

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.9 + phase: 2 /pseudo/partial
(223 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
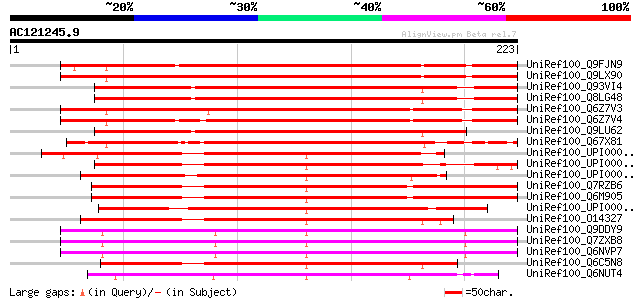
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJN9 Poly(A)-binding protein II-like [Arabidopsis th... 293 2e-78
UniRef100_Q9LX90 RNA binding protein-like [Arabidopsis thaliana] 286 4e-76
UniRef100_Q93VI4 Hypothetical protein At5g51120 [Arabidopsis tha... 258 8e-68
UniRef100_Q8LG48 Contains similarity to poly(A)-binding protein ... 258 8e-68
UniRef100_Q6Z7V3 Putative poly(A) binding protein [Oryza sativa] 253 3e-66
UniRef100_Q6Z7V4 Putative poly(A) binding protein [Oryza sativa] 253 3e-66
UniRef100_Q9LU62 Similarity to poly(A)-binding protein II [Arabi... 241 8e-63
UniRef100_Q67X81 Putative poly(A) binding protein II [Oryza sativa] 229 4e-59
UniRef100_UPI000023F69C UPI000023F69C UniRef100 entry 163 3e-39
UniRef100_UPI0000234412 UPI0000234412 UniRef100 entry 156 4e-37
UniRef100_UPI00003C1908 UPI00003C1908 UniRef100 entry 155 6e-37
UniRef100_Q7RZB6 Hypothetical protein [Neurospora crassa] 155 9e-37
UniRef100_Q6M905 Probable RRM-type RNA binding protein [Neurospo... 155 9e-37
UniRef100_UPI0000219B7F UPI0000219B7F UniRef100 entry 153 4e-36
UniRef100_O14327 Pab2 protein [Schizosaccharomyces pombe] 148 1e-34
UniRef100_Q9DDY9 Poly(A) binding protein II (PABPII protein) (Nu... 145 1e-33
UniRef100_Q7ZXB8 Pabpn1-prov protein [Xenopus laevis] 144 1e-33
UniRef100_Q6NVP7 Hypothetical protein MGC69525 [Xenopus tropicalis] 144 2e-33
UniRef100_Q6C5N8 Similar to tr|Q9P4V6 Candida boidinii RNA-bindi... 143 3e-33
UniRef100_Q6NUT4 Zgc:85979 [Brachydanio rerio] 142 5e-33
>UniRef100_Q9FJN9 Poly(A)-binding protein II-like [Arabidopsis thaliana]
Length = 220
Score = 293 bits (751), Expect = 2e-78
Identities = 152/204 (74%), Positives = 173/204 (84%), Gaps = 7/204 (3%)
Query: 23 D*DME--NDDVDMIGADNNEA-DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPAN 79
D DME N D+DM AD++ +LD+MKKRLKEMEDEAAAL+EMQAKVEKEMG+ QDPA+
Sbjct: 20 DGDMEALNPDLDMAAADDDAVKELDEMKKRLKEMEDEAAALREMQAKVEKEMGA-QDPAS 78
Query: 80 ASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVE 139
+A+Q +EEVDARS+FVGNVDYACTPE+VQQHFQ+CGTV+RVTI TDKFGQPKG+AYVE
Sbjct: 79 IAANQAGKEEVDARSVFVGNVDYACTPEEVQQHFQTCGTVHRVTILTDKFGQPKGFAYVE 138
Query: 140 FVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPF 199
FVEVEAVQEAL LNESELHGRQLKV KRTNVPG+KQFR RR NPYMG+R R P+ P+
Sbjct: 139 FVEVEAVQEALQLNESELHGRQLKVLQKRTNVPGLKQFRGRR-FNPYMGYRFRRPFMSPY 197
Query: 200 AYAPAPYGYGKIPRFRMGMRYSPY 223
Y +PYGYGK PRFR MRY PY
Sbjct: 198 MY--SPYGYGKAPRFRRPMRYMPY 219
>UniRef100_Q9LX90 RNA binding protein-like [Arabidopsis thaliana]
Length = 217
Score = 286 bits (731), Expect = 4e-76
Identities = 143/202 (70%), Positives = 168/202 (82%), Gaps = 4/202 (1%)
