

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
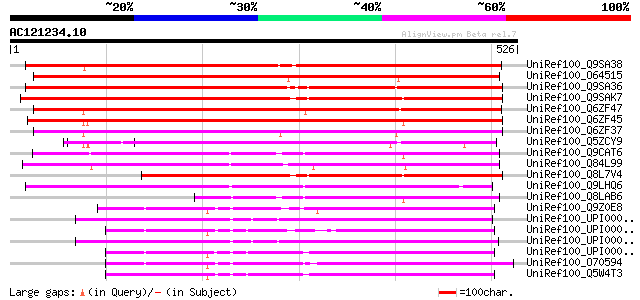
Query= AC121234.10 + phase: 0
(526 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SA38 F3O9.19 protein [Arabidopsis thaliana] 498 e-139
UniRef100_O64515 YUP8H12R.2 protein [Arabidopsis thaliana] 491 e-137
UniRef100_Q9SA36 F3O9.17 protein [Arabidopsis thaliana] 428 e-118
UniRef100_Q9SAK7 T8K14.17 protein [Arabidopsis thaliana] 421 e-116
UniRef100_Q6ZF47 Putative organic cation transporter [Oryza sativa] 398 e-109
UniRef100_Q6ZF45 Putative organic cation transporter [Oryza sativa] 382 e-104
UniRef100_Q6ZF37 Putative organic cation transporter [Oryza sativa] 380 e-104
UniRef100_Q5ZCY9 Putative solute carrier family 22 member 3 [Ory... 342 2e-92
UniRef100_Q9CAT6 Putative transporter; 29320-27598 [Arabidopsis ... 334 3e-90
UniRef100_Q84L99 Putative organic cation transport protein [Phas... 325 2e-87
UniRef100_Q8L7V4 At1g79410/T8K14_17 [Arabidopsis thaliana] 300 7e-80
UniRef100_Q9LHQ6 Organic anion transporter-like protein [Arabido... 250 8e-65
UniRef100_Q8LAB6 Hypothetical protein [Arabidopsis thaliana] 202 3e-50
UniRef100_Q9Z0E8 Organic cation/carnitine transporter 2 [Mus mus... 173 1e-41
UniRef100_UPI000036EE60 UPI000036EE60 UniRef100 entry 173 1e-41
UniRef100_UPI00003AABBA UPI00003AABBA UniRef100 entry 173 1e-41
UniRef100_UPI000036EE5E UPI000036EE5E UniRef100 entry 172 2e-41
UniRef100_UPI00003AABB9 UPI00003AABB9 UniRef100 entry 172 2e-41
UniRef100_O70594 Organic cation/carnitine transporter 2 [Rattus ... 172 2e-41
UniRef100_Q5W4T3 OCTN2 protein [Gallus gallus] 172 3e-41
>UniRef100_Q9SA38 F3O9.19 protein [Arabidopsis thaliana]
Length = 518
Score = 498 bits (1283), Expect = e-139
Identities = 238/501 (47%), Positives = 350/501 (69%), Gaps = 11/501 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTNS 76
E P S + IE+ + +FGW FLQA LV+ A FFDAQQ+FI+++TD+ P WHC NS
Sbjct: 16 ESNLPPPRSLEETIERCIGDFGWAQFLQAALVSFAWFFDAQQTFITVFTDSQPMWHCDNS 75
Query: 77 -----IC-TSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLG 130
+C TSSS++C LP +WSWD +P +IIS W L+CA +F+ G P SSFF+GCL+G
Sbjct: 76 DRVDSVCNTSSSNLCTLPNQTWSWDLNPHVSIISEWGLQCAGSFLKGFPASSFFLGCLIG 135
Query: 131 SFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVL 190
L+ LADSS+GRKNML+ SC+ MS++SML FST++W+Y+ L+FL G R++IGTC L
Sbjct: 136 GLALSTLADSSLGRKNMLLLSCLIMSLSSMLTAFSTSIWVYAFLRFLNGCGRATIGTCAL 195
Query: 191 VLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIA 250
VL TE V +WR +VG + +F FT+G++SLP YIN +SW++LY+W+S+P + Y +
Sbjct: 196 VLSTELVGKKWRGQVGAMGFFCFTLGFLSLPMLGYINEGNSWRNLYVWTSIPTLIYCCLV 255
Query: 251 YLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYS 310
FV ESPRWL+++GR++E + +L+ ++S + NL + + S +Y
Sbjct: 256 RSFVRESPRWLIVKGRKEEAVSILQSIAS-NAITMSFTNLCFE---VENDQSKSNPDVYD 311
Query: 311 SIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVA 370
++ L K W+ R++A M++G G+GMVY+GMPLA+ NL FN+YL VVF+A E P+ +
Sbjct: 312 ALKILVRKSWSFRRLLAAMVVGFGIGMVYYGMPLALTNLNFNLYLGVVFNALSEFPAFLI 371
Query: 371 T-YFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIY 429
T +F++ + R+ +++ F+ L GI AVL ++ + ++VL +V+FF ACTA+N+ LIY
Sbjct: 372 TFFFIDKINRRDALIGFTALSGISSALIAVLGQQLGSLQIVLELVSFFSACTAFNMTLIY 431
Query: 430 IIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFL 489
IE+FPTCVRN+ S+VRQA+VFG +F P +++AGR+N +SYG+FG++I L +F L
Sbjct: 432 TIEMFPTCVRNSAISMVRQALVFGGVFSPVMVAAGRENQFWSYGLFGLIIGLCGLFVFGL 491
Query: 490 PETIGIVLCDTMDQQEKKEIA 510
PET G VLCDTMD++E K +A
Sbjct: 492 PETRGSVLCDTMDEEEYKTLA 512
>UniRef100_O64515 YUP8H12R.2 protein [Arabidopsis thaliana]
Length = 527
Score = 491 bits (1263), Expect = e-137
Identities = 243/497 (48%), Positives = 349/497 (69%), Gaps = 13/497 (2%)
Query: 25 SWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCT--NSICTSS- 81
S D IE + +FGW FLQA LV+ + FDAQQ+FIS++TD+ P WHCT NSIC S
Sbjct: 19 SLDDTIESYIGSFGWAQFLQAALVSFSGVFDAQQTFISVFTDSEPTWHCTDSNSICHESI 78
Query: 82 SDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSS 141
S+IC LP+++WSWD P ++IS W L+CA +F+ GLP+SSFF+GCL+G L+ LADSS
Sbjct: 79 SNICILPKTAWSWDYSPHVSVISEWGLQCAGSFVKGLPESSFFVGCLIGGLVLSTLADSS 138
Query: 142 IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEW 201
+GRKNML SC+ M+I++ML +FS N+W+Y+ L+F+ GF R++IGTC LVL TE V +W
Sbjct: 139 LGRKNMLFLSCLVMAISTMLTVFSPNIWVYAVLRFVNGFGRATIGTCALVLSTELVGKKW 198
Query: 202 RFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWL 261
R RVGI+ +F F +G++SLP AY+NR SSW+ LY W+S+P I Y V+ FV ESPRWL
Sbjct: 199 RGRVGIMSFFGFMLGFLSLPLMAYMNRGSSWRILYAWTSIPTIIYCVLVRFFVCESPRWL 258
Query: 262 VMQGREKEILKMLKRVSSEESADDDS---VNLA-SNLPILPPKEKVSF-FQLYSSIGELF 316
++GR +E + +LKRV+S S D S ++++ S+LP +EK S +++++ L
Sbjct: 259 FVRGRREEAISILKRVASIPSTDVSSGGAISMSFSSLPFEEDEEKPSTNVNIFTTMKVLV 318
