

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
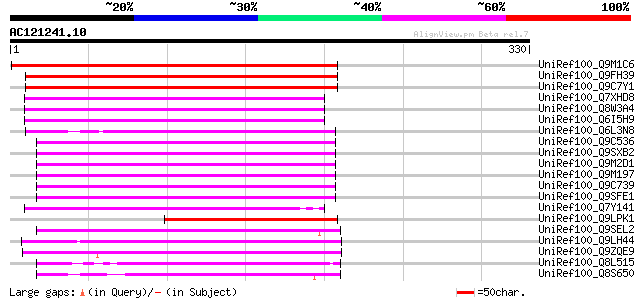
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thal... 233 5e-60
UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana] 205 1e-51
UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [... 205 1e-51
UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa] 134 3e-30
UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa] 133 8e-30
UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa] 130 6e-29
UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum] 129 8e-29
UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis t... 125 1e-27
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 125 1e-27
UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana] 125 1e-27
UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [... 125 1e-27
UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis t... 125 1e-27
UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana] 124 3e-27
UniRef100_Q7Y141 Putative polyprotein [Oryza sativa] 121 2e-26
UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana] 110 4e-23
UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera] 109 9e-23
UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidops... 107 6e-22
UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidop... 104 4e-21
UniRef100_Q8L515 P0018C10.19 protein [Oryza sativa] 100 5e-20
UniRef100_Q8S650 Putative retroelement [Oryza sativa] 93 1e-17
>UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thaliana]
Length = 1339
Score = 233 bits (594), Expect = 5e-60
Identities = 118/209 (56%), Positives = 153/209 (72%), Gaps = 2/209 (0%)
Query: 2 NDSSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQ-TEAQRKLI 60
+ S FV IP+FDG+YD W+M MENFLRS+E W L+E I + G +EAQR +
Sbjct: 2 SSSEKFVQPAIPRFDGYYDFWSMTMENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAV 61
Query: 61 KEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRD 120
+E KL +LK KN+LFQAI ++LETIL+K T K IW+SMK + GS +V RAQLQALR++
Sbjct: 62 EEAKLKDLKVKNFLFQAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKE 121
Query: 121 FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEES 180
FE+L MKEGE I+T+ RTLT+ NKMK + E+M Q+ I KILRS+ K +YVVCSIEES
Sbjct: 122 FELLAMKEGEKIDTFLGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEES 181
Query: 181 NNLDTMTIDELQSS-LVHEQRIKSNGEEE 208
N+L T++IDEL S LVHEQR+ + +EE
Sbjct: 182 NDLSTLSIDELHGSLLVHEQRLNGHVQEE 210
>UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1334
Score = 205 bits (522), Expect = 1e-51
Identities = 107/199 (53%), Positives = 138/199 (68%), Gaps = 1/199 (0%)
Query: 11 IIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKT 70
IIPKFDG Y+HWAM MEN +RSKE+WD+IE I E V T AQR + E+ + + K
Sbjct: 8 IIPKFDGDYEHWAMLMENLIRSKEWWDIIETGIPRPERNVILTGAQRTELAEKTVKDHKV 67
Query: 71 KNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
KNYLF +I +L+TIL K+T K++W+SMK + G++RV AQLQ LRR FE+L MK GE
Sbjct: 68 KNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLRRSFEVLEMKIGE 127
Query: 131 TINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDE 190
TI YFSR + I N M+ E M + + EKILR++V K YVVC+IEESNN+ +T+D
Sbjct: 128 TITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIEESNNIKELTVDG 187
Query: 191 LQSSL-VHEQRIKSNGEEE 208
LQSSL VHEQ + + EE
Sbjct: 188 LQSSLMVHEQNLSRHDVEE 206
>UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [Arabidopsis
thaliana]
Length = 1334
Score = 205 bits (522), Expect = 1e-51
Identities = 107/199 (53%), Positives = 138/199 (68%), Gaps = 1/199 (0%)
Query: 11 IIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKT 70
IIPKFDG Y+HWAM MEN +RSKE+WD+IE I E V T AQR + E+ + + K
Sbjct: 8 IIPKFDGDYEHWAMLMENLIRSKEWWDIIETGIPRPERNVILTGAQRTELAEKTVKDHKV 67
