

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
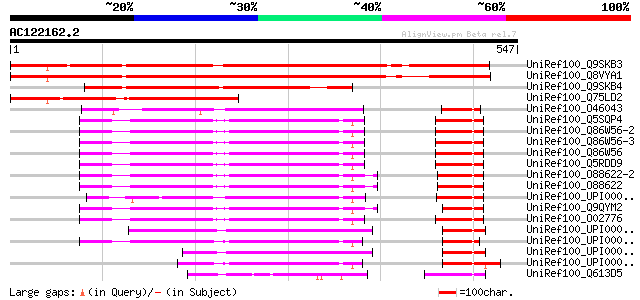
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SKB3 Poly(ADP-ribose) glycohydrolase [Arabidopsis th... 535 e-150
UniRef100_Q8VYA1 Putative poly(ADP-ribose) glycohydrolase [Arabi... 493 e-138
UniRef100_Q9SKB4 Putative poly(ADP-ribose) glycohydrolase [Arabi... 293 1e-77
UniRef100_Q75LD2 Putative poly(ADP-ribose) glycohydrolase [Oryza... 237 6e-61
UniRef100_O46043 Poly(ADP-ribose) glycohydrolase [Drosophila mel... 194 7e-48
UniRef100_Q5SQP4 Poly (ADP-ribose) glycohydrolase [Homo sapiens] 172 2e-41
UniRef100_Q86W56-2 Splice isoform 2 of Q86W56 [Homo sapiens] 172 2e-41
UniRef100_Q86W56-3 Splice isoform 3 of Q86W56 [Homo sapiens] 172 2e-41
UniRef100_Q86W56 Poly(ADP-ribose) glycohydrolase [Homo sapiens] 172 2e-41
UniRef100_Q5RDD9 Hypothetical protein DKFZp469F0213 [Pongo pygma... 172 3e-41
UniRef100_O88622-2 Splice isoform 2 of O88622 [Mus musculus] 171 5e-41
UniRef100_O88622 Poly(ADP-ribose) glycohydrolase [Mus musculus] 171 5e-41
UniRef100_UPI00002F9CB0 UPI00002F9CB0 UniRef100 entry 170 9e-41
UniRef100_Q9QYM2 Poly(ADP-ribose) glycohydrolase [Rattus norvegi... 167 7e-40
UniRef100_O02776 Poly(ADP-ribose) glycohydrolase [Bos taurus] 167 1e-39
UniRef100_UPI000027C995 UPI000027C995 UniRef100 entry 165 4e-39
UniRef100_UPI00003AE137 UPI00003AE137 UniRef100 entry 153 1e-35
UniRef100_UPI00002B206C UPI00002B206C UniRef100 entry 144 5e-33
UniRef100_UPI0000318FC5 UPI0000318FC5 UniRef100 entry 139 3e-31
UniRef100_Q613D5 Hypothetical protein CBG16441 [Caenorhabditis b... 110 1e-22
>UniRef100_Q9SKB3 Poly(ADP-ribose) glycohydrolase [Arabidopsis thaliana]
Length = 548
Score = 535 bits (1377), Expect = e-150
Identities = 287/521 (55%), Positives = 363/521 (69%), Gaps = 29/521 (5%)
Query: 1 MEKREDWRSILPYLPVVMRSSSLFWPSQVVGALRELGCG----RVDSGQLLFIFITELRN 56
ME RED SILPYLP+V+RSSSL+WP +VV AL+ + G +VDSG++L I ++R
Sbjct: 1 MENREDLNSILPYLPLVIRSSSLYWPPRVVEALKAMSEGPSHSQVDSGEVLRQAIFDMRR 60
Query: 57 SLSLSSEPLAPSAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD 116
SLS S+ L PSA++GYA FDELI +E ++WF+E++P+L LLL+ PSLLE H++NAD
Sbjct: 61 SLSFST--LEPSASNGYAFLFDELIDEKESKRWFDEIIPALASLLLQFPSLLEVHFQNAD 118
Query: 117 MVIDGKGATVRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFD 176
++ G ++TGLR+L+SQ+AGIVFL+QELI ALLACSF CLFP +R K L +NFD
Sbjct: 119 NIVSG----IKTGLRLLNSQQAGIVFLSQELIGALLACSFFCLFPDDNRGAKHLPVINFD 174
Query: 177 ELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF 236
LFASLY Y+Q QE KI CI+HYF+R S +P G+VSFERK+ + P+A F
Sbjct: 175 HLFASLYISYSQSQESKIRCIMHYFERFCSCVPIGIVSFERKIT---------AAPDADF 225
Query: 237 WSTSVKPLCRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIA 296
WS S LC F+V S GLIED +EVDFAN+YLGGG+L RGCVQEEIRFMI+PELIA
Sbjct: 226 WSKSDVSLCAFKVHSFGLIEDQPDNALEVDFANKYLGGGSLSRGCVQEEIRFMINPELIA 285
Query: 297 GMIFLPSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL 356
GM+FLP M DNEAI+IVG ERFS YTGYASSFRF+G+Y+D K +DPF RR+TRIVAIDAL
Sbjct: 286 GMLFLPRMDDNEAIEIVGAERFSCYTGYASSFRFAGEYIDKKAMDPFKRRRTRIVAIDAL 345
Query: 357 CGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQKIPQENVCSSLLVVFSMTF*HQYGLL 416
C P MR +++ LLREINKA CGFL S+ Q I + + + +V + GLL
Sbjct: 346 CTPKMRHFKDICLLREINKALCGFLNCSKAWEHQNIFMDEGDNEIQLVRNG---RDSGLL 402
Query: 417 SASLVVQLSVLNIFRFS*FDAMETSEGKYSYQEIRNSQNDYDMMENSNDIGVATGNWGCG 476
+ + +E + K + IR+ + E+ D GVATGNWGCG
Sbjct: 403 RTETTAS-------HRTPLNDVEMNREKPANNLIRDFYVEGVDNEDHEDDGVATGNWGCG 455