Query: 317 HKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-E 375
KRWA+ R+ AVM + G+G+VY+GMPLA+ NL FNIYL+ F+A M+LP+ + T FL +
Sbjct: 319 EKRWALKRLSAVMAIAFGIGLVYYGMPLALSNLDFNIYLSAAFNALMDLPANLITLFLVD 378
Query: 376 NLRRKPSILVFSILGGICCVTCAVLEN----RVPAAKVVLAMVAFFGACTAYNVFLIYII 431
L R+ +++ F+ LGG+ V L N A ++ L ++++F AC+A+N+ +IY I
Sbjct: 379 KLSRRNALIGFTALGGVSSVLIFALHNMRIGNHGALQLALELISYFSACSAFNMEMIYTI 438
Query: 432 ELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPE 491
ELFPTCVRN+ ++ RQA+V G +F P +++AGRKN +S+G+FG+ I L LPE
Sbjct: 439 ELFPTCVRNSAIAMARQALVLGGVFSPIMVAAGRKNAFWSFGLFGLAIGLLGLFAVGLPE 498
Query: 492 TIGIVLCDTMDQQEKKE 508
T G LCDTMD++E K+
Sbjct: 499 TRGSDLCDTMDEEECKD 515
>UniRef100_Q9SA36 F3O9.17 protein [Arabidopsis thaliana]
Length = 521
Score = 428 bits (1101), Expect = e-118
Identities = 222/501 (44%), Positives = 335/501 (66%), Gaps = 12/501 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN- 75
E+ + L++D I+E+SLS+FG+ F Q LV +A+ FDAQQ FI++YTD YP WHC N
Sbjct: 21 EKTRLEALTFDKIVEQSLSDFGFWQFFQISLVGLALLFDAQQIFITVYTDAYPTWHCLNH 80
Query: 76 SICT-SSSDICKLPRSSWSWDT-HPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFF 133
+IC S+SDICKLPRS+W WD ++IS + LEC+S+ + G+P S+F+IG ++G FF
Sbjct: 81 TICDPSASDICKLPRSAWEWDGGSQGKSVISEFGLECSSSLLRGMPSSAFYIGAIVGGFF 140
Query: 134 LAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLL 193
LA + D S+GRK +++FS +MSITS+ +IFSTNVWIY+ LKF+IGF RS + LVL+
Sbjct: 141 LALIPDDSLGRKKLVLFSTFAMSITSISVIFSTNVWIYTFLKFIIGFSRSQTWSYALVLI 200
Query: 194 TEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLF 253
+E+VS WR R ++ + F +G+MSL G A++ ++SSW+ LY+++SVPA+ Y + YLF
Sbjct: 201 SERVSTRWRPRATMIPFTLFVLGFMSLSGIAFLAQDSSWRYLYLYTSVPAVFYCIFLYLF 260
Query: 254 VTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIG 313
ESPRWL MQG++KE + +L ++S +E A +SV S LP+ +E Y SI
Sbjct: 261 ALESPRWLHMQGKDKEAIDVLTKMSPKEKAYLESV--VSKLPL--KQENFEQAPTY-SIK 315
Query: 314 ELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVAT-Y 372
+ F ++WA R++ VMI+ GLG+ Y+G+PLA ++ NIYL+ +A +ELP+ V T
Sbjct: 316 DFFFRKWAFRRILVVMIIMFGLGISYYGVPLAARDIDVNIYLSETLNALVELPTFVITPI 375
Query: 373 FLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLA--MVAFFGACTAYNVFLIYI 430
LE R+ S+LV ++LGG V C VL + + ++ A + FF A +N+ +++
Sbjct: 376 LLERFNRRSSVLVNTLLGGASGVLCFVL-SILGKTEIAFAFELGTFFCARIGFNLMAVFM 434
Query: 431 IELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLP 490
+E+FPTCVR++ T + RQA+V G CP + S GR S+ +FG+ + + LP
Sbjct: 435 VEMFPTCVRSSATMMFRQALVVGGACCPLIASIGRYIPSVSFAIFGIAMSGLGMFVLILP 494
Query: 491 ETIGIVLCDTMDQQEKKEIAL 511
ET G+ LCD+M++QEK++ A+
Sbjct: 495 ETKGLSLCDSMEEQEKRDQAV 515
>UniRef100_Q9SAK7 T8K14.17 protein [Arabidopsis thaliana]
Length = 515
Score = 421 bits (1083), Expect = e-116
Identities = 223/506 (44%), Positives = 329/506 (64%), Gaps = 13/506 (2%)
Query: 12 SHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKW 71
+H +++ L++D I+EKSLS+FG+ FLQ VLV +A+ FD+QQ FI+++TD YP W
Sbjct: 11 THIEEDEDTSSPLTFDKILEKSLSDFGFSQFLQIVLVGLALTFDSQQIFITVFTDAYPTW 70
Query: 72 HCTN-SICT-SSSDICKLPRSSWSWDT-HPSNTIISHWNLECASTFITGLPQSSFFIGCL 128
HC + +IC +++DICK+PRS+W WD ++IS ++LEC+S+F+ LP S+F++G +
Sbjct: 71 HCLDHTICNPATTDICKIPRSAWDWDGGFKGKSVISEFDLECSSSFLRSLPSSTFYVGSI 130
Query: 129 LGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTC 188
+G LA + D S+GRK +L FS +MS+T + I S+N+WIYS LKF+IGF RS GT
Sbjct: 131 VGGVVLAMIPDGSLGRKQLLFFSSFAMSLTGISIFLSSNIWIYSFLKFVIGFARSQTGTY 190
Query: 189 VLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSV 248
LVL++E++S +WR R +V + F +G+MSL G AY+ R++SWK LY+ +S+PA +S+
Sbjct: 191 ALVLISERISTKWRPRATMVPFTLFVLGFMSLSGIAYLVRHASWKVLYLCTSIPAGIHSI 250
Query: 249 IAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQL 308
Y F ESPRWL ++G+ KE +++LKR+S +SV+ L PKE +
Sbjct: 251 FIYFFALESPRWLHLEGKNKEAIEVLKRISPANRGYLESVSSR-----LRPKETLEQTSS 305
Query: 309 YSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSC 368
Y SI +LF +WA R+ VMI+ GLGM Y+G+PLAV ++ NIY++ +A +ELP+
Sbjct: 306 Y-SIKDLFIIKWAFRRVTLVMIIMFGLGMSYYGVPLAVRDIKVNIYMSEALNAMVELPTF 364
Query: 369 VAT-YFLENLRRKPSILVFSILGGICCVTCAV--LENRVPAAKVVLAMVAFFGACTAYNV 425
V T LE R+ S+LV ++GG V C V L R A L + +FF A +N+
Sbjct: 365 VVTPILLEQFSRRSSVLVNCLIGGASGVLCFVMSLYGRTKIA-FALELGSFFCARIGFNL 423
Query: 426 FLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFT 485
IY++ELFPTCVRN+ T ++RQA+V G CP + S GR S+ VFG +
Sbjct: 424 MAIYLVELFPTCVRNSATMMLRQALVVGGACCPLIASLGRNVPSLSFAVFGFAMSGLGLF 483
Query: 486 LFFLPETIGIVLCDTMDQQEKKEIAL 511
LPET G+ LCDTM++QE+++ AL
Sbjct: 484 ALLLPETKGLSLCDTMEEQEQRDQAL 509
>UniRef100_Q6ZF47 Putative organic cation transporter [Oryza sativa]
Length = 532
Score = 398 bits (1022), Expect = e-109
Identities = 193/498 (38%), Positives = 303/498 (60%), Gaps = 13/498 (2%)
