

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
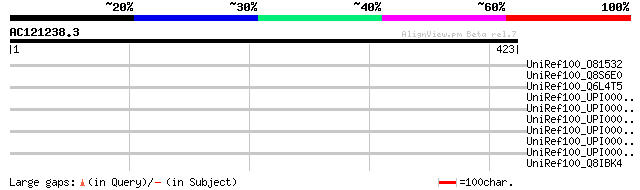
Query= AC121238.3 - phase: 0 /pseudo
(423 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81532 Mariner transposase [Glycine max] 47 0.001
UniRef100_Q8S6E0 Putative transposase [Oryza sativa] 38 0.48
UniRef100_Q6L4T5 Hypothetical protein P0478F09.8 [Oryza sativa] 35 3.1
UniRef100_UPI000046CDD7 UPI000046CDD7 UniRef100 entry 35 5.3
UniRef100_UPI00003ADC04 UPI00003ADC04 UniRef100 entry 34 9.0
UniRef100_UPI00003ADC03 UPI00003ADC03 UniRef100 entry 34 9.0
UniRef100_UPI00003ADC02 UPI00003ADC02 UniRef100 entry 34 9.0
UniRef100_UPI00003ADC01 UPI00003ADC01 UniRef100 entry 34 9.0
UniRef100_UPI00003ADC00 UPI00003ADC00 UniRef100 entry 34 9.0
UniRef100_Q8IBK4 Hypothetical protein MAL7P1.134 [Plasmodium fal... 34 9.0
>UniRef100_O81532 Mariner transposase [Glycine max]
Length = 425
Score = 46.6 bits (109), Expect = 0.001
Identities = 29/74 (39%), Positives = 44/74 (59%), Gaps = 3/74 (4%)
Query: 2 KRKRTFLDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWR--QKIETG 59
+RK L N +R I ++L+ S K+ +GV + ++SS+ V I IW+ ++ ET
Sbjct: 2 QRKVKMLSNEERITIYQLLLQKSVDGKLPQGVKESVASSFSVCRKTIDRIWKRAKESETH 61
Query: 60 DVCHKKTK-CGRKK 72
DV HKKTK GRK+
Sbjct: 62 DVSHKKTKNSGRKR 75
>UniRef100_Q8S6E0 Putative transposase [Oryza sativa]
Length = 565
Score = 38.1 bits (87), Expect = 0.48
Identities = 21/72 (29%), Positives = 34/72 (47%), Gaps = 7/72 (9%)
Query: 8 LDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQK-------IETGD 60
L N QRQ I E+L+ S K+ K + ++ + VS+ + IW++ ++
Sbjct: 144 LTNTQRQRIYELLLAKSKDGKLEKDTTRAVAQVFNVSIRTVQRIWKRAKLCIAHGVQVNV 203
Query: 61 VCHKKTKCGRKK 72
K CGRKK
Sbjct: 204 DSRKCYNCGRKK 215
>UniRef100_Q6L4T5 Hypothetical protein P0478F09.8 [Oryza sativa]
Length = 572
Score = 35.4 bits (80), Expect = 3.1
Identities = 15/58 (25%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 8 LDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKK 65
L N QR+ I ++L+ S K+ K + ++ + VS+ + IW++ +CH++
Sbjct: 151 LTNIQRRGIYQLLLQKSKDGKLEKHTTRLVAQEFHVSIRTVQRIWKR----AKICHEQ 204
>UniRef100_UPI000046CDD7 UPI000046CDD7 UniRef100 entry
Length = 178
Score = 34.7 bits (78), Expect = 5.3
Identities = 28/110 (25%), Positives = 48/110 (43%), Gaps = 24/110 (21%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKK------ 72
+L++ S KI+ +K + +++D Y ++ + + H K K + K
Sbjct: 74 ILLDGSLNAKISDFGIKEIEQCLDINIDYSYIVFPNNVIKFNNKHFKNKIKKIKIVNKGS 133
Query: 73 ---------KSHRYRKNT*--NTATQT*QSSISSRSIRVYHWTTNEVNKK 111
K+H Y+ NT N ++ T SS V+ WT+ EVNKK
Sbjct: 134 EDMLHVFSSKNHIYKYNTREINVSSNTHNSS-------VFFWTSPEVNKK 176
>UniRef100_UPI00003ADC04 UPI00003ADC04 UniRef100 entry
Length = 917
Score = 33.9 bits (76), Expect = 9.0
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 455 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 511
Query: 79 K 79
K
Sbjct: 512 K 512
>UniRef100_UPI00003ADC03 UPI00003ADC03 UniRef100 entry
Length = 911
Score = 33.9 bits (76), Expect = 9.0
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 500 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 556
Query: 79 K 79
K
Sbjct: 557 K 557
>UniRef100_UPI00003ADC02 UPI00003ADC02 UniRef100 entry
Length = 945
Score = 33.9 bits (76), Expect = 9.0
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 471 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 527
Query: 79 K 79
K
Sbjct: 528 K 528
>UniRef100_UPI00003ADC01 UPI00003ADC01 UniRef100 entry
Length = 951
Score = 33.9 bits (76), Expect = 9.0
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 476 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 532
Query: 79 K 79
K
Sbjct: 533 K 533
>UniRef100_UPI00003ADC00 UPI00003ADC00 UniRef100 entry
Length = 875
Score = 33.9 bits (76), Expect = 9.0
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 435 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 491
Query: 79 K 79
K
Sbjct: 492 K 492
>UniRef100_Q8IBK4 Hypothetical protein MAL7P1.134 [Plasmodium falciparum]
Length = 3351
Score = 33.9 bits (76), Expect = 9.0
Identities = 25/75 (33%), Positives = 35/75 (46%), Gaps = 11/75 (14%)
Query: 7 FLDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIET-----GDV 61
F D Y R+NI NS + INK ++KR + + ++ + R I DV
Sbjct: 1174 FYDAYDRENI-----NSKYVKVINKYIIKRNKNFVLIKKRIMNLMKRDNIHIRGYLRDDV 1228
Query: 62 CHKKTKCGRKKKSHR 76
K KC +KKK HR
Sbjct: 1229 LRNK-KCHKKKKDHR 1242
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.372 0.162 0.611
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 508,181,515
Number of Sequences: 2790947
Number of extensions: 15789127
Number of successful extensions: 82129
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 82124
Number of HSP's gapped (non-prelim): 13
length of query: 423
length of database: 848,049,833
effective HSP length: 130
effective length of query: 293
effective length of database: 485,226,723
effective search space: 142171429839
effective search space used: 142171429839
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC121238.3