Query: 477 AFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQNLDKI 517
FGGDPE+K IQWLAASQ RPFI+YYTFG AL+NLD++
Sbjct: 456 VFGGDPELKATIQWLAASQTRRPFISYYTFGVEALRNLDQV 496
>UniRef100_Q8VYA1 Putative poly(ADP-ribose) glycohydrolase [Arabidopsis thaliana]
Length = 522
Score = 493 bits (1269), Expect = e-138
Identities = 271/522 (51%), Positives = 341/522 (64%), Gaps = 47/522 (9%)
Query: 1 MEKREDWRSILPYLPVVMRSSSLFWPSQVVGALRELGCG----RVDSGQLLFIFITELRN 56
ME R D RSIL YLP+V +SSSL WP V L+ + G V+SG+ L + IT +R
Sbjct: 1 MELRADLRSILQYLPLVAQSSSLVWPPSVEEELQTISRGPSESMVNSGEALALHITNMRK 60
Query: 57 SLSLSSEPLAPSAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD 116
SLSL++ LAP A GY LFFD+ ISREE +F EV+P+L LLL+LPS+LE HY+ AD
Sbjct: 61 SLSLNASDLAPYALQGYGLFFDKKISREESANFFGEVVPALCRLLLQLPSMLEKHYQKAD 120
Query: 117 MVIDGKGATVRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFD 176
V+DG V++GLR+L QEAGIV L+QELIAALLACSF CLFP DR K LQ +NF
Sbjct: 121 HVLDG----VKSGLRLLGPQEAGIVLLSQELIAALLACSFFCLFPEVDRSLKNLQGINFS 176
Query: 177 ELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF 236
LF+ Y + KQE+KI C+IHYF RI MP G VSFERK+LP E +SYP A
Sbjct: 177 GLFSFPYMRHCTKQENKIKCLIHYFGRICRWMPTGFVSFERKILPLEYHPHFVSYPKADS 236
Query: 237 WSTSVKPLCRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIA 296
W+ SV PLC E+ +SG IED E +EVDFA+EY GG L +QEEIRF+I+PELIA
Sbjct: 237 WANSVTPLCSIEIHTSGAIEDQPCEALEVDFADEYFGGLTLSYDTLQEEIRFVINPELIA 296
Query: 297 GMIFLPSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL 356
GMIFLP M NEAI+IVGVERFS YTGY SF+++GDY D+K++D F RRKTR++AIDA+
Sbjct: 297 GMIFLPRMDANEAIEIVGVERFSGYTGYGPSFQYAGDYTDNKDLDIFRRRKTRVIAIDAM 356
Query: 357 CGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQKIPQENVCSSLLVVFSMTF*HQYGLL 416
PGM QY+ L+RE+NKAF G++ Q +Y D K E S + +
Sbjct: 357 PDPGMGQYKLDALIREVNKAFSGYMHQCKYNIDVKHDPEASSSHVPLTSD---------- 406
Query: 417 SASLVVQLSVLNIFRFS*FDAMETSEGKYSYQEIRNSQNDYDMMENSNDIGVATGNWGCG 476
SAS V++ S + + + IGVATGNWGCG
Sbjct: 407 SASQVIE-----------------------------SSHRWCIDHEEKKIGVATGNWGCG 437
Query: 477 AFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQNLDKIL 518
FGGDPE+K ++QWLA SQ+GRPF++YYTFG ALQNL++++
Sbjct: 438 VFGGDPELKIMLQWLAISQSGRPFMSYYTFGLQALQNLNQVI 479
>UniRef100_Q9SKB4 Putative poly(ADP-ribose) glycohydrolase [Arabidopsis thaliana]
Length = 364
Score = 293 bits (749), Expect = 1e-77
Identities = 162/291 (55%), Positives = 200/291 (68%), Gaps = 26/291 (8%)
Query: 81 ISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDSQEAGI 140
+S+EE +WF E LP++ LLLR PSLLE+HY N+D +I+G +TGLR+L +AGI
Sbjct: 1 MSKEESSRWFNEFLPAMACLLLRFPSLLESHYLNSDNLING----TKTGLRVLVPNKAGI 56
Query: 141 VFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYND-YTQKQEDKIWCIIH 199
VFL+QELI ALL+CSF CLFPV DR L +NFD+LF SL N + QE+KI CIIH
Sbjct: 57 VFLSQELIGALLSCSFFCLFPVDDRGSNHLPIINFDKLFGSLINTGRNEHQENKIKCIIH 116
Query: 200 YFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLIEDHS 259
YFQR++S++ G VSFERK+L E D S + FW S LC EV++SGLIED S
Sbjct: 117 YFQRLSSSISPGFVSFERKILSLEQDS---STLDEGFWGKSTVNLCPVEVRTSGLIEDQS 173
Query: 260 SETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGVERFS 319
E +EVDFAN+ LGGGALR+GCVQEEIRFMI+PELI GM+FLP+M EAI++VG ERFS
Sbjct: 174 VEALEVDFANKNLGGGALRKGCVQEEIRFMINPELIVGMLFLPTMEVTEAIEVVGAERFS 233
Query: 320 SYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKFLL 370
YTG F + KTRIVAIDAL PG+ QY+ + LL
Sbjct: 234 LYTGC------------------FRKAKTRIVAIDALRHPGVSQYKLESLL 266