Query: 71 KNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
KNYLF +I +L+TIL K+T K++W+SMK + G++RV AQLQ LRR FE+L MK GE
Sbjct: 68 KNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLRRSFEVLEMKIGE 127
Query: 131 TINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDE 190
TI YFSR + I N M+ E M + + EKILR++V K YVVC+IEESNN+ +T+D
Sbjct: 128 TITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIEESNNIKELTVDG 187
Query: 191 LQSSL-VHEQRIKSNGEEE 208
LQSSL VHEQ + + EE
Sbjct: 188 LQSSLMVHEQNLSRHDVEE 206
>UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa]
Length = 1350
Score = 134 bits (337), Expect = 3e-30
Identities = 72/193 (37%), Positives = 111/193 (57%), Gaps = 2/193 (1%)
Query: 10 SIIPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNL 68
S++P F G +YD W++ M L S+ WD++EN AG T Q+K + E ++ +
Sbjct: 4 SMVPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDA 63
Query: 69 KTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKE 128
K + Q ++ + I+ K WD +K F GS +V+ +LQ LRR F+ L MKE
Sbjct: 64 KALFLIQQGVAESLFPRIIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKE 123
Query: 129 GETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTI 188
E + YF R + I N+M+ + E ++ + EKIL S+ K +Y+V +IEES +L T+TI
Sbjct: 124 SEKVKDYFPRVIEIVNQMRLYGEDINDQKVVEKILISLPEKYEYIVAAIEESKDLSTLTI 183
Query: 189 DELQSSL-VHEQR 200
+L SSL HE+R
Sbjct: 184 QQLMSSLESHEER 196
>UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa]
Length = 1167
Score = 133 bits (334), Expect = 8e-30
Identities = 71/193 (36%), Positives = 112/193 (57%), Gaps = 2/193 (1%)
Query: 10 SIIPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNL 68
S++P F G +YD W++ M L S+ WD++EN AG T Q+K + E ++ +
Sbjct: 4 SMVPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDA 63
Query: 69 KTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKE 128
K + Q ++ + I+ K WD +K F GS +V+ +LQ LRR F+ L MKE
Sbjct: 64 KALFLIQQGVAESLFPRIIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKE 123
Query: 129 GETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTI 188
E + YFSR + I N+++ + E ++ + E+IL S+ K +Y+V +IEES +L T+TI
Sbjct: 124 SEKVKDYFSRVIEIVNQIRLYGEDINDQKVVEEILISLPEKYEYIVAAIEESKDLSTLTI 183
Query: 189 DELQSSL-VHEQR 200
+L SSL HE+R
Sbjct: 184 QQLMSSLESHEER 196
>UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa]
Length = 1136
Score = 130 bits (326), Expect = 6e-29
Identities = 70/193 (36%), Positives = 110/193 (56%), Gaps = 2/193 (1%)
Query: 10 SIIPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNL 68
S++P F +YD W++ M L S+ WD++EN AG T Q+K + E ++ +
Sbjct: 4 SMVPVFARENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLVEDRMSDA 63
Query: 69 KTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKE 128
K + Q ++ + I+ K WD +K F S +V+ +LQ LRR F+ L MKE
Sbjct: 64 KALFLIQQGVAESLFPRIIGAKKSKEAWDKLKEEFQASQKVLAVKLQTLRRQFQNLQMKE 123
Query: 129 GETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTI 188
E + YFSR + I N+M+ + E ++ + +KIL S+ K +Y+V +IEES +L T+TI
Sbjct: 124 SEKVKDYFSRVIEIVNQMRLYGEDINDQKVVKKILISLPEKYEYIVAAIEESKDLSTLTI 183
Query: 189 DELQSSL-VHEQR 200
+L SSL HE+R
Sbjct: 184 QQLMSSLESHEER 196
>UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum]
Length = 1333
Score = 129 bits (325), Expect = 8e-29
Identities = 71/199 (35%), Positives = 117/199 (58%), Gaps = 11/199 (5%)
Query: 11 IIPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLK 69
+IP F G +Y W++ M+ +S+E WD +VE G+ + A + ++E + + K
Sbjct: 13 LIPIFRGENYQFWSLKMKTLFKSQELWD-------IVETGIPEGNANQ--MREHRKRDSK 63
Query: 70 TKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEG 129
+ QA+ ++ I +T K W+ +K + G ++VI +LQ LRRDFE L M E
Sbjct: 64 ALFTIQQALDDEIFPRISAVETSKQAWEILKQEYFGDDKVITVKLQTLRRDFETLFMNEN 123
Query: 130 ETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTID 189
