

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.5 + phase: 1 /partial
(95 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
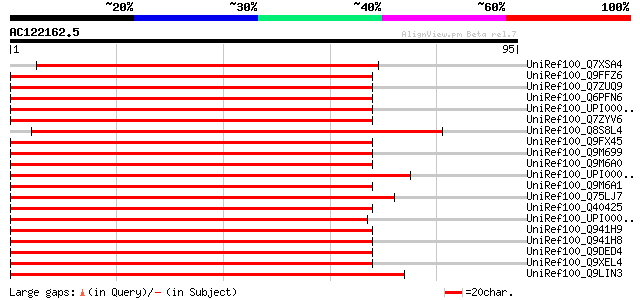
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa] 82 3e-15
UniRef100_Q9FFZ6 Similarity to RNA-binding protein [Arabidopsis ... 76 2e-13
UniRef100_Q7ZUQ9 Cold inducible RNA binding protein [Brachydanio... 72 3e-12
UniRef100_Q6PFN6 Cirbp protein [Brachydanio rerio] 72 3e-12
UniRef100_UPI00003660A9 UPI00003660A9 UniRef100 entry 70 1e-11
UniRef100_Q7ZYV6 Hyperosmotic glycine rich protein [Salmo salar] 70 1e-11
UniRef100_Q8S8L4 Putative RNA-binding protein [Arabidopsis thali... 70 1e-11
UniRef100_Q9FX45 RNA-binding glycine-rich protein, putative [Ara... 69 2e-11
UniRef100_Q9M699 Putative glycine-rich RNA-binding protein 2 [Ca... 69 3e-11
UniRef100_Q9M6A0 Putative glycine-rich RNA binding protein 3 [Ca... 68 6e-11
UniRef100_UPI00004996C0 UPI00004996C0 UniRef100 entry 67 1e-10
UniRef100_Q9M6A1 Putative glycine-rich RNA binding protein 1 [Ca... 67 1e-10
UniRef100_Q75LJ7 Putative RNA binding protein [Oryza sativa] 66 2e-10
UniRef100_Q40425 RNA-binding gricine-rich protein-1 [Nicotiana s... 66 2e-10
UniRef100_UPI000027A5FC UPI000027A5FC UniRef100 entry 65 4e-10
UniRef100_Q941H9 RNA-binding protein precursor [Nicotiana tabacum] 65 4e-10
UniRef100_Q941H8 RNA-binding protein precursor [Solanum tuberosum] 65 4e-10
UniRef100_Q9DED4 Cold-inducible RNA binding protein 2 [Xenopus l... 65 5e-10
UniRef100_Q9XEL4 Glycine-rich RNA-binding protein [Picea glauca] 65 5e-10
UniRef100_Q9LIN3 Similarity to RNA-binding protein [Arabidopsis ... 64 9e-10
>UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa]
Length = 137
Score = 82.0 bits (201), Expect = 3e-15
Identities = 40/64 (62%), Positives = 50/64 (77%)
Query: 6 TEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVL 65
TE+ LAEAFSQYG V++A IV +K R KGFG+V FA EE A KA+ MNGK+L+GRV+
Sbjct: 49 TEDSLAEAFSQYGQVLEATIVTDKMTNRPKGFGFVKFASEEAANKAKEEMNGKVLNGRVI 108
Query: 66 YVDM 69
YVD+
Sbjct: 109 YVDI 112
>UniRef100_Q9FFZ6 Similarity to RNA-binding protein [Arabidopsis thaliana]
Length = 146
Score = 76.3 bits (186), Expect = 2e-13
Identities = 36/68 (52%), Positives = 52/68 (75%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F TTE+ L+EAFS+ G VV+A IV+++ R KGFG+VTFA +EA+KA + NG+ L
Sbjct: 41 LSFCTTEQGLSEAFSKCGQVVEAQIVMDRVSDRSKGFGFVTFASADEAQKALMEFNGQQL 100
Query: 61 HGRVLYVD 68
+GR ++VD
Sbjct: 101 NGRTIFVD 108
>UniRef100_Q7ZUQ9 Cold inducible RNA binding protein [Brachydanio rerio]
Length = 184
Score = 72.0 bits (175), Expect = 3e-12
Identities = 31/68 (45%), Positives = 47/68 (68%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ TTE+ L EAFS+YG + K D++ ++ R +GFG+VTF E+A+ A MNGK +
Sbjct: 12 LSYDTTEQSLEEAFSKYGTIAKVDVIRDRETDRSRGFGFVTFENPEDAKDAMAAMNGKQV 71
Query: 61 HGRVLYVD 68
GR++ VD
Sbjct: 72 DGRMIRVD 79
>UniRef100_Q6PFN6 Cirbp protein [Brachydanio rerio]
Length = 121
Score = 72.0 bits (175), Expect = 3e-12
Identities = 31/68 (45%), Positives = 47/68 (68%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ TTE+ L EAFS+YG + K D++ ++ R +GFG+VTF E+A+ A MNGK +
Sbjct: 5 LSYDTTEQSLEEAFSKYGTIAKVDVIRDRETDRSRGFGFVTFENPEDAKDAMAAMNGKQV 64
Query: 61 HGRVLYVD 68
GR++ VD
Sbjct: 65 DGRMIRVD 72
>UniRef100_UPI00003660A9 UPI00003660A9 UniRef100 entry
Length = 169
Score = 70.1 bits (170), Expect = 1e-11
Identities = 32/68 (47%), Positives = 44/68 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T EE LAEAF +YG + K D++ +K R +GFG+V + E+A+ A MNGK L
Sbjct: 12 LSFETNEESLAEAFGKYGTIEKVDVIRDKETGRSRGFGFVKYESVEDAKDAMTAMNGKSL 71
