

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122163.4 - phase: 0
(431 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
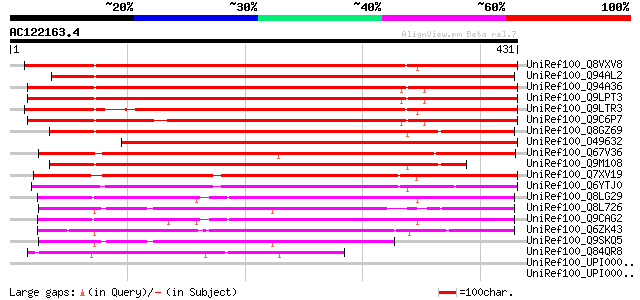
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VXV8 AT3g20300/MQC12_5 [Arabidopsis thaliana] 458 e-127
UniRef100_Q94AL2 Hypothetical protein At4g22270 [Arabidopsis tha... 443 e-123
UniRef100_Q94A36 At1g50630/F17J6_15 [Arabidopsis thaliana] 428 e-118
UniRef100_Q9LPT3 F11F12.5 protein [Arabidopsis thaliana] 428 e-118
UniRef100_Q9LTR3 Emb|CAA16777.1 [Arabidopsis thaliana] 425 e-117
UniRef100_Q9C6P7 Hypothetical protein F17J6.15 [Arabidopsis thal... 412 e-114
UniRef100_Q8GZ69 Hypothetical protein At4g03820/T7M24_8 [Arabido... 406 e-112
UniRef100_O49632 Hypothetical protein AT4g22270 [Arabidopsis tha... 379 e-103
UniRef100_Q67V36 Hypothetical protein OSJNBa0019I19.14 [Oryza sa... 344 2e-93
UniRef100_Q9M108 Hypothetical protein AT4g03820 [Arabidopsis tha... 339 1e-91
UniRef100_Q7XV19 OSJNBa0064H22.7 protein [Oryza sativa] 332 1e-89
UniRef100_Q6YTJ0 Hypothetical protein P0020D05.9 [Oryza sativa] 312 1e-83
UniRef100_Q8LG29 Hypothetical protein [Arabidopsis thaliana] 208 2e-52
UniRef100_Q8L726 Hypothetical protein At2g21080 [Arabidopsis tha... 204 3e-51
UniRef100_Q9CAG2 Hypothetical protein F12B7.12 [Arabidopsis thal... 194 4e-48
UniRef100_Q6ZK43 Hypothetical protein OJ1163_G08.36 [Oryza sativa] 193 7e-48
UniRef100_Q9SKQ5 Hypothetical protein At2g21080 [Arabidopsis tha... 165 3e-39
UniRef100_Q84QR8 Hypothetical protein OJ1081_B12.104 [Oryza sativa] 120 6e-26
UniRef100_UPI00003AC228 UPI00003AC228 UniRef100 entry 37 0.83
UniRef100_UPI00003AC227 UPI00003AC227 UniRef100 entry 37 0.83
>UniRef100_Q8VXV8 AT3g20300/MQC12_5 [Arabidopsis thaliana]
Length = 452
Score = 458 bits (1178), Expect = e-127
Identities = 235/424 (55%), Positives = 302/424 (70%), Gaps = 7/424 (1%)
Query: 13 NKENSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCST 72
NK + H EL FR YLRW+ VDQS+ A +SWS+F VVP SHF+L CS
Sbjct: 30 NKFTRSVSHAQDELHSFRKYLRWMCVDQSSPWTAVLSWSMFVVFTLVVPATSHFMLACSD 89
Query: 73 TCDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAI 132
CD+ H RPY VQ+SLS FA LSF+ LS++ KYG +FLF DK+ DES ++ GY
Sbjct: 90 -CDSHHSRPYDSVVQLSLSSFAALSFLCLSRFVSKYGLRRFLFFDKLWDESETVRLGYTN 148
Query: 133 QMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRT 192
Q+ R++K++ + PCF+ YKIWWY SGASQIP+ G V +S + C +ELCSW YRT
Sbjct: 149 QLNRSLKILSYFVSPCFLAMSSYKIWWYASGASQIPFLGNVILSDTVACLMELCSWLYRT 208
Query: 193 SIFFLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFIL 252
++ FLVCVLFRLI +LQI RL +FA VFQ +++VG+IL EHLRIRR+LR+ISHR+R FIL
Sbjct: 209 TVIFLVCVLFRLICHLQILRLQDFAQVFQMDSDVGSILSEHLRIRRHLRIISHRYRTFIL 268
Query: 253 SSLLLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSL 312
SL+LVT SQ LL+ K +A+++I + GEL L S+TLV+ L ILLRSA+KITHKAQ++
Sbjct: 269 LSLILVTGSQFYSLLITTKAYAELNIYRAGELALCSMTLVTALLILLRSASKITHKAQAV 328
Query: 313 TGLASKWHICATINSFDNIDGETPTALIASAQA---MAADISWGSSEDES-GDEEDELNN 368
T LA+KWH+CATI SF+ +DGETP L+ A D G S+ E GDEED+ +N
Sbjct: 329 TCLAAKWHVCATIESFETVDGETP-RLVDRASGHGYYPTDDDNGESDSEDYGDEEDDFDN 387
Query: 369 TKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNK 427
++P TISF KRQALV Y ENNR+GITV+GF LDR+ LH+IFGI+++L LWLL K
Sbjct: 388 NNLIPAYAYSTISFQKRQALVNYFENNRSGITVFGFTLDRSTLHTIFGIEMSLVLWLLGK 447
Query: 428 TVGI 431
T+GI
Sbjct: 448 TIGI 451
>UniRef100_Q94AL2 Hypothetical protein At4g22270 [Arabidopsis thaliana]
Length = 437
Score = 443 bits (1139), Expect = e-123
Identities = 216/395 (54%), Positives = 298/395 (74%), Gaps = 2/395 (0%)
Query: 36 VYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFAT 95
++ DQSN A +SWSVFF L +VP++SHFLL CS CD HRRPY V VQ+SLS+FA
Sbjct: 40 LWFDQSNFGTALLSWSVFFLLVVIVPLISHFLLVCSD-CDFHHRRPYDVIVQLSLSIFAG 98
Query: 96 LSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVY 155
+SF+SLS W RK+G +FLFLDK+ D S K++ Y +++R++K ++++ LP + Y
Sbjct: 99 ISFVSLSIWSRKFGMRRFLFLDKLWDVSDKVRIEYEAEIQRSLKRLMIFVLPSLTLEATY 158