E++ Y SRT I N+M+++ E + I+ K+LRS+ +K ++VV +IEES +L T + D
Sbjct: 124 ESVQGYLSRTSAIVNRMRSYGEKIDNQIVVSKVLRSLTTKFEHVVTAIEESKDLSTYSFD 183
Query: 190 ELQSSLV-HEQRIKSNGEE 207
EL SSL+ HE R+ + E+
Sbjct: 184 ELMSSLLAHEDRLNRSREK 202
>UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1320
Score = 125 bits (315), Expect = 1e-27
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 125 bits (315), Expect = 1e-27
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1352
Score = 125 bits (315), Expect = 1e-27
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [Arabidopsis
thaliana]
Length = 1272
Score = 125 bits (315), Expect = 1e-27
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 125 bits (315), Expect = 1e-27
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 124 bits (312), Expect = 3e-27
Identities = 59/190 (31%), Positives = 111/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLQMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G ++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGVDQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>UniRef100_Q7Y141 Putative polyprotein [Oryza sativa]
Length = 1335
Score = 121 bits (304), Expect = 2e-26
Identities = 68/192 (35%), Positives = 106/192 (54%), Gaps = 7/192 (3%)
Query: 10 SIIPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNL 68
S++P F G +YD W++ M L S+ WD++EN AG T Q+K + E ++ +
Sbjct: 4 SMVPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDA 63
Query: 69 KTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKE 128
K + Q ++ + I+ K WD +K F GS +V+ +LQ LRR F+ L MKE
Sbjct: 64 KALFLIQQGVAESLFPRIIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKE 123
Query: 129 GETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTI 188
E + YFSR + I N+M+ + E ++ + EKIL S+ K +Y+V + EES +L
Sbjct: 124 SEKVKDYFSRVIEIVNQMRLYGEDINDQKVVEKILISLPEKYEYIVAATEESKDLSK--- 180
Query: 189 DELQSSLVHEQR 200
D L+S HE+R
Sbjct: 181 DSLES---HEER 189
>UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana]
Length = 1207
Score = 110 bits (276), Expect = 4e-23
Identities = 60/111 (54%), Positives = 76/111 (68%), Gaps = 1/111 (0%)
Query: 99 MKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTII 158
MK + G++RV AQLQ LRR FE+L MK GETI YFSR + I N M+ E M + +
Sbjct: 1 MKRKYQGNDRVQSAQLQRLRRSFEVLEMKIGETITGYFSRVMEITNDMRNLGEDMPDSKV 60
Query: 159 TEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSL-VHEQRIKSNGEEE 208
EKILR++V K YVVC+IEESNN+ +T+D LQSSL VHEQ + + EE
Sbjct: 61 VEKILRTLVEKFTYVVCAIEESNNIKELTVDGLQSSLMVHEQNLSRHDVEE 111
>UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera]
Length = 581
Score = 109 bits (273), Expect = 9e-23
Identities = 61/195 (31%), Positives = 106/195 (54%), Gaps = 2/195 (1%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+Y+ WA+ M L++ + W+ +E V G + T AQ KL KE++ K K LF A
Sbjct: 17 NYETWAVRMTVHLQALDVWEAVEENYEVPPLGADPTVAQMKLHKERRTRKAKAKACLFAA 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+S + I+ D+ IW+ +K + G R+ Q+ L R+FE+ M+E + + Y +
Sbjct: 77 VSPSIFIKIMKIDSAAEIWEYLKEEYKGDERIKNMQVMNLIREFEMKKMRESDAVKDYAA 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSL-- 195
+ L+IANK++ + S I +KIL ++ K + + S+E S +L T+++ EL SL
Sbjct: 137 QLLSIANKVRLLGKEFSNEKIVQKILVTLPKKYEATISSLENSKDLSTISLTELLHSLEA 196
Query: 196 VHEQRIKSNGEEENG 210
V ++R+ G+ G
Sbjct: 197 VEQRRLMRQGDTAEG 211
>UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1499
Score = 107 bits (266), Expect = 6e-22
Identities = 60/206 (29%), Positives = 114/206 (55%), Gaps = 3/206 (1%)
Query: 8 VHSIIPKFDGH-YDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLM 66
+ +IP F+G Y W + M L++++ WD+IEN + + E + A + +Q +
Sbjct: 5 MQQVIPIFNGESYGFWKIKMITILKTRKLWDVIENGV-TSNSSPETSPALTRERDDQVMK 63
