

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.7 + phase: 0 /pseudo
(69 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
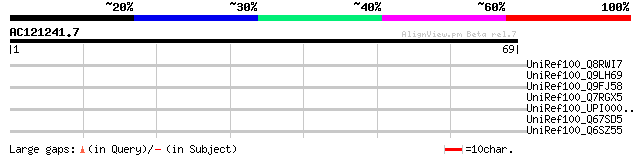
UniRef100_Q8RWI7 Hypothetical protein At3g14860 [Arabidopsis tha... 38 0.076
UniRef100_Q9LH69 Arabidopsis thaliana genomic DNA, chromosome 3,... 38 0.076
UniRef100_Q9FJ58 Arabidopsis thaliana genomic DNA, chromosome 5,... 35 0.64
UniRef100_Q7RGX5 Putative yir1 protein [Plasmodium yoelii yoelii] 31 7.1
UniRef100_UPI0000316BA7 UPI0000316BA7 UniRef100 entry 31 9.3
UniRef100_Q67SD5 Putative Fe-S oxidoreductase [Symbiobacterium t... 31 9.3
UniRef100_Q6SZ55 LPXTG anchored putative adhesin [Streptococcus ... 31 9.3
>UniRef100_Q8RWI7 Hypothetical protein At3g14860 [Arabidopsis thaliana]
Length = 492
Score = 37.7 bits (86), Expect = 0.076
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 307 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 345
>UniRef100_Q9LH69 Arabidopsis thaliana genomic DNA, chromosome 3, BAC clone: T21E2
[Arabidopsis thaliana]
Length = 511
Score = 37.7 bits (86), Expect = 0.076
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 326 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 364
>UniRef100_Q9FJ58 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MTE17
[Arabidopsis thaliana]
Length = 439
Score = 34.7 bits (78), Expect = 0.64
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 6/72 (8%)
Query: 4 QPSERDFKGEASSDKSKSTLERTKEETGWPSFRQ-VVDLLKLSLEEFTSSILKF-----M 57
Q SE + GE S +K+ ++ K E +Q +VD+ S+++FT S+ K +
Sbjct: 345 QESESEASGETSEEKTVKSVLTVKVEPESKVVQQDIVDMYMKSMQQFTDSLAKMKLPLDI 404
Query: 58 SKPHTTERSSSD 69
P +E SSSD
Sbjct: 405 DSPTKSENSSSD 416
>UniRef100_Q7RGX5 Putative yir1 protein [Plasmodium yoelii yoelii]
Length = 313
Score = 31.2 bits (69), Expect = 7.1
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 6/52 (11%)
Query: 17 DKSKSTLERTKEETGWPSFRQVV----DLLKLSLEEFTS--SILKFMSKPHT 62
D K + KE+TG+ +F++++ DLL + E+ + KF+ K HT
Sbjct: 123 DNGKDCNSKFKEKTGYNNFKEIIDKRKDLLNIKFEDISKLYDAFKFLCKMHT 174
>UniRef100_UPI0000316BA7 UPI0000316BA7 UniRef100 entry
Length = 181
Score = 30.8 bits (68), Expect = 9.3
Identities = 19/79 (24%), Positives = 39/79 (49%), Gaps = 10/79 (12%)
Query: 1 MLQQPSERDFKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSS-------- 52
ML++P K EA+ ++ STL + P+ +++++ L ++ SS
Sbjct: 85 MLKKPVTEKTKKEATKEEIVSTLNNGESSKLSPAVKKLINENDLDSDKIISSRSDGRLTK 144
Query: 53 --ILKFMSKPHTTERSSSD 69
+ FM+ P TT+++S +
Sbjct: 145 SDVFDFMNNPSTTDKTSKE 163
>UniRef100_Q67SD5 Putative Fe-S oxidoreductase [Symbiobacterium thermophilum]
Length = 636
Score = 30.8 bits (68), Expect = 9.3
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 9/62 (14%)
Query: 1 MLQQPSERDFKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKP 60
+L P+E D EA ++K + LE +EE G R+V T S+ F+ KP
Sbjct: 413 ILGLPTETDADLEAIAEKGRWALEIAEEELGRQEARKV---------RVTISVSVFVPKP 463
Query: 61 HT 62
HT
Sbjct: 464 HT 465
>UniRef100_Q6SZ55 LPXTG anchored putative adhesin [Streptococcus pyogenes]
Length = 1123
Score = 30.8 bits (68), Expect = 9.3
Identities = 20/67 (29%), Positives = 35/67 (51%), Gaps = 2/67 (2%)
Query: 3 QQPSERDFKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKPHT 62
++ SE+ + +A+ + +K LE K+E + +V+ K LE+F I K K T
Sbjct: 157 KESSEKVTELKANLESAKKDLE--KKEADYVKENALVERDKKDLEKFEKEIAKAREKKQT 214
Query: 63 TERSSSD 69
TE++ D
Sbjct: 215 TEKAIKD 221
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.306 0.122 0.322
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 93,912,267
Number of Sequences: 2790947
Number of extensions: 2629898
Number of successful extensions: 7448
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 7448
Number of HSP's gapped (non-prelim): 7
length of query: 69
length of database: 848,049,833
effective HSP length: 45
effective length of query: 24
effective length of database: 722,457,218
effective search space: 17338973232
effective search space used: 17338973232
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC121241.7