Query: 156 KIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLDE 215
+IWWYISG +QIPY +S ++ CTL+L SW YR S+F +VC+L+++ +LQ RLD+
Sbjct: 159 RIWWYISGFNQIPYIINPILSHVVACTLQLSSWLYRNSLFIIVCILYKITCHLQTLRLDD 218
Query: 216 FAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPHA 274
FA F E T+V + L EH +IRRNLR++SHRFR FIL SL+LVTA+Q + LL +
Sbjct: 219 FARCFASEITDVRSALGEHQKIRRNLRIVSHRFRRFILLSLILVTATQFMALLTTTRASV 278
Query: 275 DVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNIDGE 334
V+I + GEL L S++LV+G++I LRSATKITHKAQS+T LA+KW++CAT++SFD++DGE
Sbjct: 279 AVNIYEVGELALCSLSLVTGVFICLRSATKITHKAQSVTSLAAKWNVCATVDSFDHLDGE 338
Query: 335 TPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALVTYMENN 394
TPT I +Q + +S+DE G+ +D+L+NTK+ PI TIS+ KRQALVTY+ENN
Sbjct: 339 TPTGSIIESQVSLRGNAIETSDDEEGEGDDDLDNTKIHPIYANTISYQKRQALVTYLENN 398
Query: 395 RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
+AGITVYGF++DR+WL++IFGI+LAL LWLLNKT+
Sbjct: 399 KAGITVYGFLVDRSWLNTIFGIELALLLWLLNKTI 433
>UniRef100_Q94A36 At1g50630/F17J6_15 [Arabidopsis thaliana]
Length = 453
Score = 428 bits (1100), Expect = e-118
Identities = 222/428 (51%), Positives = 295/428 (68%), Gaps = 14/428 (3%)
Query: 16 NSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCD 75
N + H EL FR YLRW+ VD S+ A +SW++F VVP +SHFLL C+ CD
Sbjct: 27 NRCVSHQQDELHSFRKYLRWMCVDHSSPWTAILSWTMFIVFTLVVPAISHFLLACAD-CD 85
Query: 76 ADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMK 135
+ H RPY VQ+SLS AT+SF+ L+++ KYG +FLF DK+ DES ++R Y Q+
Sbjct: 86 SYHSRPYDSVVQLSLSSVATVSFLCLTRFVSKYGLRRFLFFDKLWDESETVRRNYTNQLN 145
Query: 136 RTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIF 195
++ ++ + +PCF YKIWWY SG S+IP+ G +S + C +ELCSW YRT++
Sbjct: 146 TSLHIVSYFVIPCFSAMSAYKIWWYASGGSRIPFLGNAVLSDTVACIMELCSWLYRTTVI 205
Query: 196 FLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSL 255
FLVCVLFRLI +LQI RL +FA +FQ +++VG+IL EHLRIRR+LR+ISHR+R+FIL L
Sbjct: 206 FLVCVLFRLICHLQILRLQDFAKLFQIDSDVGSILSEHLRIRRHLRIISHRYRSFILCLL 265
Query: 256 LLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGL 315
+LVT SQ LL+ K + +V+I + GEL L S+TLV+ L ILLRSA+KITHKAQ++T L
Sbjct: 266 ILVTGSQFSSLLITTKAYTEVNIYRAGELALCSMTLVTALLILLRSASKITHKAQAVTCL 325
Query: 316 ASKWHICATINSFDNI------DGETPTALIASAQAMAADIS-----WGSSEDESGDEED 364
A+KWH+CAT+ SFD ETPT L+A ++ S DE GDEED
Sbjct: 326 AAKWHVCATLESFDQTVVSFDQTVETPT-LVARNNNDNNNVHDVVTLTESDSDEYGDEED 384
Query: 365 ELNNTKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLW 423
+L+N ++P+ T+SF KRQALV+Y ENN AGITVYGF LDR LH+IFG++L+L LW
Sbjct: 385 DLDNNDIIPVYAFSTMSFQKRQALVSYFENNSAGITVYGFTLDRGTLHTIFGLELSLVLW 444
Query: 424 LLNKTVGI 431
LL KT+GI
Sbjct: 445 LLGKTIGI 452
>UniRef100_Q9LPT3 F11F12.5 protein [Arabidopsis thaliana]
Length = 453
Score = 428 bits (1100), Expect = e-118
Identities = 222/428 (51%), Positives = 295/428 (68%), Gaps = 14/428 (3%)
Query: 16 NSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCD 75
N + H EL FR YLRW+ VD S+ A +SW++F VVP +SHFLL C+ CD
Sbjct: 27 NRCVSHQQDELHSFRKYLRWMCVDHSSPWTAILSWTMFIVFTLVVPAISHFLLACAD-CD 85
Query: 76 ADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMK 135
+ H RPY VQ+SLS AT+SF+ L+++ KYG +FLF DK+ DES ++R Y Q+
Sbjct: 86 SYHSRPYDSVVQLSLSSVATVSFLCLTRFVSKYGLRRFLFFDKLWDESETVRRNYTNQLN 145
Query: 136 RTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIF 195
++ ++ + +PCF YKIWWY SG S+IP+ G +S + C +ELCSW YRT++
Sbjct: 146 TSLHIVSYFVIPCFSAMSAYKIWWYASGGSRIPFLGNAVLSDTVACIMELCSWLYRTTVI 205
Query: 196 FLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSL 255
FLVCVLFRLI +LQI RL +FA +FQ +++VG+IL EHLRIRR+LR+ISHR+R+FIL L
Sbjct: 206 FLVCVLFRLICHLQILRLQDFAKLFQIDSDVGSILSEHLRIRRHLRIISHRYRSFILCLL 265
Query: 256 LLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGL 315
+LVT SQ LL+ K + +V+I + GEL L S+TLV+ L ILLRSA+KITHKAQ++T L
Sbjct: 266 ILVTGSQFSSLLITTKAYTEVNIYRAGELALCSMTLVTALLILLRSASKITHKAQAVTCL 325
Query: 316 ASKWHICATINSFDNI------DGETPTALIASAQAMAADIS-----WGSSEDESGDEED 364