Query: 23 D*DMENDDVDMIGADNNEA-DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
D D+ + D+DM AD + +L +MK+RLKEME+EAAAL+EMQAKVEKEMG+ QDPA+ +
Sbjct: 18 DTDVPDPDIDMSAADEDAVTELAEMKRRLKEMEEEAAALREMQAKVEKEMGATQDPASMA 77
Query: 82 ASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFV 141
A+Q +EEVDARS++VGNVDYACTPE+VQ HFQ+CGTVNRVTI DKFGQPKG+AYVEFV
Sbjct: 78 ANQEGKEEVDARSVYVGNVDYACTPEEVQLHFQTCGTVNRVTILMDKFGQPKGFAYVEFV 137
Query: 142 EVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAY 201
EVEAVQEAL LNESELHGRQLKV+ KRTNVPGMKQ+ P R NP MG+R R P+ PP+ Y
Sbjct: 138 EVEAVQEALQLNESELHGRQLKVSPKRTNVPGMKQYHPGR-FNPSMGYRFRRPFVPPYFY 196
Query: 202 APAPYGYGKIPRFRMGMRYSPY 223
+PYGYGK PRFR MRY PY
Sbjct: 197 --SPYGYGKAPRFRRPMRYMPY 216
>UniRef100_Q93VI4 Hypothetical protein At5g51120 [Arabidopsis thaliana]
Length = 227
Score = 258 bits (659), Expect = 8e-68
Identities = 128/187 (68%), Positives = 151/187 (80%), Gaps = 9/187 (4%)
Query: 38 NNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFV 97
++ DL+DMKKR+KE+E+EA AL+EMQAK EK+MG+ QDP+ S +EEVD+RSI+V
Sbjct: 49 SSSRDLEDMKKRIKEIEEEAGALREMQAKAEKDMGASQDPSGG-VSAAEKEEVDSRSIYV 107
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESEL 157
GNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVEVEAVQ +L+LNESEL
Sbjct: 108 GNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEVEAVQNSLILNESEL 167
Query: 158 HGRQLKVTAKRTNVPGMKQFRPR-RPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFRM 216
HGRQ+KV+AKRTNVPGM+QFR R RP P GF P+ P PY YG++PRFR
Sbjct: 168 HGRQIKVSAKRTNVPGMRQFRGRGRPFRPMRGFMPGVPFYP-------PYAYGRVPRFRR 220
Query: 217 GMRYSPY 223
MRY PY
Sbjct: 221 PMRYRPY 227
>UniRef100_Q8LG48 Contains similarity to poly(A)-binding protein II [Arabidopsis
thaliana]
Length = 227
Score = 258 bits (659), Expect = 8e-68
Identities = 128/187 (68%), Positives = 151/187 (80%), Gaps = 9/187 (4%)
Query: 38 NNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFV 97
++ DL+DMKKR+KE+E+EA AL+EMQAK EK+MG+ QDP+ S +EEVD+RSI+V
Sbjct: 49 SSSRDLEDMKKRIKEIEEEAGALREMQAKAEKDMGASQDPSGG-VSAAEKEEVDSRSIYV 107
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESEL 157
GNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVEVEAVQ +L+LNESEL
Sbjct: 108 GNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEVEAVQNSLILNESEL 167
Query: 158 HGRQLKVTAKRTNVPGMKQFRPR-RPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFRM 216
HGRQ+KV+AKRTNVPGM+QFR R RP P GF P+ P PY YG++PRFR
Sbjct: 168 HGRQIKVSAKRTNVPGMRQFRGRGRPFRPMRGFMPGVPFYP-------PYAYGRVPRFRR 220
Query: 217 GMRYSPY 223
MRY PY
Sbjct: 221 PMRYRPY 227
>UniRef100_Q6Z7V3 Putative poly(A) binding protein [Oryza sativa]
Length = 216
Score = 253 bits (646), Expect = 3e-66
Identities = 135/205 (65%), Positives = 157/205 (75%), Gaps = 9/205 (4%)
Query: 23 D*DMENDDVDMIGADNNEA---DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPAN 79
