

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.8 + phase: 0 /pseudo
(427 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
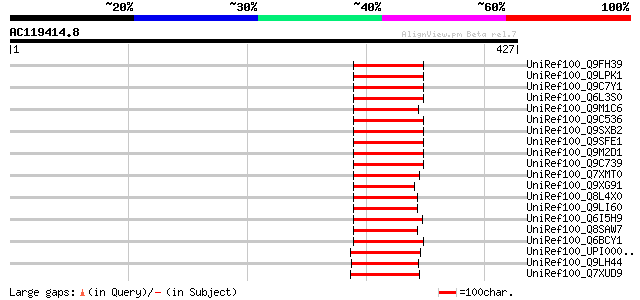
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana] 76 2e-12
UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana] 76 2e-12
UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [... 76 2e-12
UniRef100_Q6L3S0 Putative gag-pol polyprotein [Solanum demissum] 75 5e-12
UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thal... 70 1e-10
UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis t... 69 3e-10
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 69 3e-10
UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana] 69 3e-10
UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana] 69 3e-10
UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis t... 67 1e-09
UniRef100_Q7XMT0 OSJNBa0029L02.7 protein [Oryza sativa] 60 2e-07
UniRef100_Q9XG91 Tpv2-1c protein [Phaseolus vulgaris] 59 2e-07
UniRef100_Q8L4X0 Putative pol polyprotein [Oryza sativa] 59 3e-07
UniRef100_Q9LI60 Similar to Arabidopsis thaliana chromosome II B... 59 3e-07
UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa] 59 3e-07
UniRef100_Q8SAW7 Putative pol polyprotein [Oryza sativa] 59 3e-07
UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas] 59 3e-07
UniRef100_UPI000042EBFE UPI000042EBFE UniRef100 entry 58 5e-07
UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidops... 58 5e-07
UniRef100_Q7XUD9 OSJNBa0088A01.6 protein [Oryza sativa] 58 5e-07
>UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1334
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1194 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1252
>UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana]
Length = 1207
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1067 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1125
>UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [Arabidopsis thaliana]
Length = 1334
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1194 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1252
>UniRef100_Q6L3S0 Putative gag-pol polyprotein [Solanum demissum]
Length = 532
Score = 74.7 bits (182), Expect = 5e-12
Identities = 35/59 (59%), Positives = 45/59 (75%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+++G+G +SWSS K+PIVTLSTTEAE+VAA SC+ IW+R I + QE
Sbjct: 392 KSTSGYVFLLGSGVVSWSSKKQPIVTLSTTEAEYVAATSCAYQAIWVRNILGELDFKQE 450
>UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thaliana]
Length = 1339
Score = 69.7 bits (169), Expect = 1e-10
Identities = 28/55 (50%), Positives = 44/55 (79%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+STS +V+++ +GAI W+S K+P+V LSTTEAE++AAA C+C +WLR++ +G
Sbjct: 1197 RSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLG 1251
>UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1320
Score = 68.6 bits (166), Expect = 3e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1174 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1232
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 68.6 bits (166), Expect = 3e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 68.6 bits (166), Expect = 3e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1145 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1203
>UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1352
Score = 68.6 bits (166), Expect = 3e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 66.6 bits (161), Expect = 1e-09
Identities = 31/59 (52%), Positives = 40/59 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIV LST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>UniRef100_Q7XMT0 OSJNBa0029L02.7 protein [Oryza sativa]
Length = 856
Score = 59.7 bits (143), Expect = 2e-07
Identities = 28/57 (49%), Positives = 40/57 (70%), Gaps = 1/57 (1%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSH-IGE 345
KST+ +YM+G ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + IGE