A+KWH+CAT+ SFD ETPT L+A ++ S DE GDEED
Sbjct: 326 AAKWHVCATLESFDQTVESFDQTVETPT-LVARNNNDNNNVHDVVTLTESDSDEYGDEED 384
Query: 365 ELNNTKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLW 423
+L+N ++P+ T+SF KRQALV+Y ENN AGITVYGF LDR LH+IFG++L+L LW
Sbjct: 385 DLDNNDIIPVYAFSTMSFQKRQALVSYFENNSAGITVYGFTLDRGTLHTIFGLELSLVLW 444
Query: 424 LLNKTVGI 431
LL KT+GI
Sbjct: 445 LLGKTIGI 452
>UniRef100_Q9LTR3 Emb|CAA16777.1 [Arabidopsis thaliana]
Length = 430
Score = 425 bits (1092), Expect = e-117
Identities = 224/424 (52%), Positives = 288/424 (67%), Gaps = 29/424 (6%)
Query: 13 NKENSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCST 72
NK + H EL FR YLRW+ VDQS+ A +SWS+F VVP SHF+L CS
Sbjct: 30 NKFTRSVSHAQDELHSFRKYLRWMCVDQSSPWTAVLSWSMFVVFTLVVPATSHFMLACSD 89
Query: 73 TCDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAI 132
CD+ H RP F+S KYG +FLF DK+ DES ++ GY
Sbjct: 90 -CDSHHSRP----------------FVS------KYGLRRFLFFDKLWDESETVRLGYTN 126
Query: 133 QMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRT 192
Q+ R++K++ + PCF+ YKIWWY SGASQIP+ G V +S + C +ELCSW YRT
Sbjct: 127 QLNRSLKILSYFVSPCFLAMSSYKIWWYASGASQIPFLGNVILSDTVACLMELCSWLYRT 186
Query: 193 SIFFLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFIL 252
++ FLVCVLFRLI +LQI RL +FA VFQ +++VG+IL EHLRIRR+LR+ISHR+R FIL
Sbjct: 187 TVIFLVCVLFRLICHLQILRLQDFAQVFQMDSDVGSILSEHLRIRRHLRIISHRYRTFIL 246
Query: 253 SSLLLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSL 312
SL+LVT SQ LL+ K +A+++I + GEL L S+TLV+ L ILLRSA+KITHKAQ++
Sbjct: 247 LSLILVTGSQFYSLLITTKAYAELNIYRAGELALCSMTLVTALLILLRSASKITHKAQAV 306
Query: 313 TGLASKWHICATINSFDNIDGETPTALIASAQA---MAADISWGSSEDES-GDEEDELNN 368
T LA+KWH+CATI SF+ +DGETP L+ A D G S+ E GDEED+ +N
Sbjct: 307 TCLAAKWHVCATIESFETVDGETP-RLVDRASGHGYYPTDDDNGESDSEDYGDEEDDFDN 365
Query: 369 TKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNK 427
++P TISF KRQALV Y ENNR+GITV+GF LDR+ LH+IFGI+++L LWLL K
Sbjct: 366 NNLIPAYAYSTISFQKRQALVNYFENNRSGITVFGFTLDRSTLHTIFGIEMSLVLWLLGK 425
Query: 428 TVGI 431
T+GI
Sbjct: 426 TIGI 429
>UniRef100_Q9C6P7 Hypothetical protein F17J6.15 [Arabidopsis thaliana]
Length = 443
Score = 412 bits (1060), Expect = e-114
Identities = 219/428 (51%), Positives = 289/428 (67%), Gaps = 24/428 (5%)
Query: 16 NSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCD 75
N + H EL FR YLRW+ VD S+ A +SW++F VVP +SHFLL C+ CD
Sbjct: 27 NRCVSHQQDELHSFRKYLRWMCVDHSSPWTAILSWTMFIVFTLVVPAISHFLLACAD-CD 85
Query: 76 ADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMK 135
+ H RPY VQ+SLS AT+SF+ L+++ KYG +FLF DK+ DES
Sbjct: 86 SYHSRPYDSVVQLSLSSVATVSFLCLTRFVSKYGLRRFLFFDKLWDES----------ET 135
Query: 136 RTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIF 195
++ ++ + +PCF YKIWWY SG S+IP+ G +S + C +ELCSW YRT++
Sbjct: 136 TSLHIVSYFVIPCFSAMSAYKIWWYASGGSRIPFLGNAVLSDTVACIMELCSWLYRTTVI 195
Query: 196 FLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSL 255
FLVCVLFRLI +LQI RL +FA +FQ +++VG+IL EHLRIRR+LR+ISHR+R+FIL L
Sbjct: 196 FLVCVLFRLICHLQILRLQDFAKLFQIDSDVGSILSEHLRIRRHLRIISHRYRSFILCLL 255
Query: 256 LLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGL 315
+LVT SQ LL+ K + +V+I + GEL L S+TLV+ L ILLRSA+KITHKAQ++T L
Sbjct: 256 ILVTGSQFSSLLITTKAYTEVNIYRAGELALCSMTLVTALLILLRSASKITHKAQAVTCL 315
Query: 316 ASKWHICATINSFDNI------DGETPTALIASAQAMAADIS-----WGSSEDESGDEED 364
A+KWH+CAT+ SFD ETPT L+A ++ S DE GDEED
Sbjct: 316 AAKWHVCATLESFDQTVESFDQTVETPT-LVARNNNDNNNVHDVVTLTESDSDEYGDEED 374
Query: 365 ELNNTKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLW 423
+L+N ++P+ T+SF KRQALV+Y ENN AGITVYGF LDR LH+IFG++L+L LW
Sbjct: 375 DLDNNDIIPVYAFSTMSFQKRQALVSYFENNSAGITVYGFTLDRGTLHTIFGLELSLVLW 434
Query: 424 LLNKTVGI 431
LL KT+GI
Sbjct: 435 LLGKTIGI 442
>UniRef100_Q8GZ69 Hypothetical protein At4g03820/T7M24_8 [Arabidopsis thaliana]
Length = 437
Score = 406 bits (1044), Expect = e-112
Identities = 204/401 (50%), Positives = 287/401 (70%), Gaps = 9/401 (2%)