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 72 DGRAIRVD 79
>UniRef100_Q7ZYV6 Hyperosmotic glycine rich protein [Salmo salar]
Length = 205
Score = 70.1 bits (170), Expect = 1e-11
Identities = 31/68 (45%), Positives = 48/68 (70%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F TTE+ LAEAFS+YGN+ K D+++++ R +GFG+V + E+A+ A MNG+ L
Sbjct: 12 LSFDTTEQSLAEAFSKYGNISKCDVIMDRETGRPRGFGFVKYDNPEDAKDAMDAMNGQSL 71
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 72 DGRTIRVN 79
>UniRef100_Q8S8L4 Putative RNA-binding protein [Arabidopsis thaliana]
Length = 142
Score = 70.1 bits (170), Expect = 1e-11
Identities = 33/77 (42%), Positives = 50/77 (64%)
Query: 5 TTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRV 64
TT EKL +AF+ +G +V A ++ ++ R KGFG+VT+A E+A KA+ MN K L G V
Sbjct: 45 TTNEKLQDAFASFGQLVDARVITDRDSGRSKGFGFVTYATIEDAEKAKAEMNAKFLDGWV 104
Query: 65 LYVDMDPPNEQKNKIKQ 81
++VD P E + ++Q
Sbjct: 105 IFVDPARPREPRRPLQQ 121
>UniRef100_Q9FX45 RNA-binding glycine-rich protein, putative [Arabidopsis thaliana]
Length = 181
Score = 69.3 bits (168), Expect = 2e-11
Identities = 31/68 (45%), Positives = 48/68 (70%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F TTE+ L + F Q+GN++ ++V++K R KGF ++ + EEEA KA GM+GK L
Sbjct: 84 LSFRTTEDTLRDTFEQFGNLIHMNMVMDKVANRPKGFAFLRYETEEEAMKAIQGMHGKFL 143
Query: 61 HGRVLYVD 68
GRV++V+
Sbjct: 144 DGRVIFVE 151
>UniRef100_Q9M699 Putative glycine-rich RNA-binding protein 2 [Catharanthus roseus]
Length = 160
Score = 68.9 bits (167), Expect = 3e-11
Identities = 30/68 (44%), Positives = 51/68 (74%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++TT++ L+EAFSQYG V+++ ++ ++ R +GFG+VTF +E+ + A +GMNG+ L
Sbjct: 15 LAWATTDQSLSEAFSQYGEVLESKVINDRETGRSRGFGFVTFGDEKSMKDAIVGMNGQTL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRNITVN 82
>UniRef100_Q9M6A0 Putative glycine-rich RNA binding protein 3 [Catharanthus roseus]
Length = 164
Score = 67.8 bits (164), Expect = 6e-11
Identities = 31/68 (45%), Positives = 50/68 (72%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++TT++ L+EAFSQYG ++++ I+ ++ R +GFG+VTF +E+ R A GMNG+ L
Sbjct: 15 LAWATTDQSLSEAFSQYGEILESKIINDRETGRSRGFGFVTFKDEQSMRDAIEGMNGQTL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRNITVN 82
>UniRef100_UPI00004996C0 UPI00004996C0 UniRef100 entry
Length = 136
Score = 67.0 bits (162), Expect = 1e-10
Identities = 32/75 (42%), Positives = 49/75 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA+S T+E L AF ++G V +V ++ +R KGFG+VTF ++E+A+KA MN + L
Sbjct: 9 LAYSVTDESLKAAFEKFGTVTDCKVVTDRDSQRSKGFGFVTFEKDEDAKKAIEEMNEQEL 68
Query: 61 HGRVLYVDMDPPNEQ 75
GR + VD+ P E+
Sbjct: 69 EGRRIKVDVSRPREE 83
>UniRef100_Q9M6A1 Putative glycine-rich RNA binding protein 1 [Catharanthus roseus]
Length = 137
Score = 66.6 bits (161), Expect = 1e-10
Identities = 31/68 (45%), Positives = 50/68 (72%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++TT++ L+EAFSQYG V+++ I+ ++ R +GFG+VTF +E+ + A GMNG+ L
Sbjct: 15 LAWATTDQSLSEAFSQYGEVLESKIINDRETGRSRGFGFVTFGDEKSMKDAIEGMNGQTL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRNVTVN 82
>UniRef100_Q75LJ7 Putative RNA binding protein [Oryza sativa]
Length = 205
Score = 65.9 bits (159), Expect = 2e-10
Identities = 32/72 (44%), Positives = 47/72 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++STT+E L +AF ++GN+ +A +V +K R +GFG+VTF E++ A GMNG L
Sbjct: 14 LSWSTTDESLKDAFGKFGNLTEAKVVFDKYSGRSRGFGFVTFDEKKAMEDAIEGMNGLDL 73
Query: 61 HGRVLYVDMDPP 72
GR + VD P
Sbjct: 74 DGRAITVDKAQP 85
>UniRef100_Q40425 RNA-binding gricine-rich protein-1 [Nicotiana sylvestris]