Score = 37.7 bits (86), Expect = 0.87
Identities = 17/34 (50%), Positives = 25/34 (73%), Gaps = 3/34 (8%)
Query: 487 IIQWLA---ASQAGRPFIAYYTFGSGALQNLDKI 517
++ WL + QA RPF++YYTFG ALQNL+++
Sbjct: 331 VVVWLTTLHSFQARRPFMSYYTFGFEALQNLNQV 364
>UniRef100_Q75LD2 Putative poly(ADP-ribose) glycohydrolase [Oryza sativa]
Length = 242
Score = 237 bits (605), Expect = 6e-61
Identities = 131/251 (52%), Positives = 163/251 (64%), Gaps = 14/251 (5%)
Query: 1 MEKREDWRSILPYLPVVMRSSSLFWPSQVVGALRELGCG----RVDSGQLLFIFITELRN 56
ME R D RSILPYLPVV+R +LFWP AL+ L G RV SG +L +T+LR
Sbjct: 1 MEARGDLRSILPYLPVVLRGGALFWPPAAQEALKALALGPDVSRVSSGDVLADALTDLR- 59
Query: 57 SLSLSSEPLAPSAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD 116
L+L+ +PL AA G+ALFFD+L+SR + R WF+ V PSL LLLRLP+LLE HY A
Sbjct: 60 -LALNLDPLPRRAAEGFALFFDDLLSRAQARDWFDHVAPSLARLLLRLPTLLEGHYRAA- 117
Query: 117 MVIDGKGATVRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFD 176
G R GLR+L SQ+AG+V L+QEL AALLAC+ CLFP DR E L +NFD
Sbjct: 118 ------GDEAR-GLRILSSQDAGLVLLSQELAAALLACALFCLFPTADRAEACLPAINFD 170
Query: 177 ELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF 236
LFA+L + Q QE K+ C++HYF R+T++ P G VSFERKVLP + I+YP+
Sbjct: 171 SLFAALCYNSRQSQEQKVRCLVHYFDRVTASTPTGSVSFERKVLPRRPESDGITYPDMDT 230
Query: 237 WSTSVKPLCRF 247
W S PLC F
Sbjct: 231 WMKSGVPLCTF 241
>UniRef100_O46043 Poly(ADP-ribose) glycohydrolase [Drosophila melanogaster]
Length = 768
Score = 194 bits (492), Expect = 7e-48
Identities = 124/321 (38%), Positives = 176/321 (54%), Gaps = 51/321 (15%)
Query: 78 DELISREECRKWFEEVLPSLGDLLLRLPSLLEA------HYENADMVIDGKGATVRTGLR 131
DE + E R +FE++LP + L LRLP L+++ H++NA +
Sbjct: 197 DEELDESETRVFFEDLLPRIIRLALRLPDLIQSPVPLLKHHKNASLS------------- 243
Query: 132 MLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDEL-FASLYNDYTQKQ 190
L+Q+ I+ LLA +FLC FP + +++ + F ++ F LY
Sbjct: 244 -----------LSQQQISCLLANAFLCTFPRRNTLKRKSEYSTFPDINFNRLYQSTGPAV 292
Query: 191 EDKIWCIIHYFQRI------TSNMPKGVVSFERKV-LPWEDDCIHISYPNASFWSTSVKP 243
+K+ CI+HYF+R+ SN+P GVV+F R+ LP + WS S P
Sbjct: 293 LEKLKCIMHYFRRVCPTERDASNVPTGVVTFVRRSGLP----------EHLIDWSQSAAP 342
Query: 244 L--CRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFL 301
L V + G IED ++VDFAN+YLGGG L GCVQEEIRF+I PEL+ G +F
Sbjct: 343 LGDVPLHVDAEGTIEDEGIGLLQVDFANKYLGGGVLGHGCVQEEIRFVICPELLVGKLFT 402
Query: 302 PSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL-CGPG 360
+ EA+ ++G ER+S+YTGYA SF +SG++ D D GRR+T IVAIDAL
Sbjct: 403 ECLRPFEALVMLGAERYSNYTGYAGSFEWSGNFEDSTPRDSSGRRQTAIVAIDALHFAQS 462
Query: 361 MRQYREKFLLREINKAFCGFL 381
QYRE + RE+NKA+ GF+
Sbjct: 463 HHQYREDLMERELNKAYIGFV 483
Score = 63.2 bits (152), Expect = 2e-08
Identities = 27/42 (64%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 467 GVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGS 508
GVATGNWGCGAFGGD +K ++Q + +Q GRP +AYYTFG+
Sbjct: 492 GVATGNWGCGAFGGDSYLKALLQLMVCAQLGRP-LAYYTFGN 532
>UniRef100_Q5SQP4 Poly (ADP-ribose) glycohydrolase [Homo sapiens]
Length = 491
Score = 172 bits (436), Expect = 2e-41
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 91 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 132
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 133 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 192
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 193 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 241
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 242 EENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 301