Query: 25 SWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN----SICTS 80
S D IE + G +A+L+A A FDAQQ FIS++TD P+WHCT S
Sbjct: 23 SIDDAIETYIGATGAGQLFKAILLAFAWAFDAQQVFISVFTDAEPRWHCTAGADPSCSPG 82
Query: 81 SSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADS 140
++ C LP +W+WD +++S W L+CA + LP SSFF GCL G F L LADS
Sbjct: 83 AASPCALPPGAWAWDRPAETSVVSEWALKCAGPALVSLPASSFFAGCLAGGFLLTTLADS 142
Query: 141 SIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAE 200
+GR+ ML+ S SMS+ +L FS NVW Y+AL+F+ GF RS +GTC LVL TE V
Sbjct: 143 LLGRRKMLLVSLASMSVAGVLTAFSPNVWAYAALRFVCGFGRSMVGTCALVLSTELVGKR 202
Query: 201 WRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRW 260
WR V + + FT+G++SLP AY R +SW+S+Y+W+S+P++ Y+++ Y V ESPRW
Sbjct: 203 WRDTVSVAGFVCFTVGFLSLPALAYTFREASWRSMYLWTSLPSLGYAILLYFLVQESPRW 262
Query: 261 LVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSF-------FQLYSSIG 313
L+++GR+ + ++ ++++++ + + + +E + +++++
Sbjct: 263 LLVRGRKHDAIETVRQIAALNGGGGITCSFSMLHACATEREDDAAGGAGGGGGGVFATLR 322
Query: 314 ELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYF 373
++ +RWA+ R+ A+M G+GMVY+GMPL VGNLG N+YL+V ++A E PS V ++
Sbjct: 323 SMWERRWALRRLAAIMTASFGVGMVYYGMPLNVGNLGSNLYLSVTYNALAEFPSSVLSWL 382
Query: 374 L-ENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIE 432
L + R+ S++ + G+C + C + ++ +++FF CTA+N+ L+Y IE
Sbjct: 383 LMGRINRRSSVVALTAAAGVCSLACVAIPEGT-GGRMAAEVLSFFATCTAFNIILMYSIE 441
Query: 433 LFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET 492
LFPT VRN+ +VRQA+V G + P L++ GR+ + +S+GVFG+ + LPET
Sbjct: 442 LFPTSVRNSAVGMVRQALVLGGVAAPMLVALGRERSFWSFGVFGLAVGCLGLFAVCLPET 501
Query: 493 IGIVLCDTMDQQEKKEIA 510
G + DTM+++E KE A
Sbjct: 502 RGRSMSDTMEEEEHKEAA 519
>UniRef100_Q6ZF45 Putative organic cation transporter [Oryza sativa]
Length = 534
Score = 382 bits (981), Expect = e-104
Identities = 198/518 (38%), Positives = 313/518 (60%), Gaps = 25/518 (4%)
Query: 19 EKKPI--LSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN- 75
E KP LS D IE + G L A+L+A A F+AQQ F+S++TD P WHCT
Sbjct: 14 EAKPAKALSIDDAIETYIGATGARQLLTAMLLAFAWAFEAQQVFMSVFTDAEPTWHCTGV 73
Query: 76 ------SICT-----SSSDICKLPRSSWSWDTHPSNTIISHWNLECAS--TFITGLPQSS 122
S C+ +S+ C LP +W WD +++S W L+C + LP SS
Sbjct: 74 AAGDPGSFCSLAAASASASACALPPGTWEWDRPAETSVVSEWALKCGGGGPALVSLPASS 133
Query: 123 FFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWR 182
FF G L G F L LAD+ +GR+ ML+ S V+MS+ +L +FS NVW+Y+AL+F+ GF R
Sbjct: 134 FFAGNLAGGFLLTTLADTLLGRRKMLVLSLVTMSVAGVLTVFSPNVWVYAALRFVCGFCR 193
Query: 183 SSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVP 242
S+ GT +VL TE V WR V + + F++G+MSLP AY R +SW+++Y+W+S+P
Sbjct: 194 STAGTSAMVLSTELVGKWWRNTVSVAAFVFFSVGFMSLPALAYTLREASWRNMYVWTSLP 253
Query: 243 AICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPK-- 300
++CY+V+ Y V ESPRWL+++GR++E + L++++S + + + + L +
Sbjct: 254 SLCYAVLLYFLVQESPRWLLVRGRKQEAIAALRQIASLNGGEGITTSSFTKLETCAGEVG 313
Query: 301 EKVSFFQ-LYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVF 359
+ V+ + ++S++ + +RWA+ R+ A+ G+G+VY+GMPL+VG+L ++YL+V +
Sbjct: 314 DGVAGGEGMFSTLRSICERRWALRRLAAITTATFGVGVVYYGMPLSVGSLSSDLYLSVAY 373
Query: 360 SASMELPSCVATYFL-ENLRRKPSILVFSILGGICCVTCAVLENRVPAA-----KVVLAM 413
+A+ ELPS V ++ L R+ S++ + G+C + C V+ + A ++ +
Sbjct: 374 NAAAELPSSVLSWLLMGRFNRRSSLVALTAASGLCSLACVVIPDPEAGAGGSHLRLAAEL 433
Query: 414 VAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYG 473
+FF +C AY+V L+Y IELFPT VRN+ LVRQA V G + P L++ GR+ + +S+G
Sbjct: 434 ASFFASCAAYDVLLMYSIELFPTSVRNSAVGLVRQAGVLGGVVAPMLVALGRERSYWSFG 493
Query: 474 VFGVVIMLSNFTLFFLPETIGIVLCDTMDQQEKKEIAL 511
VFG+ + + FLPET G L DTM+ +E+ L
Sbjct: 494 VFGLTVGCLGLFVTFLPETKGRRLSDTMEDEEEAAAVL 531
>UniRef100_Q6ZF37 Putative organic cation transporter [Oryza sativa]
Length = 537
Score = 380 bits (975), Expect = e-104
Identities = 189/500 (37%), Positives = 303/500 (59%), Gaps = 13/500 (2%)
Query: 25 SWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN----SICTS 80
S D IE + G +A+L+A A FDAQQ FIS++TD P+WHCT S
Sbjct: 22 SIDDAIETYIGATGAGQLFKAILLAFAWAFDAQQVFISVFTDAEPRWHCTAGADPSCSPG 81
Query: 81 SSDICKLPRSSWSWDTHPSNTIISHWNLECAS--TFITGLPQSSFFIGCLLGSFFLAALA 138
++ C LP +W+WD +++S W L+C + LP SSFF G L G F L LA
Sbjct: 82 AASPCALPPGAWAWDRPAETSVVSEWALKCGGGGPALVSLPASSFFAGNLAGGFLLTTLA 141
Query: 139 DSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVS 198
D+ +GR+ ML+ S +MS+ +L FS NVW+Y+AL+F+ GF RS +GT +VL TE V
Sbjct: 142 DTHLGRRKMLVLSLATMSVAGVLTAFSPNVWVYAALRFVSGFGRSMVGTSAMVLSTELVG 201
Query: 199 AEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESP 258
WR V + + F++G+MSLP AY R +SW+++Y+W+S+P++CY+V+ Y V ESP
Sbjct: 202 KWWRNTVSVAGFVLFSVGFMSLPALAYTLREASWRTMYVWTSLPSLCYAVLLYFLVQESP 261
Query: 259 RWLVMQGREKEILKMLKRVSS---EESADDDSVNLASNLPILPPKEKVSFFQLYSSIGEL 315