Length = 165
Score = 65.9 bits (159), Expect = 2e-10
Identities = 32/68 (47%), Positives = 48/68 (70%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++TT+ L EAFSQYG V+++ I+ ++ R +GFG+VTF +E+ R A GMNG+ L
Sbjct: 13 LAWATTDRTLGEAFSQYGEVLESKIINDRETGRSRGFGFVTFGDEKSMRDAIEGMNGQDL 72
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 73 DGRNITVN 80
>UniRef100_UPI000027A5FC UPI000027A5FC UniRef100 entry
Length = 74
Score = 65.1 bits (157), Expect = 4e-10
Identities = 31/67 (46%), Positives = 45/67 (66%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L + + +E LAE FS+YG VV A I+L++ R +GFG+V + +E+A+KA G+NGK
Sbjct: 8 LPYRSRDEDLAEIFSEYGEVVSAKIILDRETNRSRGFGFVEMSSDEDAKKAIEGLNGKEF 67
Query: 61 HGRVLYV 67
GR L V
Sbjct: 68 DGRSLVV 74
>UniRef100_Q941H9 RNA-binding protein precursor [Nicotiana tabacum]
Length = 277
Score = 65.1 bits (157), Expect = 4e-10
Identities = 29/68 (42%), Positives = 47/68 (68%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ T E L EAFSQYG+V++A +++++ R +GFG+++F EEA A M+G+ L
Sbjct: 47 LSYGTDESSLKEAFSQYGDVIEARVIMDRDTGRSRGFGFISFPSSEEAASALQAMDGQDL 106
Query: 61 HGRVLYVD 68
HGR + V+
Sbjct: 107 HGRRIRVN 114
>UniRef100_Q941H8 RNA-binding protein precursor [Solanum tuberosum]
Length = 339
Score = 65.1 bits (157), Expect = 4e-10
Identities = 29/68 (42%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ T E L E FSQYG V++A ++L++ R +GFG+++F EEA A M+G+ L
Sbjct: 47 LSYGTDESSLKETFSQYGEVIEARVILDRETGRSRGFGFISFPSSEEATSAMQAMDGQDL 106
Query: 61 HGRVLYVD 68
HGR + V+
Sbjct: 107 HGRRIKVN 114
>UniRef100_Q9DED4 Cold-inducible RNA binding protein 2 [Xenopus laevis]
Length = 166
Score = 64.7 bits (156), Expect = 5e-10
Identities = 29/68 (42%), Positives = 44/68 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L F T EE L + FS+YG + + +V ++ KR +GFG+VTF ++A+ A + MNGK +
Sbjct: 12 LNFDTNEESLEQVFSKYGQISEVVVVKDRETKRSRGFGFVTFENPDDAKDAMMAMNGKAV 71
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 72 DGRQIRVD 79
>UniRef100_Q9XEL4 Glycine-rich RNA-binding protein [Picea glauca]
Length = 155
Score = 64.7 bits (156), Expect = 5e-10
Identities = 32/68 (47%), Positives = 46/68 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA+ST + L EAFS YG VV++ I+ ++ R +GFG+VTF +E+ R A MNGK+L
Sbjct: 15 LAWSTDDRSLQEAFSPYGEVVESKIISDRETGRSRGFGFVTFNDEQSMRDAIDAMNGKML 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRSITVN 82
>UniRef100_Q9LIN3 Similarity to RNA-binding protein [Arabidopsis thaliana]
Length = 245
Score = 63.9 bits (154), Expect = 9e-10
Identities = 30/74 (40%), Positives = 50/74 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T++ L +AF +YG++V+A +VL+K R +GFG++TF E++ +A MNG L
Sbjct: 14 LAWTTSDRGLRDAFEKYGHLVEAKVVLDKFSGRSRGFGFITFDEKKAMDEAIAAMNGMDL 73
Query: 61 HGRVLYVDMDPPNE 74
GR + VD P++
Sbjct: 74 DGRTITVDKAQPHQ 87
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.130 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 155,078,136
Number of Sequences: 2790947
Number of extensions: 5646619
Number of successful extensions: 18424
Number of sequences better than 10.0: 3590
Number of HSP's better than 10.0 without gapping: 2897
Number of HSP's successfully gapped in prelim test: 693
Number of HSP's that attempted gapping in prelim test: 13238
Number of HSP's gapped (non-prelim): 5522
length of query: 95
length of database: 848,049,833
effective HSP length: 71
effective length of query: 24
effective length of database: 649,892,596
effective search space: 15597422304
effective search space used: 15597422304
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC122162.5


