ E G E++ +
Sbjct: 833 YRKQVEKIRYCITKLDRELRGLDDIKDLKVGSVEEFQGQERSVI 876
>UniRef100_UPI0000452A0A UPI0000452A0A UniRef100 entry
Length = 558
Score = 34.7 bits (78), Expect = 0.46
Identities = 19/55 (34%), Positives = 30/55 (54%)
Query: 75 QQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSP 129
+QE I +R+++ K K +E D R+ R + N+R L G + TS G S+P
Sbjct: 218 RQEEILQSREEKIKYKNEELDTRLREIKRMEHILNMRILEGEKNTSFGRNANSNP 272
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 253,995,786
Number of Sequences: 2790947
Number of extensions: 10532731
Number of successful extensions: 39780
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 39697
Number of HSP's gapped (non-prelim): 233
length of query: 144
length of database: 848,049,833
effective HSP length: 120
effective length of query: 24
effective length of database: 513,136,193
effective search space: 12315268632
effective search space used: 12315268632
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC122165.8