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 302 EQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 358
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 359 ELNKAYCGFLR 369
Score = 58.2 bits (139), Expect = 6e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 372 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 422
>UniRef100_Q86W56-2 Splice isoform 2 of Q86W56 [Homo sapiens]
Length = 894
Score = 172 bits (436), Expect = 2e-41
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 494 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 535
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 536 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 595
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 596 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 644
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 645 EENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 704
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 705 EQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 761
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 762 ELNKAYCGFLR 772
Score = 58.2 bits (139), Expect = 6e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 775 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 825
>UniRef100_Q86W56-3 Splice isoform 3 of Q86W56 [Homo sapiens]
Length = 868
Score = 172 bits (436), Expect = 2e-41
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 468 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 509
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 510 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 569
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 570 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 618
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 619 EENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 678
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 679 EQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 735
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 736 ELNKAYCGFLR 746
Score = 58.2 bits (139), Expect = 6e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 749 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 799
>UniRef100_Q86W56 Poly(ADP-ribose) glycohydrolase [Homo sapiens]
Length = 976
Score = 172 bits (436), Expect = 2e-41
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 576 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 617
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 618 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 677
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 678 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 726
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 727 EENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 786
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 787 EQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 843
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 844 ELNKAYCGFLR 854
Score = 58.2 bits (139), Expect = 6e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 857 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 907
>UniRef100_Q5RDD9 Hypothetical protein DKFZp469F0213 [Pongo pygmaeus]
Length = 978
Score = 172 bits (435), Expect = 3e-41
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 578 FWDKILEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 619
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 620 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 679