RWL+++GR +E ++ L++++S E S ++ + +++S+ +
Sbjct: 262 RWLLVRGRNQEAIEALRQIASLNGGEGVTTSSFSMLDACAVEVGDGVAGGDGMFASLRLI 321
Query: 316 FHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL- 374
+ +RWA R+ A+M G+G+VY+G+PL+VG+L ++YL+V ++A+ ELPS V ++ L
Sbjct: 322 WERRWAFQRLAAMMTASFGVGVVYYGLPLSVGSLSSDLYLSVAYNAAAELPSSVLSWLLM 381
Query: 375 ENLRRKPSILVFSILGGICCVTCAVL---ENRVPAAKVVLAMVAFFGACTAYNVFLIYII 431
R+ S++ + G+C + C V+ E ++ + +FF +C A++V L+Y I
Sbjct: 382 GRFNRRSSLVALTAASGLCSLACVVIPDEEAGTGGLRLAAELASFFASCAAFDVMLMYSI 441
Query: 432 ELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPE 491
ELFPT VRN+ LVR+A V G + P L++ GR+ + +S+GVFG+ + + +LPE
Sbjct: 442 ELFPTSVRNSAVGLVRKAAVLGGVVAPMLVALGRERSYWSFGVFGLAVGCLGLFVTWLPE 501
Query: 492 TIGIVLCDTMDQQEKKEIAL 511
T G L DTM+++E+ A+
Sbjct: 502 TKGRRLSDTMEEEEEAAAAI 521
>UniRef100_Q5ZCY9 Putative solute carrier family 22 member 3 [Oryza sativa]
Length = 583
Score = 342 bits (876), Expect = 2e-92
Identities = 180/476 (37%), Positives = 283/476 (58%), Gaps = 31/476 (6%)
Query: 57 QQSFISIYTDNYPKWHCTN-------------SICT---SSSDICKLPRSSWSWDTHPSN 100
+Q F+S++TD P WHCT S C+ +S+ C LP +W WD
Sbjct: 93 RQVFMSVFTDAEPPWHCTGVVDAAAAAAADSGSSCSPPAASASPCALPPGTWEWDRPAET 152
Query: 101 TIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSM 160
+++S W L C ++ LP SSFF G L G F LA LAD+ +GR+ ML+ S V+MS+ +
Sbjct: 153 SVVSDWALNCGPALVS-LPASSFFAGNLAGGFLLATLADTHLGRRKMLLLSLVTMSVAAA 211
Query: 161 LIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSL 220
L FS NVW+YSAL+F+ GF RS +GT +VL TE WR V + F++G++SL
Sbjct: 212 LTAFSPNVWVYSALRFVSGFGRSMVGTSAMVLSTELDGKRWRNTVSAAGFVFFSVGFVSL 271
Query: 221 PGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSE 280
P AY R +SW+++Y+W+S+P++CY+V+ YL V ESPRWL+++GR++E ++ +++++S
Sbjct: 272 PALAYTFREASWRNMYVWTSLPSLCYAVLLYLLVQESPRWLLVRGRKQEAIEAVRQIASL 331
Query: 281 ESADDD-SVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVY 339
+ + S L + +++++ ++ +R A+ R+ A+ G+GMVY
Sbjct: 332 NGGGGGITTSSFSMLDACAVELGDGGEGMFATLHSIWERRRALRRLAAITAASFGVGMVY 391
Query: 340 FGMPLAVGNLG-FNIYLAVVFSASMELPSCVATYFLEN--LRRKPSILVFSILGGIC--- 393
+GMPL VG+L N+YL+V ++A ELPS + + L R+ S++ + G+C
Sbjct: 392 YGMPLNVGSLSPSNLYLSVAYNAVAELPSSILAWLLMGRWFNRRGSVVALTTASGLCSLA 451
Query: 394 -CVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVF 452
CV VL + A++ + +FF +CTAY++ L+Y IELFPT VRN+ LVRQA+
Sbjct: 452 ACVPAVVLPD---GARMAAEVASFFASCTAYDMMLMYTIELFPTSVRNSAVGLVRQAVAL 508
Query: 453 GCIFCPFLISAGRKNNIY---SYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQQE 505
G + P L++ GR+ Y S+GVFG+ + + LPET G L DTM+++E
Sbjct: 509 GGVVAPVLVALGRETTSYWSSSFGVFGLAVGCLGLLVTCLPETRGRRLSDTMEEEE 564
Score = 56.2 bits (134), Expect = 2e-06
Identities = 30/87 (34%), Positives = 42/87 (47%), Gaps = 18/87 (20%)
Query: 61 ISIYTDNYPKWHCTN-------------SICTS---SSDICKLPRSSWSWDTHPSNTIIS 104
+S++TD P WHCT S C+S S+ C LP +W WD +++S
Sbjct: 1 MSVFTDAEPPWHCTGVVDAVEAAAGDSGSSCSSPATSASPCALPLGTWEWDRLAKTSVVS 60
Query: 105 HWNLECA--STFITGLPQSSFFIGCLL 129
W L C+ + LP SSFF G L+
Sbjct: 61 DWALNCSGGGPALVSLPASSFFTGNLV 87
>UniRef100_Q9CAT6 Putative transporter; 29320-27598 [Arabidopsis thaliana]
Length = 539
Score = 334 bits (857), Expect = 3e-90
Identities = 196/496 (39%), Positives = 286/496 (57%), Gaps = 20/496 (4%)
Query: 24 LSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYP--KWHCTNSICTSS 81
L+ D +IE+ + G+ L A+LV++A FDAQ + ISI++D P + T +I +
Sbjct: 43 LTVDEVIEQHIGALGFAQILHALLVSIAWIFDAQTTLISIFSDAQPAARLLATGAIVEGA 102
Query: 82 SDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSS 141
S +C L W W S+T++S WNL C F+ +P + FFIG L GS LADS
Sbjct: 103 S-LCGLASGEWEWIGPKSDTVVSEWNLICQHKFLVAVPSTLFFIGSLFGSGVYGYLADSW 161
Query: 142 IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEW 201
GRK L+ SCV +T+ I FS NVW+Y+ L+F GF+RS IG+C +VL TE V +W
Sbjct: 162 FGRKKTLLLSCVLTFVTAFAISFSPNVWVYAFLRFANGFFRSGIGSCCIVLATEIVGKKW 221
Query: 202 RFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWL 261
R +VG +F FT+G++SLP AY+ R SW++LY S + Y+V F ESPRWL
Sbjct: 222 RGQVGQYGFFFFTLGFLSLPLMAYLER-KSWRNLYRIISFLPLGYAVCLLPFAYESPRWL 280
Query: 262 VMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWA 321
+++GR KE + +LK++ A + L ++L ++ P + SS + + +WA
Sbjct: 281 LVKGRNKEAMVVLKKL-----ARLNGKQLPADLSLVDPIPERD--DQTSSSEKFWKTKWA 333
Query: 322 VIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSC-VATYFLENLRRK 380
V R+I VM+ G G G VY+G+ L NL FN+YL V +A ME P+ + ++ L + R+
Sbjct: 334 VKRIIMVMMAGFGSGFVYYGIQLNAENLNFNLYLTVAVNALMEFPAVFIGSFLLGVMNRR 393
Query: 381 PSILVFSILGGICCVTCAVLE-NRVPAA-------KVVLAMVAFFGACTAYNVFLIYIIE 432
P S L G C+ CAVL +RV A ++ + V F + TAY+V +Y +E
Sbjct: 394 PLFSNSSYLAGFACLLCAVLSIHRVIRAISVAKWLQLAVEAVGFMASSTAYDVLYVYCVE 453
Query: 433 LFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET 492