Sbjct: 715 KSTTGVLYMLGKSLISWQSQKQKVVALSSCEAEYIAATTAACQGIWLTRLLAELIGE 771
>UniRef100_Q9XG91 Tpv2-1c protein [Phaseolus vulgaris]
Length = 374
Score = 59.3 bits (142), Expect = 2e-07
Identities = 29/52 (55%), Positives = 34/52 (64%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITS 341
KSTS YV+ +GN A SW S K+PIVTLST E E+V C IWLR + S
Sbjct: 234 KSTSGYVFFMGNTAFSWLSKKQPIVTLSTCEREYVEHLGVVCHAIWLRNLLS 285
>UniRef100_Q8L4X0 Putative pol polyprotein [Oryza sativa]
Length = 1426
Score = 58.9 bits (141), Expect = 3e-07
Identities = 25/54 (46%), Positives = 39/54 (71%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST+ +YM+G+ ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + +
Sbjct: 1285 KSTTGVLYMLGDSLISWQSQKQKVVALSSCEAEYIAATTGACQGIWLNRLLAEL 1338
>UniRef100_Q9LI60 Similar to Arabidopsis thaliana chromosome II BAC F9O13 genomic
sequence [Oryza sativa]
Length = 1330
Score = 58.9 bits (141), Expect = 3e-07
Identities = 25/54 (46%), Positives = 39/54 (71%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST+ +YM+G+ ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + +
Sbjct: 1189 KSTTGVLYMLGDSLISWQSQKQKVVALSSCEAEYIAATTGACQGIWLNRLLAEL 1242
>UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa]
Length = 1136
Score = 58.9 bits (141), Expect = 3e-07
Identities = 29/58 (50%), Positives = 39/58 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
KSTS Y + +G+G SWS+ K+ IV LS+ EAE+VAA+ +WLRRI +GE Q
Sbjct: 998 KSTSGYAFSLGSGMWSWSTKKQNIVALSSAEAEYVAASKAVSQVVWLRRIMEDLGEKQ 1055
>UniRef100_Q8SAW7 Putative pol polyprotein [Oryza sativa]
Length = 1283
Score = 58.9 bits (141), Expect = 3e-07
Identities = 24/54 (44%), Positives = 38/54 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KSTS ++ V G I+W S K+ +V LS+ EAE++AAA+ +C G+WL R+ + +
Sbjct: 1139 KSTSGQIFFVNGGPITWQSSKQKVVALSSCEAEYIAAAAATCQGVWLARLLAEV 1192
>UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas]
Length = 1298
Score = 58.5 bits (140), Expect = 3e-07
Identities = 28/59 (47%), Positives = 37/59 (62%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KST+ YV+ V GA+SW S + +V STTEAE+VAA S IWL+ + +G QE
Sbjct: 1161 KSTTGYVFKVAGGAVSWVSKLQAVVATSTTEAEYVAATQASKEAIWLKMLLEELGHKQE 1219
>UniRef100_UPI000042EBFE UPI000042EBFE UniRef100 entry
Length = 868
Score = 58.2 bits (139), Expect = 5e-07
Identities = 26/59 (44%), Positives = 38/59 (64%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEG 346
T +ST+ YV+ + G + W S ++P V LSTTEAE++A+A + IWLR + IG G
Sbjct: 709 TRRSTTGYVFKIHGGVVGWKSKRQPTVALSTTEAEYMASADAAKQAIWLRLLLEDIGLG 767
>UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1499
Score = 58.2 bits (139), Expect = 5e-07
Identities = 26/56 (46%), Positives = 38/56 (67%)
Query: 289 EKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+KSTS YV+ +G+GA W+S K+ V ST EAE++A S + IWL+R+ + IG
Sbjct: 1208 KKSTSGYVFTIGSGAFCWNSSKQKTVAQSTAEAEYIAVCSAANQAIWLQRLVNEIG 1263
>UniRef100_Q7XUD9 OSJNBa0088A01.6 protein [Oryza sativa]
Length = 1403
Score = 58.2 bits (139), Expect = 5e-07
Identities = 23/58 (39%), Positives = 39/58 (66%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGE 345
T KST+ V+ +G +SW S+K+ +V LS+ EAE++AA + +C G+WL ++ I +
Sbjct: 1257 TRKSTTGVVFFLGTSPVSWQSVKQKVVALSSCEAEYIAATTAACQGVWLSQLLGEINQ 1314
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.372 0.166 0.671
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 548,825,391
Number of Sequences: 2790947
Number of extensions: 17410155
Number of successful extensions: 62674
Number of sequences better than 10.0: 482
Number of HSP's better than 10.0 without gapping: 473
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 62121
Number of HSP's gapped (non-prelim): 558
length of query: 427
length of database: 848,049,833
effective HSP length: 130
effective length of query: 297
effective length of database: 485,226,723
effective search space: 144112336731
effective search space used: 144112336731
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC119414.8