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 680 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 728
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 729 EENGRGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 788
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 789 EQYSEYTGYAETYRWSRSHEDGSERDGWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 845
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 846 ELNKAYCGFLR 856
Score = 58.2 bits (139), Expect = 6e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 859 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 909
>UniRef100_O88622-2 Splice isoform 2 of O88622 [Mus musculus]
Length = 920
Score = 171 bits (433), Expect = 5e-41
Identities = 110/326 (33%), Positives = 170/326 (51%), Gaps = 43/326 (13%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 569 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 610
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ V ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 611 KMNHSVTMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 670
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 671 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 719
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 720 EGNGRGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 779
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 780 EQYSEYTGYAETYRWARSHEDGSEKDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKVRR 836
Query: 372 EINKAFCGFLQQSQYQRDQKIPQENV 397
E+NKA+CGFL+ +P EN+
Sbjct: 837 ELNKAYCGFLRPG-------VPSENL 855
Score = 57.8 bits (138), Expect = 8e-07
Identities = 28/50 (56%), Positives = 34/50 (68%), Gaps = 1/50 (2%)
Query: 462 NSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 852 SENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 900
>UniRef100_O88622 Poly(ADP-ribose) glycohydrolase [Mus musculus]
Length = 969
Score = 171 bits (433), Expect = 5e-41
Identities = 110/326 (33%), Positives = 170/326 (51%), Gaps = 43/326 (13%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 569 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 610
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ V ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 611 KMNHSVTMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 670
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 671 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 719
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 720 EGNGRGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 779
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 780 EQYSEYTGYAETYRWARSHEDGSEKDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKVRR 836
Query: 372 EINKAFCGFLQQSQYQRDQKIPQENV 397
E+NKA+CGFL+ +P EN+
Sbjct: 837 ELNKAYCGFLRPG-------VPSENL 855
Score = 57.8 bits (138), Expect = 8e-07
Identities = 28/50 (56%), Positives = 34/50 (68%), Gaps = 1/50 (2%)
Query: 462 NSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 852 SENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 900
>UniRef100_UPI00002F9CB0 UPI00002F9CB0 UniRef100 entry
Length = 463
Score = 170 bits (431), Expect = 9e-41
Identities = 110/313 (35%), Positives = 171/313 (54%), Gaps = 44/313 (14%)
Query: 84 EECRK--WFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLR-----MLDSQ 136
E+C K +FE VLP++ +L ++ P L T+ + LR +L ++
Sbjct: 30 EDCEKQHFFEVVLPAMVELAVQAPFL----------------CTMASILRIFPIPLLKAR 73