Query: 35 WVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFA 94
+++ DQSN K +SWS+FF LA +VP++SHF+L C+ CD HRRPY VQ+SLS+FA
Sbjct: 36 FLWFDQSNRIKTLLSWSIFFLLAVIVPMISHFVLICAD-CDFKHRRPYDGLVQLSLSIFA 94
Query: 95 TLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCV 154
+SF+SLS W +KYG +FLF DK+ D S K++ GY ++++R+MKL+ ++ LP Q +
Sbjct: 95 GISFVSLSDWSKKYGIRRFLFFDKLKDVSDKVRIGYEVKIQRSMKLLAIFVLPSTTLQAI 154
Query: 155 YKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLD 214
Y+IWWY SG +QIPY +S ++ CTL+L SW YRTS+F + C+L++ I +LQ+ RLD
Sbjct: 155 YRIWWYASGFNQIPYIINPTLSHVLACTLQLSSWLYRTSLFIIACILYQNICHLQVLRLD 214
Query: 215 EFAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPH 273
EFA F E + +IL EHL+IRR L+++SHRFR FIL SL VTA+Q + LL I+
Sbjct: 215 EFARCFASEIKDFSSILAEHLKIRRELKIVSHRFRRFILLSLFFVTATQFMALLTTIRAS 274
Query: 274 ADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNI-D 332
+I + GEL L S +LVSGL+I L+SAT++THKAQS+T +A+KW++CA++++FD + D
Sbjct: 275 VPFNIYEVGELALCSTSLVSGLFICLKSATQMTHKAQSVTSIATKWNVCASLDTFDVLYD 334
Query: 333 GETP----TALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALV 388
GETP T + + ++ S +DE G+ +D N+ ++ PI R IS KRQALV
Sbjct: 335 GETPKCPTTTQHSQILSRRRNVVQSSDDDEEGEGDD--NDLEIHPIFARAISSQKRQALV 392
Query: 389 TYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
TY+ENNRAGITVYGF++D+TWL IF I+LAL LWLL KT+
Sbjct: 393 TYLENNRAGITVYGFLVDKTWLRMIFSIELALLLWLLKKTI 433
>UniRef100_O49632 Hypothetical protein AT4g22270 [Arabidopsis thaliana]
Length = 387
Score = 379 bits (972), Expect = e-103
Identities = 182/337 (54%), Positives = 256/337 (75%), Gaps = 1/337 (0%)
Query: 96 LSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVY 155
+SF+SLS W RK+G +FLFLDK+ D S K++ Y +++R++K ++++ LP + Y
Sbjct: 49 ISFVSLSIWSRKFGMRRFLFLDKLWDVSDKVRIEYEAEIQRSLKRLMIFVLPSLTLEATY 108
Query: 156 KIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLDE 215
+IWWYISG +QIPY +S ++ CTL+L SW YR S+F +VC+L+++ +LQ RLD+
Sbjct: 109 RIWWYISGFNQIPYIINPILSHVVACTLQLSSWLYRNSLFIIVCILYKITCHLQTLRLDD 168
Query: 216 FAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPHA 274
FA F E T+V + L EH +IRRNLR++SHRFR FIL SL+LVTA+Q + LL +
Sbjct: 169 FARCFASEITDVRSALGEHQKIRRNLRIVSHRFRRFILLSLILVTATQFMALLTTTRASV 228
Query: 275 DVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNIDGE 334
V+I + GEL L S++LV+G++I LRSATKITHKAQS+T LA+KW++CAT++SFD++DGE
Sbjct: 229 AVNIYEVGELALCSLSLVTGVFICLRSATKITHKAQSVTSLAAKWNVCATVDSFDHLDGE 288
Query: 335 TPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALVTYMENN 394
TPT I +Q + +S+DE G+ +D+L+NTK+ PI TIS+ KRQALVTY+ENN
Sbjct: 289 TPTGSIIESQVSLRGNAIETSDDEEGEGDDDLDNTKIHPIYANTISYQKRQALVTYLENN 348
Query: 395 RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI 431
+AGITVYGF++DR+WL++IFGI+LAL LWLLNKT+GI
Sbjct: 349 KAGITVYGFLVDRSWLNTIFGIELALLLWLLNKTIGI 385
>UniRef100_Q67V36 Hypothetical protein OSJNBa0019I19.14 [Oryza sativa]
Length = 452
Score = 344 bits (883), Expect = 2e-93
Identities = 182/411 (44%), Positives = 257/411 (62%), Gaps = 11/411 (2%)
Query: 25 ELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHV 84
EL+ FR+ LRWV +D S A +SW +F LA VP +HFLL + RRP+
Sbjct: 46 ELQWFRSCLRWVCMDHSGPWGAALSWLLFLLLAVAVPAAAHFLLAFRAS-----RRPFSA 100
Query: 85 PVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLW 144
VQ+SLS + F+ LS R+ G + L+LDK+ +S +++ Y ++ + +L+
Sbjct: 101 VVQVSLSAASAAGFLCLSSSFRRIGLRRLLYLDKLRTKSDRVRLNYTARLSFSFRLLASL 160
Query: 145 GLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRL 204
PCF + YK+WWY + ++P++G +S + C++E+ +W YR++I+ L CVLFRL
Sbjct: 161 VAPCFAAEAAYKVWWYATSGDRVPFFGNDVLSNAVACSVEMAAWMYRSAIYLLTCVLFRL 220
Query: 205 IGYLQIQRLDEFAPVFQRETEVG-----TILLEHLRIRRNLRVISHRFRAFILSSLLLVT 259
I +LQ RL++FA E E G +L EHL IR+ L+VISHRFR FI++SLL+ T
Sbjct: 221 ICHLQGLRLEDFAGTLLVEVEEGRAGVERVLREHLDIRKQLKVISHRFRKFIVASLLIAT 280
Query: 260 ASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKW 319
ASQ LL+ + + D+L EL L S+ L+SGL I+L SA KITH+AQ+LTG +KW