D DM+ DVDM ++ A +LD MK+RLKEME+EAAAL++MQAKV KEM +
Sbjct: 16 DGDMDGADVDMASGGDDAAKLQELDQMKRRLKEMEEEAAALRDMQAKVAKEMQGGPPGGD 75
Query: 80 ASASQIN-REEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYV 138
SAS +E+VDARS++VGNVDYACTPE+VQQHFQ+CGTVNRVTI TDKFGQPKG+AYV
Sbjct: 76 PSASTAEAKEQVDARSVYVGNVDYACTPEEVQQHFQACGTVNRVTILTDKFGQPKGFAYV 135
Query: 139 EFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPP 198
EF+E EAVQEAL LNESELHGRQ+KV KRTNVPGMKQ RP R NPY G+ R+ AP
Sbjct: 136 EFLEQEAVQEALNLNESELHGRQIKVAPKRTNVPGMKQ-RPPRGYNPYHGYPYRSYGAPY 194
Query: 199 FAYAPAPYGYGKIPRFRMGMRYSPY 223
F PYGYG++PRFR MRY PY
Sbjct: 195 F----PPYGYGRVPRFRRPMRYRPY 215
>UniRef100_Q6Z7V4 Putative poly(A) binding protein [Oryza sativa]
Length = 212
Score = 253 bits (645), Expect = 3e-66
Identities = 135/204 (66%), Positives = 159/204 (77%), Gaps = 11/204 (5%)
Query: 23 D*DMENDDVDMIGADNNEA---DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPAN 79
D DM+ DVDM ++ A +LD MK+RLKEME+EAAAL++MQAKV KEM DP+
Sbjct: 16 DGDMDGADVDMASGGDDAAKLQELDQMKRRLKEMEEEAAALRDMQAKVAKEMQG-GDPSA 74
Query: 80 ASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVE 139
++A +E+VDARS++VGNVDYACTPE+VQQHFQ+CGTVNRVTI TDKFGQPKG+AYVE
Sbjct: 75 STAEA--KEQVDARSVYVGNVDYACTPEEVQQHFQACGTVNRVTILTDKFGQPKGFAYVE 132
Query: 140 FVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPF 199
F+E EAVQEAL LNESELHGRQ+KV KRTNVPGMKQ RP R NPY G+ R+ AP F
Sbjct: 133 FLEQEAVQEALNLNESELHGRQIKVAPKRTNVPGMKQ-RPPRGYNPYHGYPYRSYGAPYF 191
Query: 200 AYAPAPYGYGKIPRFRMGMRYSPY 223
PYGYG++PRFR MRY PY
Sbjct: 192 ----PPYGYGRVPRFRRPMRYRPY 211
>UniRef100_Q9LU62 Similarity to poly(A)-binding protein II [Arabidopsis thaliana]
Length = 267
Score = 241 bits (616), Expect = 8e-63
Identities = 118/165 (71%), Positives = 140/165 (84%), Gaps = 2/165 (1%)
Query: 38 NNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFV 97
++ DL+DMKKR+KE+E+EA AL+EMQAK EK+MG+ QDP+ S +EEVD+RSI+V
Sbjct: 49 SSSRDLEDMKKRIKEIEEEAGALREMQAKAEKDMGASQDPSGG-VSAAEKEEVDSRSIYV 107
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESEL 157
GNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVEVEAVQ +L+LNESEL
Sbjct: 108 GNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEVEAVQNSLILNESEL 167
Query: 158 HGRQLKVTAKRTNVPGMKQFRPR-RPSNPYMGFRGRTPYAPPFAY 201
HGRQ+KV+AKRTNVPGM+QFR R RP P GF P+ PP+AY
Sbjct: 168 HGRQIKVSAKRTNVPGMRQFRGRGRPFRPMRGFMPGVPFYPPYAY 212
>UniRef100_Q67X81 Putative poly(A) binding protein II [Oryza sativa]
Length = 204
Score = 229 bits (584), Expect = 4e-59
Identities = 133/204 (65%), Positives = 156/204 (76%), Gaps = 21/204 (10%)
Query: 26 MENDDVDMIGADNNEA----DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
++ +DVDM GA +EA +LD+MK+RLKEME+EA AL+EMQ KV KEM + DP NAS
Sbjct: 15 LDGEDVDM-GAPGDEAAKMQELDEMKRRLKEMEEEANALREMQTKVAKEMQGL-DP-NAS 71
Query: 82 ASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFV 141