Query: 137 EAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDEL-FASLYNDYTQKQEDKIW 195
+ L+QE IA LLA +F C FP R ++ + N+ E+ F L+ + ++ +K+
Sbjct: 74 MNHSLTLSQEQIACLLANAFFCTFP--RRNSRKSEYGNYPEINFYRLFEGSSMRKIEKLK 131
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
++ YF+R+T N PKG+V+F R+ L W +S K L R + G I
Sbjct: 132 TLLCYFRRVTQNRPKGLVTFTRQTL-----------NEPPKWESSQKQLTRLHITCQGTI 180
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
ED ++VDFAN ++GGG G VQEEIRF+I+PELI +F ++ +E + I G
Sbjct: 181 EDDGYGMLQVDFANRFVGGGVTGHGLVQEEIRFLINPELIVSRLFTEALEHDECLIITGT 240
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
ER+S Y+GYA S+++ ++D+ D + RR T IVAIDAL R Y E+F + R
Sbjct: 241 ERYSKYSGYAESYKWQDCHIDETPSDEWQRRCTEIVAIDAL---KFRHYLEQFVPQKIRR 297
Query: 372 EINKAFCGFLQQS 384
E+ KA+CGF + +
Sbjct: 298 ELYKAYCGFFRNN 310
Score = 60.1 bits (144), Expect = 2e-07
Identities = 28/51 (54%), Positives = 37/51 (71%), Gaps = 1/51 (1%)
Query: 461 ENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
E+ + VATGNWGCGAFGGD +K +IQ +AA++AGR +AY+TF L
Sbjct: 312 ESKHLSAVATGNWGCGAFGGDTRLKALIQLMAAAEAGRD-VAYFTFRDAQL 361
>UniRef100_Q9QYM2 Poly(ADP-ribose) glycohydrolase [Rattus norvegicus]
Length = 972
Score = 167 bits (423), Expect = 7e-40
Identities = 109/326 (33%), Positives = 169/326 (51%), Gaps = 43/326 (13%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 572 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 613
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ V ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 614 KMNHSVTMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 673
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 674 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCDKPLTRLHVTYEGTI 722
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 723 EGNGRGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 782
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + R T IVAIDAL R+Y ++F + R
Sbjct: 783 EQYSEYTGYAETYRWARSHEDGSEKDDWQRCCTEIVAIDAL---HFRRYLDQFVPEKVRR 839
Query: 372 EINKAFCGFLQQSQYQRDQKIPQENV 397
E+NKA+CGFL+ +P EN+
Sbjct: 840 ELNKAYCGFLRPG-------VPPENL 858
Score = 57.8 bits (138), Expect = 8e-07
Identities = 27/44 (61%), Positives = 32/44 (72%), Gaps = 1/44 (2%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 861 VATGNWGCGAFGGDARLKALIQLLAAAAAERD-VVYFTFGDSEL 903
>UniRef100_O02776 Poly(ADP-ribose) glycohydrolase [Bos taurus]
Length = 977
Score = 167 bits (422), Expect = 1e-39
Identities = 105/311 (33%), Positives = 164/311 (51%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 577 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 618
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 619 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 678
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W K L R V G I
Sbjct: 679 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKLLTRLHVTYEGTI 727
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 728 EGNGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 787
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 788 EQYSEYTGYAETYRWARSHEDRSERDDWQRRTTEIVAIDAL---HFRRYLDQFVPEKIRR 844
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 845 ELNKAYCGFLR 855
Score = 57.8 bits (138), Expect = 8e-07
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 858 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAVAERD-VVYFTFGDSEL 908
>UniRef100_UPI000027C995 UPI000027C995 UniRef100 entry