LFPT VRNT SL+RQA + G P L++ GR++ + S+ VFGV +LS +L ET
Sbjct: 454 LFPTNVRNTAVSLLRQAFMLGASAAPLLVALGRESAMMSFIVFGVASVLSGIVSLWLRET 513
Query: 493 IGIVLCDTMDQQEKKE 508
L +T+ QQ K E
Sbjct: 514 RNAPLYETLAQQGKAE 529
>UniRef100_Q84L99 Putative organic cation transport protein [Phaseolus vulgaris]
Length = 547
Score = 325 bits (833), Expect = 2e-87
Identities = 185/524 (35%), Positives = 289/524 (54%), Gaps = 34/524 (6%)
Query: 14 SNIE-QEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPK-W 71
+N+E E K L+ D ++E+ + + G+ + +LV++A FDAQ + ++I++D P W
Sbjct: 13 TNLEGSEAKLELTVDEVVEEYVGSLGFSQLVHVLLVSLAWIFDAQSTLVTIFSDAQPSAW 72
Query: 72 HCTNSICTSSSD------ICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFI 125
C + C +SS +C+L +W W +++II+ WNL C F+ +P S +F+
Sbjct: 73 RCKSGFCNNSSSSSSTGSVCELVPGTWEWVGGHTSSIIAEWNLICDRRFLAAVPASVYFL 132
Query: 126 GCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSI 185
G L+GS L+D+ +GRK + SC+ SIT+ S N+W Y+ +F GF RS I
Sbjct: 133 GSLIGSGVYGHLSDTWLGRKRSVQLSCILTSITAFATSLSPNIWTYAFFRFTNGFARSGI 192
Query: 186 GTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAIC 245
G C LVL TE V +WR +VG +F FT+G+++LP AY R + W+SLY S+ +
Sbjct: 193 GICCLVLTTESVGRKWRGQVGQYGFFFFTIGFLTLPLVAYPTR-THWRSLYKLLSLLPLA 251
Query: 246 YSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSF 305
YSV+ V+ESPRWL+++GR KE L++L R + L +NL + P +
Sbjct: 252 YSVLILPLVSESPRWLLIRGRTKEALQVLDRFARLNGKQ----KLPTNLTLTNPSGSSNV 307
Query: 306 FQLYSSIG---------ELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLA 356
+S G L+ +WA IRM+ VM+ G G+G VY+G+ L V NL FN+Y++
Sbjct: 308 APCTTSEGGERTPNNKENLWTTKWAAIRMVTVMLAGFGVGFVYYGVQLNVENLNFNLYVS 367
Query: 357 VVFSASMELPSCV-ATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKV------ 409
V +A ME+P+ V T+ L R+ + V S + + + C +R +KV
Sbjct: 368 VALNALMEIPAVVIGTFLLGFTNRRLLLSVSSYIAAVSSILCTFFSHRGKTSKVHSNHGG 427
Query: 410 -----VLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAG 464
++ + F GA T +++ IY +ELFPT VRN S++RQAI+ G P L+ G
Sbjct: 428 SWAQLIIEAIGFMGASTTFDILYIYCVELFPTNVRNFAVSMLRQAIMLGASVAPLLVVLG 487
Query: 465 RKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQQEKKE 508
R + S+ VFG + S +LPET L +T+ QQE++E
Sbjct: 488 RLSPSLSFFVFGAFSISSGVLSLWLPETRNAPLYETLKQQEEEE 531
>UniRef100_Q8L7V4 At1g79410/T8K14_17 [Arabidopsis thaliana]
Length = 378
Score = 300 bits (768), Expect = 7e-80
Identities = 165/378 (43%), Positives = 237/378 (62%), Gaps = 10/378 (2%)
Query: 137 LADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEK 196
+ D S+GRK +L FS +MS+T + I S+N+WIYS LKF+IGF RS GT LVL++E+
Sbjct: 2 IPDGSLGRKQLLFFSSFAMSLTGISIFLSSNIWIYSFLKFVIGFARSQTGTYALVLISER 61
Query: 197 VSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTE 256
+S +WR R +V + F +G+MSL G AY+ R++SWK LY+ +S+PA +S+ Y F E
Sbjct: 62 ISTKWRPRATMVPFTLFVLGFMSLSGIAYLVRHASWKVLYLCTSIPAGIHSIFIYFFALE 121
Query: 257 SPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELF 316
SPRWL ++G+ KE +++LKR+S +SV+ L PKE + Y SI +LF
Sbjct: 122 SPRWLHLEGKNKEAIEVLKRISPANRGYLESVSSR-----LRPKETLEQTSSY-SIKDLF 175
Query: 317 HKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVAT-YFLE 375
+WA R+ VMI+ GLGM Y+G+PLAV ++ NIY++ +A +ELP+ V T LE
Sbjct: 176 IIKWAFRRVTLVMIIMFGLGMSYYGVPLAVRDIKVNIYMSEALNAMVELPTFVVTPILLE 235
Query: 376 NLRRKPSILVFSILGGICCVTCAV--LENRVPAAKVVLAMVAFFGACTAYNVFLIYIIEL 433
R+ S+LV ++GG V C V L R A L + +FF A +N+ IY++EL
Sbjct: 236 QFSRRSSVLVNCLIGGASGVLCFVMSLYGRTKIA-FALELGSFFCARIGFNLMAIYLVEL 294
Query: 434 FPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETI 493
FPTCVRN+ T ++RQA+V G CP + S GR S+ VFG + LPET
Sbjct: 295 FPTCVRNSATMMLRQALVVGGACCPLIASLGRNVPSLSFAVFGFAMSGLGLFALLLPETK 354
Query: 494 GIVLCDTMDQQEKKEIAL 511
G+ LCDTM++QE+++ AL
Sbjct: 355 GLSLCDTMEEQEQRDQAL 372
>UniRef100_Q9LHQ6 Organic anion transporter-like protein [Arabidopsis thaliana]
Length = 526
Score = 250 bits (638), Expect = 8e-65
Identities = 148/489 (30%), Positives = 241/489 (49%), Gaps = 10/489 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTNS 76
E E + L D ++++ FG VL +A +A + + I+ D P+W C S
Sbjct: 25 EAEGEERLCIDEMLQRYCGEFGRWQLKHFVLTCIAWALEAFHTMVMIFADQEPEWRCVGS 84
Query: 77 ICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAA 136
C S C+L SSW W ++ +S W L C + GL Q+ FF GC++G+
Sbjct: 85 DCRVGSLNCELDPSSWEWTAGKGSSTVSEWGLICGDKYKVGLVQALFFAGCMIGAGVFGH 144
Query: 137 LADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEK 196
L+DS +GRK L C+ +I + FS N W Y L+FL GF +G VL TE
Sbjct: 145 LSDSKLGRKGSLTVVCIINAIFGIATAFSPNYWTYVVLRFLTGFSTGGVGLTAFVLATEP 204
Query: 197 VSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTE 256
+ R G+ ++ F+ G L G AY+ R SW+ L+I SS+P++ + +I F++E
Sbjct: 205 IGPSKRGVAGMSTFYFFSAGIAVLSGIAYVFR--SWRELFIVSSLPSLLFLLIVIPFISE 262