+S+ ++EE+DARS++VGNVDYACTPE+VQQHF SCGTVNRVTI TDKFGQPKG+AYVEF+
Sbjct: 72 SSE-SKEEMDARSVYVGNVDYACTPEEVQQHFNSCGTVNRVTILTDKFGQPKGFAYVEFL 130
Query: 142 EVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRR--PSNPYMGFRGRTPYAPPF 199
EVEAVQEA+ LNESELHGRQ+KV KRTNVPGMKQ R R +PYM PY PF
Sbjct: 131 EVEAVQEAVKLNESELHGRQIKVAPKRTNVPGMKQPRGGRGFGGHPYM-----RPYGAPF 185
Query: 200 AYAPAPYGYGKIPRFRMGMRYSPY 223
PYGYG PRFR R PY
Sbjct: 186 Y---NPYGYG-YPRFRRPRR--PY 203
>UniRef100_UPI000023F69C UPI000023F69C UniRef100 entry
Length = 208
Score = 163 bits (413), Expect = 3e-39
Identities = 93/187 (49%), Positives = 119/187 (62%), Gaps = 25/187 (13%)
Query: 15 LDDGVPFC------D*DMENDDVDMIGAD---NNEADLDDMKKRLKEMEDEAAALKEMQA 65
+ DG P D D E+D+ G + E ++ MK+R+ EME+EA L+EMQA
Sbjct: 6 IQDGKPMVENQHEIDHDREHDEEQEQGTEMSATEEEEITAMKRRVAEMEEEAKKLREMQA 65
Query: 66 KVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIR 125
+E++ + D ++E +DARSIFVGNVDY+ +PED+Q HFQSCG++NRVTI
Sbjct: 66 TLEQQSAELAD---------DKESIDARSIFVGNVDYSASPEDIQGHFQSCGSINRVTIL 116
Query: 126 TDKF-GQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSN 184
DKF GQPKGYAYVEF E V +AL+LNES GR +KVT KRTNVPGM + R R
Sbjct: 117 LDKFTGQPKGYAYVEFTEPSLVAQALVLNESMFKGRSIKVTPKRTNVPGMSRGRGRG--- 173
Query: 185 PYMGFRG 191
GFRG
Sbjct: 174 ---GFRG 177
>UniRef100_UPI0000234412 UPI0000234412 UniRef100 entry
Length = 187
Score = 156 bits (394), Expect = 4e-37
Identities = 92/194 (47%), Positives = 121/194 (61%), Gaps = 36/194 (18%)
Query: 38 NNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFV 97
++E +++ MK+R+ EME EAA L+EMQA ++++ S+++ ++E++DARSIFV
Sbjct: 22 DDEEEIEAMKRRVAEMESEAAKLREMQATLDQQSESLKE---------DKEDIDARSIFV 72
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNESE 156
GNVDY +PE++Q HFQSCG++NRVTI DKF GQPKGYAYVEF E V +AL+LNES
Sbjct: 73 GNVDYGASPEEIQAHFQSCGSINRVTILLDKFTGQPKGYAYVEFAEPSLVAQALVLNESV 132
Query: 157 LHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPR--F 214
GR LKV KRTN+PGM R RGR YG G PR +
Sbjct: 133 FRGRNLKVVPKRTNLPGMSSRGRGRG-------RGR------------GYGRGGFPRGGY 173
Query: 215 RMGMR-----YSPY 223
R G R Y+PY
Sbjct: 174 RGGYRGRGRGYAPY 187
>UniRef100_UPI00003C1908 UPI00003C1908 UniRef100 entry
Length = 204
Score = 155 bits (393), Expect = 6e-37
Identities = 86/165 (52%), Positives = 112/165 (67%), Gaps = 10/165 (6%)
Query: 32 DMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVD 91
D+ + +A+++ MK+R+ EME EAA L+E+Q + G+ P ++ REEVD
Sbjct: 39 DVTAMTDEDAEIEAMKQRVAEMEAEAAKLRELQQAAGEAGGAGLHP-----TEEEREEVD 93
Query: 92 ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEAL 150
+RSI+VGNVDY TPE+VQQHFQSCGT+NRVTI DKF G PKG+AYVEF E V A+
Sbjct: 94 SRSIYVGNVDYGATPEEVQQHFQSCGTINRVTILCDKFTGHPKGFAYVEFAEPSLVANAM 153
Query: 151 LLNESELHGRQLKVTAKRTNVPGMK---QFRPRRPSNPYMGFRGR 192
+LNES GR +KVTAKRTN+PG+ + R R P+ G RGR
Sbjct: 154 VLNESLFRGRLIKVTAKRTNMPGLSARGRGRGRGRGRPFRG-RGR 197