Sbjct: 281 ASQFASLLLTTRHDSVDDLLNTSELALCSVVLMSGLIIILSSAAKITHQAQALTGQTTKW 340
Query: 320 HICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTI 379
H C TI + + E + + + S G S +E+GD ED L NTK+M Q I
Sbjct: 341 HACCTIEPVPDEEAEPGSNHSSMLEVEPVSDSDGESSEETGD-EDLLENTKIMLPQAHVI 399
Query: 380 SFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVG 430
SF KRQALVTY+ENNRAGITV+GF LDR++LH+IF ++ L LWLL KT+G
Sbjct: 400 SFQKRQALVTYLENNRAGITVFGFTLDRSYLHTIFMLEWTLFLWLLGKTIG 450
>UniRef100_Q9M108 Hypothetical protein AT4g03820 [Arabidopsis thaliana]
Length = 528
Score = 339 bits (869), Expect = 1e-91
Identities = 174/360 (48%), Positives = 250/360 (69%), Gaps = 9/360 (2%)
Query: 35 WVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFA 94
+++ DQSN K +SWS+FF LA +VP++SHF+L C+ CD HRRPY VQ+SLS+FA
Sbjct: 169 FLWFDQSNRIKTLLSWSIFFLLAVIVPMISHFVLICAD-CDFKHRRPYDGLVQLSLSIFA 227
Query: 95 TLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCV 154
+SF+SLS W +KYG +FLF DK+ D S K++ GY +++R+MKL+ ++ LP Q +
Sbjct: 228 GISFVSLSDWSKKYGIRRFLFFDKLKDVSDKVRIGYEAKIQRSMKLLAIFVLPSTTLQAI 287
Query: 155 YKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLD 214
Y+IWWY SG +QIPY +S ++ CTL+L SW YRTS+F + C+L++ I +LQ+ RLD
Sbjct: 288 YRIWWYASGFNQIPYIINPTLSHVLACTLQLSSWLYRTSLFIIACILYQNICHLQVLRLD 347
Query: 215 EFAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPH 273
EFA F E + +IL EHL+IRR L+++SHRFR FIL SL VTA+Q + LL I+
Sbjct: 348 EFARCFASEIKDFSSILAEHLKIRRELKIVSHRFRRFILLSLFFVTATQFMALLTTIRAS 407
Query: 274 ADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNI-D 332
+I + GEL L S +LVSGL+I L+SAT++THKAQS+T +A+KW++CA++++FD + D
Sbjct: 408 VPFNIYEVGELALCSTSLVSGLFICLKSATQMTHKAQSVTSIATKWNVCASLDTFDVLYD 467
Query: 333 GETP----TALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALV 388
GETP T + + ++ S +DE G+ +D N+ ++ PI R IS KRQALV
Sbjct: 468 GETPKCPTTTQHSQILSRRRNVVQSSDDDEEGEGDD--NDLEIHPIFARAISSQKRQALV 525
>UniRef100_Q7XV19 OSJNBa0064H22.7 protein [Oryza sativa]
Length = 442
Score = 332 bits (852), Expect = 1e-89
Identities = 183/423 (43%), Positives = 257/423 (60%), Gaps = 28/423 (6%)
Query: 21 HDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRR 80
H EL FR+ LRW+ V+ S+ A SW VF LA VP + L RR
Sbjct: 35 HARDELGSFRSCLRWMCVEHSDGSSAVASWLVFTLLAVAVPAAARAALP---------RR 85
Query: 81 PYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKL 140
Y VQ SL++ A L++++LS+ R+ G + L+LD++ +S ++ GY +++ + +L
Sbjct: 86 AYDGQVQASLTLSAALAYLTLSRLVRRRGLRRLLYLDRLRHDSQDVRAGYTVELAGSFRL 145
Query: 141 ILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCV 200
+ + LPCF+ YK++WY + ++ C LE+ SW YRT++FF+ CV
Sbjct: 146 LACFVLPCFLADAAYKVYWYCANRPFPLWWSAA------ACALEMASWMYRTAMFFMACV 199
Query: 201 LFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTA 260
LFR+I +LQI R+ FA F + +V +L +H RIR LR ISHR+R FI+S LLLVTA
Sbjct: 200 LFRIICFLQILRMTGFARDFGQCADVADVLRQHRRIREQLRRISHRYRKFIVSCLLLVTA 259
Query: 261 SQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWH 320
SQ LL +PHA V+I GEL L S++LV+GL I L SA KITHK Q++T +A++WH
Sbjct: 260 SQFSALLAATRPHAQVNIATSGELALCSLSLVTGLLICLHSAAKITHKTQAITSVAAQWH 319
Query: 321 ICATINSFDNIDGETPTALIASAQ------------AMAADISWGSSEDESGDEEDELNN 368
ATINS + D E P I ++ A A S SEDE+ +D L+
Sbjct: 320 ADATINSQER-DHENPRTPIKASSYLHAAGPVVPQPAPNASSSGDESEDETSPSDDGLDG 378
Query: 369 TKMMPIQTRTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKT 428
TK++ ISF KRQALVTY+ENNRAGITV+GF++DRTWLH++F I+ +L +WLL KT
Sbjct: 379 TKIVSFHATHISFQKRQALVTYLENNRAGITVFGFVVDRTWLHALFMIEFSLVMWLLGKT 438
Query: 429 VGI 431
+GI
Sbjct: 439 IGI 441
>UniRef100_Q6YTJ0 Hypothetical protein P0020D05.9 [Oryza sativa]
Length = 449
Score = 312 bits (799), Expect = 1e-83
Identities = 173/426 (40%), Positives = 252/426 (58%), Gaps = 24/426 (5%)
Query: 19 IYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADH 78
+ H G EL FR+ L W+ VD S + SW+ F LA P + L
Sbjct: 34 VSHAGDELHSFRSCLAWMCVDHSTRARGAASWAAFLLLAVAAPSAATLALPSPVGGGGS- 92