Query: 257 SPRWLVMQGREKEILKMLKRVSSEESADDDS-VNLASNLPILPPKEKVSFFQLYSSIGEL 315
SPRW +++G+ E +K++ ++ + V LA + + + + + S+ ++
Sbjct: 263 SPRWYLVRGKVDEAMKLMHSIAKTNGRHIPAGVTLALDDDVENNNGERNT-AVEGSLKDV 321
Query: 316 FHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPS-CVATYFL 374
+R++ + + + +VY+G+ L VGNL N+YL V +A E+P+ + L
Sbjct: 322 ILSPLMRMRLVISVAISFTVSIVYYGLSLNVGNLKTNLYLNVFVNAVSEMPAFAITAVLL 381
Query: 375 ENLRRKPSILVFSILGGICCVTCAVLENRVP--AAKVVLAMVAFFGACTAYNVFLIYIIE 432
+ RKP + + C+ + P + ++V ++ FG YN+ IYI E
Sbjct: 382 DKYGRKPLSIGTQWFSCVFCLVGFSVWGAGPWKSVRMVSGVLGIFGMAGTYNLLFIYIAE 441
Query: 433 LFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET 492
LFPT VRN QA G I PF++ G + +GVF V ++ F+LPET
Sbjct: 442 LFPTVVRNAALGCATQAAQMGAILAPFVVVLGEE---LPFGVFAVCGLVGGGLAFYLPET 498
Query: 493 IGIVLCDTM 501
+ L DTM
Sbjct: 499 LNKPLYDTM 507
>UniRef100_Q8LAB6 Hypothetical protein [Arabidopsis thaliana]
Length = 328
Score = 202 bits (513), Expect = 3e-50
Identities = 125/326 (38%), Positives = 185/326 (56%), Gaps = 17/326 (5%)
Query: 192 LLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAY 251
L TE V +WR +VG +F FT+G++SLP AY+ R S W++LY S + Y+V
Sbjct: 1 LATEIVGKKWRGQVGQYGFFFFTLGFLSLPLMAYLERKS-WRNLYRIISFLPLGYAVCLL 59
Query: 252 LFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSS 311
F ESPRWL+++GR KE + +LK++ A + L ++L ++ P + SS
Sbjct: 60 PFAYESPRWLLVKGRNKEAMVVLKKL-----ARLNGKQLPADLSLVDPIPERD--DQTSS 112
Query: 312 IGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSC-VA 370
+ + +WAV R+I VM+ G G G VY+G+ L NL FN+YL V +A ME P+ +
Sbjct: 113 SEKFWKTKWAVKRIIMVMMAGFGSGFVYYGIQLNAENLNFNLYLTVAVNALMEFPAVFIG 172
Query: 371 TYFLENLRRKPSILVFSILGGICCVTCAVLE-NRVPAA-------KVVLAMVAFFGACTA 422
++ L + R+P S L G C+ CAVL +RV A ++ + V F + TA
Sbjct: 173 SFLLGVMNRRPLFSNSSYLAGFACLLCAVLSIHRVIRAISVAKWLQLAVEAVGFMASSTA 232
Query: 423 YNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLS 482
Y+V +Y +ELFPT VRNT SL+RQA + G P L++ GR++ + S+ VFGV +LS
Sbjct: 233 YDVLYVYCVELFPTNVRNTAVSLLRQAFMLGASAAPLLVALGRESAMMSFIVFGVASVLS 292
Query: 483 NFTLFFLPETIGIVLCDTMDQQEKKE 508
+L ET L +T+ QQ K E
Sbjct: 293 GIVSLWLRETRNAPLYETLAQQGKAE 318
>UniRef100_Q9Z0E8 Organic cation/carnitine transporter 2 [Mus musculus]
Length = 557
Score = 173 bits (439), Expect = 1e-41
Identities = 130/423 (30%), Positives = 208/423 (48%), Gaps = 27/423 (6%)
Query: 92 WSWDTHPS-NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIF 150
W +D +TI++ W+L C + L S FF+G L+GSF L+D GRKN+L
Sbjct: 117 WEYDKDVFLSTIVTEWDLVCKDDWKAPLTTSLFFVGVLMGSFISGQLSDR-FGRKNVLFL 175
Query: 151 SCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRF---RVGI 207
+ + S L +FS N +++ L L+G + S VL TE +S R +G+
Sbjct: 176 TMGMQTGFSFLQVFSVNFEMFTVLFVLVGMGQISNYVAAFVLGTEILSKSIRIIFATLGV 235
Query: 208 VEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGRE 267
++ F G+M LP FAY R+ W+ L + +VP + + + F+ ESPRWL+ QGR
Sbjct: 236 CIFYAF--GFMVLPLFAYFIRD--WRMLLLALTVPGVLCGAL-WWFIPESPRWLISQGRI 290
Query: 268 KEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHK----RWAVI 323
KE ++++ + + + I P E L S+ +L H R I
Sbjct: 291 KEAEVIIRKAAKING-------IVAPSTIFDPSE---LQDLNSTKPQLHHIYDLIRTRNI 340
Query: 324 RMIAVM--ILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-ENLRRK 380
R+I +M IL + + + YFG+ L NL +IY+ A++E+P+ V + L + L R+
Sbjct: 341 RVITIMSIILWLTISVGYFGLSLDTPNLHGDIYVNCFLLAAVEVPAYVLAWLLLQYLPRR 400
Query: 381 PSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRN 440
SI LGG + ++ + + L MV FG +AY++ +Y EL+PT VRN
Sbjct: 401 YSISAALFLGGSVLLFMQLVPSELFYLSTALVMVGKFGITSAYSMVYVYTAELYPTVVRN 460
Query: 441 TTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDT 500
+ A G I P+ + G + Y + G + +L+ F PE+ G+ L DT
Sbjct: 461 MGVGVSSTASRLGSILSPYFVYLGAYDRFLPYILMGSLTILTAILTLFFPESFGVPLPDT 520
Query: 501 MDQ 503
+DQ
Sbjct: 521 IDQ 523
>UniRef100_UPI000036EE60 UPI000036EE60 UniRef100 entry
Length = 540
Score = 173 bits (438), Expect = 1e-41
Identities = 127/438 (28%), Positives = 218/438 (48%), Gaps = 10/438 (2%)
Query: 69 PKWHCT--NSICTSSSDICKLPR-SSWSWDTHP-SNTIISHWNLECASTFITGLPQSSFF 124
P+W N+ TS S+ P W +D ++TI++ W+L C+S + L QS F
Sbjct: 91 PQWQLLDPNATATSWSEADTEPCVDGWVYDRSVFTSTIVAKWDLVCSSQGLKPLSQSIFM 150
Query: 125 IGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSS 184
G L+GSF L+ GRK ML + C+ +++ IF+ IY L+F+ F +
Sbjct: 151 SGILVGSFIWGLLS-YRFGRKPMLSWCCLQLAVAGTSTIFAPTFVIYCGLRFVAAFGMAG 209
Query: 185 IGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAI 244
I L L+ E + R V F+ G +L G A+ R+ W++L + +SVP
Sbjct: 210 IFLSSLTLMVEWTTTSRRAVTMTVVGCAFSAGQAALGGLAFALRD--WRTLQLAASVPFF 267
Query: 245 CYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVS 304
S+I++ ++ ES RWL+++G+ + L+ L++V+ + ++ NL + + KE+V+
Sbjct: 268 AISLISW-WLPESARWLIIKGKPDQALQELRKVA-RINGHKEAKNLTIEVLMSSVKEEVA 325