Length = 327
Score = 165 bits (417), Expect = 4e-39
Identities = 93/267 (34%), Positives = 150/267 (55%), Gaps = 13/267 (4%)
Query: 129 GLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDEL-FASLYNDYT 187
G+ +L A + L+Q I+ LLA +F C FP + + + F L++ ++
Sbjct: 5 GIPLLQQGRAAAITLSQAQISCLLANAFFCTFPHRNSTSFHSDYHTYPSINFTRLFSHWS 64
Query: 188 QKQEDKIWCIIHYFQRITSNMPK--GVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLC 245
+++ +K+ I+HYF T K G+V+FER+ L D A WS + +
Sbjct: 65 ERKMEKLKAIVHYFHVATDEKTKLDGLVTFERRCLANTD---------ARTWSCCKEEMN 115
Query: 246 RFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMA 305
+ V S G IE S ++VDFA+ +LGGG L G +QEEI F++SPELI +F +
Sbjct: 116 KLYVSSCGAIETEGSGLLQVDFASSWLGGGVLDSGLLQEEILFLMSPELIVSRLFTEKLQ 175
Query: 306 DNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL-CGPGMRQY 364
DNE + + G ++FS+Y+GY +FR+ G Y D D + RR+ +++A+DAL G QY
Sbjct: 176 DNECVIVTGCQQFSTYSGYGDTFRWKGPYADPTGRDGWARRQRQVLAMDALRFTHGRDQY 235
Query: 365 REKFLLREINKAFCGFLQQSQYQRDQK 391
K ++RE+NKA+CGF + + +R ++
Sbjct: 236 SMKLVVRELNKAYCGFKRCEEERRGEE 262
Score = 55.8 bits (133), Expect = 3e-06
Identities = 24/46 (52%), Positives = 36/46 (78%), Gaps = 1/46 (2%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQN 513
+ATG WGCGAF GDP++K +IQ +AA++AGR +A++TF L++
Sbjct: 265 IATGKWGCGAFKGDPQLKAVIQLMAAARAGRG-LAFFTFKDEKLEH 309
>UniRef100_UPI00003AE137 UPI00003AE137 UniRef100 entry
Length = 893
Score = 153 bits (387), Expect = 1e-35
Identities = 100/310 (32%), Positives = 157/310 (50%), Gaps = 37/310 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F D+++ E + F+ +LP + L L LP++ + +L
Sbjct: 500 FCDKVLEDAEAQHLFQSILPDMVKLALCLPTICTQP------------------IPLLKQ 541
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+ LA +F C FP + K D F L+ + ++ +K+
Sbjct: 542 KMNHSITMSQEQIASFLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSPRKPEKLK 601
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L +P+ W S K L R V G I
Sbjct: 602 TLFCYFRRVTEKKPTGLVTFTRQCLQ--------EFPD---WERSQKKLSRLHVTYEGTI 650
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI + + NE + I G
Sbjct: 651 ESNGQGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLITEVLDHNECLIITGT 710
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D D + RR T IVAIDA R++ ++F + R
Sbjct: 711 EQYSEYTGYAETYRWARSHEDKTPRDEWQRRYTEIVAIDAF---HFRRFLDQFGPEKIRR 767
Query: 372 EINKA-FCGF 380
E+NKA +CGF
Sbjct: 768 ELNKASYCGF 777
Score = 59.3 bits (142), Expect = 3e-07
Identities = 26/40 (65%), Positives = 33/40 (82%), Gaps = 1/40 (2%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFG 507
+ATGNWGCGAFGGD +K +IQ LAA+++GR I Y+TFG
Sbjct: 790 IATGNWGCGAFGGDSRLKALIQILAAAESGRD-IVYFTFG 828
>UniRef100_UPI00002B206C UPI00002B206C UniRef100 entry
Length = 264
Score = 144 bits (364), Expect = 5e-33
Identities = 78/208 (37%), Positives = 123/208 (58%), Gaps = 12/208 (5%)
Query: 187 TQKQEDKIWCIIHYFQRITSNMPK--GVVSFERKVLPWEDDCIHISYPNASFWSTSVKPL 244
++++ +K+ I+HYF T K G+V+FER+ L D A WS + +
Sbjct: 1 SERKMEKLKAIVHYFHVATDEKTKLDGLVTFERRCLANTD---------ARTWSCCKEEM 51
Query: 245 CRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSM 304
+ V S G IE S ++VDFA+ +LGGG L G +QEEI F++SPELI +F +
Sbjct: 52 NKLYVSSCGAIETEGSGLLQVDFASSWLGGGVLDSGLLQEEILFLMSPELIVSRLFTEKL 111
Query: 305 ADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL-CGPGMRQ 363
DNE + + G ++FS+Y+GY +FR+ G Y D D + RR+ +++A+DAL G Q
Sbjct: 112 QDNECVIVTGCQQFSTYSGYGDTFRWKGPYADPTGRDGWARRQRQVLAMDALRFTHGRDQ 171