Query: 67 NLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHM 126
++ L A+S + I + W++++ F GS++V LQ LRR++E L M
Sbjct: 64 DMMALQILQSAVSDSIFPRIAPASSATEAWNALEMEFQGSSQVKMINLQTLRREYENLKM 123
Query: 127 KEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTM 186
+EGETIN + ++ + ++N+++ H E S + +KIL S+ + D +V +E++ +L T+
Sbjct: 124 EEGETINDFTTKLINLSNQLRVHGEEKSDYQVVQKILISVPQQFDSIVGVLEQTKDLSTL 183
Query: 187 TIDELQSSL-VHEQRIKSNGEEENGG 211
++ EL +L HE+R+ + N G
Sbjct: 184 SVTELIGTLKAHERRLNLREDRINEG 209
>UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1347
Score = 104 bits (259), Expect = 4e-21
Identities = 61/208 (29%), Positives = 110/208 (52%), Gaps = 5/208 (2%)
Query: 9 HSIIPKFDGH-YDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTE--AQRKLIKEQKL 65
H +IP FDG YD W++ M R+++ W ++E + V E+T A+ K ++E+ +
Sbjct: 6 HQVIPIFDGEKYDFWSIKMATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAV 65
Query: 66 MNLKTKNYLFQ-AISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEIL 124
N + Q A++ + I + K WD +K + GS +V +LQ+LRR++E L
Sbjct: 66 TNDTMALQILQTAVTDQIFSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENL 125
Query: 125 HMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLD 184
M + + I T+ + + + ++ H E + T + +KIL S+ +K D +V +E++ +LD
Sbjct: 126 KMYDNDNIKTFTDKLIVLEIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLD 185
Query: 185 TMTIDELQSSL-VHEQRIKSNGEEENGG 211
+T+ EL L E R+ + E G
Sbjct: 186 ALTMSELLGILKAQEARVTAREESTKEG 213
>UniRef100_Q8L515 P0018C10.19 protein [Oryza sativa]
Length = 278
Score = 100 bits (249), Expect = 5e-20
Identities = 65/194 (33%), Positives = 107/194 (54%), Gaps = 17/194 (8%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+Y WA+ ME + ++ WD IE A E + IK+ K + + LF A
Sbjct: 37 NYSTWAIKMEANMEAQGIWDAIE------PAADEAVD-----IKKDK----QARACLFGA 81
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ DVL+ I K T K +W+S+KT F G RV +A++Q L+ DF+ LHMK+ E+I+ +
Sbjct: 82 VPEDVLQQIAKKKTAKEVWESLKTRFLGVERVKKARVQTLKSDFKALHMKDTESIDEFAG 141
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSL-V 196
+ +ANK+ M +K+L S+ K +V+ +IE+ ++LD+M DE L
Sbjct: 142 KISGLANKLSDLGVTMDDDEQVKKLLDSVPDKFLHVIAAIEQFSDLDSMPFDEAIGRLKA 201
Query: 197 HEQRIKSNGEEENG 210
+E+RI+ +++NG
Sbjct: 202 YEERIRKR-DDKNG 214
>UniRef100_Q8S650 Putative retroelement [Oryza sativa]
Length = 299
Score = 92.8 bits (229), Expect = 1e-17
Identities = 55/196 (28%), Positives = 100/196 (50%), Gaps = 21/196 (10%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+Y W++ M+ ++++ WD VE G + R + + QA
Sbjct: 56 NYIEWSLLMKVNMQAQGVWD-------AVELGNTEHRIDRMALAA-----------ILQA 97
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ ++L T+ +KDT K W++++T G RV A+ Q+LR++FE++ MKEGE+++ +
Sbjct: 98 VPSEMLATLASKDTAKEAWEAVRTMRMGVERVREAKAQSLRKEFELIRMKEGESVDDFAM 157
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQ---SS 194
R + N + +M + ++ +K LR + SK V SIE+ +L T++++EL S
Sbjct: 158 RLTGLVNNIHTLGAMMEEEMVVKKFLRVVPSKYTQVAISIEQLIDLKTLSVEELTGRLKS 217
Query: 195 LVHEQRIKSNGEEENG 210
+ I NG E G
Sbjct: 218 VEERYEIDGNGSEAYG 233
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 485,678,358
Number of Sequences: 2790947
Number of extensions: 18376734
Number of successful extensions: 84411
Number of sequences better than 10.0: 242
Number of HSP's better than 10.0 without gapping: 153
Number of HSP's successfully gapped in prelim test: 89
Number of HSP's that attempted gapping in prelim test: 84148
Number of HSP's gapped (non-prelim): 266
length of query: 330
length of database: 848,049,833
effective HSP length: 128
effective length of query: 202
effective length of database: 490,808,617
effective search space: 99143340634
effective search space used: 99143340634
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC121241.10