Query: 305 FFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASME 364
+ S+ +LF R A++++ G +FG+ L + LG NI+L +F ++
Sbjct: 326 SAKEPRSVLDLFCMPVLRWRSCAMLVVKFAFGFTFFGLALDLQALGSNIFLLQMFIGVVD 385
Query: 365 LPSCV-ATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAY 423
+P+ + A L L R+P++ +L G+C + ++ + + A + LA++ G A+
Sbjct: 386 IPAKMGALLLLSRLGRRPTLAASLLLAGLCILANTLVPHEMGALRSALAVLGLGGVGAAF 445
Query: 424 NVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSN 483
IY ELFPT +R T L + A G I P + G V+G V +LS
Sbjct: 446 TCITIYSSELFPTVLRMTAVGLGQMAARGGAILGPLVRLLGVHGPWLPLLVYGTVPVLSG 505
Query: 484 FTLFFLPETIGIVLCDTM 501
LPET + L DT+
Sbjct: 506 LAALLLPETQSLPLPDTI 523
>UniRef100_UPI00003AABBA UPI00003AABBA UniRef100 entry
Length = 506
Score = 173 bits (438), Expect = 1e-41
Identities = 124/408 (30%), Positives = 196/408 (47%), Gaps = 32/408 (7%)
Query: 100 NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITS 159
+TI+S WNL C + L S FF+G L+GSF L+D GRKN+L + +I S
Sbjct: 126 STIVSEWNLVCEDDWKVPLTTSLFFVGVLIGSFISGQLSDR-FGRKNVLFSTLAMQTIFS 184
Query: 160 MLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRF---RVGIVEYFTFTMG 216
+ IFST+ ++S L+G + S VL TE + R +G+ ++ F G
Sbjct: 185 FIQIFSTSWEMFSVFFLLVGMGQISNYVAAFVLGTEILGKSIRLLFCTLGVCIFYAF--G 242
Query: 217 YMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKR 276
YM LP FAY R+ W+ L + ++P + + + + ESPRWL+ QGR +E ++++
Sbjct: 243 YMLLPLFAYFIRD--WRMLLLALTLPGLL-CIPLWWIIPESPRWLISQGRFQEAEDIIRK 299
Query: 277 VSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGLG 336
+ V I P E V L F MI+ +G
Sbjct: 300 AAKINKVTAPDV-------IFDPSENVLKSNLTDGSCSSFR-----------MIISVG-- 339
Query: 337 MVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-ENLRRKPSILVFSILGGICCV 395
YFG+ L NL ++YL SA +E+P+ + ++ L NL R+ S+ LGG +
Sbjct: 340 --YFGLSLDTPNLHGDVYLNCFLSAVIEVPAYIISWLLLRNLPRRYSMAAALFLGGCVLL 397
Query: 396 TCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCI 455
++ + + A ++L M+ FG +A+++ +Y EL+PT VRN A G I
Sbjct: 398 FIQLVPSHLRALSILLVMLGKFGITSAFSMVYVYTAELYPTVVRNMGVGASSMASRLGSI 457
Query: 456 FCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQ 503
P+ G + Y + G + +LS FLPE+ G+ L DT++Q
Sbjct: 458 LSPYFAYLGAYDRFLPYILMGSLTVLSGILTLFLPESYGMPLPDTIEQ 505
>UniRef100_UPI000036EE5E UPI000036EE5E UniRef100 entry
Length = 529
Score = 172 bits (437), Expect = 2e-41
Identities = 126/444 (28%), Positives = 216/444 (48%), Gaps = 10/444 (2%)
Query: 69 PKWHCT--NSICTSSSDICKLPR-SSWSWDTHP-SNTIISHWNLECASTFITGLPQSSFF 124
P+W N+ TS S+ P W +D ++TI++ W+L C+S + L QS F
Sbjct: 91 PQWQLLDPNATATSWSEADTEPCVDGWVYDRSVFTSTIVAKWDLVCSSQGLKPLSQSIFM 150
Query: 125 IGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSS 184
G L+GSF L+ GRK ML + C+ +++ IF+ IY L+F+ F +
Sbjct: 151 SGILVGSFIWGLLS-YRFGRKPMLSWCCLQLAVAGTSTIFAPTFVIYCGLRFVAAFGMAG 209
Query: 185 IGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAI 244
I L L+ E + R V F+ G +L G A+ R+ W++L + +SVP
Sbjct: 210 IFLSSLTLMVEWTTTSRRAVTMTVVGCAFSAGQAALGGLAFALRD--WRTLQLAASVPFF 267
Query: 245 CYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVS 304
S+I++ ++ ES RWL+++G+ + L+ L++V+ + ++ NL + + +E++S
Sbjct: 268 AISLISW-WLPESARWLIIKGKPDQALQELRKVA-RINGHKEAKNLTIEVLLSAMREELS 325
Query: 305 FFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASME 364
Q +S+G L R + G +FG+ L + LG NI+L +F ++
Sbjct: 326 MGQPPASLGTLLRMPGLRFRTCISTLCWFAFGFTFFGLALDLQALGSNIFLLQMFIGVVD 385
Query: 365 LPSCV-ATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAY 423
+P+ + A L L R+P++ +L G+C + ++ + + A + LA++ G A+
Sbjct: 386 IPAKMGALLLLSRLGRRPTLAASLLLAGLCILANTLVPHEMGALRSALAVLGLGGVGAAF 445
Query: 424 NVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSN 483
IY ELFPT +R T L + A G I P + G V+G V +LS
Sbjct: 446 TCITIYSSELFPTVLRMTAVGLGQMAARGGAILGPLVRLLGVHGPWLPLLVYGTVPVLSG 505
Query: 484 FTLFFLPETIGIVLCDTMDQQEKK 507
LPET + L DT+ + +
Sbjct: 506 LAALLLPETQSLPLPDTIQDVQNQ 529
>UniRef100_UPI00003AABB9 UPI00003AABB9 UniRef100 entry
Length = 529
Score = 172 bits (437), Expect = 2e-41
Identities = 122/409 (29%), Positives = 208/409 (50%), Gaps = 16/409 (3%)
Query: 100 NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITS 159
+TI+S WNL C + L S FF+G L+GSF L+D GRK++L + + S
Sbjct: 126 STIVSEWNLVCEDDWKVPLTTSLFFVGVLIGSFISGQLSDR-FGRKSILFLTMAVQTGFS 184
Query: 160 MLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRF---RVGIVEYFTFTMG 216
L IFST+ +++ L ++G + S +L TE + R +G+ +F +G
Sbjct: 185 FLQIFSTSWEMFTVLFLIVGMGQISNYVVAFILGTEILGKSVRILFSTLGVCVFFA--IG 242
Query: 217 YMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKR 276
YM LP FAY R+ W+ L + ++P + V + + ESPRWL+ QGR KE ++++
Sbjct: 243 YMLLPLFAYFIRD--WRMLLLALTLPGLL-CVPLWWIIPESPRWLISQGRYKEAEVIIRK 299
Query: 277 VSSEESADDDSVNL-ASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGL 335