Query: 79 RRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTM 138
P+ VQ+SL++ A L++++L+ + G + L+LD++ D+S +++ GY ++ +
Sbjct: 93 --PFDGQVQVSLTLAAALAYLTLTALLQGRGLRRLLYLDRLRDDSEEVRSGYIEELAGSF 150
Query: 139 KLILLWGLPCFICQCVYKIWWYISGAS-QIPYYGEVYVSGIILCTLELCSWFYRTSIFFL 197
+++ + LPC + + YK +WY++ + P++ C +E+ SW YRT++FF+
Sbjct: 151 RVLACFLLPCTLAEAAYKAYWYLAAPPFRSPWWSAA------ACAVEVASWAYRTAVFFM 204
Query: 198 VCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLL 257
VCVLFR I YLQI R+ FA F R +V +L H RIR+ L ISHR+R FIL L+L
Sbjct: 205 VCVLFRTICYLQILRMKGFAREFCRFADVAAVLESHRRIRKQLHRISHRYRRFILCCLVL 264
Query: 258 VTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLAS 317
VT SQ LL +PHA +++ GEL L S++LV+GL + L+SA KITHK Q++T +A+
Sbjct: 265 VTTSQFAALLATTRPHAQINLATAGELALCSLSLVAGLLVCLQSAAKITHKTQAITSVAA 324
Query: 318 KWHICATINSFDNIDGETP-----------TALIASAQAMAADISWGSSEDESGDEEDEL 366
WH ATIN+FDN D E P + A A S SS S D++D
Sbjct: 325 GWHADATINAFDN-DQEDPNPDLPRIVGYLVPVNAYWMASGESSSDSSSSSSSSDDDDSG 383
Query: 367 N-NTKMMPIQTRTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLL 425
+ +K +P Q F +RQALVTY+ENNRAGITVYGF++DRTWLH++F I+ +L +WLL
Sbjct: 384 HPKSKYIPFQNNH-CFQQRQALVTYLENNRAGITVYGFVVDRTWLHALFMIEFSLVMWLL 442
Query: 426 NKTVGI 431
KTVGI
Sbjct: 443 GKTVGI 448
>UniRef100_Q8LG29 Hypothetical protein [Arabidopsis thaliana]
Length = 456
Score = 208 bits (530), Expect = 2e-52
Identities = 125/412 (30%), Positives = 225/412 (54%), Gaps = 15/412 (3%)
Query: 24 KELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYH 83
+ L +L + +QS+ +SW VF ++ V+P+ L C C+ + +
Sbjct: 49 RTLEWLETFLTLLGFNQSSKQSLVLSWIVFLSIGLVLPVTVLELGHC-LGCERYQYKSFE 107
Query: 84 VPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILL 143
+ + +S ++ A +S + +S RK+G KFLF+D++S +++ Y Q+ +++L+ +
Sbjct: 108 LNIVVSQALLAGVSLLCVSHNLRKHGIRKFLFVDQLSGRMDRLKAQYIQQILNSVRLLAV 167
Query: 144 WGLPCFICQCVYKI--WWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVL 201
W LPCF + V +I +Y+ + + ++S IL ++ L SW Y ++IF +
Sbjct: 168 WSLPCFALKGVREIIRMYYVP-------HDQPWLSVAILLSMIL-SWTYLSTIFLAASAM 219
Query: 202 FRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTAS 261
F L+ LQ+ +++A + + E+E+ + EH+R+R L ISHRFR F+L L+VTAS
Sbjct: 220 FHLVCNLQVIHFEDYAKLLEGESEISLFIYEHMRLRHYLSKISHRFRIFLLLQFLVVTAS 279
Query: 262 QLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHI 321
Q L + + GG+ + ++ V G+ + L +ATKI+H+AQ++ +AS+WH
Sbjct: 280 QFTTLFQTTAYSGRITYINGGDFAVSAVVQVVGIILCLHAATKISHRAQAIASVASRWHA 339
Query: 322 CATINSFDNID-GETPTALIASAQA---MAADISWGSSEDESGDEEDELNNTKMMPIQTR 377
+ +S D+ +P+ + A ++ IS S+ ES D + T P
Sbjct: 340 MMSCSSTDSTQIRASPSGVHLEATTNPPISFPISRSDSDVESMDHYMRMPVTNQFPSYMS 399
Query: 378 TISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
S+HKRQA V Y++ N GIT++G+ +DR +++IF I+L+L ++L KTV
Sbjct: 400 MSSYHKRQAFVLYLQMNPGGITIFGWTVDRHLINTIFFIELSLVTFVLGKTV 451
>UniRef100_Q8L726 Hypothetical protein At2g21080 [Arabidopsis thaliana]
Length = 414
Score = 204 bits (520), Expect = 3e-51
Identities = 124/413 (30%), Positives = 215/413 (52%), Gaps = 40/413 (9%)
Query: 25 ELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDC-----STTCDADHR 79
+LR FR L+W +D S+ C +S+ +F +VP++S + S DA+
Sbjct: 31 DLRNFRLLLKWCALDHSSSCGKAVSYMMFVVFTLLVPLISCLFIKTPRNRPSAVMDANS- 89
Query: 80 RPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMK 139
++V VQ S A + F++L + R Y +K LFLD +S ++ GY+ ++ + ++
Sbjct: 90 --FNVLVQFPESGLAVIGFLTLICFFRIYSLTKLLFLD----DSTLVRLGYSRELDKALR 143
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVS-GIILCTLELCSWFYRTSIFFLV 198
+ +P F+ + V+K ++ S P+ + ++ L L SW YRT +F LV
Sbjct: 144 YLAYILVPSFLVELVHKSIFFYSAEVSFPFIKSSCAALNFVMFFLVLFSWVYRTGVFLLV 203
Query: 199 CVLFRLIGYLQIQRLDEFAPVFQR--ETEVGTILLEHLRIRRNLRVISHRFRAFILSSLL 256
C+LFRL LQI R +F R + + EH+RI++ L SHR+R FI+++ +
Sbjct: 204 CILFRLTCELQILRFRGLHKLFDRCGSDTIEDVCKEHVRIKKQLSATSHRYRFFIITAFV 263
Query: 257 LVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLA 316
+++ SQ + LL+V+ ++ L G+L + S +SG ++ L A +ITH+AQ + +A