>UniRef100_Q7RZB6 Hypothetical protein [Neurospora crassa]
Length = 419
Score = 155 bits (391), Expect = 9e-37
Identities = 88/188 (46%), Positives = 118/188 (61%), Gaps = 12/188 (6%)
Query: 37 DNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIF 96
+++E ++ MK+R+ EME+EAA L+EMQA ++++ + + N+E++D+RSIF
Sbjct: 243 NDDEEEISAMKRRVAEMEEEAAKLREMQASLDQQHHELTE---------NKEDIDSRSIF 293
Query: 97 VGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNES 155
VGNVDY+ +PE++Q HFQSCG++NRVTI DKF GQPKGYAYVEF E V +AL+LNES
Sbjct: 294 VGNVDYSASPEEIQAHFQSCGSINRVTILLDKFTGQPKGYAYVEFTEPSLVAQALVLNES 353
Query: 156 ELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFR 215
GR +KV KRTN+PGM R R GR Y P GY R R
Sbjct: 354 VFKGRNIKVVPKRTNIPGMS--RGGRGGRGSFRGGGRGGYFGGRGGFPPRGGYRGGYRGR 411
Query: 216 MGMRYSPY 223
Y+PY
Sbjct: 412 GRGGYAPY 419
>UniRef100_Q6M905 Probable RRM-type RNA binding protein [Neurospora crassa]
Length = 215
Score = 155 bits (391), Expect = 9e-37
Identities = 88/188 (46%), Positives = 118/188 (61%), Gaps = 12/188 (6%)
Query: 37 DNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIF 96
+++E ++ MK+R+ EME+EAA L+EMQA ++++ + + N+E++D+RSIF
Sbjct: 39 NDDEEEISAMKRRVAEMEEEAAKLREMQASLDQQHHELTE---------NKEDIDSRSIF 89
Query: 97 VGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNES 155
VGNVDY+ +PE++Q HFQSCG++NRVTI DKF GQPKGYAYVEF E V +AL+LNES
Sbjct: 90 VGNVDYSASPEEIQAHFQSCGSINRVTILLDKFTGQPKGYAYVEFTEPSLVAQALVLNES 149
Query: 156 ELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFR 215
GR +KV KRTN+PGM R R GR Y P GY R R
Sbjct: 150 VFKGRNIKVVPKRTNIPGMS--RGGRGGRGSFRGGGRGGYFGGRGGFPPRGGYRGGYRGR 207
Query: 216 MGMRYSPY 223
Y+PY
Sbjct: 208 GRGGYAPY 215
>UniRef100_UPI0000219B7F UPI0000219B7F UniRef100 entry
Length = 213
Score = 153 bits (386), Expect = 4e-36
Identities = 84/172 (48%), Positives = 112/172 (64%), Gaps = 12/172 (6%)
Query: 40 EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGN 99
+ ++ MKKR+ EME+EAA L+EMQA ++ + + S +R++VD+RSIFVGN
Sbjct: 48 QQEISAMKKRVAEMEEEAAKLREMQASLDSQ--------SQELSTESRDDVDSRSIFVGN 99
Query: 100 VDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNESELH 158
VDY+ +PE++Q HFQSCG++NRVTI DKF GQPKG+AYVEF E V +AL+LNES
Sbjct: 100 VDYSASPEEIQAHFQSCGSINRVTILLDKFTGQPKGFAYVEFTEPSLVAQALVLNESVFK 159
Query: 159 GRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGYGK 210
GR +KV+ KRTN PGM + R R G GR + P Y G G+
Sbjct: 160 GRNIKVSPKRTNYPGMSRGRGRGRGR---GGFGRGGFPPRGGYRGGYRGRGR 208
>UniRef100_O14327 Pab2 protein [Schizosaccharomyces pombe]
Length = 166
Score = 148 bits (373), Expect = 1e-34
Identities = 85/173 (49%), Positives = 110/173 (63%), Gaps = 18/173 (10%)
Query: 32 DMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVD 91
D D E +L +MK+R+ EME EAA L+ MQ +++ E ++++ ++E +D
Sbjct: 3 DQDALDTQEKELLEMKERVAEMEAEAAKLRAMQEQLDNETEALRN---------DKESID 53