Query: 364 YREKFLLREINKAFCGFLQQSQYQRDQK 391
Y K ++RE+NKA+CGF + + +R ++
Sbjct: 172 YSMKLVVRELNKAYCGFKRCEEERRGEE 199
Score = 55.8 bits (133), Expect = 3e-06
Identities = 24/46 (52%), Positives = 36/46 (78%), Gaps = 1/46 (2%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQN 513
+ATG WGCGAF GDP++K +IQ +AA++AGR +A++TF L++
Sbjct: 202 IATGKWGCGAFKGDPQLKAVIQLMAAARAGRG-LAFFTFRDEKLEH 246
>UniRef100_UPI0000318FC5 UPI0000318FC5 UniRef100 entry
Length = 370
Score = 139 bits (349), Expect = 3e-31
Identities = 78/203 (38%), Positives = 119/203 (58%), Gaps = 18/203 (8%)
Query: 182 LYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSV 241
L+ + ++ +K+ ++ YF+ IT MP G+V+F RK L PN W +S
Sbjct: 59 LFEGSSSRKIEKLKTLMCYFKSITEQMPTGLVTFTRKRLD--------KPPN---WRSSQ 107
Query: 242 KPLCRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFL 301
L + + G IED ++VDFAN+++GGG G VQEEIRF+I+PELI +F
Sbjct: 108 MRLTKLHITCEGTIEDDGYGMLQVDFANQFVGGGVTSAGLVQEEIRFLINPELIVSRLFT 167
Query: 302 PSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGM 361
++ DN+ + I G +++S YTGYA ++++ G + D D + RR T IVA+DAL
Sbjct: 168 EALDDNDCLIITGTQQYSKYTGYAQTYQWRGGHHDTTPRDGWQRRCTEIVAMDAL---QF 224
Query: 362 RQYREKF----LLREINKAFCGF 380
R + E+F + RE+NKA+CGF
Sbjct: 225 RNFLEQFHPLKINRELNKAYCGF 247
Score = 64.7 bits (156), Expect = 7e-09
Identities = 34/72 (47%), Positives = 44/72 (60%), Gaps = 11/72 (15%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQ----------NLDKI 517
VATGNWGCG FGGD +K ++Q LAA++AGR +AY+TFG L L+ I
Sbjct: 260 VATGNWGCGVFGGDTRLKALLQMLAAAEAGRD-VAYFTFGDSQLMTDVHNMHSFLTLNNI 318
Query: 518 LVGVVAGLVLDF 529
VG V GL+ +
Sbjct: 319 SVGEVYGLLQQY 330
>UniRef100_Q613D5 Hypothetical protein CBG16441 [Caenorhabditis briggsae]
Length = 331
Score = 110 bits (274), Expect = 1e-22
Identities = 74/204 (36%), Positives = 105/204 (51%), Gaps = 19/204 (9%)
Query: 192 DKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKS 251
+K+ + YF RI +N PKG VSF K + E+ + P+ +
Sbjct: 72 EKLRFLFAYFDRIATNPPKGCVSFRLKTVRNEEF-------KEEWIKNKCNPMPDVRIYD 124
Query: 252 SGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAID 311
+IE+ + T ++DFANEYLGGG L+ G EEIRF++ PE+I GM+ M D +AI
Sbjct: 125 QMVIEETALCT-QIDFANEYLGGGVLKSGA--EEIRFLMCPEMIVGMLLTDKMDDTQAIS 181
Query: 312 IVGVERFSSYTGYASSFRF---SGDYV---DDKEVDPFGRRKTRIVAIDAL---CGPGMR 362
IVG +S YTGYA+ ++ SG Y D + D +GR + VAIDA+
Sbjct: 182 IVGAYVYSGYTGYANKLKWRPLSGRYARQNDARLRDKYGRLRVETVAIDAMYFKTKAIAE 241
Query: 363 QYREKFLLREINKAFCGFLQQSQY 386
Q L RE+ KA+ GF Q +
Sbjct: 242 QVTSDSLTREMKKAYTGFAQSEHF 265
Score = 51.2 bits (121), Expect = 8e-05
Identities = 25/66 (37%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 448 QEIRNSQNDYDMMENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFG 507
+E++ + + E+ I + TG WGCGAF G +K +IQ +AAS AGRP + + FG
Sbjct: 250 REMKKAYTGFAQSEHFGQIPIVTGFWGCGAFNGHKPLKFLIQLMAASIAGRP-LQFCIFG 308
Query: 508 SGALQN 513
+ N
Sbjct: 309 DTVMGN 314
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 902,598,979
Number of Sequences: 2790947
Number of extensions: 38205638
Number of successful extensions: 106511
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 106341
Number of HSP's gapped (non-prelim): 91
length of query: 547
length of database: 848,049,833
effective HSP length: 132
effective length of query: 415
effective length of database: 479,644,829
effective search space: 199052604035
effective search space used: 199052604035
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC122162.2