Sbjct: 264 VISTSQFVALLLVLASKSEKSFLSSGDLVVCSAVQLSGFFLCLLGAARITHRAQGVVCIA 323
Query: 317 SKWHICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQT 376
++WH+ AL +++A+ S + D D + + P
Sbjct: 324 TRWHM----------------ALTCASEAV--------SPESDTDSSDNI-YINVSPSLD 358
Query: 377 RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
+ F RQALV Y+ +N GIT+YG+ LDR LH++F + +L +W+L+K V
Sbjct: 359 LSSFFQARQALVEYLRHNNKGITLYGYALDRGLLHTLFAFEFSLVMWILSKVV 411
>UniRef100_Q9CAG2 Hypothetical protein F12B7.12 [Arabidopsis thaliana]
Length = 486
Score = 194 bits (493), Expect = 4e-48
Identities = 125/442 (28%), Positives = 226/442 (50%), Gaps = 45/442 (10%)
Query: 24 KELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYH 83
+ L +L + +QS+ +SW VF ++ V+P+ L C C+ + +
Sbjct: 49 RTLEWLETFLTLLGFNQSSKQSLVLSWIVFLSIGLVLPVTVLELGHC-LGCERYQYKSFE 107
Query: 84 VPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQ---------- 133
+ + +S ++ A +S + +S RK+G KFLF+D++S +++ Y Q
Sbjct: 108 LNIVVSQALLAGVSLLCVSHNLRKHGIRKFLFVDQLSGRMDRLKAQYIQQILVIYSTFSS 167
Query: 134 --------------------MKRTMKLILLWGLPCFICQCVYKI--WWYISGASQIPYYG 171
++ +++L+ +W LPCF + V +I +Y+ +
Sbjct: 168 FLRLQCDDDTYFKVCIFLIVLQNSVRLLAVWSLPCFALKGVREIIRMYYVP-------HD 220
Query: 172 EVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILL 231
+ ++S IL ++ L SW Y ++IF +F L+ LQ+ +++A + + E+E+ +
Sbjct: 221 QPWLSVAILLSMIL-SWTYLSTIFLAASAMFHLVCNLQVIHFEDYAKLLEGESEISLFIY 279
Query: 232 EHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPHADVDILKGGELGLVSITL 291
EH+R+R L ISHRFR F+L L+VTASQ L + + GG+ + ++
Sbjct: 280 EHMRLRHYLSKISHRFRIFLLLQFLVVTASQFTTLFQTTAYSGRITYINGGDFAVSAVVQ 339
Query: 292 VSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNID-GETPTALIASAQA---MA 347
V G+ + L +ATKI+H+AQ++ +AS+WH + +S D+ +P+ + A ++
Sbjct: 340 VVGIILCLHAATKISHRAQAIASVASRWHAMMSCSSTDSTQIRASPSGVHLEATTNPPIS 399
Query: 348 ADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALVTYMENNRAGITVYGFMLDR 407
IS S+ ES D + T P S+HKRQA V Y++ N GIT++G+ +DR
Sbjct: 400 FPISRSDSDVESMDHYMRMPVTNQFPSYMSMSSYHKRQAFVLYLQMNPGGITIFGWTVDR 459
Query: 408 TWLHSIFGIQLALCLWLLNKTV 429
+++IF I+L+L ++L KTV
Sbjct: 460 HLINTIFFIELSLVTFVLGKTV 481
>UniRef100_Q6ZK43 Hypothetical protein OJ1163_G08.36 [Oryza sativa]
Length = 483
Score = 193 bits (491), Expect = 7e-48
Identities = 132/415 (31%), Positives = 213/415 (50%), Gaps = 19/415 (4%)
Query: 24 KELRRFRAYLRWVYVDQSN-LCKAGISWSVFFTLAYVVPILSHFLLDC---STTCDADHR 79
K LRR A+L + S+ L KAG + S L +P L+ L C CD
Sbjct: 64 KWLRRLEAFLSAAGLAASSPLGKAGAA-SALAVLGVALPALAVALSPCRGRGRGCDEFEV 122
Query: 80 RPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMK 139
+ V V +S + A ++ +S+ YG KFLF+D ++ Q+ Y ++K
Sbjct: 123 EVFEVCVLLSQAAAAAVALACVSRKMAMYGLRKFLFVDPELGMRIRFQKEYVAKIKDFFS 182
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVC 199
IL W LPCF+ + +++ + S I + E + + SW Y T+I C
Sbjct: 183 TILWWILPCFVVKVTREMFRF----SHI--FQESTWRSCAVLFASIMSWMYLTTIILTSC 236
Query: 200 VLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVT 259
+LF L+ LQ+ D++ + +++++ L EHL++R NL ISHRFR F+L VT
Sbjct: 237 MLFNLVCNLQVIHFDDYGKLLEQDSDPLVYLKEHLQLRHNLSKISHRFRMFLLLLFFSVT 296
Query: 260 ASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKW 319
ASQ L ++ GG++ + S+ V GL + L +A KI+H+AQ++ LAS+W
Sbjct: 297 ASQFAILFKTTAYTGPINFTNGGDIAVSSVVQVVGLVLCLHAAAKISHRAQNIASLASRW 356
Query: 320 HICATINSFDNIDGETPTA----LIASAQAMAADISWGSSED-ESGDEEDELNNTKMMPI 374
H T +S D+ TP + + A D S E ESG + + T +
Sbjct: 357 HALVTCSS-DSTYVSTPNSSGNLVPFPAHMFLRDFSESDLESLESGSVQGNSHGTAQ--L 413
Query: 375 QTRTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
+ S+HKR++LV Y+ N GIT++G+++DRT+L++I ++L L L++L+KTV
Sbjct: 414 ASYMSSYHKRESLVLYLLGNPGGITIFGWIVDRTFLNTILMLELTLVLFVLSKTV 468
>UniRef100_Q9SKQ5 Hypothetical protein At2g21080 [Arabidopsis thaliana]
Length = 414
Score = 165 bits (417), Expect = 3e-39
Identities = 95/311 (30%), Positives = 167/311 (53%), Gaps = 15/311 (4%)