Query: 92 ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEAL 150
A+S++VGNVDY+ TPE++Q HF SCG+VNRVTI DKF G PKG+AY+EF E V AL
Sbjct: 54 AQSVYVGNVDYSVTPEELQSHFASCGSVNRVTILCDKFTGHPKGFAYIEFSEPSLVPNAL 113
Query: 151 LLNESELHGRQLKVTAKRTNVPGMKQFRPR-------RPSNPYMG-FRGRTPY 195
LLN S LH R LKVT KRTNVPGM + R R R Y G RG PY
Sbjct: 114 LLNGSMLHERPLKVTPKRTNVPGMSRGRGRGRGRGRGRGRGGYRGRARGFAPY 166
>UniRef100_Q9DDY9 Poly(A) binding protein II (PABPII protein) (Nuclear poly(A)
binding protein 2) [Xenopus laevis]
Length = 296
Score = 145 bits (365), Expect = 1e-33
Identities = 87/209 (41%), Positives = 119/209 (56%), Gaps = 8/209 (3%)
Query: 23 D*DMENDDVDMIGADNN--EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANA 80
D ++E ++ + D + +L+ +K R++EME+EA LKE+Q +VEK+M P NA
Sbjct: 88 DEELEEEEPGELTGDQTIEDPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNA 147
Query: 81 SASQINREE---VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYA 136
++ EE DARSI+VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+A
Sbjct: 148 GPVIMSVEEKMEADARSIYVGNVDYGATAEELEAHFHGCGSVNRVTILCDKFTGHPKGFA 207
Query: 137 YVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYA 196
Y+EF + E+V+ +L L+ES GRQ+KV KRTN PG+ P Y
Sbjct: 208 YIEFCDKESVRTSLALDESLFRGRQIKVVPKRTNRPGISTTDRGFPRARYRARASSYSSR 267
Query: 197 PPF--AYAPAPYGYGKIPRFRMGMRYSPY 223
F Y P P G R R Y+PY
Sbjct: 268 SRFYSGYTPRPRGRVYRGRARATSWYTPY 296
>UniRef100_Q7ZXB8 Pabpn1-prov protein [Xenopus laevis]
Length = 295
Score = 144 bits (364), Expect = 1e-33
Identities = 87/209 (41%), Positives = 119/209 (56%), Gaps = 8/209 (3%)
Query: 23 D*DMENDDVDMIGADNN--EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANA 80
D ++E ++ + D + +L+ +K R++EME+EA LKE+Q +VEK+M P NA
Sbjct: 87 DEELEEEEPGELTGDQTIEDPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNA 146
Query: 81 SASQINREE---VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYA 136
++ EE DARSI+VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+A
Sbjct: 147 GPVIMSVEEKMEADARSIYVGNVDYGATAEELEAHFHGCGSVNRVTILCDKFTGHPKGFA 206
Query: 137 YVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYA 196
Y+EF + E+V+ +L L+ES GRQ+KV KRTN PG+ P Y
Sbjct: 207 YIEFSDKESVRTSLALDESLFRGRQIKVVPKRTNRPGISTTDRGFPRARYRARASSYSSR 266
Query: 197 PPF--AYAPAPYGYGKIPRFRMGMRYSPY 223
F Y P P G R R Y+PY
Sbjct: 267 SRFYSGYTPRPRGRVYRGRARATSWYTPY 295
>UniRef100_Q6NVP7 Hypothetical protein MGC69525 [Xenopus tropicalis]
Length = 296
Score = 144 bits (363), Expect = 2e-33
Identities = 86/209 (41%), Positives = 120/209 (57%), Gaps = 8/209 (3%)
Query: 23 D*DMENDDVDMIGADNN--EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANA 80
D ++E ++ + D + +L+ +K R++EME+EA LKE+Q +VEK+M P NA
Sbjct: 88 DEELEEEEPGELTGDQTIEDPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNA 147
Query: 81 SASQINREE---VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYA 136