+ + ++ A+ L L + + ++ +I +L R + I +IL + +
Sbjct: 300 AAKINNTPAPAMLFDAAELQDLSSQRQQTY-----TILDLMRTRNILTITIMSVILWMII 354
Query: 336 GMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-ENLRRKPSILVFSILGGICC 394
+ YFG+ L NL ++YL SA +E+P+ + ++ L NL R+ S+ LGG
Sbjct: 355 SVGYFGLSLDTPNLHGDVYLNCFLSAVIEVPAYIISWLLLRNLPRRYSMAAALFLGGCVL 414
Query: 395 VTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGC 454
+ ++ + + A ++L M+ FG +A+++ +Y EL+PT VRN A G
Sbjct: 415 LFIQLVPSHLRALSILLVMLGKFGITSAFSMVYVYTAELYPTVVRNMGVGASSMASRLGS 474
Query: 455 IFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQ 503
I P+ G + Y + G + +LS FLPE+ G+ L DT++Q
Sbjct: 475 ILSPYFAYLGAYDRFLPYILMGSLTVLSGILTLFLPESYGMPLPDTIEQ 523
>UniRef100_O70594 Organic cation/carnitine transporter 2 [Rattus norvegicus]
Length = 557
Score = 172 bits (436), Expect = 2e-41
Identities = 131/430 (30%), Positives = 211/430 (48%), Gaps = 20/430 (4%)
Query: 100 NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITS 159
+TI++ W+L C + L S FF+G L+GSF L+D GRKN+L + + S
Sbjct: 126 STIVTEWDLVCKDDWKAPLTTSLFFVGVLMGSFISGQLSDR-FGRKNVLFLTMGMQTGFS 184
Query: 160 MLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRF---RVGIVEYFTFTMG 216
L +FS N +++ L L+G + S VL TE +S R +G+ ++ F G
Sbjct: 185 FLQLFSVNFEMFTVLFVLVGMGQISNYVAAFVLGTEILSKSIRIIFATLGVCIFYAF--G 242
Query: 217 YMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKR 276
+M LP FAY R+ W+ L + +VP + + + F+ ESPRWL+ QGR KE ++++
Sbjct: 243 FMVLPLFAYFIRD--WRMLLLALTVPGVLCGAL-WWFIPESPRWLISQGRVKEAEVIIRK 299
Query: 277 VSSEESADDDSVNL-ASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVM--ILGI 333
+ S S L L K+ S +Y + R IR+I +M IL +
Sbjct: 300 AAKFNGIVAPSTIFDPSELQDLNSKKPQSH-HIYDLV------RTRNIRIITIMSIILWL 352
Query: 334 GLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-ENLRRKPSILVFSILGGI 392
+ + YFG+ L NL +IY+ A++E+P+ V + L ++L R+ SI LGG
Sbjct: 353 TISVGYFGLSLDTPNLHGDIYVNCFLLAAVEVPAYVLAWLLLQHLPRRYSISAALFLGGS 412
Query: 393 CCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVF 452
+ ++ + + L MV FG +AY++ +Y EL+PT VRN + A
Sbjct: 413 VLLFIQLVPSELFYLSTALVMVGKFGITSAYSMVYVYTAELYPTVVRNMGVGVSSTASRL 472
Query: 453 GCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQQEKKEIALC 512
G I P+ + G + Y + G + +L+ F PE+ G L DT+DQ + +
Sbjct: 473 GSILSPYFVYLGAYDRFLPYILMGSLTILTAILTLFFPESFGAPLPDTIDQMLRVKGIKQ 532
Query: 513 DAMNQQVRTE 522
+ Q RT+
Sbjct: 533 WQIQSQTRTQ 542
>UniRef100_Q5W4T3 OCTN2 protein [Gallus gallus]
Length = 557
Score = 172 bits (435), Expect = 3e-41
Identities = 122/409 (29%), Positives = 207/409 (49%), Gaps = 16/409 (3%)
Query: 100 NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITS 159
+TI++ WNL C + + + S FF+G LLGSF L+ GRKN+L + +I S
Sbjct: 126 STIVTEWNLVCDNDWKGPMSTSLFFVGVLLGSFISGQLS-YKFGRKNVLFSTLAMQTIFS 184
Query: 160 MLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRF---RVGIVEYFTFTMG 216
+ IFST+ ++S L+G + S VL TE + R +G+ ++ F G
Sbjct: 185 FIQIFSTSWEMFSVFFLLVGMGQISNYVAAFVLGTEILGKSIRLLFCTLGVCIFYAF--G 242
Query: 217 YMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKR 276
YM LP FAY R+ W+ L + ++P + + + + ESPRWL+ QGR +E ++++
Sbjct: 243 YMLLPLFAYFIRD--WRMLLLALTLPGLL-CIPLWWIIPESPRWLISQGRFQEAEDIIRK 299
Query: 277 VSS-EESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGL 335
+ + D + S L L + + ++ +I +L R + I +IL + +
Sbjct: 300 AAKINKVTAPDVIFDPSELQDLSSQRQQTY-----TILDLMRTRNILTITIMSVILWMII 354
Query: 336 GMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-ENLRRKPSILVFSILGGICC 394
+ YFG+ L NL ++YL SA +E+P+ + ++ L NL R+ S+ LGG
Sbjct: 355 SVGYFGLSLDTPNLHGDVYLNCFLSAVIEVPAYIISWLLLRNLPRRYSMAAALFLGGCVL 414
Query: 395 VTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGC 454
+ ++ + + A ++L M+ FG +A+++ +Y EL+PT VRN A G
Sbjct: 415 LFIQLVPSHLRALSILLVMLGKFGITSAFSMVYVYTAELYPTVVRNMGVGASSMASRLGS 474
Query: 455 IFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQ 503
I P+ G + Y + G + +LS FLPE+ G+ L DT++Q
Sbjct: 475 ILSPYFAYLGAYDRFLPYILMGSLTVLSGILTLFLPESYGMPLPDTIEQ 523
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.138 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 847,476,396
Number of Sequences: 2790947
Number of extensions: 34259543
Number of successful extensions: 127892
Number of sequences better than 10.0: 2088
Number of HSP's better than 10.0 without gapping: 566
Number of HSP's successfully gapped in prelim test: 1523
Number of HSP's that attempted gapping in prelim test: 124015
Number of HSP's gapped (non-prelim): 2645
length of query: 526
length of database: 848,049,833
effective HSP length: 132
effective length of query: 394
effective length of database: 479,644,829
effective search space: 188980062626
effective search space used: 188980062626
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC121234.10