Query: 25 ELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDC-----STTCDADHR 79
+LR FR L+W +D S+ C +S+ +F +VP++S + S DA+
Sbjct: 31 DLRNFRLLLKWCALDHSSSCGKAVSYMMFVVFTLLVPLISCLFIKTPRNRPSAVMDANS- 89
Query: 80 RPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMK 139
++V VQ S A + F++L + R Y +K LFLD +S ++ GY+ ++ + ++
Sbjct: 90 --FNVLVQFPESGLAVIGFLTLICFFRIYSLTKLLFLD----DSTLVRLGYSRELDKALR 143
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVS-GIILCTLELCSWFYRTSIFFLV 198
+ +P F+ + V+K ++ S P+ + ++ L L SW YRT +F LV
Sbjct: 144 YLAYILVPSFLVELVHKSIFFYSAEVSFPFIKSSCAALNFVMFFLVLFSWVYRTGVFLLV 203
Query: 199 CVLFRLIGYLQIQRLDEFAPVFQR--ETEVGTILLEHLRIRRNLRVISHRFRAFILSSLL 256
C+LFRL LQI R +F R + + EH+RI++ L SHR+R FI+++ +
Sbjct: 204 CILFRLTCELQILRFRGLHKLFDRCGSDTIEDVCKEHVRIKKQLSATSHRYRFFIITAFV 263
Query: 257 LVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLA 316
+++ SQ + LL+V+ ++ L G+L + S +SG ++ L A +ITH+AQ + +A
Sbjct: 264 VISTSQFVALLLVLASKSEKSFLSSGDLVVCSAVQLSGFFLCLLGAARITHRAQGVVCIA 323
Query: 317 SKWHICATINS 327
++WH+ T S
Sbjct: 324 TRWHMALTCAS 334
>UniRef100_Q84QR8 Hypothetical protein OJ1081_B12.104 [Oryza sativa]
Length = 427
Score = 120 bits (302), Expect = 6e-26
Identities = 87/280 (31%), Positives = 136/280 (48%), Gaps = 15/280 (5%)
Query: 16 NSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLL--DCSTT 73
+ + HD ELR FR +L+W +D S+ S++ F LA VP L D +
Sbjct: 39 SKSVIHD--ELRSFRVFLQWCALDHSSRAARAASYAAFLALALAVPAAVSLSLRADAGAS 96
Query: 74 CDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYG-FSKFLFLDK-VSDESLKIQRGYA 131
+ ++ Q + A +SF +L+ + R+ G + LFLD + D++ ++RGYA
Sbjct: 97 PVSASAITFNRVAQAPATGLAAISFAALASFFRRGGGLRQLLFLDGGLRDDTAFVRRGYA 156
Query: 132 IQMKRTMKLILLWGLPCFICQCVYKIWWYISGAS-----QIPYYGEVYVSGIILCTLELC 186
++ R +L+ LP + +K ++ + +P G + + ++ T+
Sbjct: 157 RELDRAFRLLAALLLPSLCVEAAHKAVFFFATVRVEPPLPLPGVGVPWRAVALVATV--A 214
Query: 187 SWFYRTSIFFLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGT--ILLEHLRIRRNLRVIS 244
SW YRT +F LVCVLFRL LQI R + +F E I EH RIR L S
Sbjct: 215 SWVYRTGVFLLVCVLFRLTCELQILRFEGIYHMFDVEARAAAAEIFAEHRRIRTQLLATS 274
Query: 245 HRFRAFILSSLLLVTASQLIFLLMVIKPHADVDILKGGEL 284
HR+RAFILS L+ +T SQL L++ + G+L
Sbjct: 275 HRYRAFILSCLVTITVSQLGALVVALSSKDGKSFANSGDL 314
>UniRef100_UPI00003AC228 UPI00003AC228 UniRef100 entry
Length = 346
Score = 37.4 bits (85), Expect = 0.83
Identities = 35/116 (30%), Positives = 48/116 (41%), Gaps = 14/116 (12%)
Query: 115 FLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIP---YYG 171
FL S + Q+G + +K++L+W L + W YISG S IP Y
Sbjct: 121 FLSVTRAVSYRAQQGNT--RRAVLKMVLVWVLAFLLYGPAIISWEYISGESIIPTGECYA 178
Query: 172 EVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQ-----RLDEFAPVFQR 222
E + + L T +F F+ + F L YL IQ RLD F V R
Sbjct: 179 EFFYNWYFLMTASTLEFFTP----FISVMFFNLSIYLNIQKRTKLRLDVFHEVHNR 230
>UniRef100_UPI00003AC227 UPI00003AC227 UniRef100 entry
Length = 415
Score = 37.4 bits (85), Expect = 0.83
Identities = 35/116 (30%), Positives = 48/116 (41%), Gaps = 14/116 (12%)
Query: 115 FLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIP---YYG 171
FL S + Q+G + +K++L+W L + W YISG S IP Y
Sbjct: 121 FLSVTRAVSYRAQQGNT--RRAVLKMVLVWVLAFLLYGPAIISWEYISGESIIPTGECYA 178
Query: 172 EVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQ-----RLDEFAPVFQR 222
E + + L T +F F+ + F L YL IQ RLD F V R
Sbjct: 179 EFFYNWYFLMTASTLEFFTP----FISVMFFNLSIYLNIQKRTKLRLDVFHEVHNR 230
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 693,561,688
Number of Sequences: 2790947
Number of extensions: 27594976
Number of successful extensions: 82223
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 82150
Number of HSP's gapped (non-prelim): 48
length of query: 431
length of database: 848,049,833
effective HSP length: 130
effective length of query: 301
effective length of database: 485,226,723
effective search space: 146053243623
effective search space used: 146053243623
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC122163.4