++ EE DARSI+VGNVDY T E+++ HF CG+VNRVTI DK+ G PKG+A
Sbjct: 148 GPVIMSIEEKMEADARSIYVGNVDYGATAEELEAHFHGCGSVNRVTILCDKYTGHPKGFA 207
Query: 137 YVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYA 196
Y+EF + E+V+ +L L+ES GRQ+KV KRTN PG+ P Y
Sbjct: 208 YIEFSDKESVRTSLALDESLFRGRQIKVVPKRTNRPGISTTDRGFPRARYRARASSYSSR 267
Query: 197 PPF--AYAPAPYGYGKIPRFRMGMRYSPY 223
F Y P P G R R+ Y+PY
Sbjct: 268 SRFYSGYTPRPRGRVYRGRARVTSWYTPY 296
>UniRef100_Q6C5N8 Similar to tr|Q9P4V6 Candida boidinii RNA-binding protein [Yarrowia
lipolytica]
Length = 205
Score = 143 bits (361), Expect = 3e-33
Identities = 76/168 (45%), Positives = 109/168 (64%), Gaps = 20/168 (11%)
Query: 41 ADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNV 100
A+++ M+ RL+++++EA+ L+ MQ K +++ ++EE+DARSI+VGNV
Sbjct: 46 AEMEAMQARLQQLQEEASKLESMQQDATKSSEDLKE---------DQEEIDARSIYVGNV 96
Query: 101 DYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNESELHG 159
DY TP+++++HF+ CG+V RVTI DKF G PKGYAY+EF E + V +A+ LN+SE
Sbjct: 97 DYGATPQELEKHFEKCGSVQRVTILMDKFTGNPKGYAYIEFAEPKDVAQAVTLNDSEFRQ 156
Query: 160 RQLKVTAKRTNVPGMKQFRPR----------RPSNPYMGFRGRTPYAP 197
RQLKV+AKRTN+PG K R R R + G RGR YAP
Sbjct: 157 RQLKVSAKRTNIPGFKGGRGRGRGRGGRGGFRGRGGFRGGRGRGGYAP 204
>UniRef100_Q6NUT4 Zgc:85979 [Brachydanio rerio]
Length = 284
Score = 142 bits (359), Expect = 5e-33
Identities = 87/192 (45%), Positives = 115/192 (59%), Gaps = 14/192 (7%)
Query: 35 GADNNEADLDD-----MKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE- 88
G D EA ++D +K R++EME+EA LKE+Q +VEK+M PA I +
Sbjct: 34 GLDEGEAAIEDPELEAIKARVREMEEEAEKLKELQNEVEKQMNLSPPPAGPVIMSIEEKI 93
Query: 89 EVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQ 147
E D RSI+VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+AY+EF + E+V+
Sbjct: 94 EADGRSIYVGNVDYGATAEELEAHFHGCGSVNRVTILCDKFTGHPKGFAYIEFADKESVR 153
Query: 148 EALLLNESELHGRQLKVTAKRTNVPGM----KQFRPRRPSNPYMGFRGRTPYAPPFAYAP 203
A+ L+ES GRQ+KV AKRTN PG+ + F R + F R Y Y P
Sbjct: 154 TAMALDESLFRGRQIKVGAKRTNRPGISTTDRGFPRARFRSRGGNFSSRARYYS--GYTP 211
Query: 204 APYGYGKIPRFR 215
P G G+ RF+
Sbjct: 212 -PRGRGRAFRFQ 222
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 376,103,898
Number of Sequences: 2790947
Number of extensions: 15327761
Number of successful extensions: 75198
Number of sequences better than 10.0: 2975
Number of HSP's better than 10.0 without gapping: 861
Number of HSP's successfully gapped in prelim test: 2116
Number of HSP's that attempted gapping in prelim test: 72465
Number of HSP's gapped (non-prelim): 4339
length of query: 223
length of database: 848,049,833
effective HSP length: 123
effective length of query: 100
effective length of database: 504,763,352
effective search space: 50476335200
effective search space used: 50476335200
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC121245.9


